Submitted:
06 June 2025
Posted:
09 June 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
3. Results
3.1. COX-1 interactions
3.2. COX-2 interactions
3.4. PLA2 interactions
3.5. Blind docking results
- Active site sequence: A list of amino acid residues forming the catalytic site of the corresponding enzyme.
- Binding residues: Specific residues through which the terpene establishes contact with the enzyme during docking.
- Matches: Residues that appear most frequently across simulations, indicating potentially stable interactions and playing a key role in stabilizing the complex.
4. Discussion
5. Conclusions
Author Contributions
Funding
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Wound Repair: Results of a Pre-clinical Study to Evaluate Electropsun Collagen-Elastin-PCL Scaffolds as Dermal Substitutes. Burns 45 2019, 1639–1648. [CrossRef]
- Takeuchi, O.; Akira, S. Pattern Recognition Receptors and Inflammation. Cell. 2010, 140, 805–820. [Google Scholar] [CrossRef] [PubMed]
- Ferrero-Miliani, L.; Nielsen, O.; Andersen, P.; Girardin, S. Chronic inflammation: importance of NOD2 and NALP3 in interleukin-1β generation. Clin Exp Immunol. 2007, 147, 227–235. [Google Scholar] [CrossRef] [PubMed]
- Chertov, O.; Yang, D.; Howard, O.; Oppenheim, J.J. Leukocyte granule proteins mobilize innate host defenses and adaptive immune responses. Immunol Rev. 2000, 177, 68–78. [Google Scholar] [CrossRef] [PubMed]
- Gu, J.H.; Ge, J.B.; Li, M.; Wu, F.; Zhang, W.; Qin, Z.H. Inhibition of NF-Κb Activation Is Associated with Anti-inflammatory and Anti-Apoptotic Effects of Ginkgolide B in a Mouse Model of Cerebral Ischemia/Reperfusion Injury. Eur. J. Pharm. Sci. 2012, 47, 652–660. [Google Scholar] [CrossRef] [PubMed]
- Lawrence, T. The Nuclear Factor NF-κB Pathway in Inflammation. CSH Perspect Biol. 2009.
- Libby, P. Inflammatory mechanisms: the molecular basis of inflammation and disease. Nutr Rev. 2007, 65, 140–146. [Google Scholar] [CrossRef]
- Del Prado-Audelo, ML.; Cortés, H.; Caballero-Florán, I.H.; González-Torres, M.; Escutia-Guadarrama, L.; Bernal-Chávez, SA.; Giraldo-Gomez, D.M.; Magaña, J.J.; Leyva-Gómez, G. Therapeutic Applications of Terpenes on Inflammatory Diseases. Front Pharmacol. 2021, 12, 704197. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Prakash, V. Terpenoids as Source of Anti-Inflammatory Compounds. Asian J. Pharm. Clin. Res. 2017, 10, 68–76. [Google Scholar] [CrossRef]
- Serrano Vega, R. J.; Campos Xolalpa, N.; Castro Alonso, J. A.; González Pérez, C.; Pérez Ramos, J.; Pérez Gutiérrez, S. “Terpenes from from Natural Natural Products Products with with Potential Potential Anti- Activity Inflammatory Activity”. in Terpenes from Natural Products with Potential Anti-Inflammatory Activity. Perveen, S., Al-Taweel, A., Eds.; London: Intech Open, 2018; pp 59–85. [CrossRef]
- Miciaccia, M.; Belviso, B.D.; Iaselli, M.; Cingolani, G.; Ferorelli, S.; Cappellari, M.;, Loguercio Polosa,.; P.; Perrone, M.G.; Caliandro, R.; Scilimati, A. Three-dimensional structure of human cyclooxygenase (hCOX)-1. Sci Rep. 2021, 11(1), 4312. [CrossRef] [PubMed] [PubMed Central]
- Vecchio, A.J.; Simmons, D.M.; Malkowski, M.G. Structural basis of fatty acid substrate binding to cyclooxygenase-2. J Biol Chem. 2010, 285(29), 22152–63. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Gilbert, N.C.; Bartlett, S.G.; Waight, M, T.; Neau, D.B.; Boeglin, W.E.; Brash, A.R.; Newcomer, M.E. The structure of human 5-lipoxygenase. Science 2011, 14, 331(6014), 217-9. PMID: 21233389; PMCID: PMC3245680. [CrossRef]
- Cha, S.S.; Lee, D.; Adams, J.; Kurdyla, J.T.; Jones, C.S.; Marshall, L.A.; Bolognese, B.; Abdel-Meguid, S.S.; Oh, B.H. ; High-resolution X-ray crystallography reveals precise binding interactions between human nonpancreatic secreted phospholipase A2 and a highly potent inhibitor (FPL67047XX). J Med Chem. 1996, 39(20), 3878–81. [Google Scholar] [CrossRef] [PubMed]
- O’Boyle, N.M.; Banck, M.; James, C.A.; Morley, C.; Vandermeersch, T.; Hutchison, G.R. Open Babel: An open chemical toolbox. J Cheminform. 2011, 3, 1–14. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Hanwell, M.D.; Curtis, D. E.; Lonie, D. C.; Vandermeersch, T.; Zurek, E.; Hutchison, G. R. Avogadro: an advanced semantic chemical editor, visualization, and analysis platform. J Cheminform. 2012, 4, 1–17. [Google Scholar] [CrossRef] [PubMed]
- Read, R.J.; Pettersen, E.F.; McCoy, A.J.; Croll, TI.; Terwilliger, T.C.; Poon, B.K.; Meng, E.C.; Liebschner, D.; Adams, P.D. Likelihood-based interactive local docking into cryo-EM maps in ChimeraX. Acta Crystallogr D Struct Biol. 2024, 80(8), 588–598. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Morris, G.M.; Huey, R.; Lindstrom, W.; Sanner, M.F.; Belew, R.K.; Goodsell., D.S.; Olson, A.J. AutoDock4 and AutoDockTools4: Automated docking with selective receptor flexibility. J Comput Chem. 2009, 30, 2785–2791. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Torres, P.H.M.; Sodero, A.C.R.; Jofily, P.; Silva-Jr, F.P. Key Topics in Molecular Docking for Drug Design. Int J Mol Sci. 2019, 20(18), 4574. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Yuriev, E.; Ramsland, P.A. Latest developments in molecular docking: 2010-2011 in review. J Mol Recognit. 2013, 26(5), 215–39. [Google Scholar] [CrossRef] [PubMed]
- Sulimov, V.B.; Kutov, D.C.; Sulimov, A.V. Advances in Docking. Curr Med Chem. 2019, 26(42), 7555–7580. [Google Scholar] [CrossRef] [PubMed]
- Fuhrmann, J.; Rurainski, A.; Lenhof, H.P.; Neumann, D. A new Lamarckian genetic algorithm for flexible ligand-receptor docking. J Comput Chem. 2010, 31(9), 1911–8. [Google Scholar] [CrossRef] [PubMed]
| COX-1 | Limonene | Camphor | Linalool | Menthol | α-pinene | β-pinene | AA |
| Conformations | 2 | 13 | 33 | 20 | 54 | 60 | 9 |
| RMSD from reference structure | 59.121 A | 67.643 A | 66.924 A | 67.481 A | 66.995 A | 67.775 A | 57.255 A |
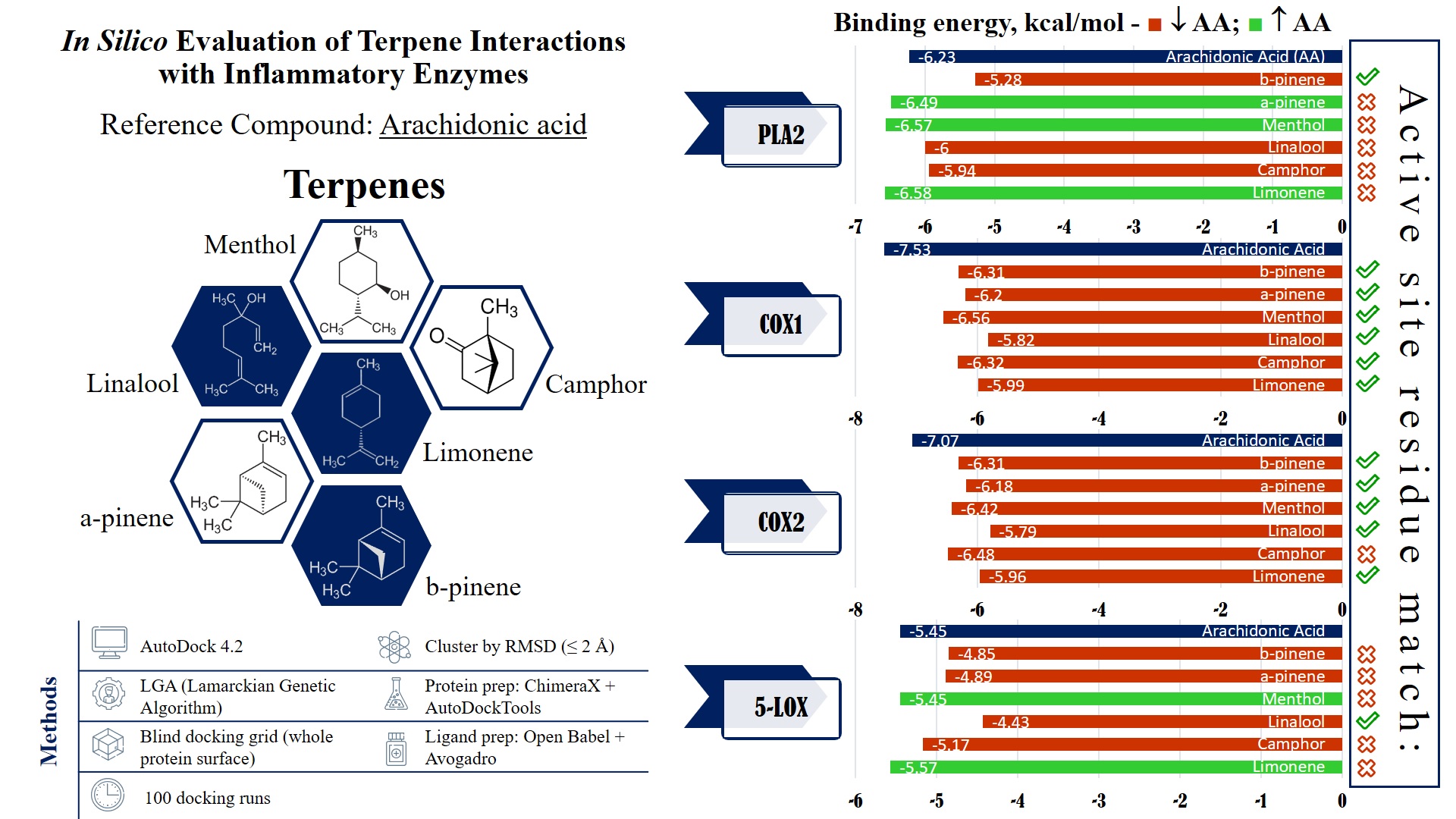
| Binding energy | -5.99 kcal/mol | -6.32 kcal/mol | -5.82 kcal/mol | -6.56 kcal/mol | -6.20 kcal/mol | -6.31 kcal/mol | -7.53 kcal/mol |
| Inhibition Constant, Ki | 40.60 µM | 23.37 µM | 54.64 µM | 15.50 µM | 28.74 µM | 23.61 µM | 3.02 µM |
| Total Internal energy | -0.13 kcal/mol | +0.00 kcal/mol | -0.33 kcal/mol | -0.17 kcal/mol | +0.00 kcal/mol | +0.00 kcal/mol | -1.61 kcal/mol |
| Torsional Free Energy | +0.30 kcal/mol | +0.00 kcal/mol | +1.49 kcal/mol | +0.60 kcal/mol | +0.00 kcal/mol | +0.00 kcal/mol | +4.47 kcal/mol |
| Unbound Energy | -0.13 kcal/mol | +0.00 kcal/mol | -0.33 kcal/mol | -0.17 kcal/mol | +0.00 kcal/mol | +0.00 kcal/mol | -1.61 kcal/mol |
| COX-2 | Limonene | Camphor | Linalol | Menthol | α-pinene | β-pinene | AA |
| Conformations | 3 | 13 | 19 | 26 | 36 | 46 | 7 |
| RMSD from reference structure | 59.204 A | 67.674 A | 66.971 A | 67.752 A | 67.087 A | 67.770 A | 58.016 A |
| Binding energy | -5.96 kcal/mol | -6.48 kcal/mol | -5.79 kcal/mol | -6.42 kcal/mol | -6.18 kcal/mol | -6.31 kcal/mol | -7.07 kcal/mol |
| Inhibition Constant, Ki | 43.00 µM | 17.84 µM | 56.61 µM | 19.80 µM | 29.44 µM | 23.61 µM | 6.55 µM |
| Total Internal energy | -0.13 kcal/mol | +0.00 kcal/mol | -0.37 kcal/mol | -0.16 kcal/mol | +0.00 kcal/mol | +0.00 kcal/mol | -1.18 kcal/mol |
| Torsional Free Energy | +0.30 kcal/mol | +0.00 kcal/mol | +1.49 kcal/mol | +0.60 kcal/mol | +0.00 kcal/mol | +0.00 kcal/mol | +4.47 kcal/mol |
| Unbound Energy | -0.13 kcal/mol | +0.00 kcal/mol | -0.37 kcal/mol | -0.16 kcal/mol | +0.00 kcal/mol | +0.00 kcal/mol | -1.18 kcal/mol |
| 5-LOX | Limonene | Camphor | Linalool | Menthol | α-pinene | β-pinene | AA |
| Conformations | 15 | 13 | 1 | 14 | 32 | 26 | 1 |
| RMSD from reference structure | 64.854 A | 77.022 A | 23.527 A | 64.372 A | 19.328 A | 18.577 A | 28.948 A |
| Binding energy | -5.57 kcal/mol | -5.17 kcal/mol | -4.43 kcal/mol | -5.45 kcal/mol | -4.89 kcal/mol | -4.85 kcal/mol | -5.45 kcal/mol |
| Inhibition Constant, Ki | 82.28 µM | 161.15 µM | 564.63 µM | 102.00 µM | 261.53 µM | 280.35 µM | 101.13 µM |
| Total Internal energy | -0.13 kcal/mol | +0.00 kcal/mol | -0.39 kcal/mol | -0.16 kcal/mol | +0.00 kcal/mol | +0.00 kcal/mol | -1.08 kcal/mol |
| Torsional Free Energy | +0.30 kcal/mol | +0.00 kcal/mol | +1.49 kcal/mol | +0.60 kcal/mol | +0.00 kcal/mol | +0.00 kcal/mol | +4.47 kcal/mol |
| Unbound Energy | -0.13 kcal/mol | +0.00 kcal/mol | -0.39 kcal/mol | -0.16 kcal/mol | +0.00 kcal/mol | +0.00 kcal/mol | -1.08 kcal/mol |
| PLA2 | Limonene | Camphor | Linalool | Menthol | α-pinene | β-pinene | AA |
| Conformations | 19 | 6 | 11 | 6 | 8 | 25 | 2 |
| RMSD from reference structure | 89.108 A | 96.944 A | 96.905 A | 98.205 A | 110.729 A | 100.997 A | 98.253 A |
| Binding energy | -6.58 kcal/mol | -5.94 kcal/mol | -6.00 kcal/mol | -6.57 kcal/mol | -6.49 kcal/mol | -5.28 kcal/mol | -6.23 kcal/mol |
| Inhibition Constant, Ki | 14.99 µM | 44.43 µM | 39.77 µM | 15.22 µM | 17.50 µM | 134.07 µM | 27.27 µM |
| Total Internal energy | -0.13 kcal/mol | +0.00 kcal/mol | +0.00 kcal/mol | -0.17 kcal/mol | -0.26 kcal/mol | +0.00 kcal/mol | -1.91 kcal/mol |
| Torsional Free Energy | +0.30 kcal/mol | +0.00 kcal/mol | +0.00 kcal/mol | +0.60 kcal/mol | +1.49 kcal/mol | +0.00 kcal/mol | +4.47 kcal/mol |
| Unbound Energy | -0.13 kcal/mol | +0.00 kcal/mol | +0.00 kcal/mol | -0.17 kcal/mol | -0.26 kcal/mol | +0.00 kcal/mol | -1.91 kcal/mol |
| Enzyme | Active Site | Bibliography |
| COX-1 | LEU117, ARG120, PHE205, PHE209, VAL344, ILE345, TYR348, VAL349, LEU352, SER353, TYR355, PHE381, TYR385, TRP387, ILE523, HIS207, HIS388, GLN203 | [11] |
| COX-2 | TYR385, SER530, LEU531, VAL116, ALA527, VAL349, TYR355, LEU352, VAL523, PHE518, PHE205, PHE209, TYR348, VAL349, LEU352, PHE381, TYR385, TRP387, VAL523, ALA527, LEU534 | [12] |
| 5-LOX | HIS367, HIS372, HIS550, PHE177, TYR181, LEU368, LEU414, ILE406, ALA603, ALA606, TRP599, ASN407, HIS432 | [13] |
| PLA2 | PHE5, HIS6, ILE9, ALA18, TYR21, GLY22, GLY29, GLY32, CYS44, HIS47, TYR51 | [14] |
| Terpene | Enzyme | Interaction | Match |
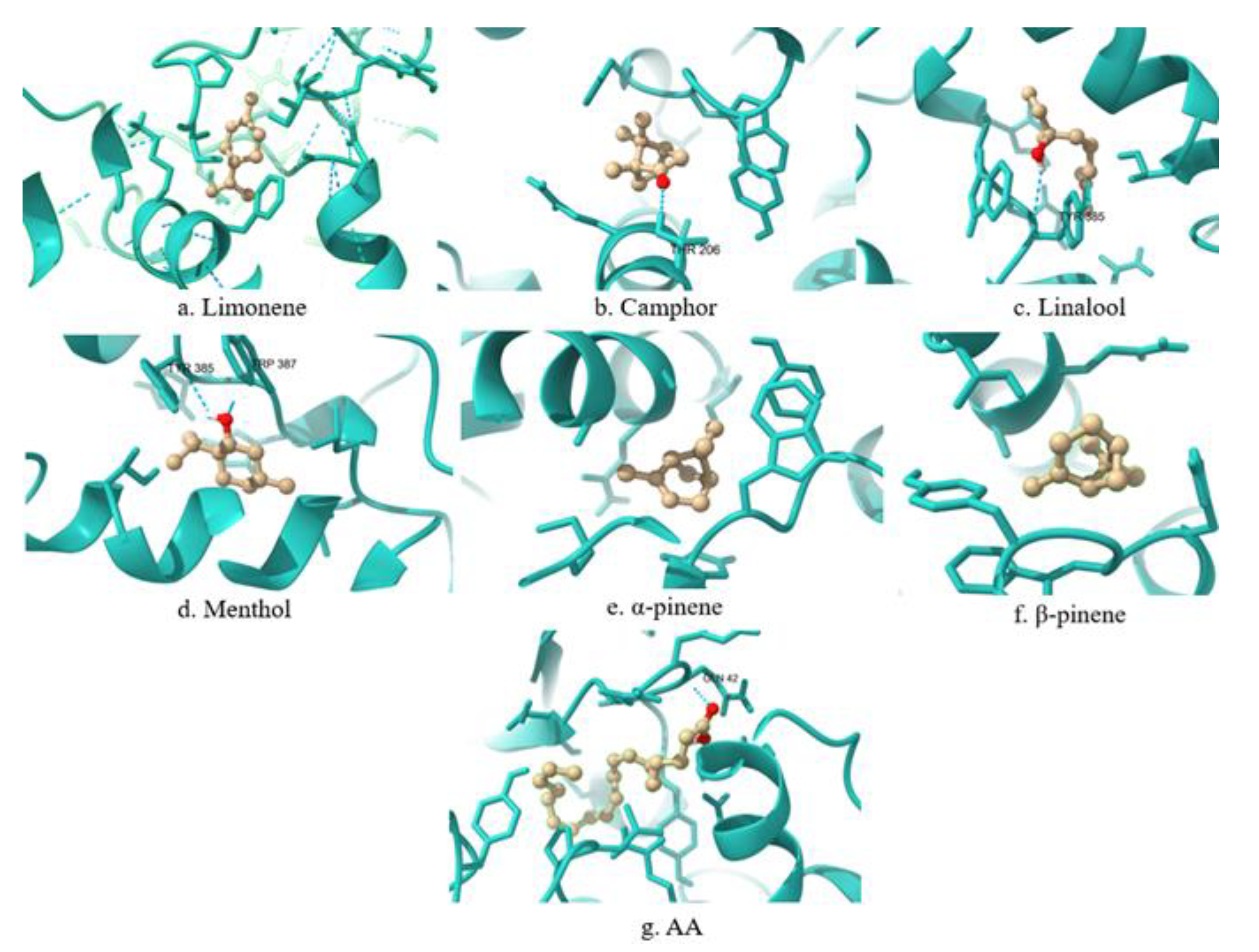
| Limonene | COX-1 | PRO128, ARG376, PRO125, ALA378, ILE124, ILE377, PHE529, PHE381, LYS532 | PHE381 |
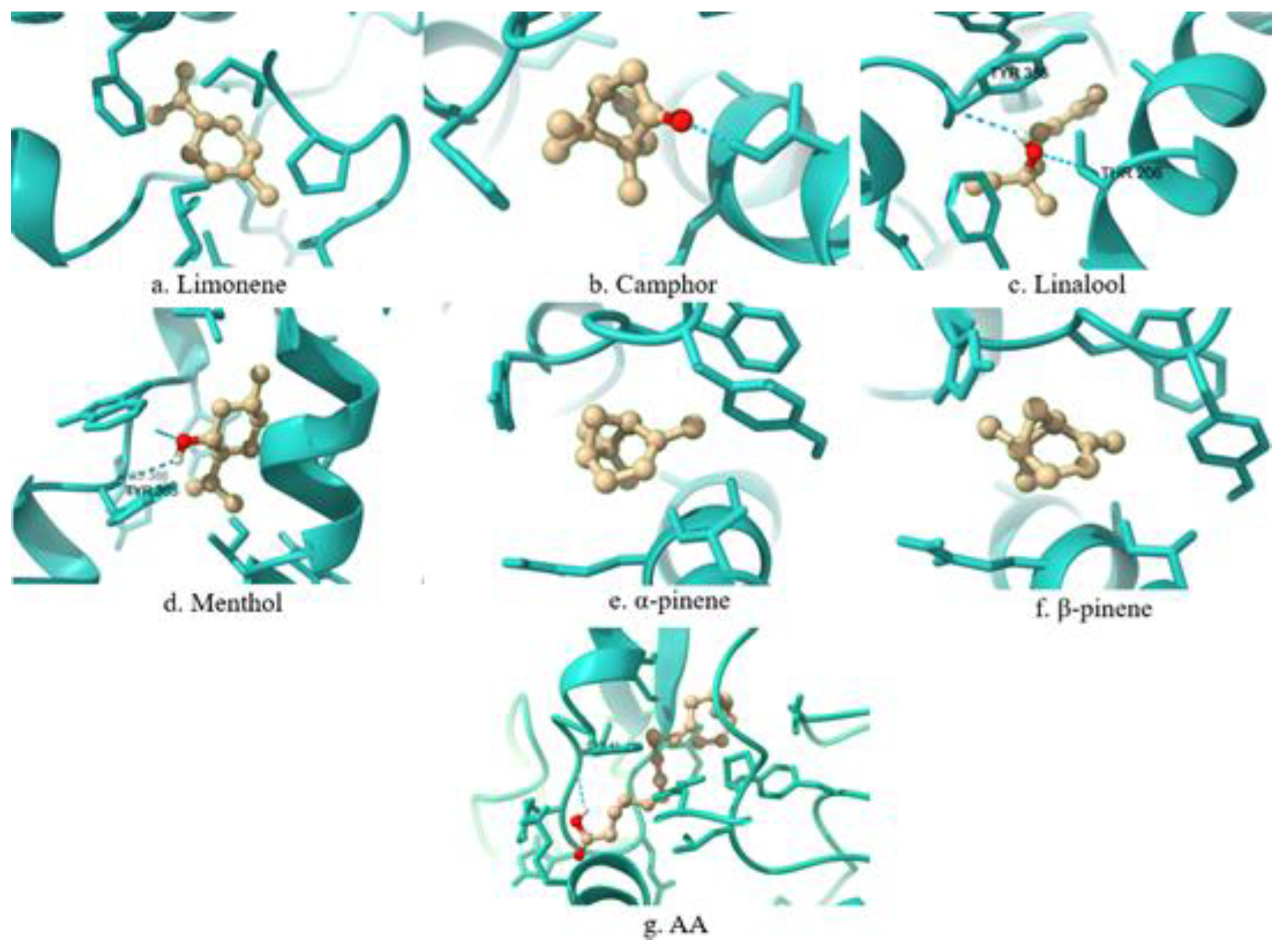
| COX-2 | PRO125, PRO128, ARG376, ILE124, ILE377, PHE529, LYS532, ALA378, PHE381 | PHE381 | |
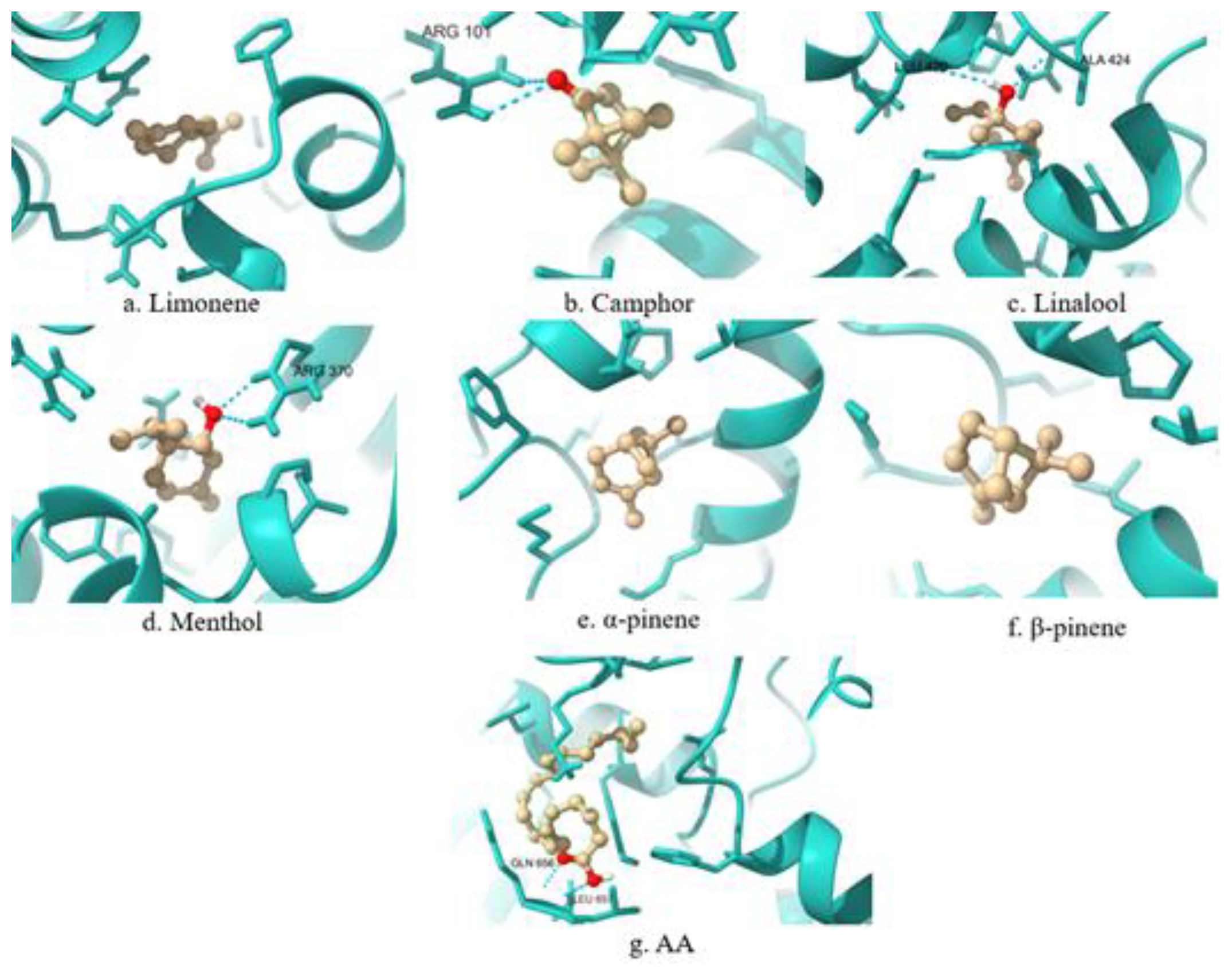
| 5-LOX | LEU448, TYR470, ILE454, PHE450, VAL243, ALA453 | No match. | |
| PLA2 | PHE63, LYS62, PHE63 | No match. | |
| Camphor | COX-1 | ALA202, LEU390, MET391, THR206, HIS388 | HIS388 |
| COX-2 | ALA202, HIS388, THR206, MET391, LEU390 | No match. | |
| 5-LOX | HIS130, VAL110, VAL109, ARG101, LYS133 | No match. | |
| PLA2 | LYS62, LEU64, PHE63, PHE63 | No match. | |
| Linalool | COX-1 | HIS388, HIS386, LEU390, TYR385, ALA202, PHE21 | HIS388, TYR385 |
| COX-2 | LEU390, MET391, ALA202, HIS388, HIS207, PHE210, HIS386, THR206, TYR385 | TYR385 | |
| 5-LOX | PHE359, PHE412, LEU414, TRP599, VAL604, HIS600, ALA603, TYR181, LEU420, ALA424 | LEU414, TRP599, ALA603, TYR181 | |
| PLA2 | LYS62, LEU64, LEU64, PHE63 | No match. | |
| Menthol | COX-1 | HIS388, TYR385, ALA202, MET391, LEU390, ALA199 | HIS388, TYR385 |
| COX-2 | HIS388, TYR385, ALA202, MET391, LEU390, ALA199 | TYR385 | |
| 5-LOX | PHE544, PHE450, SER447, ARG370, ALA453, VAL243 | No match. | |
| PLA2 | LEU64, THR61, LYS62, PHE63, PHE63 | No match. | |
| α-pinene | COX-1 | TYR385, LEU390, HIS388, MET391, ALA199, ALA202, TRP387 | TYR385, HIS388, TRP387 |
| COX-2 | ALA199, HIS388, TYR385, TRP387, LEU390, MET391, ALA202 | TYR385, TRP387 | |
| 5-LOX | VAL645, LEU641, PRO664, LEU617, MET619, ARG638 | No match. | |
| PLA2 | PHE63, THR61, PHE63, LYS62, LEU64 | No match. | |
| β-pinene | COX-1 | HIS388, LEU390, TYR385, TRP387, ALA202, MET391 | TYR385, HIS388, TRP387 |
| COX-2 | HIS388, LEU390, TYR385, TRP387, ALA202, MET391 | TYR385, TRP387 | |
| 5-LOX | LEU641, LEU617, VAL645, PRO664, MET619, ARG638 | No match. | |
| PLA2 | PHE98, PHE5, ALA94, HIS47, CYS28, CYS44 | PHE5, CYS44, HIS47 | |
| AA | COX-1 | CYS47, CYS36, PRO156, PRO153, TYR130, HIS43, ILE46, LYS468, CYS41, LEU152, ARG469, PRO40 | No match. |
| COX-2 | LEU152, TYR39, CYS47, CYS36, PRO156, PRO153, TYR130, ILE46, HIS43, CYS41 | No match. | |
| 5-LOX | MET231, LEU230, LYS319, LEU237, GLN656, TYR234, LEU657 | No match. | |
| PLA2 | HIS47, CYS28, HIS27, ASP48, GLY29, TYR51, LEU2, ALA17, PHE5, ILE9, TYR21, PHE98, CYS44 | PHE5, ILE9, TYR21, GLY29, CYS44, HIS47, TYR51 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




