Submitted:
04 June 2025
Posted:
05 June 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
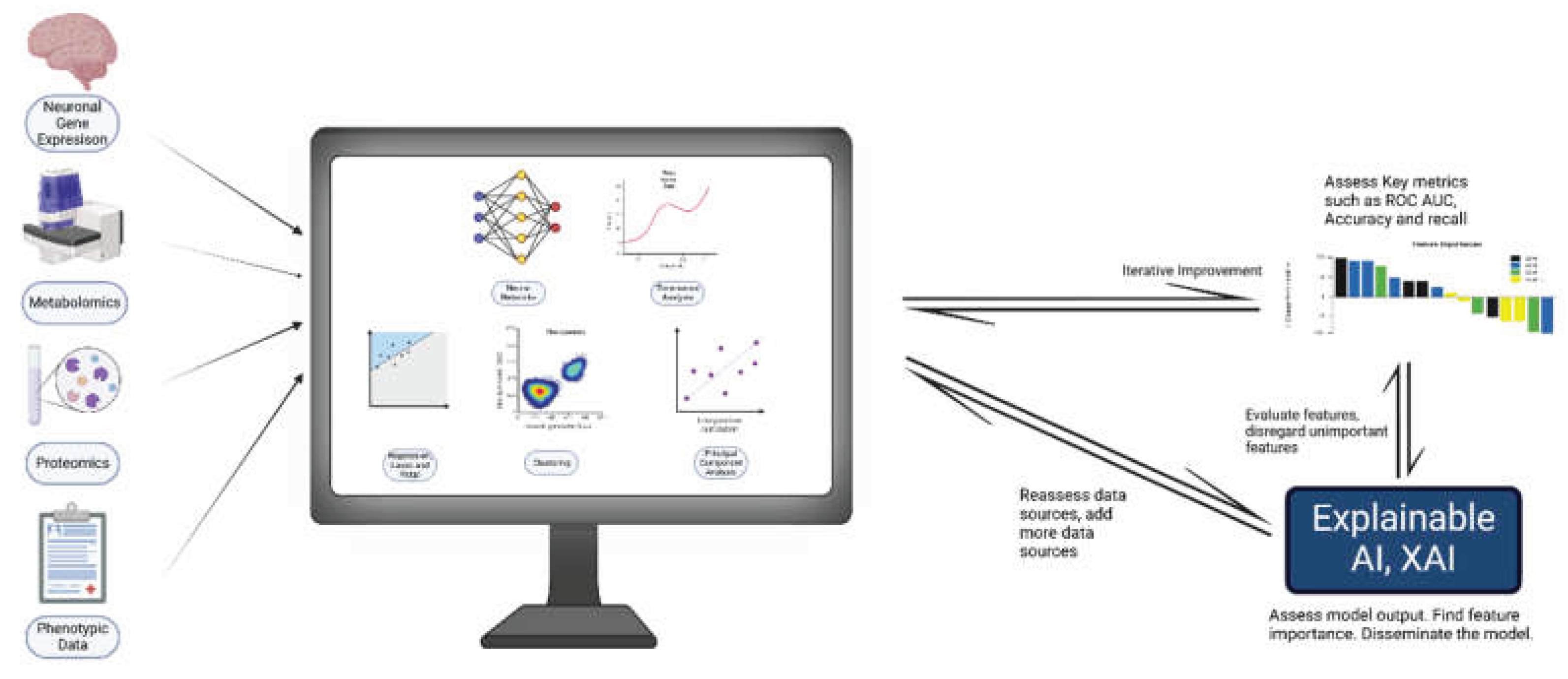
2. Integrative Statistical and Computational Approaches in Neurobiology
3. Dimensionality Reduction and Biological Insights
4. Feature Selection in Functional Genomics
5. Temporal Analysis of Neuronal Regulation
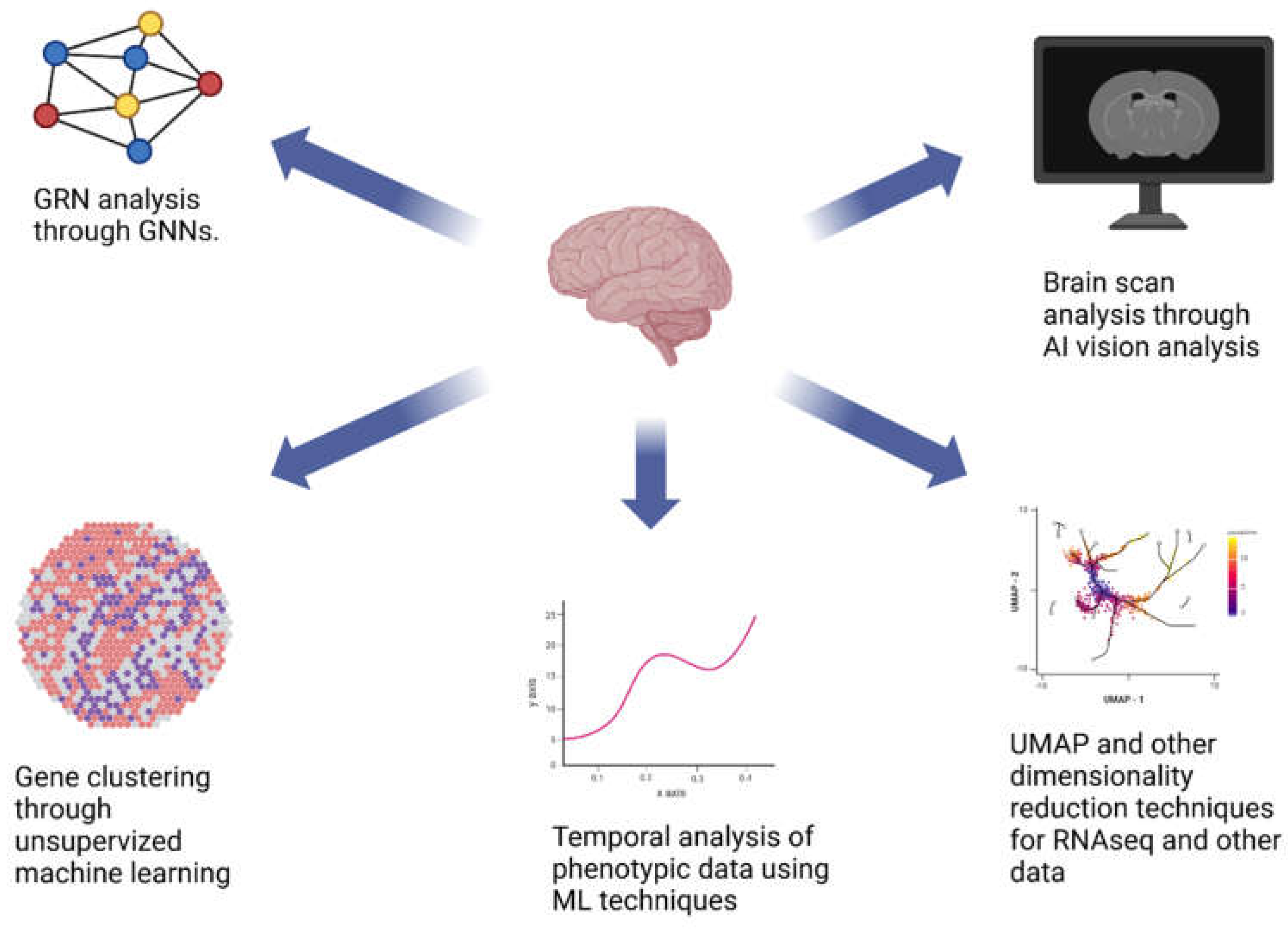
6. Network Analysis for Gene Regulation
7. Hidden Markov Models
8. Explainable AI for Model Interpretability
9. Case Studies Demonstrating Biological Applications
10. Limitations
11. Conclusion
| Datasets | ||||
| Dataset Name | Data Type | Purpose | Scale and Resolution | Key Application |
| Allen Brain Atlas (Vries, Siegle, & Koch, 2023) | Imaging, transcriptomics | Mapping neuronal activity | More than 100,000 neurons, cellular resolution | Understanding cortical structure functions |
| Tabula Muris ( (The Tabula Muris Consortium, 2020)) | Single-cell RNA-seq | Clustering neuronal subtypes | Approx. 100,000 cells, single-cell resolution | Exploring immune-neuronal interactions |
| Human Cell Atlas (Rood, et al., 2025) (Rozenblatt-Rosen, Stubbington, Regev, & Teichmann, 2017) | Multi-omics | Integrating transcriptomics & epigenomics | More than ten million cells, single cell & spatial resolution | Tracking developmental trajectories |
| Roadmap Epigenomics (Satterlee, et al., 2019) | Epigenomics | Studying histone modifications | Genome-wide, high-resolution | Identifying histone plasticity markers |
| Computational Techniques | ||||
| Methodology | Function | Example Use Case | Dataset Used | Strength |
| LASSO Regression | Feature selection with sparsity | Identifying early Alzheimer’s biomarkers | Alzheimer’s Disease National Initiative | Select the most relevant gene markers (Cui, et al., 2021) |
| Graph Neural Networks | Modeling gene regulatory networks | Infer regulatory pathways | Dream5 network initiative dataset | Captures complex interactions (Feng, Jiang, Yin, & Sun, 2023) |
| SHAP | Identifying key biological features | Understanding transcriptional mechanisms in ASD | Various | Improves AI model interpretability (Castro-Martinez, Vargas, Diaz-Beltran, & Esteban, 2024) |
| Challenges in Multi-omics Analysis | ||||
| Challenge | Issue | Suggested Approach | Impact on Research | |
| Batch Effects | Bias in multi-omics integration | Standardizing data pipelines | Improves reproducibility | |
| Data Sparsity | Incomplete feature representation | Robust imputation methods | Enables deduction of rare neuronal subtypes | |
| Deep learning models | Lack of interpretability in AI models | Implementation of explainable AI frameworks | Enhances AI-driven biomarker validation. | |
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Alatrany, A. S., Khan, W., Hussain, A., Kolivand, H., & Al-Jumeily, D. (2024). An explainable machine learning approach for Alzheimer’s disease classification. 14(4637). [CrossRef]
- Booij, M. M., van Noorden, M. S., van Vliet, I. M., Ottenheim, N. R., van der Wee, N. J., van Hemert, A. M., & Giltay, E. J. (2021). Dynamic time warp analysis of individual symptom trajectories in depressed patients treated with electroconvulsive therapy. Journal of Affective Disorder, 293, 435-443. [CrossRef]
- Castro-Martinez, J., Vargas, E., Diaz-Beltran, L., & Esteban, F. J. (2024). Enhancing Transcriptomic Insights into Neurological Disorders Through the Comparative Analysis of Shapley Values. Curr. Issues Mol. Biol., 46(12). [CrossRef]
- Citri, A., & Malenka, R. C. (2008). Synaptic Plasticity: Multiple Forms, Functions, and Mechanisms. Neuropsychopharmacology, 18-41.
- Cui, X., Xiao, R., Liu, X., Ziao, H., Zheng, X., Zhang, Y., & Du, J. (2021). Adaptive LASSO logistic regression based on particle swarm optimization for Alzheimer's disease early diagnosis. Chemometrics and Intelligent Laboratory Systems(104316). [CrossRef]
- Feng, K., Jiang, H., Yin, C., & Sun, H. (2023). Gene regulatory network inference based on causal discovery integrating with graph neural network. Quantitative Biology. [CrossRef]
- Guyon, I., Weston, J., Barnhill, S., & Vapnik, V. (2002). Gene Selection for Cancer Classification using Support Vector Machines. Machine Learning, 46, 389-422. [CrossRef]
- Han, Y., Huan, L., & Zhou, F. (2021). A dynamic recursive feature elimination framework (dRFE) to further refine a set of OMIC biomarkers. Bioinformatics, 37(15), 2183-2189. [CrossRef]
- James, G., Witten, D., Hastie, T., & Tibshirani, R. (2013). An Introduction to Statistical Learning: with Applications in R (Springer Texts in Statistics). Springer.
- Jeong, Y., Ronen, J., Kopp, W., Lutsik, P., & Akalin, A. (2024). scMaui: a widely applicable deep learning framework for single-cell multiomics integration in the presence of batch effects and missing data. BMC Bioinformatics, 25(275). [CrossRef]
- Ji, R., Geng, Y., & Quan, X. (2024). Inferring gene regulatory networks with graph convolutional network based on causal feature reconstruction. Scientific Reports, 14(21342). [CrossRef]
- Khemani, B., Patil, S., Kotecha, K., & Tanwar, S. (2024). A review of graph neural networks: concepts, architectures, techniques, challenges, datasets, applications, and future directions. Journal of Big Data, 11(18). [CrossRef]
- Landry, L. G., Ali, N., Williams, D. R., Rehm, H. L., & Bonham, V. L. (2017). Lack Of Diversity In Genomic Databases Is A Barrier To Translating Precision Medicine Research Into Practice. Health Affairs. [CrossRef]
- Lundberg, S. M., & Lee, S.-I. (2017). A unified approach to interpreting model predictions. NIPS'17: Proceedings of the 31st International Conference on Neural Information Processing Systems (pp. 4468-4777). Red Hook, NY: Curran Associates, Inc.
- McInnes, L., Healy, J., Saul, N., & Grossberger, L. (2018). UMAP: Uniform Manifold Approximation and Projection. Journal of Open Source Software, 3(29). [CrossRef]
- Mesbah, R., Koenders, M. A., Spiijker, A. T., de Leeuw, M., van Hemert, A. M., & Giltray, E. J. (2023). Dynamic time warp analysis of individual symptom trajectories in individuals with bipolar disorder. Bipolar Disord. [CrossRef]
- Methi, A., Islam, M., Kaurani, L., Sakib, M. S., Kruger, D. M., Pena, T., . . . Fischer, A. (2024). A Single-Cell Transcriptomic Analysis of the Mouse Hippocampus After Voluntary Exercise. Mol. Neurobiol., 61(8), 5628-5645. [CrossRef]
- Muller, M. (2007). Information Retrieval for Music and Motio. Springer, Berlin, Heidelberg. [CrossRef]
- Obi-Nagata, K., Temma, Y., & Hayashi-Takagi, A. (2019). Synaptic functions and their disruption in schizophrenia: From clinical evidence to synaptic optogenetics in an animal model. Proceedings of the Japan Academy, Series B Physical and Biological Sciences, (pp. 179-197). [CrossRef]
- Ouwerkerk, J., Feleus, S., van der Zwaan, K. F., Li, Y., Roos, M., van Roon-Mom, W. M., . . . Mina, E. (2023). Machine learning in Huntington’s disease: exploring the Enroll-HD dataset for prognosis and driving capability prediction. Orphanet Journal of Rare Diseases, 18(218). [CrossRef]
- Qin, L., Zhou, Q., Sun, Y., Pang, X., Chen, Z., & Zheng, J. (2024). Dynamic functional connectivity and gene expression correlates in temporal lobe epilepsy: insights from hidden markov models. Journal of Translational Medicine, 22(736). [CrossRef]
- Ribiero, M. T., Singh, S., & Guestrin, C. (2016). "Why Should I Trust You?": Explaining the Predictions of Any Classifier. KDD '16: Proceedings of the 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining. Association for Computing MachineryNew YorkNYUnited States. [CrossRef]
- Rodriguez, S., Hug, C., Todorov, P., Moret, N., Boswell, S. A., Evans, K., . . . Sokolov, A. (2021). Machine learning identifies candidates for drug repurposing in Alzheimer’s disease. Nature Communications, 12(1033). [CrossRef]
- Rood, J. E., Wynne, S., Robson, L., Hupalowska, A., Randell, J., Teichmann, S. A., & Regev, A. (2025). The Human Cell Atlas from a cell census to a unified foundation model. Nature, 637, 1065-1071. [CrossRef]
- Rozenblatt-Rosen, O., Stubbington, M. J., Regev, A., & Teichmann, S. A. (2017). The Human Cell Atlas: from vision to reality. Nature, 550, 451-453. [CrossRef]
- Sabhawal, A., Grover, G., Kaushik, S., & Unni, S. K. (2019). Modelling and forecasting Positive and Negative Syndrome Scale scores to achieve remission using time series analysis. International Journal Of Methods in Psychiatric Research, 28(1). [CrossRef]
- Satterlee, J. S., Chadwich, L. H., Tysin, F. S., McAllister, K., Beaver, J., Birnbaum, L., . . . Roy, A. L. (2019). The NIH Common Fund/Roadmap Epigenomics Program: Successes of a comprehensive consortium. Science, 5(7). [CrossRef]
- Strauss, G. P., Esfahlani, F. Z., Raugh, I. M., Luther, L., & Sayama, H. (2023). Markov chain analysis indicates that positive and negative emotions have abnormal temporal interactions during daily life in schizophrenia. Journal of Psychiatric Research. [CrossRef]
- Sunil, G., Gowtham, S., Bose, A., Harish, S., & Srinivasa, G. (2024). Graph neural network and machine learning analysis of functional neuroimaging for understanding schizophrenia. 25(2). [CrossRef]
- The Tabula Muris Consortium. (2020). A single-cell transcriptomic atlas characterizes ageing tissues in the mouse. Nature, 583, 590-595. [CrossRef]
- The Tabula Sapiens Consortium. (2022). The Tabula Sapiens: A multiple-organ, single-cell transcriptomic atlas of humans. Science, 376(eabl4896). [CrossRef]
- Vries, S. E., Siegle, J. H., & Koch, C. (2023). Sharing neurophysiology data from the Allen Brain Observatory. eLife, 12(e85550). [CrossRef]
- Wang, J., Ma, A., Chang, Y., Gong, J., Jiang, Y., Qi, R., . . . Xu, D. (2021). scGNN is a novel graph neural network framework for single-cell RNA-Seq analyses. Nature Communications, 12(1882). [CrossRef]
- Yimin, F., Li, Y., Zhong, Y., Hong, L., Li, L., & Li, Y. (2024). Learning meaningful representation of single-neuron morphology via large-scale pre-training. Bioinformatics, 40(2), ii128-ii136. [CrossRef]
- Yu, Y., Zhang, N., Mai, Y., Ren, L., Chen, Q., Cao, Z., . . . Zheng, Y. (2023). Correcting batch effects in large-scale multiomics studies using a reference-material-based ratio method. Genome Biology, 24. [CrossRef]
- Zhang, L., Yang, Q., Yuan, R., Li, M., Lv, M., Zhang, L., . . . Chen, X. (2023). Single-nucleus transcriptomic mapping of blast-induced traumatic brain injury in mice hippocampus. Scientific Data, 10(638). [CrossRef]
- Zhang, X., Yang, L., Lu, J., Yuan, Y., Li, D., Zhang, H., . . . Wang, B. (2024). Reconfiguration of brain network dynamics in bipolar disorder: a hidden Markov model approach. Translational Psychiatry, 14(507). [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




