Submitted:
25 June 2025
Posted:
26 June 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Eligibility Criteria
2.2. Information Sources and Search Strategy
2.3. Study Selection
2.4. Data Extraction
2.5. Quality Assessment
2.6. Statistical Analysis
2.7. Assessment of Publication Bias
2.8. Ethical Considerations
3. Results
3.1. Study Selection
3.2. Study Characteristics
3.3. Risk of Bias Within Studies
| Study | Year | Case Def. | Case Rep. | Control Sel. | Control Def. | Confounding | Exposure Asc. | Same Method | Non-Resp. | Total Score | Quality |
|---|---|---|---|---|---|---|---|---|---|---|---|
| Li et al. | 2012 | 1 | 0 | 1 | 1 | 2 | 1 | 1 | 0 | 7 | High |
| Loo et al. | 2011 | 1 | 0 | 1 | 1 | 2 | 1 | 1 | 0 | 7 | High |
| Xie et al. | 2009 | 1 | 0 | 0 | 1 | 2 | 1 | 1 | 0 | 6 | Moderate |
| Prakash et al. | 2015 | 1 | 0 | 0 | 1 | 2 | 1 | 1 | 0 | 5 | Low |
| Dahash et al. | 2022 | 1 | 0 | 0 | 1 | 1 | 1 | 1 | 0 | 5 | Low |
| Daing et al. | 2012 | 1 | 0 | 0 | 1 | 1 | 1 | 1 | 0 | 5 | Low |
| Dienha et al. | 2023 | 1 | 0 | 0 | 0 | 0 | 1 | 1 | 1 | 4 | Low |
4. Results of Individual Studies
5. Synthesis of Results—Meta-Analysis
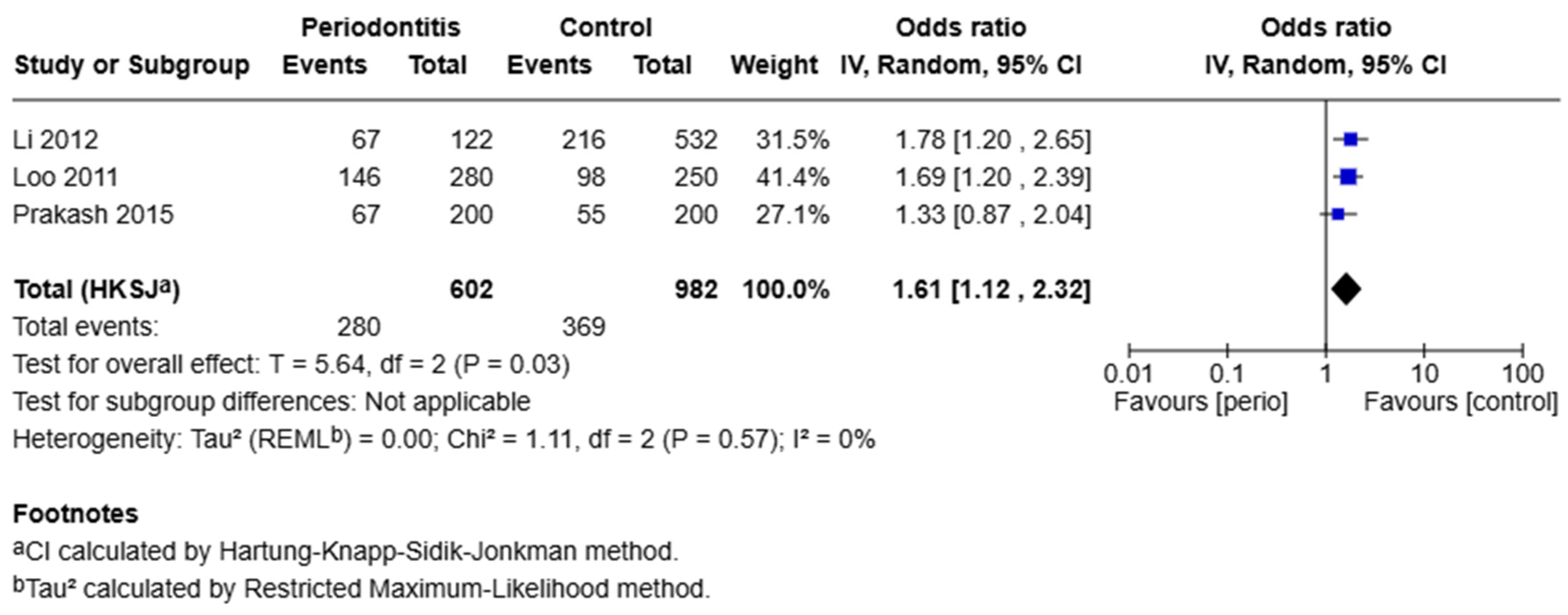
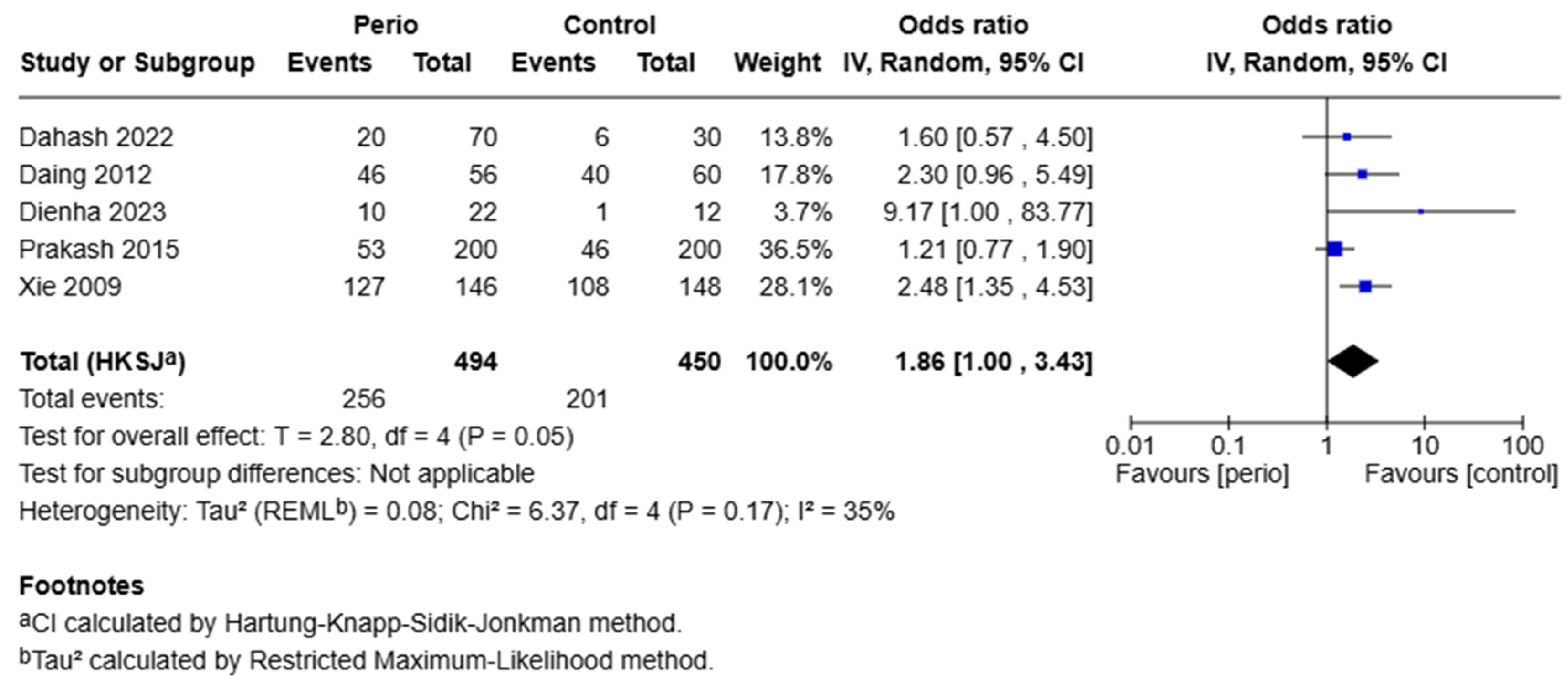
Publication Bias
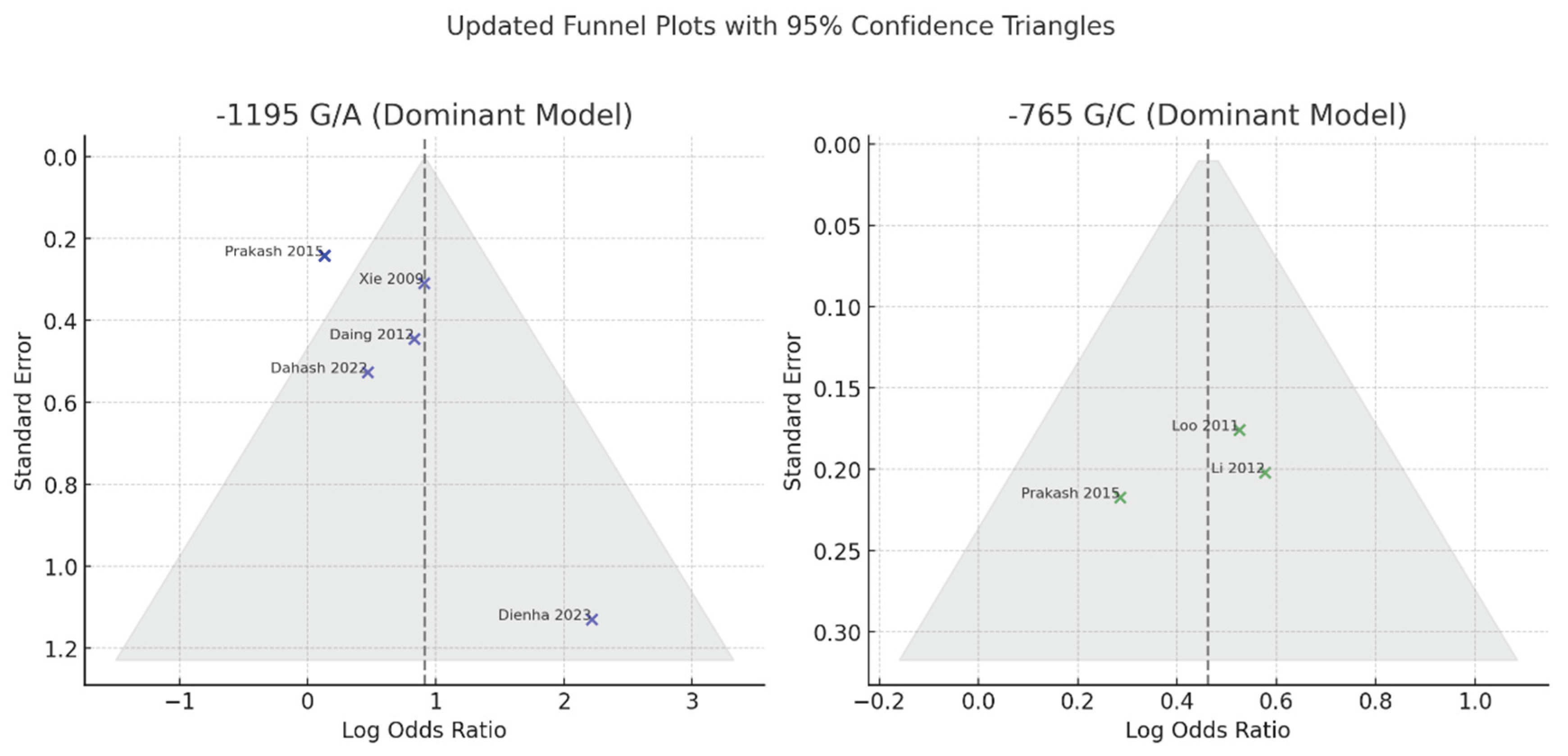
6. Discussion
7. Conclusion
Author Contributions
Conflicts of Interest
Appendix
| SECTION | ITEM | REPORTED DETAIL |
| INFORMATION SOURCES | Name of databases and platforms used | PubMed (MEDLINE), Scopus, Web of Science, EMBASE (via Ovid), Cochrane Library; supplementary: Google Scholar, manual reference screening |
| Dates of the search | Initial search: 11–15 March 2025; supplementary (gray literature and manual reference list screening): 17–20 March 2025 | |
| Timeframe covered by the search | From database inception to March 20, 2025 | |
| Restrictions (language, publication status, etc.) | Included only peer-reviewed, full-text studies published in English; excluded animal studies, reviews, conference abstracts, and studies with no genotype data | |
| SEARCH STRATEGY FOR EACH DATABASE | Boolean operators, MeSH terms, filters | See below for full structured search examples per database |
| Databases adapted for syntax | Yes – adjusted syntax for each platform (e.g., MeSH in PubMed, Emtree in EMBASE, TITLE-ABS-KEY in Scopus) |
References
- Bostanci, N.; Bao, K.; Greenwood, D.; Silbereisen, A.; Belibasakis, G.N. Periodontal disease: From the lenses of light microscopy to the specs of proteomics and next-generation sequencing. Advances in Clinical Chemistry 2019, 93, 263–290. [Google Scholar] [CrossRef]
- Nazir, M.; Al-Ansari, A.; Al-Khalifa, K.; Alhareky, M.; Gaffar, B.; Almas, K. Global Prevalence of Periodontal Disease and Lack of Its Surveillance. Scientific World Journal 2020, 2020, 2146160. [Google Scholar] [CrossRef] [PubMed]
- Tsuchida, S.; Satoh, M.; Takiwaki, M.; Nomura, F. Current status of proteomic technologies for discovering and identifying gingival crevicular fluid biomarkers for periodontal disease. Int J Mol Sci. 2019, 20, 86. [Google Scholar] [CrossRef] [PubMed]
- Kinane, D.F.; Stathopoulou, P.G.; Papapanou, P.N. Periodontal diseases. Nat Rev Dis Primers. 2017, 3, 17038. [Google Scholar] [CrossRef] [PubMed]
- PPa Ap Pe Er, R.A.L.; Lazăr, L.; Loghin, A.; et al. O OR RI IG GI IN NA Cyclooxygenase-2 and matrix metalloproteinase-9 expressions correlate with tissue inflammation degree in periodontal disease. Rom J Morphol Embryol. 2015, 56, 1441–1446. [Google Scholar]
- Prakash, G.; Umar, M.; Ajay, S.; et al. COX-2 gene polymorphisms and risk of chronic periodontitis: A case-control study and meta-analysis. Oral Dis. 2015, 21, 38–45. [Google Scholar] [CrossRef]
- Dubois, R.N.; Abramson, S.B.; Crofford, L.; et al. Cyclooxygenase in Biology and Disease. FASEB J. 1998. [Google Scholar] [CrossRef]
- Båge, T.; Kats, A.; Silva Lopez, B.; et al. Expression of prostaglandin E synthases in periodontitis: Immunolocalization and cellular regulation. American Journal of Pathology. 2011, 178, 1676–1688. [Google Scholar] [CrossRef]
- Zhang, F.; Engebretson, S.P.; Morton, R.S.; Cavanaugh PFJr Subbaramaiah, K.; Dannenberg, A.J. The overexpression of cyclo-oxygenase-2 in chronic periodontitis. J Am Dent Assoc. 2003, 134, 861–7. [Google Scholar] [CrossRef] [PubMed]
- Tai, H.; Miyaura, C.; Pilbeam, C.C.; et al. Transcriptional Induction of Cyclooxygenase-2 in Osteoblasts Is Involved in Interleukin-6-Induced Osteoclast Formation. Endocrinology 1997. [Google Scholar] [CrossRef]
- Offenbacher, S.; Salvi, G.E. Induction of Prostaglandin Release from Macrophages by Bacterial Endotoxin.
- Jiang, L.; Weng, H.; Chen, M.Y.; Zhang, C.; Zeng, X.T. Association between cyclooxygenase-2 gene polymorphisms and risk of periodontitis: A meta-analysis involving 5653 individuals. Mol Biol Rep. 2014, 41, 4795–4801. [Google Scholar] [CrossRef]
- Dahash, S.A.; Sh, M.; Mahmood, B.D.S. Association of a Genetic Variant (Rs689466) of Cyclooxygenase-2 Gene with Chronic Periodontitis in a Sample of Iraqi Population. Journal of Baghdad College of Dentistry 2019. [Google Scholar] [CrossRef]
- C-j, X.; L-m, X.; W-h, F.; D-y, X.; J-c, Z. Common single nucleotide polymorphisms in cyclooxygenase-2 and risk of severe chronic periodontitis in a Chinese population. J Clin Periodontol. 2009, 36, 198–203. [Google Scholar] [CrossRef]
- Li, G.; Yue, Y.; Tian, Y.; et al. Association of matrix metalloproteinase (MMP)-1, 3, 9, interleukin (IL)-2, 8 and cyclooxygenase (COX)-2 gene polymorphisms with chronic periodontitis in a Chinese population. Cytokine. 2012, 60, 552–560. [Google Scholar] [CrossRef]
- Loo, W.T.Y.; Wang, M.; Jin, L.J.; Cheung, M.N.B.; Li, G.R. Association of matrix metalloproteinase (MMP-1, MMP-3 and MMP-9) and cyclooxygenase-2 gene polymorphisms and their proteins with chronic periodontitis. Arch Oral Biol. 2011, 56, 1081–1090. [Google Scholar] [CrossRef]
- Dienha, O.V.; Pyndus, V.B.; Litovkin, K.V.; et al. DEFB1, MMP9 AND COX2 GENE POLYMORPHISMS AND PERIODONTITIS. World of Medicine and Biology. 2023, 19, 53. [Google Scholar] [CrossRef]
- Daing, A.; Singh, S.V.; Saimbi, C.S.; Khan, M.A.; Rath, S.K. Cyclooxygenase 2 gene polymorphisms and chronic periodontitis in a North Indian population: A pilot study. J Periodontal Implant Sci. 2012, 42, 151–157. [Google Scholar] [CrossRef]
- Fourmousis, I.; Vlachos, M. Genetic Risk Factors for the Development of Periimplantitis. Implant Dent. 2019, 28, 103–114. [Google Scholar] [CrossRef]
- Fritsche, E.; Baek, S.J.; King, L.M.; Zeldin, D.C.; Eling, T.E.; Bell, D.A. Functional characterization of cyclooxygenase-2 polymorphisms. J Pharmacol Exp Ther. 2001, 299, 468–76. [Google Scholar] [CrossRef] [PubMed]
- Armitage, G.C. Development of a classification system for periodontal diseases and conditions. Ann Periodontol. 1999, 4, 1–6. [Google Scholar]
- Tonetti, M.S.; Greenwell, H.; Kornman, K.S. Staging and grading of periodontitis: Framework and proposal of a new classification and case definition. J Clin Periodontol. 2018, 45, S149–S161. [Google Scholar] [CrossRef] [PubMed]
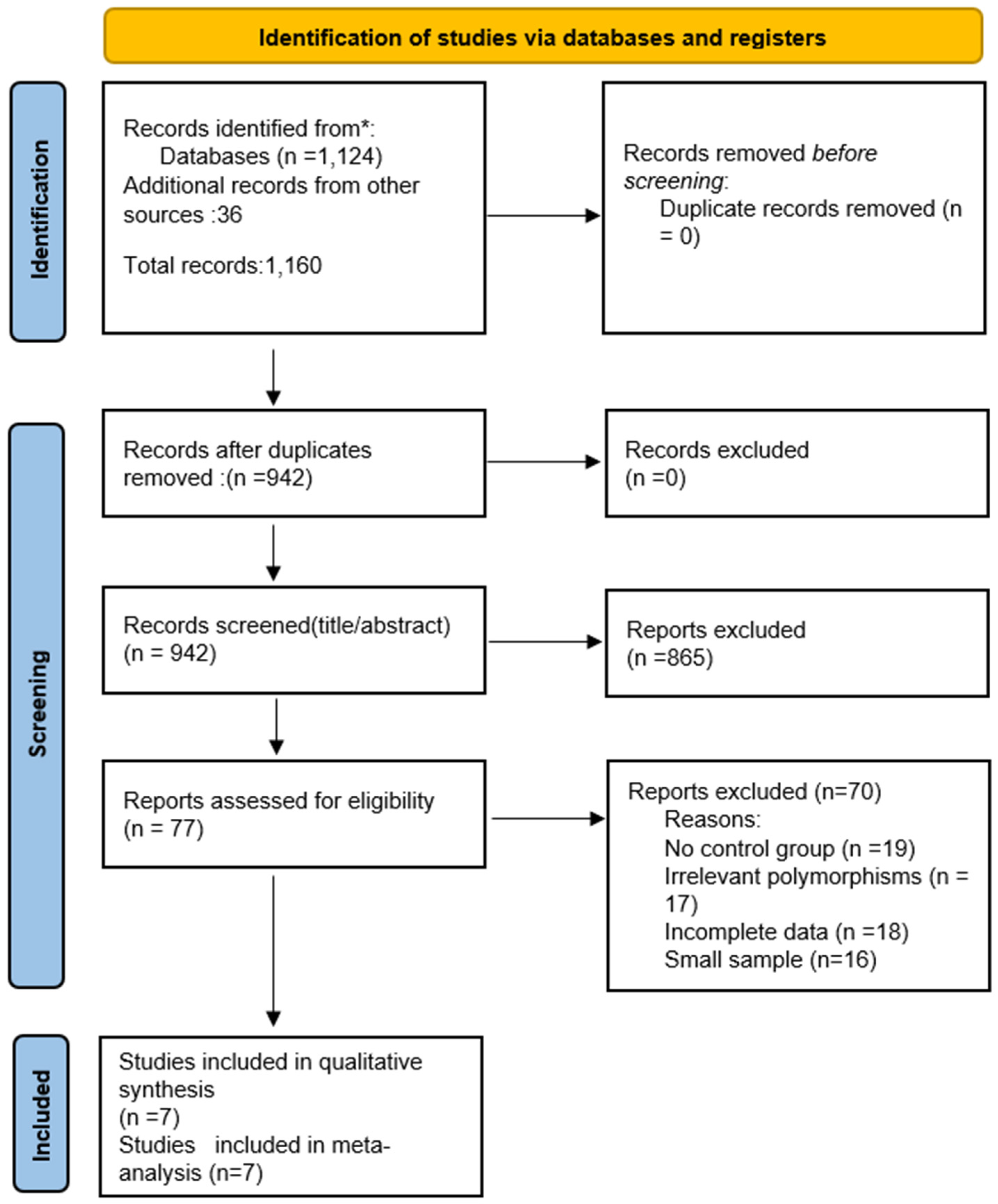
| Author | Year | Country | Ethnicity | Sample Size (Cases/Controls) | COX-2 Polymorphism(s) |
|---|---|---|---|---|---|
| Xie | 2009 | China | Han Chinese | 146 / 108 | -1195 G/A |
| Daing | 2012 | India | South Asian | 122 / 120 | -1195 G/A |
| Prakash | 2015 | India | South Asian | 200 / 200 | -765 G/C, -1195 G/A |
| Loo | 2011 | Malaysia | Southeast Asian | 146 / 85 | -765 G/C |
| Li | 2012 | China | Han Chinese | 122 / 53 | -765 G/C |
| Dahash | 2022 | Iraq | Middle Eastern | 70 / 30 | -1195 G/A |
| Dienha | 2023 | China | Han Chinese | 56 / 20 | -1195 G/A |
| Study | Polymorphism | Genotypes (Cases) | Genotypes (Controls) | Dominant Model Comparison | OR (95% CI) | p-value |
|---|---|---|---|---|---|---|
| Li et al., 2012 | -765 G/C | GG: 55, GC: 33, CC: 34 | GG: 316, GC: 204, CC: 12 | GC+CC vs. GG | 2.59 (1.93–3.47) | 0.000 |
| Loo et al., 2011 | -765 G/C | GG: 134, GC: 68, CC: 78 | GG: 152, GC: 90, CC: 8 | GC+CC vs. GG | 2.48 (1.89–3.26) | 0.0001 |
| Prakash et al., 2015 | -765 G/C | GG: 133, GC: 58, CC: 9 | GG: 145, GC: 47, CC: 8 | GC+CC vs. GG | 1.42 (0.90–2.24) | 0.132 |
| Xie et al., 2009 | -1195 G/A | GG: 19, GA: 77, AA: 50 | GG: 40, GA: 64, AA: 44 | GA+AA vs. GG | 2.49 (1.33–4.69) | 0.033 |
| Dienha et al., 2023 | -1195 G/A | GG: 2, GA: 8, AA: 12 | GG: 0, GA: 1, AA: 11 | GA+AA vs. GG | 6.00 (1.05–71.08) | 0.024 |
| Daing et al., 2012 | -1195 G/A | GG: 10, GA: 27, AA: 19 | GG: 20, GA: 25, AA: 15 | GA+AA vs. GG | 1.41 (0.89–2.21) | — |
| Prakash et al., 2015 | -1195 G/A | GG: 147, GA: 46, AA: 7 | GG: 154, GA: 42, AA: 4 | GA+AA vs. GG | 1.14 (0.71–1.84) | 0.578 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




