Submitted:
25 June 2025
Posted:
26 June 2025
Read the latest preprint version here
Abstract
Keywords:
1. Introduction
2. Dataset Description
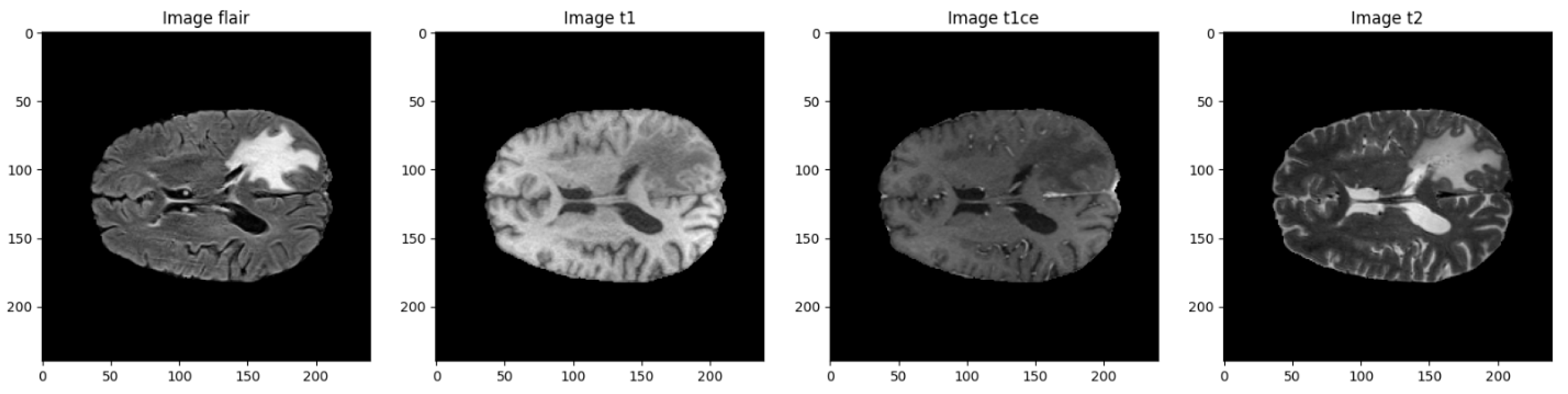
Segmentation Labels:
| Label | Description |
|---|---|
| 0 | Background |
| 1 | Necrotic/Non-enhancing Tumor Core (NCR/NET) |
| 2 | Edema (ED) |
| 4→3 | Enhancing |
Sub-region combinations:
| Label | Sub-region |
|---|---|
| 1 | Tumor Core (TC) |
| 1, 2, 3 | Whole Tumor (WT) |
| 3 | Enhancing Tumor (ET) |
3. Data and Image Preprocessing
3.1. Image Preprocessing
Intensity Normalization
Slice Selection
Cropping and Resizing
Data Augmentation
Input Modalities
3.2. Label Preprocessing
Label Remapping
| Label | Description |
|---|---|
| 0 | Background |
| 1 | Necrotic/Non-enhancing Tumor Core (NCR/NET) |
| 2 | Edema (ED) |
| 3 | Enhancing Tumor (ET) |
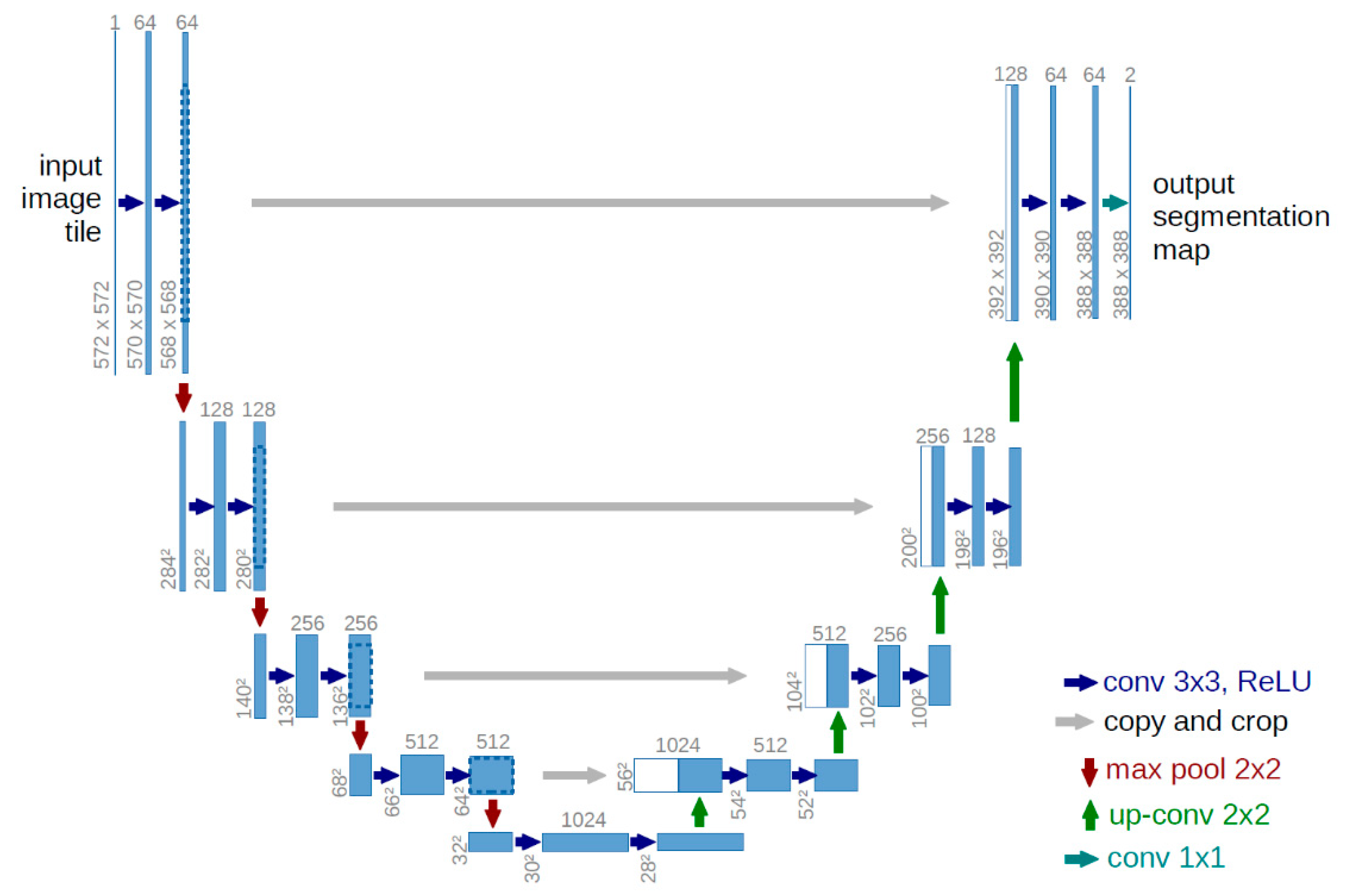
3.3. Model Architecture
U-Net Structure
- Encoder Path:
- Bottleneck:
- Decoder Path:
- Output Layer:
Input and Output Specifications
- Input Shape: Each input consists of two channels—FLAIR and T1CE—resulting in a shape of (128, 128, 2).
- Output Shape: The model produces a segmentation map of shape (128, 128, 4), with class-wise probabilities for each tumor sub-region.
Training Configuration
- Frameworks: TensorFlow 2.12 and Keras
- Loss Function: Categorical Crossentropy (suitable for multi-class segmentation)
- Optimizer: Adam optimizer with a learning rate of 0.001
- Regularization: Dropout (rate = 0.2) in the bottleneck layer
Evaluation Metrics
- Mean Intersection over Union (Mean IoU):
- Dice Similarity Coefficient (DSC):
- o Overall Dice: Aggregates segmentation performance across all classes.
- o Class-specific Dice: Computed separately for necrotic core, edema, and enhancing tumor regions.
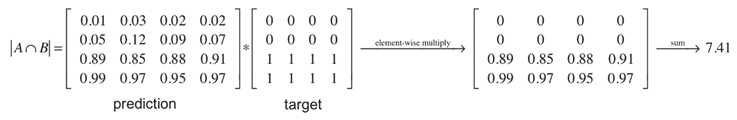
- Precision:
- Sensitivity (Recall):
- Specificity:
3.4. Training Strategy
- Train/Val/Test split: 70% / 15% / 15%
- Epochs: 30
- Optimizer: Adam (learning rate = 0.001)
- Callbacks: ReduceLROnPlateau, EarlyStopping, CSVLogger
- Batch size: 1 (due to memory constraints)
- Custom DataGenerator for real-time augmentation and loading
4. Results
| Metric | Value |
| Validation loss | 0.0284 |
| Validation accuracy | 98.84% |
| Global Dice Coefficient | 0.5139 |
| Precision | 99.08% |
| Specificity | 99.69% |
| Tumor Sub-Region | Validation Dice Score |
|---|---|
| Necrotic Core (NCR/NET) | 0.4292 |
| Edema (ED) | 0.4644 |
| Enhancing Tumor (ET) | 0.5895 |
- Training per-class Dice:
- NCR/NET: 0.4920
- ED: 0.6751
- ET: 0.6251
| Tumor Sub-Region | Training Dice Score |
|---|---|
| Necrotic Core (NCR/NET) | 0.4920 |
| Edema (ED) | 0.6751 |
| Enhancing Tumor (ET) | 0.6251 |
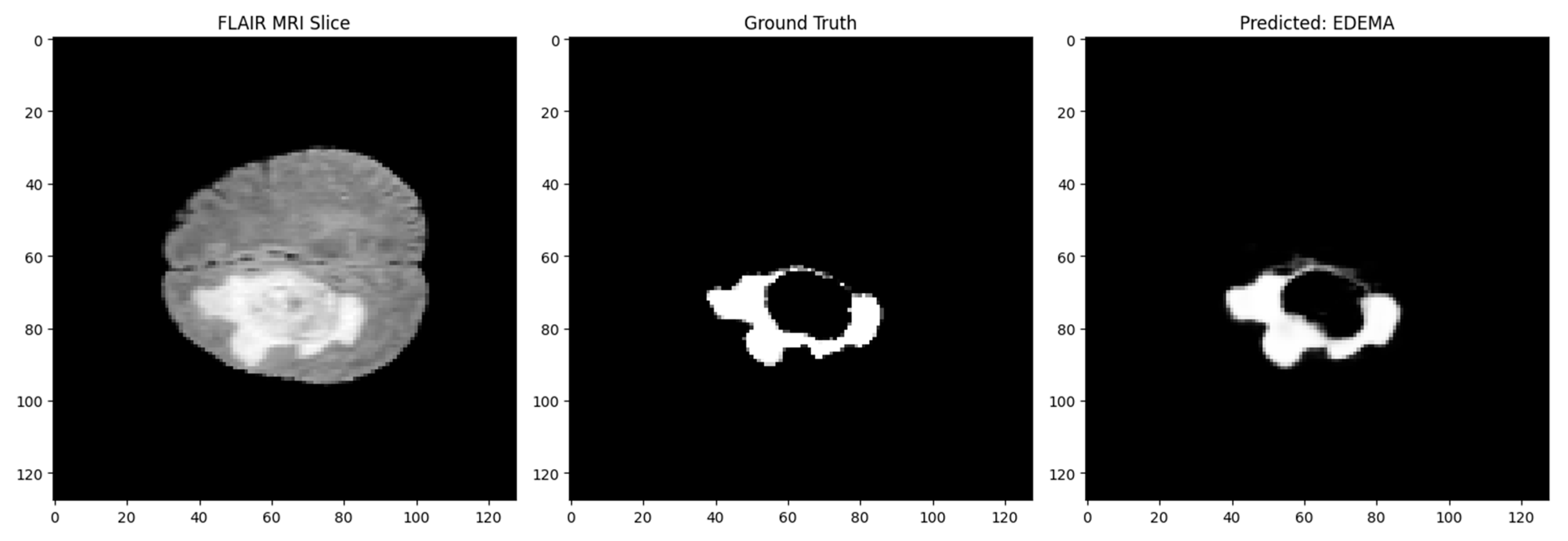
5. Discussion
Performance Observations
Model Advantages
6. Conclusion
References
- W. Stupp et al., “Radiotherapy plus concomitant and adjuvant temozolomide for glioblastoma,” New England Journal of Medicine, vol. 352, no. 10, pp. 987–996, 2005.
- S. Bauer, R. Wiest, L.-P. Nolte, and M. Reyes, “A survey of MRI-based medical image analysis for brain tumor studies,” Physics in Medicine and Biology, vol. 58, no. 13, pp. R97–R129, 2013.
- B. H. Menze, A. Jakab, S. Bauer, J. Kalpathy-Cramer, K. Farahani, J. Kirby, et al. "The Multimodal Brain Tumor Image Segmentation Benchmark (BRATS)", IEEE Transactions on Medical Imaging 34(10), 1993-2024 (2015). [CrossRef]
- U.Baid, et al., The RSNA-ASNR-MICCAI BraTS 2021 Benchmark on Brain Tumor Segmentation and Radiogenomic Classification, arXiv:2107.02314, 2021.
- S. Bakas, H. Akbari, A. Sotiras, M. Bilello, M. Rozycki, J.S. Kirby, et al., "Advancing The Cancer Genome Atlas glioma MRI collections with expert segmentation labels and radiomic features", Nature Scientific Data, 4:170117 (2017). [CrossRef]
- M. Havaei et al., “Brain tumor segmentation with deep neural networks,” Medical Image Analysis, vol. 35, pp. 18–31, Jan. 2017.
- N. Otsu, “A threshold selection method from gray-level histograms,” IEEE Transactions on Systems, Man, and Cybernetics, vol. 9, no. 1, pp. 62–66, 1979.
- R. Adams and L. Bischof, “Seeded region growing,” IEEE Transactions on Pattern Analysis and Machine Intelligence, vol. 16, no. 6, pp. 641–647, 1994.
- J. Canny, “A computational approach to edge detection,” IEEE Transactions on Pattern Analysis and Machine Intelligence, vol. 8, no. 6, pp. 679–698, 1986.
- J. C. Bezdek, R. Ehrlich, and W. Full, “FCM: The fuzzy c-means clustering algorithm,” Computers & Geosciences, vol. 10, no. 2–3, pp. 191–203, 1984.
- M. Kass, A. Witkin, and D. Terzopoulos, “Snakes: Active contour models,” International Journal of Computer Vision, vol. 1, no. 4, pp. 321–331, 1988.
- S. Osher and J. A. Sethian, “Fronts propagating with curvature-dependent speed: Algorithms based on Hamilton–Jacobi formulations,” Journal of Computational Physics, vol. 79, no. 1, pp. 12–49, 1988.
- O. Ronneberger, P. Fischer, and T. Brox, “U-Net: Convolutional networks for biomedical image segmentation,” in *Proc. Medical Image Computing and Computer-Assisted Intervention (MICCAI)*, 2015, pp. 234–241.
- K. He, G. Gkioxari, P. Dollár, and R. Girshick, “Mask R-CNN,” in *Proc. IEEE International Conference on Computer Vision (ICCV)*, 2017, pp. 2961–2969.
- J. Chen et al., “TransUNet: Transformers make strong encoders for medical image segmentation,” arXiv preprint arXiv:2102.04306, 2021.
- Elaff, I. EL-KEMANY, A. and KHOLIF, M. "Universal and stable medical image generation for tissue segmentation (The unistable method)" Turkish Journal of Electrical Engineering and Computer Sciences: Vol. 25: No. 2, Article 32, 2017. [CrossRef]
- Elaff, I. “Medical Image Enhancement Based on Volumetric Tissue Segmentation Fusion (Uni-Stable 3D Method)”, Journal of Science, Technology and Engineering Research, vol. 4, no. 2, pp. 78–89, 2023. [CrossRef]
- Elaff I "Brain Tissue Classification Based on Diffusion Tensor Imaging: A Comparative Study Between Some Clustering Algorithms and Their Effect on Different Diffusion Tensor Imaging Scalar Indices". Iran J Radiol. 2016 Feb 28;13(2):e23726. [CrossRef]
- El-Aff I "Human brain tissues segmentation based on DTI data," 2012 11th International Conference on Information Science, Signal Processing and their Applications (ISSPA), Montreal, QC, Canada, 2012, pp. 876-881. [CrossRef]
- El-Dahshan, E.-S. A., Hosny, T., & Salem, A.-B. M. (2010). Hybrid intelligent techniques for MRI brain images classification. Digital Signal Processing, 20(2), 433–441.
- Kapur, T., Grimson, W. E. L., Wells, W. M., & Kikinis, R. (1996). Segmentation of brain tissue from magnetic resonance images. Medical Image Analysis, 1(2), 109–127.
- Ronneberger, O., Fischer, P., & Brox, T. (2015). U-Net: Convolutional networks for biomedical image segmentation. In MICCAI (pp. 234–241). Springer.
- Milletari, F., Navab, N., & Ahmadi, S. A. (2016). V-Net: Fully convolutional neural networks for volumetric medical image segmentation. In 3DV (pp. 565–571). IEEE.
- Jiang, Z., Zhang, H., Wang, Y., & Ko, H. (2021). Mask R-CNN with DenseNet for brain tumor segmentation. Computers, Materials & Continua, 67(2), 1941–1955.
- I. Elaff, “Comparative study between spatio-temporal models for brain tumor growth,” *Biochemical and Biophysical Research Communications*, vol. 496, no. 4, pp. 1263–1268, 2018. [CrossRef]
- The BraTS Consortium, “BraTS: Brain Tumor Segmentation dataset,” The Cancer Imaging Archive. [Online]. Available: https://www.synapse.org/#!Synapse:syn36640318.
- CBICA/UPenn, “BraTS: Brain Tumor Segmentation dataset,” Synapse, 2023. [Online]. Available: https://www.synapse.org/#!Synapse:syn51156910/wiki/622351.
- Nyúl, L. G., Udupa, J. K., & Zhang, X. (2000). New variants of a method of MRI scale standardization. IEEE Transactions on Medical Imaging, 19(2), 143–150.
- Isensee, F., Petersen, J., Kohl, S., Jäger, P. F., & Maier-Hein, K. H. (2021). nnU-Net: a self-configuring method for deep learning-based biomedical image segmentation. Nature Methods, 18, 203–211.
- Shorten, C., & Khoshgoftaar, T. M. (2019). A survey on image data augmentation for deep learning. Journal of Big Data, 6(1), 60.
- Bakas, S., Reyes, M., Jakab, A., Bauer, S., Rempfler, M., Crimi, A., ... & Menze, B. H. (2018). Identifying the best machine learning algorithms for brain tumor segmentation, progression assessment, and overall survival prediction in the BRATS challenge. arXiv preprint arXiv:1811.02629.
- Havaei, M., Davy, A., Warde-Farley, D., Biard, A., Courville, A., Bengio, Y., ... & Larochelle, H. (2017). Brain tumor segmentation with deep neural networks. Medical Image Analysis, 35, 18–31.
- Ronneberger, O., Fischer, P., & Brox, T. (2015). U-Net: Convolutional Networks for Biomedical Image Segmentation. In Medical Image Computing and Computer-Assisted Intervention (MICCAI), pp. 234–241.
- O. Ronneberger, P. Fischer, and T. Brox, "U-Net: Convolutional Networks for Biomedical Image Segmentation," Proc. MICCAI, vol. 9351, pp. 234–241, 2015.
- Taha, A. A., & Hanbury, A. (2015). Metrics for evaluating 3D medical image segmentation: analysis, selection, and tool. BMC Medical Imaging, 15(1), 1–28. [CrossRef]
- Zou, K. H., et al. (2004). Statistical validation of image segmentation quality based on a spatial overlap index: scientific reports. Academic Radiology, 11(2), 178–189. [CrossRef]
- Bakas, S., et al. (2018). "Identifying the best machine learning algorithms for brain tumor segmentation, progression assessment, and overall survival prediction in the BRATS challenge." Medical Image Analysis, 55, 115–142.
- Isensee, F., et al. (2021). "Nnu-net: Self-adapting framework for U-Net-based medical image segmentation." Nature Methods, 18(2), 203–211.
- Shorten, C., & Khoshgoftaar, T. M. (2019). "A survey on image data augmentation for deep learning." Journal of Big Data, 6(1), 1–48.
- Zhou, Z., et al. (2018). "UNet++: A nested U-Net architecture for medical image segmentation." Deep Learning in Medical Image Analysis and Multimodal Learning for Clinical Decision Support, 3–11.
| Modality | Description |
|---|---|
| T1 | T1-weighted structural MRI |
| T1CE | T1-weighted with contrast enhancement (gadolinium) |
| T2 | T2-weighted imaging, useful for fluid detection |
| FLAIR | Fluid-Attenuated Inversion Recovery, suppresses CSF to highlight lesions |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




