Submitted:
25 July 2025
Posted:
27 July 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Preparation of Proteins and Nucleic Acids
| Name | Sequence | Application |
|---|---|---|
| GT-crRNA | UAAUUUCUACUGUUGUAGAUCUCUCUUCACUGUCACGUAC | crRNA for the B1 gene |
| Act-crRNA | UAAUUUCUACUCUUGUAGAUUUCCGCAAUACUCCCCCAGGU | crRNA for Act DNA |
| Act-ssDNA | ACC TGG GGG AGT ATT GCG GAG GAA GGT | single-stranded Act DNA |
| Act-ssDNA-C | ACCTTCCTCCGCAATACTCCCCCAGGT | Complementary strand of Act DNA |
| T7-crRNA-F | GAAATTAATACGACTCACTATAGGG | In vitro transcription of crRNA targeting the hly gene of L. monocytogenes |
| T7-LM2-R2 | CAGGGAGAACATCTGGTTGAATCTACAACAGTAGAAATTCCCTATAGTGAGTCGTATTAATTTC | |
| ssDNA | 5’FAM-TCTACTCTC-3’BHQ1 | Reporter probe |
| Act-H-2p | GGA ACC TGG GGG AGT ATT GCG GAG GAATTTTC CTC CGC AAT ACT CCC CCA GGT TCC | Screening of hairpin DNA loop length |
| Act-H-4p | GGA ACC TGG GGG AGT ATT GCG GAG GAATTTTTTC CTC CGC AAT ACT CCC CCA GGT TCC | |
| Act-H-6p | GGA ACC TGG GGG AGT ATT GCG GAG GAATTTTTTTTC CTC CGC AAT ACT CCC CCAGGT TCC | |
| Act-H-8p | GGA ACC TGG GGG AGT ATT GCG GAG GAATTTTTTTTTTC CTC CGC AAT ACT CCC CCAGGT TCC | |
| Act-H-10p | GGA ACC TGG GGG AGT ATT GCG GAG GAATTTTTTTTTT TTCC TCC GCA ATA CTC CCCCAG GTT CC | |
| Act-H-8p-C | GGA ACC TGG GGG AGT ATT GCG GAG GAACCCCCCCCTTC CTC CGC AAT ACT CCC CCAGGT TCC | Screening of hairpin loop compositions |
| Act-H-8p-A | GGA ACC TGG GGG AGT ATT GCG GAG GAAAAAAAAAATTC CTC CGC AAT ACT CCC CCAGGT TCC | |
| Act-H-8p-G | GGA ACC TGG GGG AGT ATT GCG GAG GAAGGGGGGGGTTC CTC CGC AAT ACT CCC CCAGGT TCC | |
| GT-95DNA-T | GGCGGACCTCTCTTGTCTCGAATACACGAACGAGATCTGCTGGATCTCTTCCCTTGAATGCGATGTCGTACGTGACAGTGAAGAGAGGAAACAGG | B1 gene fragment |
| Act-H-2Loop-Bio | Bio-TGGA ACC TGG GGG AGT ATT GCG GAG GATTTATTT TCC TCC GCA ATACTTATTTC CCC CAG GTTCCA | Screening of labeled hairpin loops |
| Act-H-2Loop-N | TGGA ACC TGG GGG AGT ATT GCG GAG GATTTATTT TCC TCC GCA ATACTTATTTC CCC CAG GTTCCA |
2.2. Polyacrylamide Gel Electrophoresis (PAGE) Analysis
2.3. Validation of Hairpin DNA Effects on Cas12a Trans-Cleavage Activity
2.4. Single Cas12a Assay
2.5. CRISPR- Cascade Assay
2.6. Evaluation of Reaction System Specificity and Sensitivity
2.7. Detection of Artificially Spiked Samples
3. Results
3.1. One-Tube Amplification-Free CRISPR/Cas12a Detection Workflow
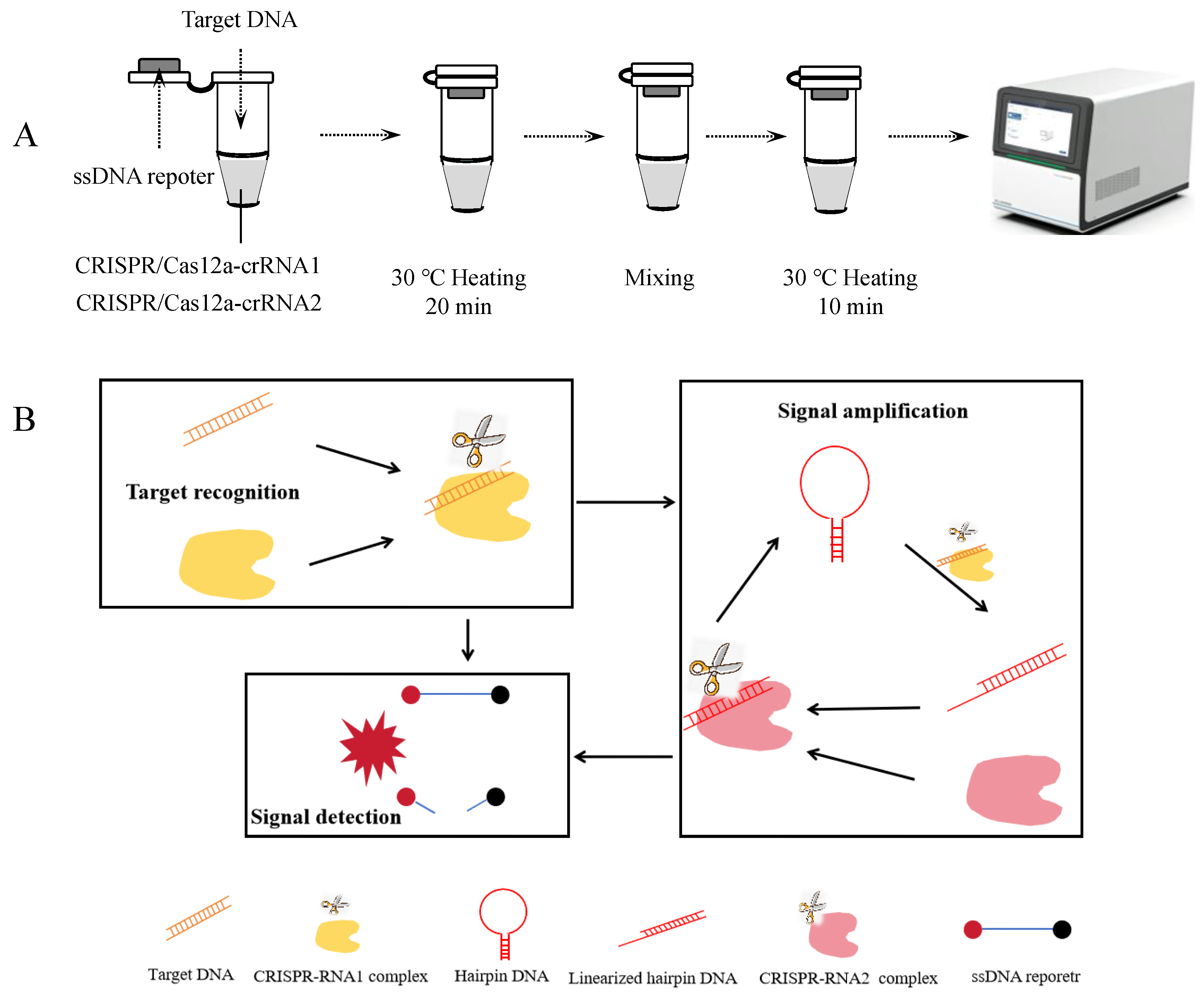
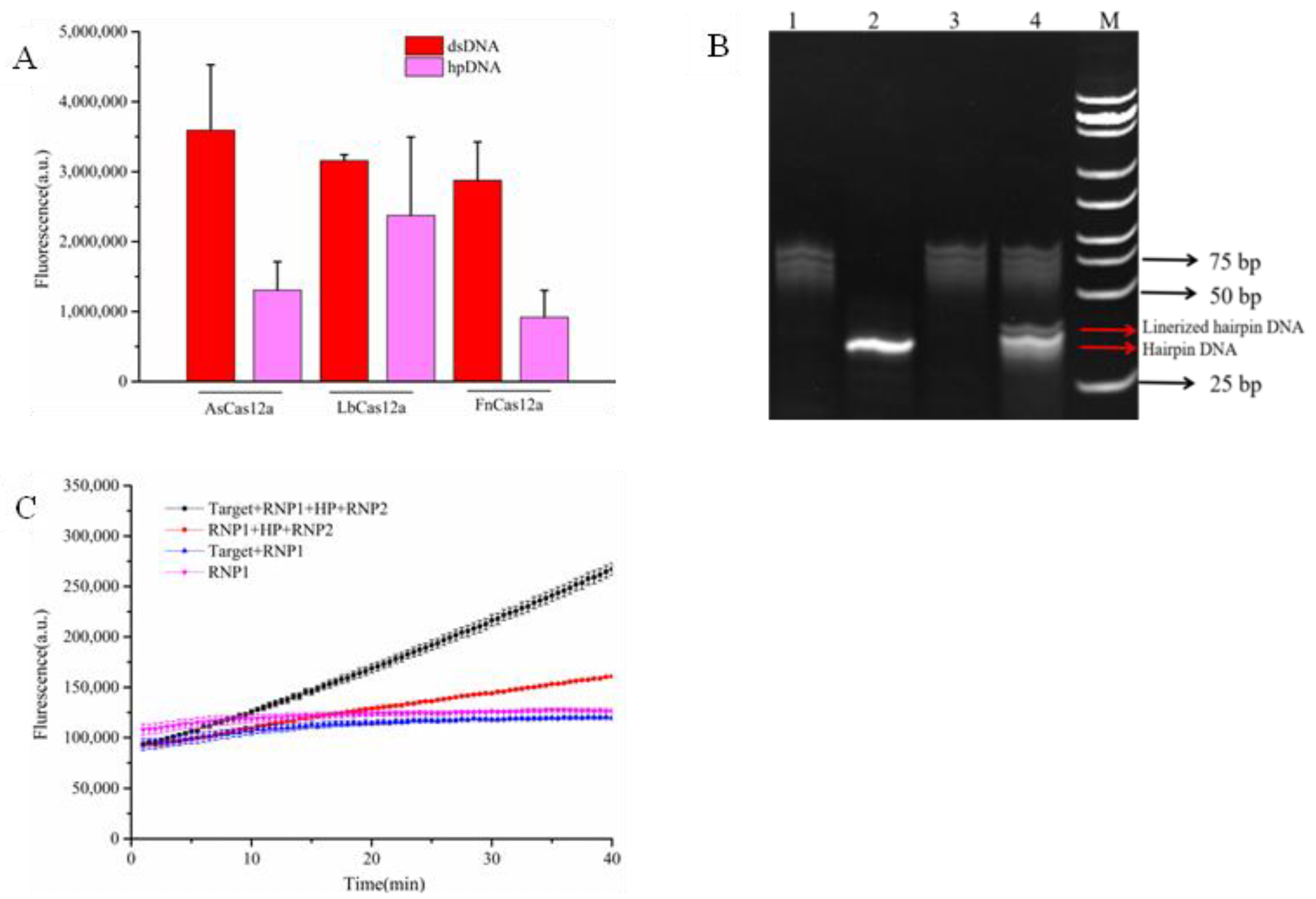
3.2. Feasibility Analysis of the Amplification-Free Detection System
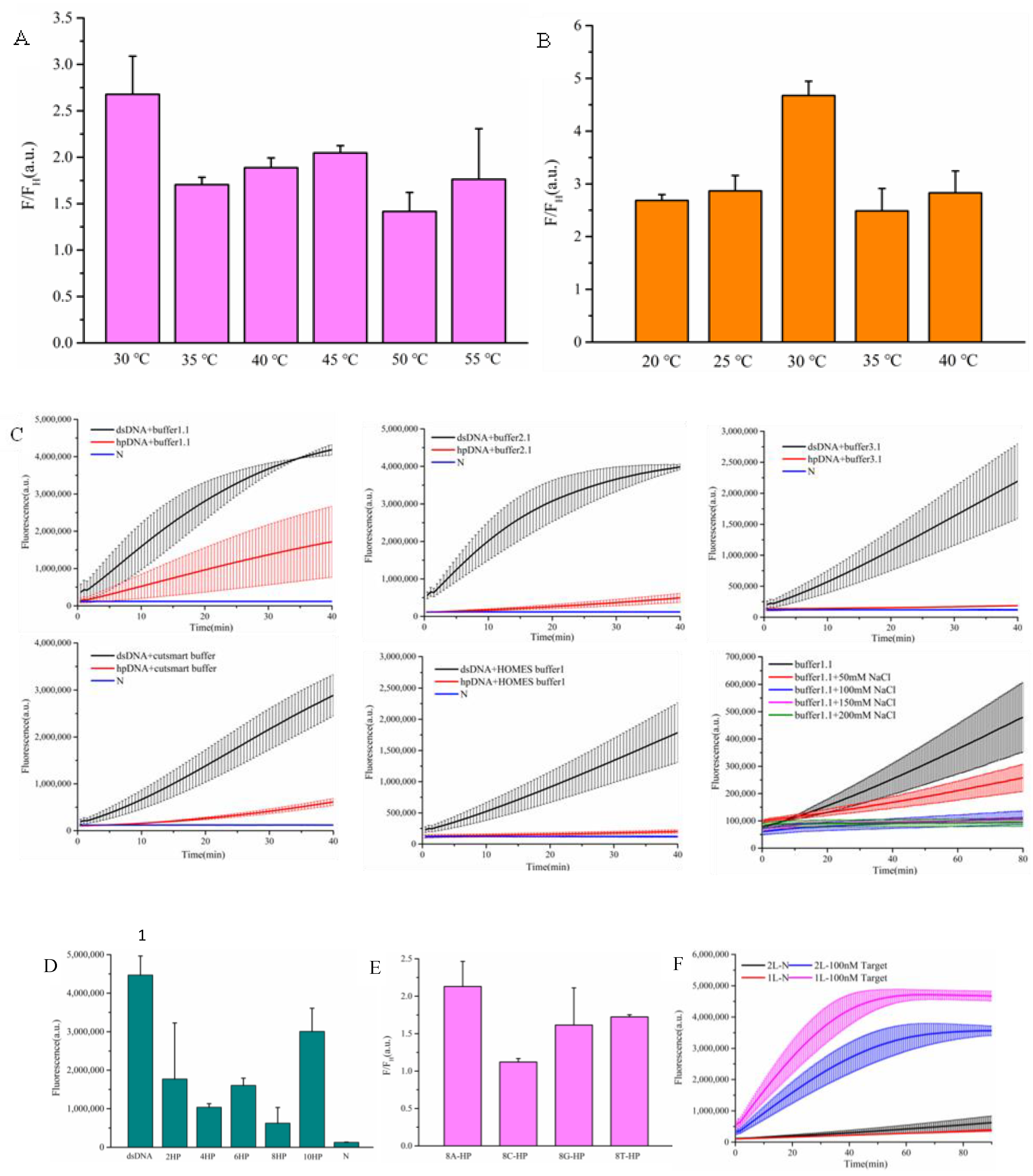
3.3. Optimization of Amplification-Free Detection System
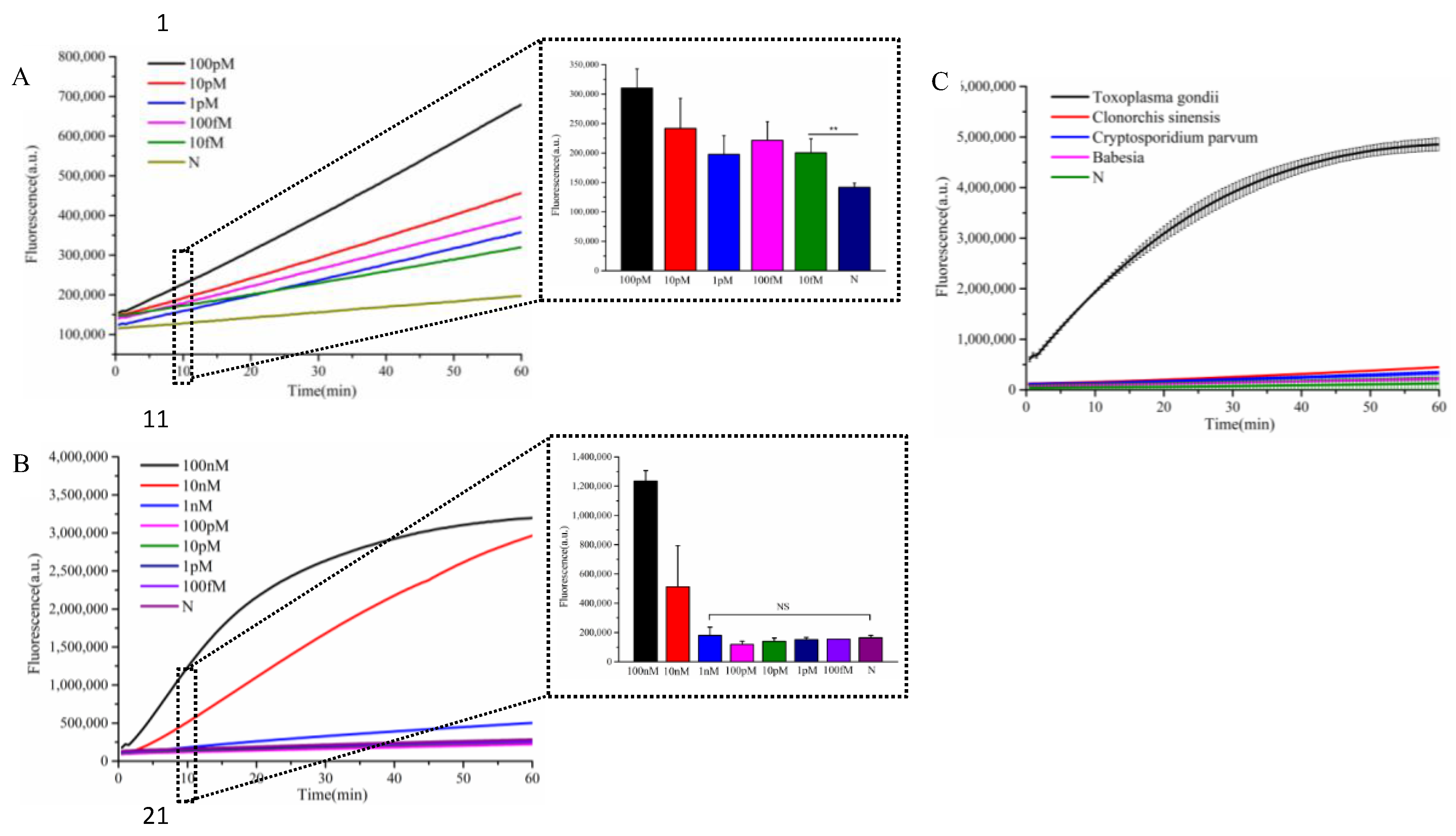
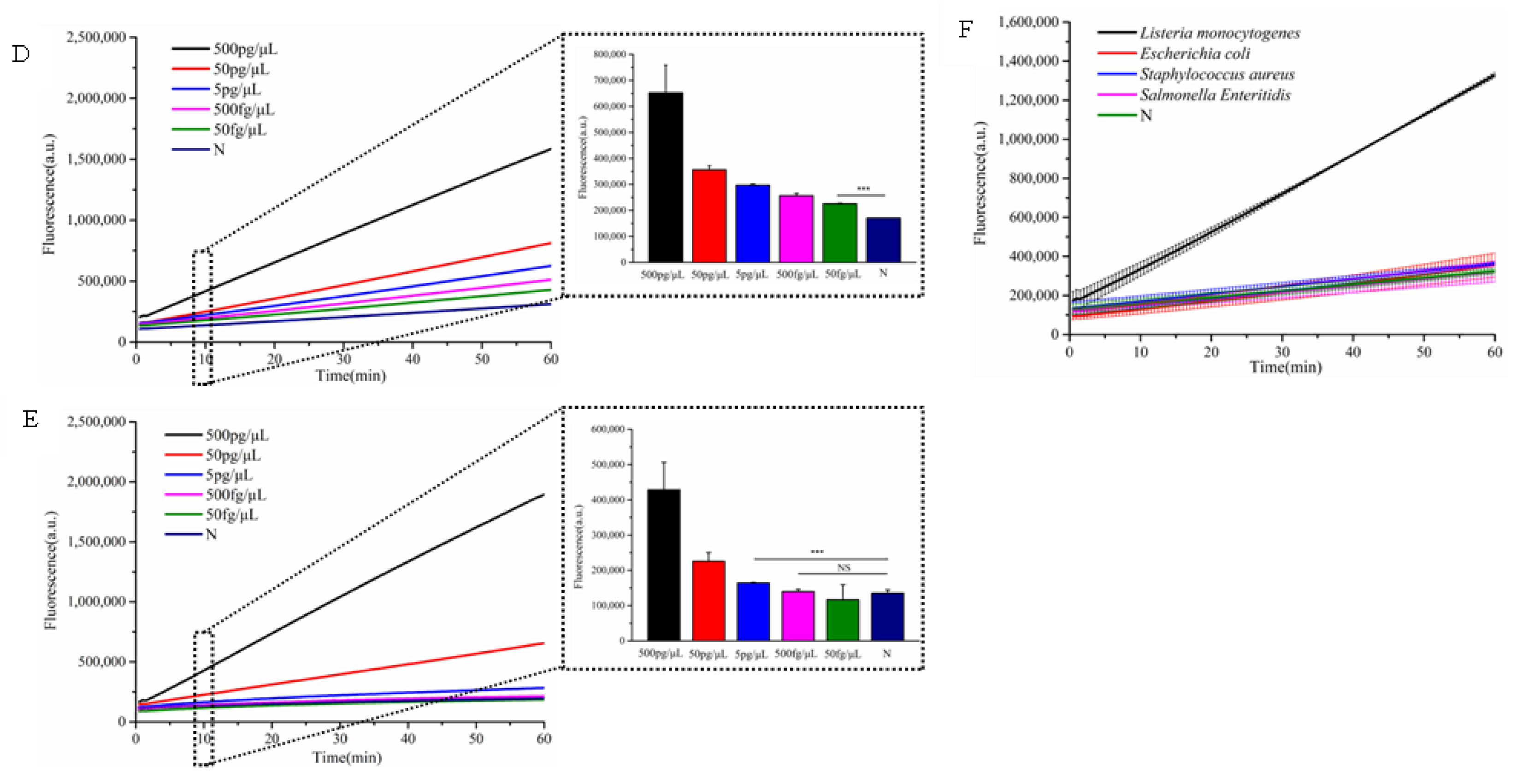
3.4. Evaluation of Detection Sensitivity and Specificity
3.5. Detection of Artificially Spiked Samples
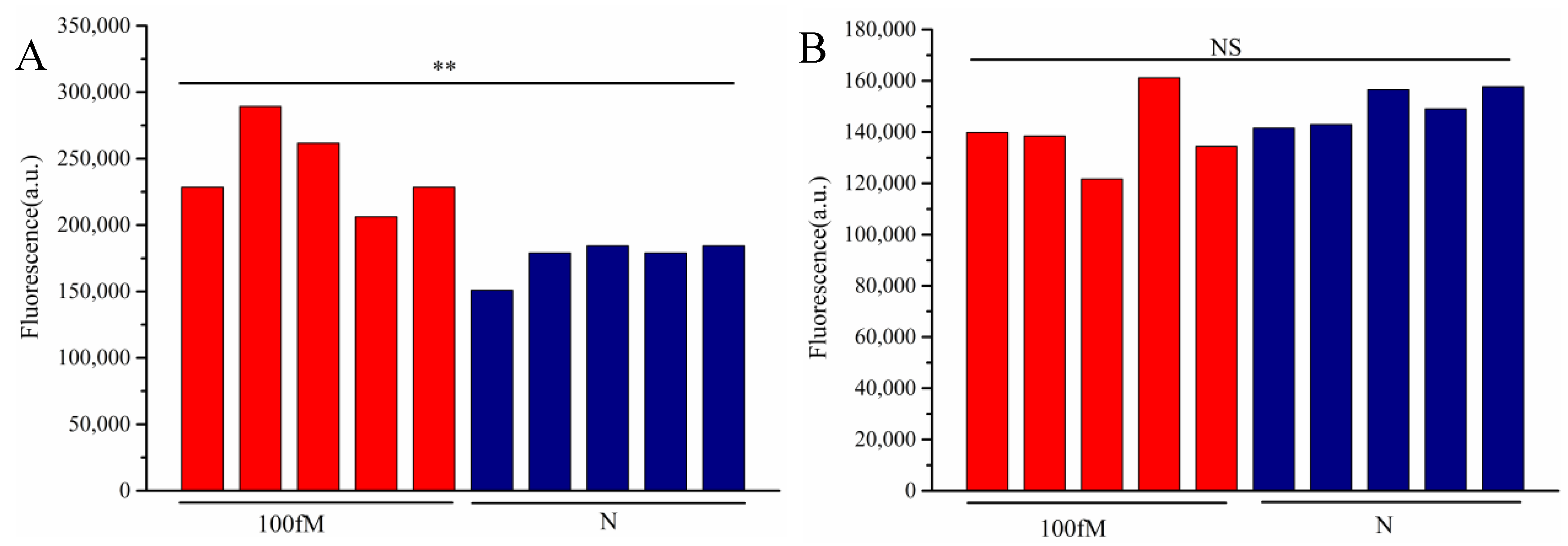
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Gootenberg, J. S.; Abudayyeh, O. O.; Lee, J. W.; Essletzbichler, P.; Dy, A. J.; Joung, J.; Verdine, V.; Donghia, N.; Daringer, N. M.; Freije, C. A.; et al. Nucleic acid detection with CRISPR-Cas13a/C2c2. Science. 2017, 356, 438–442. [Google Scholar] [CrossRef] [PubMed]
- Li, S. Y.; Cheng, Q. X.; Wang, J. M.; Li, X. Y.; Zhang, Z. L.; Gao, S.; Cao, R. B.; Zhao, G. P.; Wang, J. CRISPR-Cas12a-assisted nucleic acid detection. Cell Discov. 2018, 4, 20. [Google Scholar] [CrossRef] [PubMed]
- Harrington, L. B.; Burstein, D.; Chen, J. S.; Paez-Espino, D.; Ma, E.; Witte, I. P.; Cofsky, J. C.; Kyrpides, N. C.; Banfield, J. F.; Doudna, J. A. Programmed DNA destruction by miniature CRISPR-Cas14 enzymes. Science 2018, 362, 839–842. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Chen, Y.; Huang, L.; Lin, X.; Chen, H.; Xiang, W.; Liu, L. Trans-nuclease activity of Cas9 activated by DNA or RNA target binding. Nat Biotechnol. 2025, 43, 558–568. [Google Scholar] [CrossRef]
- Xu, H.; Tang, H.; Li, R.; Xia, Z.; Yang, W.; Zhu, Y.; Liu, Z.; Lu, G.; Ni, S.; Shen, J. A New Method Based on LAMP-CRISPR-Cas12a-Lateral Flow Immunochromatographic Strip for Detection. Infect Drug Resist. 2022, 15, 685–696. [Google Scholar] [CrossRef]
- Chen, H.; Zhang, H.; Guo, J.; Meng, X.; Yao, M.; He, L.; Nie, X. , Xu; H., Liu, C.; Sun, J.; et al. Rapid detection of feline parvovirus using RAA-CRISPR/Cas12a-based lateral flow strip and fluorescence. Front Microbiol. 2025, 16, 1501635. [Google Scholar]
- Wang, J.; Zhu, X.; Yin, D.; Cai, C.; Liu, H.; Yang, Y.; Guo, Z.; Yin, L.; Shen, X.; Dai, Y.; et al. Rapid and Easy-Read Porcine Circovirus Type 4 Detection with CRISPR-Cas13a-Based Lateral Flow Strip. Microorganisms. 2023, 11, 354. [Google Scholar] [CrossRef]
- Shen, J.; Chen, Z.; Xie, R.; Li, J.; Liu, C.; He, Y.; Ma, X.; Yang, H.; Xie, Z. CRISPR/Cas12a-Assisted isothermal amplification for rapid and specific diagnosis of respiratory virus on an microfluidic platform. Biosens Bioelectron. 2023, 237, 115523. [Google Scholar] [CrossRef]
- Xu, T.; Cao, F.; Dai, T.; Liu, T. RPA-CRISPR/Cas12a-Mediated Isothermal Amplification for Rapid Detection of Phytopythium helicoides. Plant Dis. 2024, 108, 3463–3472. [Google Scholar] [CrossRef]
- Xu, D.; Zeng, H.; Wu, W.; Liu, H.; Wang, J. Isothermal Amplification and CRISPR/Cas12a-System-Based Assay for Rapid, Sensitive and Visual Detection of Staphylococcus aureus. Foods 2023, 12, 4432. [Google Scholar] [CrossRef]
- Lu, S.; Tong, X.; Han, Y.; Zhang, K.; Zhang, Y.; Chen, Q.; Duan, J.; Lei, X.; Huang, M.; Qiu, Y. Fast and sensitive detection of SARS-CoV-2 RNA using suboptimal protospacer adjacent motifs for Cas12a. Nat Biomed Eng. 2022, 6, 286–297. [Google Scholar] [CrossRef] [PubMed]
- Broughton, J. P.; Deng, X.; Yu, G.; Fasching, C. L.; Servellita, V.; Singh, J.; Miao, X.; Streithorst, J. A.; Granados, A.; Sotomayor-Gonzalez, A.; et al. CRISPR-Cas12-based detection of SARS-CoV-2. Nat Biotechnol. 2020, 38, 870–874. [Google Scholar] [CrossRef] [PubMed]
- Ding, X.; Yin, K.; Li, Z.; Lalla, R. V.; Ballesteros, E.; Sfeir, M. M.; Liu, C. Ultrasensitive and visual detection of SARS-CoV-2 using all-in-one dual CRISPR-Cas12a assay. Nat Commun. 2020, 11, 4711. [Google Scholar] [CrossRef] [PubMed]
- Ye, X.; Wu, H.; Liu, J.; Xiang, J.; Feng, Y.; Liu, Q. One-pot diagnostic methods based on CRISPR/Cas and Argonaute nucleases: strategies and perspectives. Trends Biotechnol. 2024, 42, 1410–1426. [Google Scholar] [CrossRef]
- Sha, Y.; Huang, R.; Huang, M.; Yue, H.; Shan, Y.; Hu, J.; Xing, D. Cascade CRISPR/cas enables amplification-free microRNA sensing with fM-sensitivity and single-base-specificity. Chem Commun. 2021, 57, 247–250. [Google Scholar] [CrossRef]
- Shi, K.; Xie, S.; Tian, R.; Wang, S.; Lu, Q.; Gao, D.; Lei, C.; Zhu, H.; Nie, Z. A CRISPR-Cas autocatalysis-driven feedback amplification network for supersensitive DNA diagnostics. Sci Adv. 2021, 7, eabc7802. [Google Scholar] [CrossRef]
- Lim, J.; Van, A. B.; Koprowski, K.; Wester, M.; Valera, E.; Bashir, R. Amplification-free, OR-gated CRISPR-Cascade reaction for pathogen detection in blood samples. Proc Natl Acad Sci USA 2025, 122, e2420166122. [Google Scholar] [CrossRef]
- Jeung, J. H.; Han, H.; Lee, C. Y.; Ahn, J. K. CRISPR/Cas12a Collateral Cleavage Activity for Sensitive 3'-5' Exonuclease Assay. Biosensors 2023, 13, 963. [Google Scholar] [CrossRef]
- Li, L.; Li, S.; Wu, N.; Wu, J.; Wang, G.; Zhao, G.; Wang, J. HOLMESv2: A CRISPR-Cas12b-Assisted Platform for Nucleic Acid Detection and DNA Methylation Quantitation. ACS Synth Biol. 2019, 8, 2228–2237. [Google Scholar] [CrossRef]
- Aladhadh, M. A Review of Modern Methods for the Detection of Foodborne Pathogens. Microorganisms 2023, 11, 1111. [Google Scholar] [CrossRef]
- Bai, W.; Chen, J.; Chen, D.; Zhu, Y.; Hu, K.; Lin, X.; Chen, J.; Song, D. Sensitive and rapid detection of three foodborne pathogens in meat by recombinase polymerase amplification with lateral flow dipstick (RPA-LFD). Int J Food Microbiol. 2024, 422, 110822. [Google Scholar] [CrossRef] [PubMed]
- Ndraha, N.; Lin, H. Y.; Wang, C. Y.; Hsiao, H. I.; Lin, H. J. Rapid detection methods for foodborne pathogens based on nucleic acid amplification: Recent advances, remaining challenges, and possible opportunities. Food Chem. 2023, 7, 100183. [Google Scholar] [CrossRef]
- Li, X.; Zhang, X.; Shi, X.; Shi, H.; Wang, Z.; Peng, C. Review in isothermal amplification technology in food microbiological detection. Food Sci Biotechnol. 2022, 31, 1501–1511. [Google Scholar] [CrossRef]
- Deng, F.; Li, Y.; Yang, B.; Sang, R.; Deng, W.; Kansara, M.; Lin, F.; Thavaneswaran, S.; Thomas, D. M.; Goldys, E. M. Topological barrier to Cas12a activation by circular DNA nanostructures facilitates autocatalysis and transforms DNA/RNA sensing. Nat Commun. 2024, 15, 1818. [Google Scholar] [CrossRef]
- Sun, K.; Pu, L.; Chen, C.; Chen, M.; Li, K.; Li, X.; Li, H.; Geng, J. An autocatalytic CRISPR-Cas amplification effect propelled by the LNA-modified split activators for DNA sensing. Nucleic Acids Res. 2024, 52, e39. [Google Scholar] [CrossRef]
- Zhou, Z.; Lau, C. H.; Wang, J.; Guo, R.; Tong, S.; Li, J.; Dong, W.; Huang, Z.; Wang, T.; Huang, X.; et al. Rapid and Amplification-free Nucleic Acid Detection with DNA Substrate-Mediated Autocatalysis of CRISPR/Cas12a. ACS Omega 2024, 9, 28866–28878. [Google Scholar] [CrossRef] [PubMed]
- Hwang, I.; Song, Y. H.; Lee, S. Enhanced trans-cleavage activity using CRISPR-Cas12a variant designed to reduce steric inhibition by cis-cleavage products. Biosens Bioelectron. 2025, 267, 116859. [Google Scholar] [CrossRef]
| Sample number | Detection of L. monocytogenes | ||
|---|---|---|---|
| Single Cas12a | Cascade-CRISPR | qPCR | |
| 1 | + | + | + |
| 2 | + | + | + |
| 3 | + | + | + |
| 4 | - | + | + |
| 5 | - | + | + |
| 6 | + | + | + |
| 7 | + | + | + |
| 8 | + | + | + |
| 9 | - | + | + |
| 10 | - | + | + |
| 11 | - | - | - |
| 12 | - | - | - |
| 13 | - | - | - |
| 14 | - | - | - |
| 15 | - | - | - |
| 16 | - | - | - |
| 17 | - | - | - |
| 18 | - | - | - |
| 19 | - | - | - |
| 20 | - | - | - |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




