Submitted:
01 July 2025
Posted:
04 July 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
- C: Code complexity (e.g., – in genome analysis)
- : Operating system overhead (context switches, thread management)
- M: Memory hierarchy inefficiencies (cache miss rates, RAM/disk access latency)
- B: Binary execution inefficiencies (pipeline stalls, branch mispredictions)
2. Methods
2.1. 1. Complexity of the Data Structure
Sparsity.
Power-law Distribution.
Dynamic Topology.
Theorem 1: Runtime Scaling Bound
- : Algorithmic complexity
- : Operating system overhead
- : Memory hierarchy latency
- : Binary execution inefficiency
Algorithmic Lower Bound.
- Genome assembly requires pairwise overlap comparisons.
- Variant calling on n variants requires genotype likelihood operations.
- Thus, .
OS Overhead Lower Bound.
- For n processes on p processors, context switches .
- Scheduling latency grows as .
- Thus, .
Memory Access Lower Bound.
- Genomic graphs have working set size .
- For any fixed cache size , there exists such that for all , .
- Thus, .
Execution Stalls Lower Bound.
- Branch prediction is bounded by input-dependent variation.
- Pipeline stalls scale linearly with n, so .
Compositions
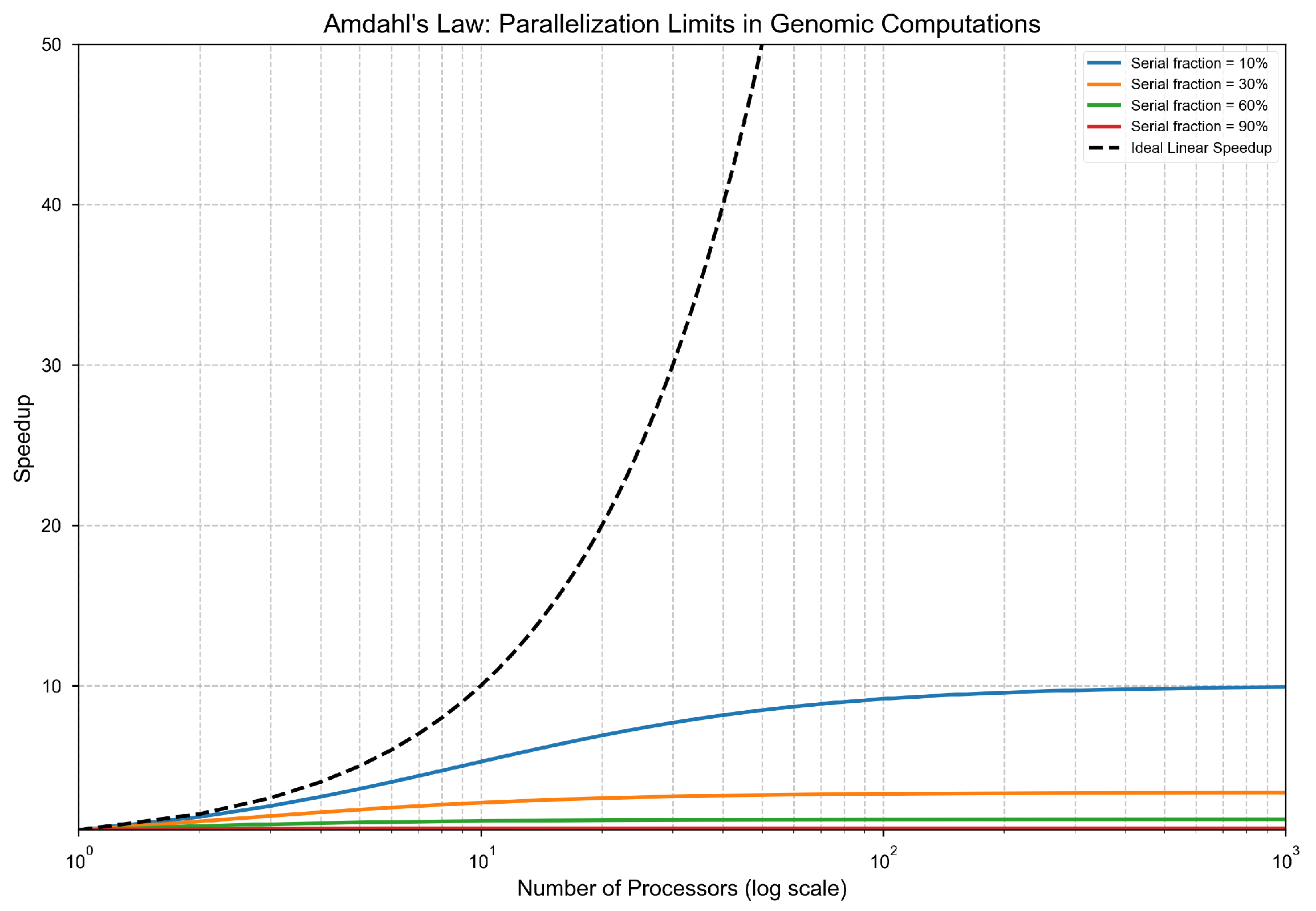
Theorem 2: Parallelization Ineffectiveness

Serial Dependency Analysis
- Genome assembly involves graph construction and consensus phases that are inherently sequential.
- For all assembly algorithms, topological ordering and consistency checking require serial operations.
- Thus, serial fraction .
Amdahl’s Law Application.
- The theoretical speedup is bounded as:
- For , we obtain:
- Therefore, the parallel execution time is lower bounded by:
Asymptotic Preservation.
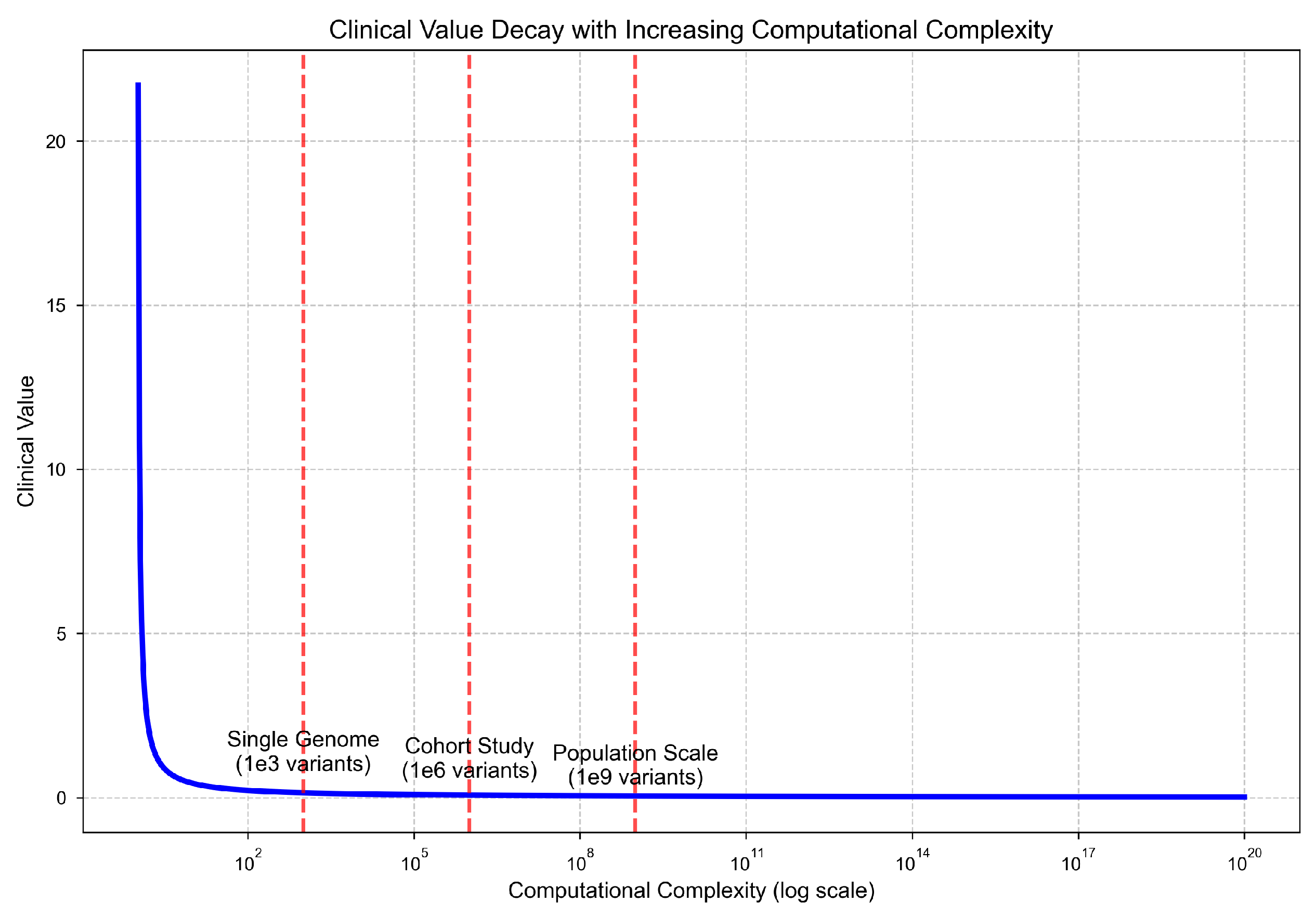
Theorem 3: Clinical Interpretability Decay
Cognitive Capacity Bound.
- Human interpretability is bounded by Miller’s Law: the brain can process approximately elements.
- Define actionability as , where .
- Hence, .
Signal Degradation.
- For computational steps, error accumulation follows .
- Given , signal-to-noise ratio is bounded by:
Time-to-Insight Penalty.
- Longer runtime delays clinical relevance: utility diminishes as .
- With , temporal discounting becomes:
Composite Clinical Value.
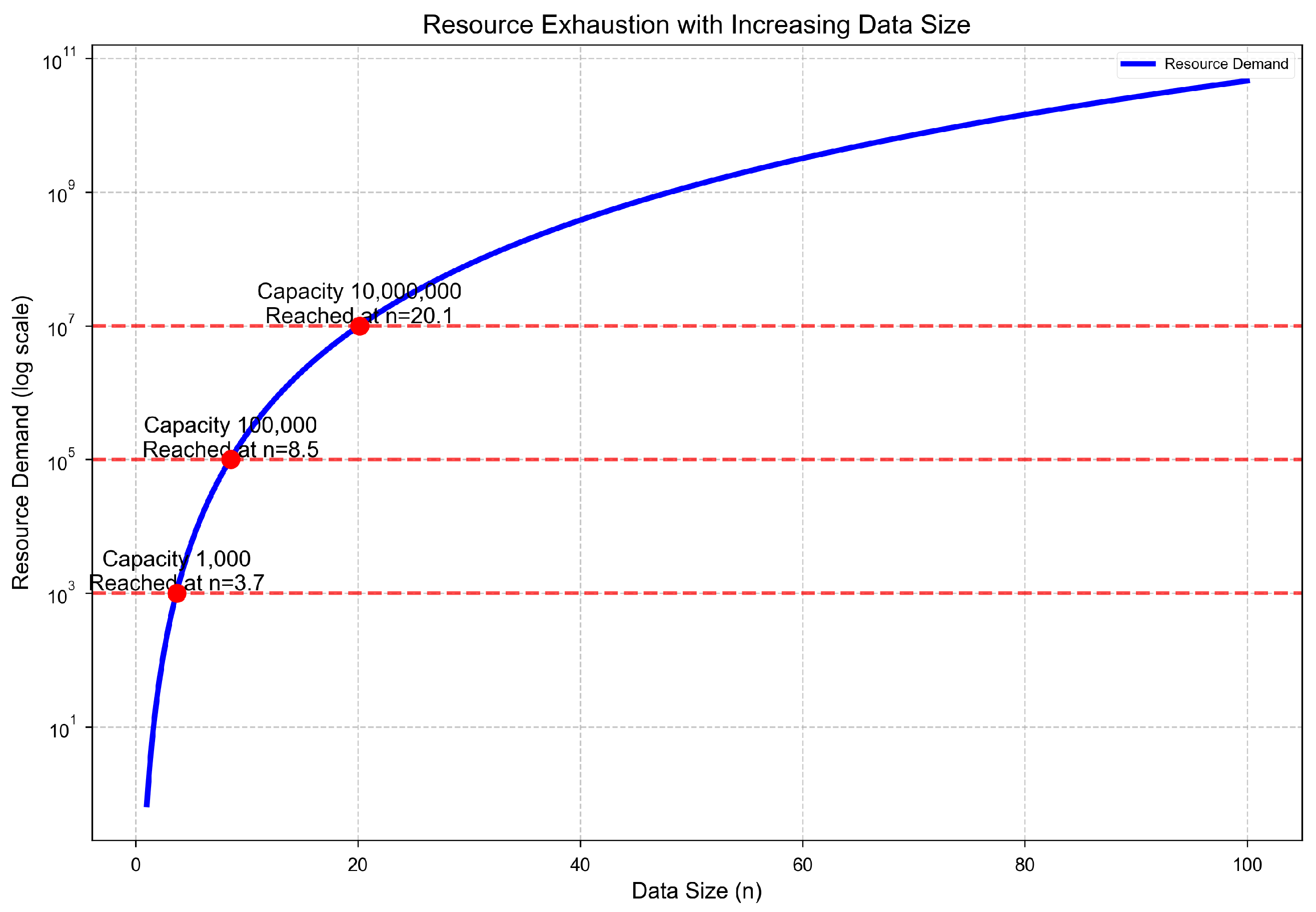
Theorem 4: Resource Exhaustion Inevitability
Base Cases.
Inductive Hypothesis.
Inductive Step.
Divergence.
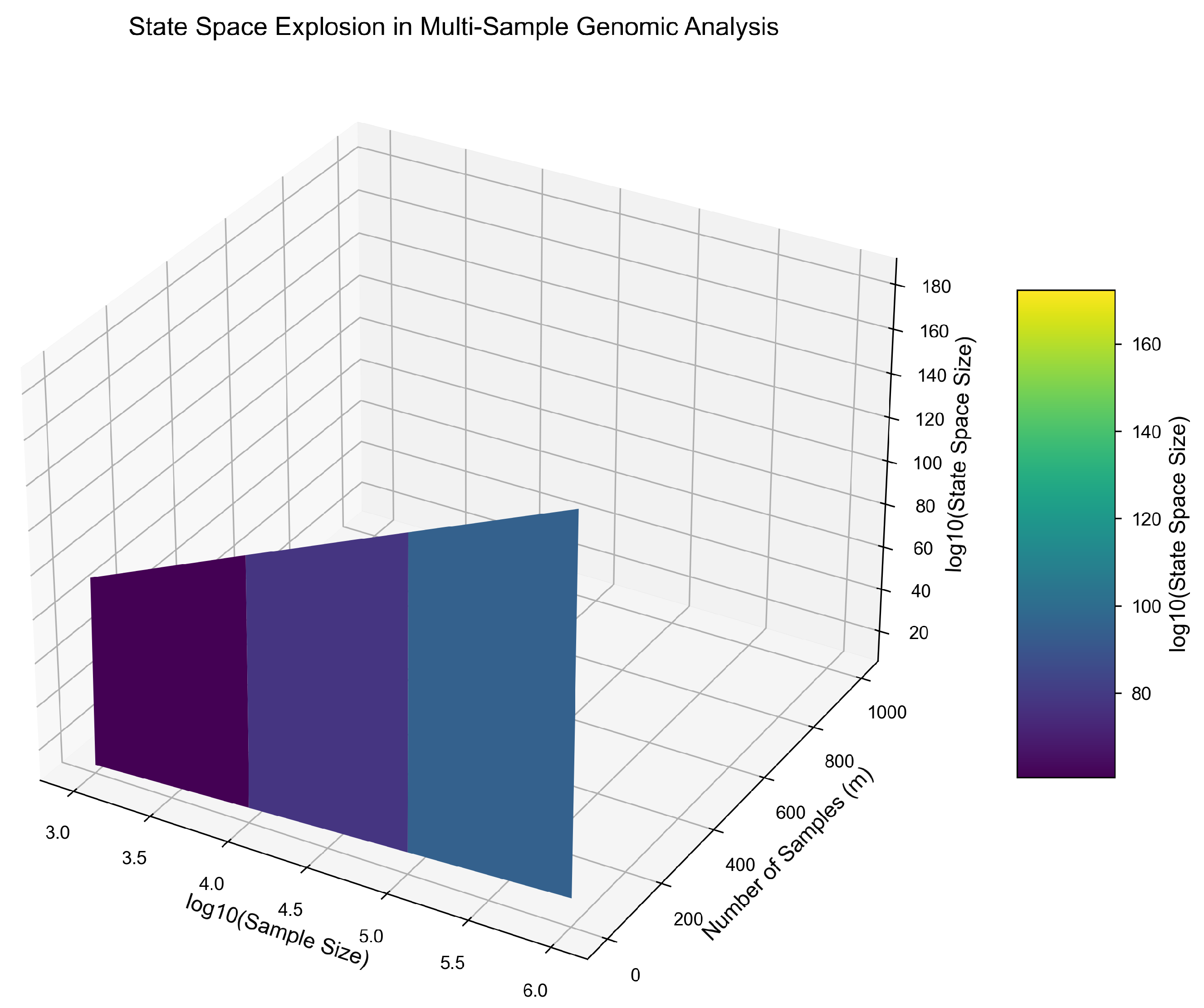
Theorem 5: State Space Explosion
Single-Sample State Bound.
- Processing states: from pairwise feature interactions.
- Memory states: from addressable memory or configuration space.
- Error states: due to position-specific mutations or uncertainty.
Cross-Sample Dependencies.
Intractability Bound.
- (typical variant set per sample)
- (population-scale cohort)
Corollary: Fundamental Intractability
- Runtime Bound:
- Parallelization Limit: Asymptotic complexity is preserved under parallel execution
- Clinical Value Decay:
- Resource Ceiling: Resource demand exceeds any fixed capacity
- State Space Explosion:
Numerical Verification
Single Genome Analysis.
- Input size: base pairs
- Estimated operations:
- At 1 THz compute rate ( ops/sec):
Population Analysis.
- Input size: data points
- Estimated operations:
- Conclusion: Infeasible under any realistic computational infrastructure
3. Bio-AI Implications
Mathematical Correlation: Software, Compute, and Clinical ROI
Model Definition
- CI: Clinical Insight — number of actionable findings.
- TC: Total Cost — includes software, compute, and personnel expenses.
3.0.0.25. Clinical Insight (CI).
- k: Scaling factor (e.g., 100 actionable variants)
- SE: Software Efficiency (optimal / actual runtime)
- CU: Computational Utilization (effective / peak resources)
- IQ: Insight Quality (reproducibility and clinical value)
Total Cost (TC).
- : Cost per unit software efficiency
- : Cost per unit compute utilization
- : Fixed personnel cost
Component Models
Software Efficiency (SE).
- Current tools:
- Advanced tools:
Computational Utilization (CU).
- GPU workloads:
- CPU workloads:
- Specialized hardware:
Insight Quality (IQ).
Total Cost Approximation.
Full ROI Model
Numerical Example: Cancer Genomics
Current Pipeline.
- , ,
- , ,
Optimized Pipeline.
- , ,
Break-even Analysis
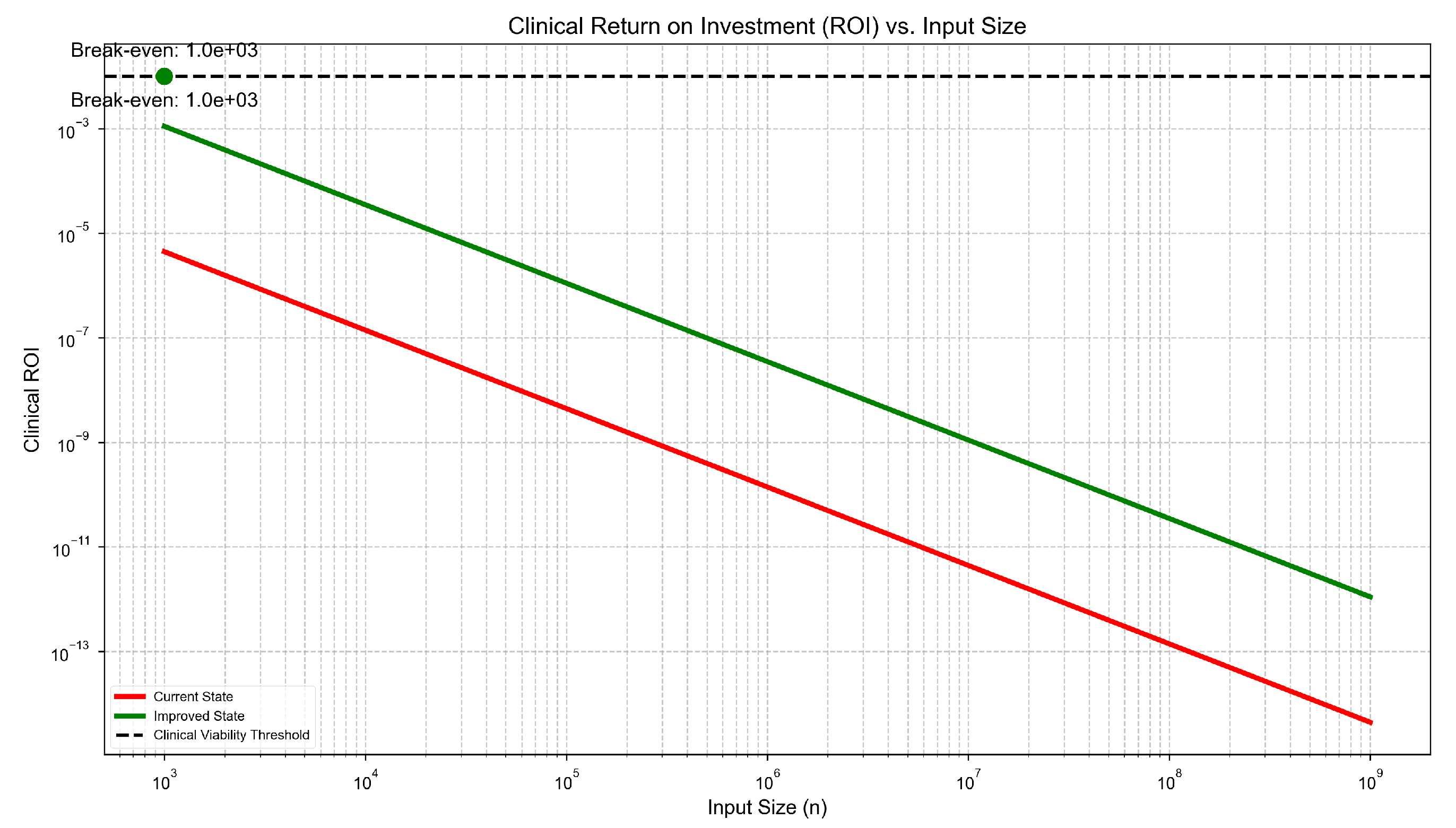
4. Results
Cross-Domain Implications
Liquid Biopsy.
Cancer Stratification.
Single-Cell Lineage Tracing.
Clinical ROI Saturation
5. Discussion
- Runtime scaling:
- Parallelization bound: Speedup limited to (Amdahl’s Law)
- State space: growth in multi-sample analyses
- Clinical ROI:
Software Efficiency as the Dominant Lever
Toward Structural Realignment
- Prioritize irregular memory access and graph-native operations
- Align software logic with data sparsity and power-law distributions
- Emphasize interpretability alongside throughput
Conclusions
References
- Puckelwartz, M.J.; et al. Supercomputing for the parallelization of whole genome analysis. Nature Communications 2014, 5, 1–9. [Google Scholar]
- Deng, Y.; Zhang, H.; Wang, H. A high-performance computing framework for genome variant analysis based on a network-on-chip architecture. IEEE Transactions on Computers 2019, 68, 578–591. [Google Scholar]
- Zhang, Y.; Zhang, H. A Network-on-Chip Accelerator for Genome Variant Analysis. ResearchGate 2019. [Google Scholar]
- Blaauw, D.; Huang, J. GenAx: A Genome Sequencing Accelerator for High-Performance Computing Systems. University of Michigan 2019. [Google Scholar]
- Isik, M.; et al. Graph-Based Processing Architectures for Next-Generation Genomic Analysis. Stanford University 2024. [Google Scholar]
- Hughes, T. A Study on Data Management in Genome Analysis. Clemson University 2017. [Google Scholar]
- Wang, H.; Zhang, Y. Efficient data management for genome sequencing in cloud computing environments. ACM Transactions on Architecture and Code Optimization 2020, 17, 1–25. [Google Scholar]
- Ghose, S.; Karamcheti, S. SEGRAM: A Scalable Framework for Genome Analysis on Multi-Core Architectures. Illinois Research 2022. [Google Scholar]
- Koonin, E.; Wolf, Y.; Karev, G. The dominance of the population by a selected few: power-law behaviour applies to a wide variety of genomic properties. Genome Biology 2002, 3, research0036. [Google Scholar]
- Furusawa, C.; Kaneko, K. Power-law distribution of gene expression fluctuations. Physical Review E 2003, 68, 011909. [Google Scholar]
- Karev, G.; Wolf, Y.; Koonin, E. The power-law distribution of gene family size is driven by the pseudogenisation rate’s heterogeneity between gene families. Gene 2003, 311, 43–51. [Google Scholar]
- Paten, B.; Novak, A.; Eizenga, J.; Garrison, E. Genome graphs and the evolution of genome inference. Nature Reviews Genetics 2017, 18, 679–692. [Google Scholar]
- Li, H.; Durbin, R. Graph genomes. Nature Methods 2020, 17, 759–760. [Google Scholar]
- Garrison, E.; Novak, A.; Hickey, G.; et al. Fast and accurate genomic analyses using genome graphs. Nature Biotechnology 2018, 36, 354–362. [Google Scholar]
- Andrews, T.; Hemberg, M. Handling sparsity: Analysis of single cell RNA-seq data. Current Opinion in Biotechnology 2018, 52, 18–25. [Google Scholar]
- Denti, L.; Rizzi, R.; Beretta, S.; et al. Taming large-scale genomic analyses via sparsified genomics. Bioinformatics 2021, 37, 4655–4663. [Google Scholar]
- Medvedev, P.; Brudno, M. Computational complexity of algorithms for sequence comparison, short-read assembly and genome alignment. BMC Bioinformatics 2009, 10, S5. [Google Scholar]
- Schatz, M.; Langmead, B.; Salzberg, S. Computational Strategies for Scalable Genomics Analysis. Genome Biology 2010, 11, R121. [Google Scholar]
- Pevzner, P.; Tang, H.; Waterman, M. Modeling biological problems in computer science: a case study in genome assembly. Communications of the ACM 2011, 44, 73–80. [Google Scholar]
- Fisher, J.; Henzinger, T. Computational methods for understanding complexity: the use of formal methods in biology. Nature 2007, 447, 879–886. [Google Scholar]
- Ashley, E. Computational Genomics in the Era of Precision Medicine: Applications to Variant Analysis and Gene Therapy. Annual Review of Genomics and Human Genetics 2015, 16, 33–61. [Google Scholar]
- Alser, M.; Rotman, J.; Bertels, K.; et al. Systems Challenges and Opportunities for Genomics. Nature Reviews Genetics 2020, 21, 563–576. [Google Scholar]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



