1. Introduction
Congenital disorders of glycosylation (CDG) are a group of rare, multisystemic genetic conditions caused by defects in glycosylation pathways. With over 160 different CDG types currently identified, these disorders exhibit broad clinical and genetic heterogeneity [
1]. Clinical glycomics refers to the application of glycan analysis to investigate disease-related glycosylation changes for diagnostic, prognostic, and therapeutic purposes. It translates complex glycan profiles into clinically actionable insights, particularly in disorders such as CDG where glycosylation defects are central to disease pathophysiology. By utilizing advanced mass spectrometry (MS) technologies, clinical glycomics delivers deep insights into patient-specific glycan profiles, enabling earlier diagnosis and more targeted therapeutic strategies. Two major analytical focuses include N-glycan profiling, which can be performed globally through total serum or plasma N-glycan analysis, or more specifically by analyzing transferrin glycosylation, a well-established marker for the majority of CDG types, particularly those affecting the N-glycosylation pathway. In contrast, O-glycan analysis, often targeting apolipoprotein C-III (ApoCIII), provides diagnostic information for mucin-type glycosylation defects, such as those observed in subtypes like COG6-CDG and B4GALT1-CDG, where abnormalities in O-glycosylation have been reported [
2].
In recent years, whole exome sequencing (WES) has emerged as a valuable tool for diagnosing CDG, especially in patients with unclear or atypical clinical presentations. It has successfully identified pathogenic mutations in genes associated with glycosylation and helped resolve previously undiagnosed cases [
3,
4,
5]. However, while WES provides detailed genetic information, it does not offer insight into the functional consequences of these mutations at the glycan level. For instance, in PGM1-CDG, glycomics revealed abnormal transferrin glycoforms and total plasma N-glycan profiles, which confirmed pathogenicity and guided therapeutic decisions, and offered additional insight into structural defects in glycosylation. It also helped speed up the diagnosis. Therefore, biochemical characterization through glycomics and glycoproteomics remains crucial for confirming pathogenicity and for understanding structural defects in glycosylation [
6].
The clinical heterogeneity of CDG requires a comprehensive diagnostic strategy. Biochemical screening using glycomics has been shown to expedite diagnosis and improve prognosis, as demonstrated in PGM1-CDG using intact transferrin and total plasma glycoprofiling [
6]. In parallel, advances in glycoproteomics, particularly in site-specific glycosylation profiling, have linked specific genetic mutations to altered glycan structures and associated phenotypes [
7]. Furthermore, consensus guidelines, such as those established for phosphoglucomutase 1 deficiency (PGM1 CDG), have supported the harmonization of diagnostic and therapeutic approaches, thereby enhancing consistency in patient management [
8].
The integration of glycomics with other omics technologies, such as genomics, transcriptomics, and metabolomics, has expanded our understanding of CDG pathophysiology. This review provides an updated overview of key advancements in clinical glycomics and glycoproteomics, with emphasis on emerging biomarkers, multi-omics integration, and diagnostic strategies that are shaping the future of clinical practice in CDG.
2. Technological Advancements
Over the past decade, continuous innovations in MS technologies have significantly advanced the diagnostic capabilities for CDG, particularly in resolving complex glycan structures. High-sensitivity methods such as porous graphitized carbon (PGC) liquid chromatography (LC) MS (PGC-LC-MS) and quadrupole time-of-flight (QTOF) MS (QTOF-MS) have provided researchers and clinicians with enhanced capabilities to perform detailed glycan and glycoprotein profiling, including the structural resolution of isomeric forms [
9,
10]. In particular, the adoption of soft ionization methods, such as electrospray ionization (ESI), for intact transferrin glycoprofiling has improved structural resolution and diagnostic accuracy [
11,
12]. A targeted LC tandem mass spectrometry (MS/MS) (LC-MS/MS) approach using dried blood spots (DBS) has also been developed for intact transferrin glycoform analysis, offering a high-throughput and clinically robust alternative to traditional isoelectric focusing methods for CDG diagnosis and newborn screening [
13].
Automation with high throughput workflows has streamlined glycomic diagnostics by reducing manual processing time and improving reproducibility. This aligns with recent technical standards established by the American College of Medical Genetics and Genomics (ACMG), which emphasize the implementation of validated workflows for transferrin glycoform analysis and broader biochemical testing in CDG [
14]. Advances in sample preparation, such as solid-phase extraction, glycan derivatization, and glycan labeling, have further increased detection sensitivity and reliability. These methods combined with high-throughput analytical platforms have enabled large-scale glycomic studies in clinical and research settings and improved both reproducibility and diagnostic utility in glycosylation disorders [
15]. Matrix assisted laser desorption ionization time of flight MS (MALDI TOF MS) remains a valuable tool for rapid screening, particularly useful in resource-limited settings or for high-throughput population studies due to its minimal sample preparation and cost efficiency [
16].
Significant strides have also been made in the analysis of O-glycans. For example, PGC-nanoLC-MS now enables high-resolution profiling of O-glycan structures without derivatization, eliminating the need for labor-intensive chemical labeling and simplifying analysis [
17]. ApoCIII MS profiling remains a practical diagnostic marker for mucin-type O-glycosylation defects, such as those observed in COG6-CDG, while modern LC-MS/MS platforms facilitate concurrent quantification of multiple apolipoproteins, including ApoCIII and others relevant to glycosylation status, supporting broader clinical biomarker panels [
18,
19].
3. Integration with Multi-Omics
Recent advances have highlighted the value of integrating glycomics and glycoproteomics with other omics platforms, including transcriptomics, proteomics, and metabolomics, to enhance our understanding of CDG mechanisms and improve clinical interpretation. These integrative approaches have been applied in disease models to investigate glycosylation remodeling at the systems level, revealing unique glyco-transcriptomic signatures, protein glycosylation changes, and metabolic shifts associated with disease progression and therapy response. Such studies have identified novel glycoproteins and glycan alterations that contribute to cellular dysfunction and resistance mechanisms, offering insights that may be translatable to CDG research and biomarker development [
20,
21].
Although historically less emphasized, metabolomics is now increasingly recognized as a complementary tool in CDG research. Targeted metabolomic profiling of patient samples has been applied to map disease-specific metabolic signatures, such as in PGM1-CDG and PMM2-CDG, offering insight into underlying biochemical disturbances [
22,
23]. These strategies support variant interpretation and therapy monitoring. Isotopic tracing of nucleotide sugar metabolism has also emerged as a useful method to study glycosylation pathways by tracking the biosynthesis and utilization of sugar nucleotides in cellular systems, with potential application to CDG [
24]. In addition, metabolomics has revealed broader systemic effects in CDG and can be particularly informative in cases with combined metabolic and glycosylation defects. A recent study demonstrated complex metabolomic signatures in a patient with both PGM1-CDG and mitochondrial dysfunction, reflecting altered glycan synthesis and energy metabolism [
25]. When integrated with glycoproteomics, metabolomic data enhances interpretation of biochemical phenotypes and support variant classification [
26,
27].
Systems biology approaches integrating glycomics with transcriptomics, proteomics, and metabolomics offer a powerful means to reconstruct disease pathways and guide individualized therapy [
28]. For example, transcriptomic data combined with N-glycomics has been used to predict glycan biosynthesis and tissue-specific expression patterns, offering a basis for structure-informed diagnostics [
29]. This has been demonstrated in studies of ALG1-CDG and PGM1-CDG. These strategies can be applied to CDG to identify affected biosynthetic pathways, classify overlapping phenotypes, and support precision diagnostics, particularly where genetic findings are inconclusive or phenotypic overlap complicates diagnosis.
Glycoproteomics provides site-specific glycosylation data that complements metabolomics by enabling deeper interpretation of disrupted pathways and supporting therapy monitoring. This integration strengthens personalized diagnostics, particularly in CDG patients receiving dietary or enzyme replacement therapies, where glycan remodeling is treatment responsive [
30]. Recent studies have extended the application of glycoproteomics to both patient-derived cells and clinical biofluids. In ALG1-CDG fibroblasts, glycoproteomic profiling has revealed a dysregulated glycoproteome that supports both diagnostic refinement and insight into disease mechanisms at the cellular level [
31]. Similarly, in SRD5A3-CDG fibroblasts, combined glycoproteomic and proteomic analysis revealed distinct N-glycosylation defects that affect protein function, localization, and stability [
32]. Glycoproteomic profiling of cerebrospinal fluid has uncovered brain-specific glycosylation changes that may help explain the neurological features observed in certain CDG subtypes and offer potential as fluid-based biomarkers to monitor neurological involvement in CDG [
33].
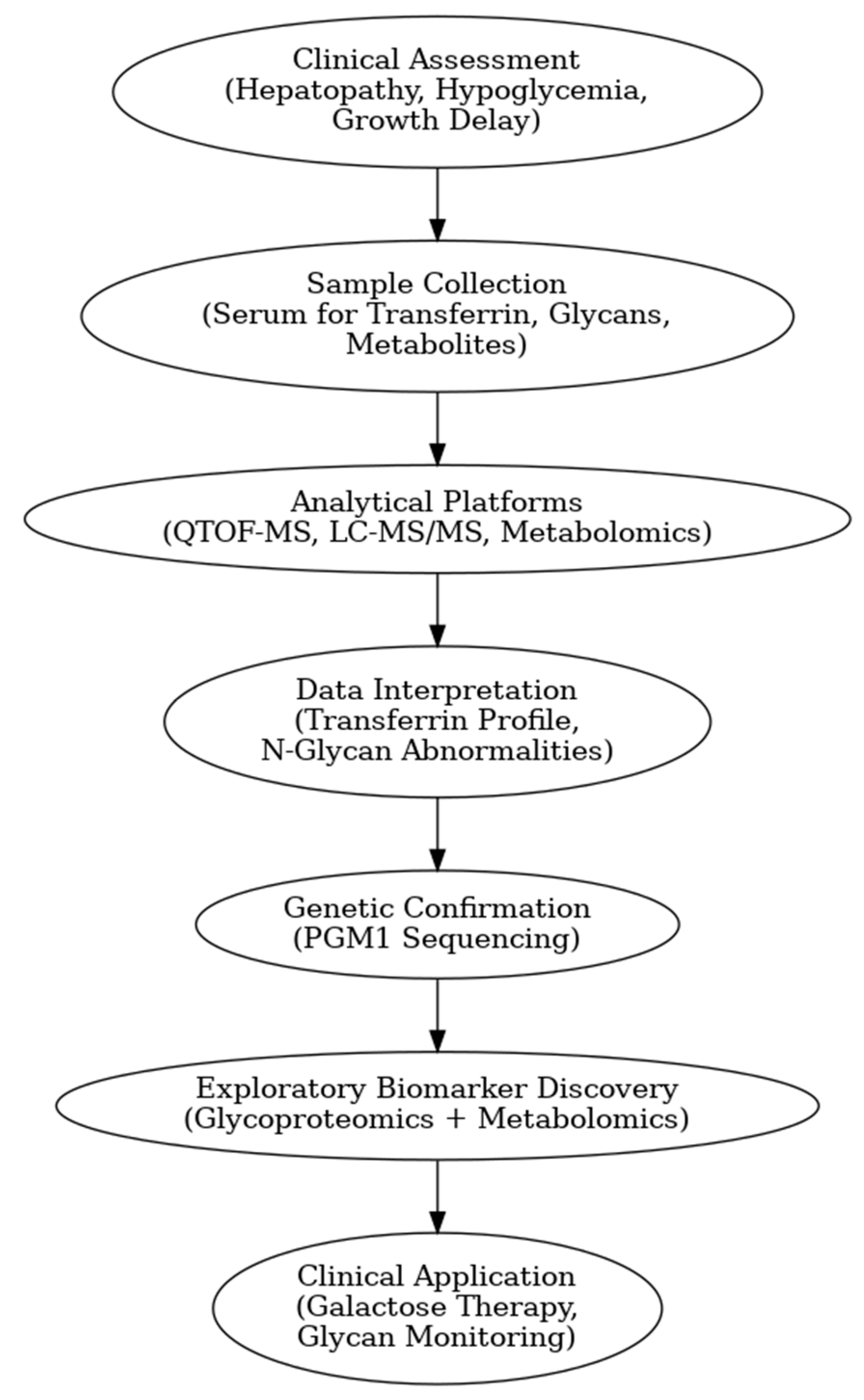
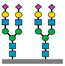
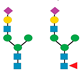
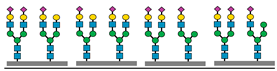
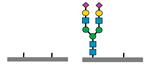
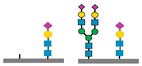
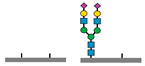
Figure 1 presents a clinical and exploration multi-omics workflow for PGM1-CDG, showing how glycomics, glycoproteomics, and metabolomics contribute from diagnosis to biomarker discovery. This case highlights how integrative omics approaches applied to well characterized subtypes like PGM1 CDG can bridge clinical evaluation with exploratory biomarker discovery and therapy monitoring. This framework captures both current diagnostic practice and the future potential of research-driven biomarker development in CDG.
The figure illustrates an integrated clinical and research workflow for PGM1-CDG. The process begins with clinical assessment based on characteristic features such as hepatopathy, hypoglycemia, and growth delay. This is followed by sample collection for transferrin glycoprofiling and N-glycan analysis using platforms such as QTOF-MS and LC-MS/MS. Genetic confirmation through PGM1 sequencing establishes the molecular diagnosis. In parallel, research-level integration of glycoproteomics and metabolomics facilitates exploration of biomarker discovery. This comprehensive strategy not only enhances diagnostic precision and therapy monitoring but also paves the way for novel biomarker identification and individualized care in CDG.
4. Biomarker Discovery
The identification of SLC10A7 as a regulator of bone development and Golgi glycosylation demonstrates how integrating glycomics with genomics can reveal gene–phenotype relationships and elucidate disease mechanisms [
34]. Similarly, the use of single-molecule molecular inversion probes combined with glycomics has improved the classification of CDG type I subtypes, such as PMM2-CDG and ALG1-CDG. PMM2-CDG, being the most prevalent subtype, benefits from early glycomic markers for timely diagnosis. Furthermore, ALG1-CDG presents diagnostic challenges due to mild biochemical phenotypes and uncertain variant interpretation, where glycomic analysis helps resolve functional ambiguity [
35].
Table 1 summarizes glycomics-derived biomarkers (glycomarkers) reported in selected CDG subtypes, including PGM1-CDG, SLC10A7-CDG, PMM2-CDG, and ALG1-CDG. These glycomarkers have demonstrated diagnostic utility and potential application in therapy monitoring.
Quantitative glycomics has also enabled the discovery of subtype-specific markers, such as a diagnostic tetrasaccharide in MOGS-CDG, expanding the clinical spectrum and diagnostic reach [
36]. The identification of this glycomarker by high-throughput profiling not only aids early diagnosis but also broadens our understanding of glycosylation defects in CDG subtypes beyond the most common N-glycan profiles. This marker demonstrates how quantitative glycomics can uncover disease-specific glycan signatures that complement genetic and enzymatic testing in challenging diagnostic cases.
Recent studies have strengthened the position of glycoproteomics as a powerful platform for biomarker discovery in CDG. Plasma glycoproteomics has demonstrated high specificity in detecting site-specific glycosylation abnormalities that distinguish CDG from other metabolic disorders, supporting its utility for differential diagnosis and patient stratification [
37]. The ability of glycoproteomics to quantify site-specific glycosylation has been demonstrated in CDG patient cells, where unique glycan abnormalities were linked to underlying genetic defects [
31]. Moreover, recent work has identified a complement C4-derived glycopeptide as a diagnostic biomarker for PMM2-CDG, further establishing the clinical relevance of glycoproteomics in biomarker development [
38].
The profiling of isomeric glycan structures adds an additional layer of diagnostic resolution. For example, alpha 2,3-linked sialylated isomers have been associated with immune-mediated disorders such as Behcet’s disease, providing a model for how linkage-specific glycan analysis may benefit CDG diagnostics [
39]. Complementary to this, high-sensitivity capillary electrophoresis and hydrophilic interaction chromatography with ultra-performance liquid chromatography (HILIC-UPLC) MS platforms now enable detailed mapping and high-throughput separation of glycan isomers. The application of HILIC-UPLC-MS for analyzing isomeric N-glycans in CDG patient samples has confirmed its diagnostic utility in resolving complex glycan structures relevant to glycosylation disorders [
40]. In addition, LC-MS/MS has been used to profile isomeric N-glycans derived from low-abundance serum glycoproteins, demonstrating the sensitivity of this method in detecting subtle glycan variations that may support early disease detection or patient stratification [
41].
In addition to diagnosis, glycoproteomic biomarkers serve as tools for therapy monitoring. Longitudinal glycan profiling in patients with PGM1-CDG receiving galactose supplementation has shown measurable glycosylation changes correlating with clinical improvement [
42]. These markers function as surrogate endpoints and help guide treatment decisions over time. The adoption of high-throughput LC-MS/MS workflows allows for consistent and reproducible biomarker quantification across larger patient cohorts, bridging the gap between research discoveries and clinical implementation. Together, these advancements further establish glycoproteomics as a key platform in personalized diagnostics and therapeutic monitoring in CDG.
5. Standardization of Diagnostic Practices
International consensus guidelines have played a pivotal role in unifying diagnostic practices for CDG. For instance, the PGM1-CDG guideline emphasizes a multi-pronged approach that integrates clinical features, biochemical analyses, and genetic testing to achieve accurate diagnosis. It also provides detailed guidance on long-term follow-up, including monitoring of glycosylation markers, liver and endocrine function, and nutritional status. These recommendations help standardize patient care and support early therapeutic intervention, particularly in response to galactose supplementation [
8].
Among the most reliable tools is intact transferrin glycoprofiling using high-resolution MS, which supports both diagnosis and longitudinal therapy monitoring, as shown in phosphoglucomutase 1 deficiency [
6]. In addition, serum glycoprotein profiling allows the tracking of site-specific glycosylation changes after treatment, offering insights into how therapies influence glycan processing and trafficking. A transferrin-based treatment index has also been proposed to evaluate response to D-galactose supplementation in PGM1-CDG, showing how glycosylation changes can serve as surrogate markers of therapeutic efficacy [
42].
To promote diagnostic accessibility, especially in resource-limited settings, non-invasive sampling methods such as DBS for transferrin and ApoCIII profiling are increasingly used. These methods allow stable storage and easy transport, enabling remote testing and earlier detection of glycosylation defects [
12]. As a result, they are now widely adopted in clinical workflows to support both diagnosis and therapy monitoring.
Recent technical standards by the ACMG have emphasized the need for validated laboratory protocols in CDG diagnostics. Such protocols cover quality-assured transferrin glycoform analysis, result interpretation, and standardized reporting aimed at harmonizing laboratory practices [
14]. As new therapies emerge, high-throughput workflows such as data-independent acquisition (DIA) MS offer scalable, reproducible solutions for longitudinal monitoring of glycosylation dynamics, supporting consistent and reproducible measurements across patient cohorts [
46].
Together, these developments highlight the close relationship between diagnostic standardization and therapeutic monitoring in CDG, underscoring the importance of integrating glycomics-based strategies into routine clinical care. Such integration is essential for ensuring diagnostic accuracy and equitable patient care worldwide.
6. Challenges in Clinical Translation
Despite significant progress, several challenges remain in translating glycomics and glycoproteomics into routine clinical diagnostics for CDG. A key barrier is limited access to advanced MS platforms and skilled personnel, including bioinformatics specialists, particularly in low-resource settings. High costs associated with instrumentation, reagents, and bioinformatics infrastructure hinder widespread clinical adoption.
Another major issue is the lack of standardized protocols across laboratories. Differences in sample handling, glycan derivatization, data acquisition, and interpretation often result in inconsistent findings and reduced reproducibility. As emphasized in a recent study, harmonizing workflows and developing certified reference materials, such as standardized transferrin glycoform controls are critical for transforming glycomics from a research-intensive approach to a validated clinical diagnostic platform [
26].
Informatics integration presents additional complexity. Clinical glycomics generates high-dimensional datasets that require advanced analytical pipelines for accurate interpretation due to glycan isomerism, structural branching, and linkage variability. The integration of glycomics with genomic, proteomic, and clinical metadata is essential but remains technically demanding. A perspective by Van der Burgt and Wuhrer emphasized the need for interoperable data standards, shared ontologies, and cross-disciplinary collaboration to effectively incorporate glycoproteomic data into precision medicine frameworks [
30].
Moreover, translating multi-omics tools into clinical diagnostics requires thoughtful validation and clinical-grade implementation. A review by Hertzog and colleagues emphasized that integrating emerging omics platforms, including metabolomics and glycoproteomics, into clinical practice for inborn errors of metabolism requires not only technical readiness but also support across regulatory, educational, and infrastructural domains. Their review underscores the importance of multidisciplinary cooperation to enable clinical laboratories to adopt advanced -omics tools for diagnostic and therapeutic purposes [
47].
Collectively, these challenges highlight the importance of coordinated efforts among researchers, clinicians, and policy makers to ensure that the benefits of glycomics and glycoproteomics reach patients through validated, scalable, and equitable clinical applications.
7. Role of Glycan Databases and Bioinformatics
The interpretation of complex glycomic and glycoproteomic data in CDG heavily depends on specialized bioinformatics platforms and curated databases. These tools support accurate glycan annotation, data integration, and structure-function correlation, thereby enhancing both biomarker discovery and clinical diagnostics.
UniCarbKB serves as a central glycoproteomics knowledge platform, consolidating experimentally determined glycan structures and glycoprotein data, and enabling consistent annotation and searchability across studies [
48]. GlyConnect, with its interactive analytical interface, facilitates the exploration of glycoprotein–glycan relationships, helping researchers map glycosylation patterns to disease phenotypes [
49]. Meanwhile, GlyTouCan functions as a global glycan structure repository that assigns unique identifiers to glycan compositions, promoting data interoperability and international collaboration [
50].
In addition, platforms such as GlycoWorkbench provide computational tools for MS-based annotation and structural elucidation of glycans, streamlining data interpretation in clinical and research laboratories [
51]. Collectively, these platforms enhance diagnostic precision and increase analytical throughput by offering curated resources and automated pipelines for structure annotation, pathway mapping, and visualization.
From a broader perspective, the field of glycoinformatics has matured into a core component of glycoscience, integrating data from genomics, proteomics, and clinical phenotypes. The application of glyco-bioinformatics is crucial to support functional interpretation of glycan-related changes and enable discovery-driven diagnostics. Recent perspectives highlight the importance of data standardization, computational interoperability, and multi-omics integration in advancing clinical glycoscience [
52].
Furthermore, as noted by Packer and colleagues, the development and application of bioinformatics in glycomics have opened new avenues for biomarker discovery by linking structural glycan data with disease mechanisms [
53]. These informatics tools not only support diagnosis of known CDG subtypes but also for discovering novel variants. By mapping observed glycan abnormalities to biosynthetic pathways, researchers can prioritize candidate genes and assess the pathogenic relevance of uncertain variants.
Together, these databases and computational tools form the bioinformatics backbone of clinical glycomics, providing essential infrastructure to interpret complex datasets, guide diagnostics, and advance biomarker discovery in CDG. As tools and databases continue to evolve, their integration with clinical decision support systems may further streamline diagnostics in rare glycosylation disorders.
8. Future Directions and Perspectives
The field of clinical glycomics and glycoproteomics in CDG is entering a new era marked by rapid technological growth, computational advancements, and a more integrative understanding of disease biology. Artificial intelligence (AI) and machine learning (ML) are beginning to play a prominent role by enabling the analysis of high-dimensional glycomic datasets. These tools can identify subtle and complex glycosylation patterns that are characteristic of specific disease states. Such capability could greatly improve the sensitivity and specificity of biomarker discovery, especially for ultra-rare CDG subtypes that may not be easily identified by conventional methods. Glycoinformatics is evolving into a powerful data science field, offering significant contributions to diagnostics, precision medicine, and translational research in glycosylation disorders [
54,
55].
Next-generation MS platforms such as DIA are anticipated to substantially improve the resolution, sensitivity, and throughput of glycoproteomic analyses. This approach enables the systematic capture of all ionized peptides within a sample, regardless of their abundance, thus providing more comprehensive and reproducible glycopeptide datasets. DIA-based workflows have already shown promise in clinical proteomics and are expected to enhance biomarker discovery and therapy monitoring in CDG by improving the quantification of site-specific glycosylation changes across larger patient cohorts [
46].
One promising direction is the expansion of glycomic and glycoproteomic profiling to various biological fluids and tissues. While plasma and serum remain the primary matrices, cerebrospinal fluid, urine, and tissue biopsies can provide insight into organ-specific glycosylation patterns. This is particularly valuable for CDG subtypes with dominant neurological, hepatic, or muscular symptoms, where localized biomarkers may guide precision therapy and improve understanding of disease pathophysiology.
Building international glycoprofile reference libraries and adopting harmonized annotation systems will be instrumental in facilitating global research collaboration and comparative studies. Standardized databases and consistent terminology will help overcome current barriers in data sharing, allowing researchers to better interpret glycomic profiles across populations and disease subtypes. This infrastructure is essential to support the clinical utility of glycomics and to promote its integration into routine diagnostic workflows.
Importantly, the integration of glycomics with genomics, transcriptomics, proteomics, and clinical phenotyping is paving the way for systems-level diagnostics. These systems-level approaches enhance diagnostic resolution in genetically unresolved cases by linking genetic variations to their biochemical and functional consequences. For example, transcriptomic data can be used to infer glycosylation capacity in specific tissues. When combined with N-glycomic profiles, these approaches can explain phenotypic variability and help uncover the functional consequences of genetic variants. This level of integration supports the identification of novel disease genes, clarifies the biochemical basis of disease, and enhances diagnostic yield, particularly in genetically unresolved cases.
A novel and emerging approach is the use of stable isotope-labeled sugars to study nucleotide sugar metabolism. This technique allows researchers to functionally assess biosynthetic pathways involved in glycosylation. In CDG, it can be used to pinpoint metabolic bottlenecks and evaluate the efficiency of therapeutic interventions. For example, isotopic tracing in pluripotent stem cell models has successfully mapped sugar nucleotide biosynthesis, offering valuable information about upstream defects in glycosylation pathways [
24].
The development of site-specific glycopeptide biomarkers is another area showing great promise. These markers provide detailed molecular information on glycosylation changes at specific protein sites and may be used to monitor disease activity or therapeutic response. For instance, complement C4-derived glycopeptides have been proposed as specific markers for PMM2-CDG, and transferrin glycopeptides are currently used to assess treatment outcomes in PGM1-CDG. Expanding this strategy to other CDG subtypes could lead to highly personalized monitoring tools for patient management [
38,
42].
To ensure broad clinical adoption, there is an urgent need for standardization across analytical protocols. This includes harmonizing sample preparation methods, MS acquisition parameters, and data processing workflows. The development of validated bioinformatics pipelines and interoperable databases is equally important for improving reproducibility and enabling inter-laboratory comparisons.
Looking ahead, as personalized medicine continues to evolve, clinical glycoproteomics is likely to become a fundamental tool in individualized patient care. Future clinical workflows may include newborn screening programs that utilize glycan-based biomarkers, patient-specific glycosylation profiles for therapeutic decision-making, and dynamic glycoproteomic indices to monitor treatment efficacy over time. These future applications align with the goals of precision medicine and early intervention. These advancements will help shift CDG management from a reactive model to a proactive, precision-guided approach. This evolution prioritizes early detection, targeted intervention, and improved quality of life for affected individuals.
9. Conclusions
Clinical glycomics and glycoproteomics have matured into indispensable tools for the diagnosis, subclassification, and monitoring of CDG. This review highlights how advancements in MS, informatics, and multi-omics integration have expanded the clinical utility of glycan-based biomarkers. Through the analysis of transferrin glycoforms, total plasma N-glycome, site-specific glycopeptides, and emerging biosynthetic and functional readouts, glycomics now offers a multifaceted approach to understanding and managing CDG.
While challenges in standardization, accessibility, and bioinformatics integration remain, the field is moving rapidly toward solutions supported by international guidelines, collaborative databases, and automated platforms. The use of DBS testing, transferrin-based treatment indices, and isotopic tracing exemplifies the translational impact of glycomics in both diagnosis and therapeutic monitoring.
Looking ahead, glycomics is poised to play a central role in precision medicine frameworks. From newborn screening to individualized therapy monitoring, future workflows will integrate glycoprofiling alongside genomic and clinical data to optimize patient care. Continued investment in harmonized standards, clinician education, and global research collaboration will be essential to realize the full diagnostic and therapeutic potential of glycomics and glycoproteomics in CDG.
In summary, the convergence of technological innovation, systems biology, and clinical translation marks a promising future for glycomics in rare disease diagnostics. The field is now approaching broader clinical adoption, which will transform how we diagnose, monitor, and treat CDG.
Author Contributions
Conceptualization, N.A.B.; methodology, N.A.B. and N.I.H; writing—original draft preparation, N.A.B.; writing—review and editing, N.A.B. and N.I.H; visualization, N.A.B; supervision, N.A.B. and N.I.H; project administration, N.A.B. and N.I.H; funding acquisition, N.I.H. All authors have read and agreed to the published version of the manuscript.
Funding
This research was funded by THE NATIONAL INSTITUTES OF HEALTH (NIH) MALAYSIA, under registration number NMRR ID-21-02426-WU9.
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
Not applicable.
Acknowledgments
The authors thank the Director General of Health Malaysia and the Director of the Institute of Medical Research (IMR), Malaysia, for permitting us to publish this article. During the preparation of this manuscript, the authors used ChatGPT (OpenAI, GPT-4,
https://chat.openai.com/, accessed on 1 June 2025) to assist with language editing and phrasing. The authors have reviewed and edited the AI-generated content and take full responsibility for the final version of the manuscript.
Conflicts of Interest
The authors declare no conflicts of interest.
Abbreviations
The following abbreviations are used in this manuscript:
| CDG |
congenital disorders of glycosylation |
| MS |
mass spectrometry |
| ApoCIII |
apolipoprotein C-III |
| WES |
whole exome sequencing |
| PGC |
porous graphitized carbon |
| LC |
liquid chromatography |
| QTOF |
quadrupole time-of-flight |
| ESI |
electrospray ionization |
| MS/MS |
tandem mass spectrometry |
| DBS |
dried blood spots |
| ACMG |
American College of Medical Genetics and Genomics |
| MALDI TOF |
matrix assisted laser desorption ionization time of flight |
| HILIC |
hydrophilic interaction chromatography |
| UPLC |
ultra-performance liquid chromatography |
| DIA |
data-independent acquisition |
References
- Francisco, R.; Brasil, S.; Poejo, J.; Jaeken, J.; Pascoal, C.; Videira, P.A.; dos Reis Ferreira, V. Congenital Disorders of Glycosylation (CDG): State of the Art in 2022. Orphanet J Rare Dis 2023, 18, 329. [Google Scholar] [CrossRef] [PubMed]
- Abu Bakar, N.; Lefeber, D.J.; van Scherpenzeel, M. Clinical Glycomics for the Diagnosis of Congenital Disorders of Glycosylation. J Inherit Metab Dis 2018, 41. [Google Scholar] [CrossRef] [PubMed]
- Papi, A.; Zamani, M.; Shariati, G.; Sedaghat, A.; Seifi, T.; Negahdari, S.; Sedighzadeh, S.S.; Zeighami, J.; Saberi, A.; Hamid, M.; et al. Whole Exome Sequencing Reveals Several Novel Variants in Congenital Disorders of Glycosylation and Glycogen Storage Diseases in Seven Patients from Iran. Mol Genet Genomic Med 2023, 11. [Google Scholar] [CrossRef]
- Zhang, Z.; Huang, T.L.; Ma, J.; He, W.J.; Gu, H. Clinical and Whole-Exome Sequencing Findings in Two Siblings from Hani Ethnic Minority with Congenital Glycosylation Disorders. BMC Med Genet 2019, 20. [Google Scholar] [CrossRef]
- Ng, B.G.; Freeze, H.H. Perspectives on Glycosylation and Its Congenital Disorders. Trends in Genetics 2018, 34. [Google Scholar] [CrossRef]
- Abu Bakar, N.; Voermans, N.C.; Marquardt, T.; Thiel, C.; Janssen, M.C.H.; Hansikova, H.; Crushell, E.; Sykut-Cegielska, J.; Bowling, F.; MØrkrid, L.; et al. Intact Transferrin and Total Plasma Glycoprofiling for Diagnosis and Therapy Monitoring in Phosphoglucomutase-I Deficiency. Translational Research 2018, 199. [Google Scholar] [CrossRef]
- He, K.; Baniasad, M.; Kwon, H.; Caval, T.; Xu, G.; Lebrilla, C.; Hommes, D.W.; Bertozzi, C. Decoding the Glycoproteome: A New Frontier for Biomarker Discovery in Cancer. J Hematol Oncol 2024, 17. [Google Scholar] [CrossRef]
- Altassan, R.; Radenkovic, S.; Edmondson, A.C.; Barone, R.; Brasil, S.; Cechova, A.; Coman, D.; Donoghue, S.; Falkenstein, K.; Ferreira, V.; et al. International Consensus Guidelines for Phosphoglucomutase 1 Deficiency (PGM1-CDG): Diagnosis, Follow-up, and Management. J Inherit Metab Dis 2021, 44. [Google Scholar] [CrossRef]
- Young, C.; Condina, M.R.; Briggs, M.T.; Moh, E.S.X.; Kaur, G.; Oehler, M.K.; Hoffmann, P. In-House Packed Porous Graphitic Carbon Columns for Liquid Chromatography-Mass Spectrometry Analysis of N-Glycans. Front Chem 2021, 9. [Google Scholar] [CrossRef]
- Chen, C.H.; Lin, Y.P.; Ren, C.T.; Shivatare, S.S.; Lin, N.H.; Wu, C.Y.; Chen, C.H.; Lin, J.L. Enhancement of Fucosylated N-Glycan Isomer Separation with an Ultrahigh Column Temperature in Porous Graphitic Carbon Liquid Chromatography-Mass Spectrometry. J Chromatogr A 2020, 1632. [Google Scholar] [CrossRef]
- Van Scherpenzeel, M.; Steenbergen, G.; Morava, E.; Wevers, R.A.; Lefeber, D.J. High-Resolution Mass Spectrometry Glycoprofiling of Intact Transferrin for Diagnosis and Subtype Identification in the Congenital Disorders of Glycosylation. Translational Research 2015, 166. [Google Scholar] [CrossRef] [PubMed]
- Wada, Y.; Kadoya, M.; Okamoto, N. Mass Spectrometry of Transferrin and Apolipoprotein CIII from Dried Blood Spots for Congenital Disorders of Glycosylation. Mass Spectrometry 2022, 11. [Google Scholar] [CrossRef] [PubMed]
- Casetta, B.; Malvagia, S.; Funghini, S.; Martinelli, D.; Dionisi-Vici, C.; Barone, R.; Fiumara, A.; Donati, M.A.; Guerrini, R.; La Marca, G. A New Strategy Implementing Mass Spectrometry in the Diagnosis of Congenital Disorders of N-Glycosylation (CDG). Clin Chem Lab Med 2020, 59. [Google Scholar] [CrossRef] [PubMed]
- Hall, P.L.; Lam, C.; Wolfe, L.; Edmondson, A. ; ACMG Laboratory Quality Assurance Committee Biochemical Testing for Congenital Disorders of Glycosylation: A Technical Standard of the American College of Medical Genetics and Genomics (ACMG). Genetics in Medicine 2025, 27, 101328. [Google Scholar] [CrossRef]
- Trbojević-Akmačić, I.; Lageveen-Kammeijer, G.S.M.; Heijs, B.; Petrović, T.; Deriš, H.; Wuhrer, M.; Lauc, G. High-Throughput Glycomic Methods. Chem Rev 2022, 122. [Google Scholar] [CrossRef]
- Wada, Y.; Okamoto, N. Apolipoprotein C-III O-Glycoform Profiling of 500 Serum Samples by Matrix-Assisted Laser Desorption/Ionization Mass Spectrometry for Diagnosis of Congenital Disorders of Glycosylation. Journal of Mass Spectrometry 2021, 56. [Google Scholar] [CrossRef]
- Zhang, T.; Wang, W.; Wuhrer, M.; de Haan, N. Comprehensive O-Glycan Analysis by Porous Graphitized Carbon Nanoliquid Chromatography-Mass Spectrometry. Anal Chem 2024, 96, 8942–8948. [Google Scholar] [CrossRef]
- Wada, Y.; Okamoto, N. Electrospray Ionization Mass Spectrometry of Apolipoprotein CIII to Evaluate O-Glycan Site Occupancy and Sialylation in Congenital Disorders of Glycosylation. Mass Spectrometry 2022, 11. [Google Scholar] [CrossRef]
- Kim, H.; Yang, W.S.; An, D.; Lee, S.G.; Baek, J.H. Fast and Straightforward Simultaneous Quantification of Multiple Apolipoproteins in Human Serum on a High-Throughput LC-MS/MS Platform. Proteomics Clin Appl 2023, 17. [Google Scholar] [CrossRef]
- Cao, L.; Zhou, Y.; Li, X.; Lin, S.; Tan, Z.; Guan, F. Integrating Transcriptomics, Proteomics, Glycomics and Glycoproteomics to Characterize Paclitaxel Resistance in Breast Cancer Cells. J Proteomics 2021, 243. [Google Scholar] [CrossRef]
- Cui, D.; Li, W.; Jiang, D.; Wu, J.; Xie, J.; Wu, Y. Advances in Multi-Omics Applications in HBV-Associated Hepatocellular Carcinoma. Front Med (Lausanne) 2021, 8. [Google Scholar] [CrossRef] [PubMed]
- Radenkovic, S.; Bird, M.J.; Emmerzaal, T.L.; Wong, S.Y.; Felgueira, C.; Stiers, K.M.; Sabbagh, L.; Himmelreich, N.; Poschet, G.; Windmolders, P.; et al. The Metabolic Map into the Pathomechanism and Treatment of PGM1-CDG. Am J Hum Genet 2019, 104. [Google Scholar] [CrossRef] [PubMed]
- Mangione, R.; Cirnigliaro, L.; Saab, M.W.; Pettinato, F.; Barbato, A.; Distefano, A.; Spina, E. La; Lazzarino, G.; Volti, G.L.; Longhitano, L.; et al. Targeted Metabolomic Evaluation of Peripheral Blood Mononucleated Cells from Patients with PMM2-CDG. Sci Rep 2025, 15, 15929. [Google Scholar] [CrossRef]
- Conte, F.; Noga, M.J.; van Scherpenzeel, M.; Veizaj, R.; Scharn, R.; Sam, J. El; Palumbo, C.; van den Brandt, F.C.A.; Freund, C.; Soares, E.; et al. Isotopic Tracing of Nucleotide Sugar Metabolism in Human Pluripotent Stem Cells. Cells 2023, 12. [Google Scholar] [CrossRef]
- Radenkovic, S.; Adant, I.; Bird, M.J.; Swinnen, J.V.; Cassiman, D.; Kozicz, T.; Gruenert, S.C.; Ghesquière, B.; Morava, E. Complex Metabolomic Changes in a Combined Defect of Glycosylation and Oxidative Phosphorylation in a Patient with Pathogenic Variants in PGM1 and NDUFA13. Cells 2025, 14, 638. [Google Scholar] [CrossRef]
- Ferreira, E.A.; Veenvliet, A.R.J.; Engelke, U.F.H.; Kluijtmans, L.A.J.; Huigen, M.C.D.G.; Hoegen, B.; de Boer, L.; de Vries, M.C.; van Bon, B.W.; Leenders, E.; et al. Diagnosing, Discarding, or de-VUSsing: A Practical Guide to (Un)Targeted Metabolomics as Variant-Transcending Functional Tests. Genetics in Medicine 2023, 25. [Google Scholar] [CrossRef]
- Lin, C.; Tian, Q.; Guo, S.; Xie, D.; Cai, Y.; Wang, Z.; Chu, H.; Qiu, S.; Tang, S.; Zhang, A. Metabolomics for Clinical Biomarker Discovery and Therapeutic Target Identification. Molecules 2024, 29, 2198. [Google Scholar] [CrossRef]
- Gutierrez Reyes, C.D.; Alejo-Jacuinde, G.; Perez Sanchez, B.; Chavez Reyes, J.; Onigbinde, S.; Mogut, D.; Hernández-Jasso, I.; Calderón-Vallejo, D.; Quintanar, J.L.; Mechref, Y. Multi Omics Applications in Biological Systems. Curr Issues Mol Biol 2024, 46, 5777–5793. [Google Scholar] [CrossRef]
- Alvarez, M.R.S.; Holmes, X.A.; Oloumi, A.; Grijaldo-Alvarez, S.J.; Schindler, R.; Zhou, Q.; Yadlapati, A.; Silsirivanit, A.; Lebrilla, C.B. Integration of RNAseq Transcriptomics and N -Glycomics Reveal Biosynthetic Pathways and Predict Structure-Specific N -Glycan Expression. Chem Sci 2025, 16, 7155–7172. [Google Scholar] [CrossRef]
- Van Der Burgt, Y.; Wuhrer, M. The Role of Clinical Glyco(Proteo)Mics in Precision Medicine. Molecular and Cellular Proteomics 2023, 22. [Google Scholar] [CrossRef]
- Budhraja, R.; Joshi, N.; Radenkovic, S.; Kozicz, T.; Morava, E.; Pandey, A. Dysregulated Proteome and N-Glycoproteome in ALG1-Deficient Fibroblasts. Proteomics 2024, 24. [Google Scholar] [CrossRef] [PubMed]
- Garapati, K.; Ranatunga, W.; Joshi, N.; Budhraja, R.; Sabu, S.; Kantautas, K.A.; Preston, G.; Perlstein, E.O.; Kozicz, T.; Morava, E.; et al. N-Glycoproteomic and Proteomic Alterations in SRD5A3-Deficient Fibroblasts. Glycobiology 2024, 34. [Google Scholar] [CrossRef] [PubMed]
- Baerenfaenger, M.; Post, M.A.; Langerhorst, P.; Huijben, K.; Zijlstra, F.; Jacobs, J.F.M.; Verbeek, M.M.; Wessels, H.J.C.T.; Lefeber, D.J. Glycoproteomics in Cerebrospinal Fluid Reveals Brain-Specific Glycosylation Changes. Int J Mol Sci 2023, 24. [Google Scholar] [CrossRef] [PubMed]
- Ashikov, A.; Bakar, N.A.; Wen, X.Y.; Niemeijer, M.; Osorio, G.R.P.; Brand-Arzamendi, K.; Hasadsri, L.; Hansikova, H.; Raymond, K.; Vicogne, D.; et al. Integrating Glycomics and Genomics Uncovers SLC10A7 as Essential Factor for Bone Mineralization by Regulating Post-Golgi Protein Transport and Glycosylation. Hum Mol Genet 2018, 27. [Google Scholar] [CrossRef]
- Abu Bakar, N.; Ashikov, A.; Brum, J.M.; Smeets, R.; Kersten, M.; Huijben, K.; Keng, W.T.; Speck-Martins, C.E.; de Carvalho, D.R.; de Rizzo, I.M.P.O.; et al. Synergistic Use of Glycomics and Single-Molecule Molecular Inversion Probes for Identification of Congenital Disorders of Glycosylation Type-1. J Inherit Metab Dis 2022, 45. [Google Scholar] [CrossRef]
- Post, M.A.; de Wit, I.; Zijlstra, F.S.M.; Engelke, U.F.H.; van Rooij, A.; Christodoulou, J.; Tan, T.Y.; Le Fevre, A.; Jin, D.; Yaplito-Lee, J.; et al. MOGS-CDG: Quantitative Analysis of the Diagnostic Glc3Man Tetrasaccharide and Clinical Spectrum of Six New Cases. J Inherit Metab Dis 2023, 46. [Google Scholar] [CrossRef]
- Wessels, H.J.C.T.; Kulkarni, P.; van Dael, M.; Suppers, A.; Willems, E.; Zijlstra, F.; Kragt, E.; Gloerich, J.; Schmit, P.-O.; Pengelley, S.; et al. Plasma Glycoproteomics Delivers High-Specificity Disease Biomarkers by Detecting Site-Specific Glycosylation Abnormalities. J Adv Res 2024, 61, 179–192. [Google Scholar] [CrossRef]
- Garapati, K.; Budhraja, R.; Saraswat, M.; Kim, J.; Joshi, N.; Sachdeva, G.S.; Jain, A.; Ligezka, A.N.; Radenkovic, S.; Ramarajan, M.G.; et al. A Complement C4–Derived Glycopeptide Is a Biomarker for PMM2-CDG. JCI Insight 2024, 9. [Google Scholar] [CrossRef]
- Seo, N.; Lee, H.; Oh, M.J.; Kim, G.H.; Lee, S.G.; Ahn, J.K.; Cha, H.S.; Kim, K.H.; Kim, J.; An, H.J. Isomer-Specific Monitoring of Sialylated N-Glycans Reveals Association of A2,3-Linked Sialic Acid Epitope With Behcet’s Disease. Front Mol Biosci 2021, 8. [Google Scholar] [CrossRef]
- Messina, A.; Palmigiano, A.; Esposito, F.; Fiumara, A.; Bordugo, A.; Barone, R.; Sturiale, L.; Jaeken, J.; Garozzo, D. HILIC-UPLC-MS for High Throughput and Isomeric N-Glycan Separation and Characterization in Congenital Disorders Glycosylation and Human Diseases. Glycoconj J 2021, 38. [Google Scholar] [CrossRef]
- Gutierrez Reyes, C.D.; Hakim, M.A.; Atashi, M.; Goli, M.; Gautam, S.; Wang, J.; Bennett, A.I.; Zhu, J.; Lubman, D.M.; Mechref, Y. LC-MS/MS Isomeric Profiling of N-Glycans Derived from Low-Abundant Serum Glycoproteins in Mild Cognitive Impairment Patients. Biomolecules 2022, 12. [Google Scholar] [CrossRef]
- Perales-Clemente, E.; Liedtke, K.; Studinski, A.; Radenkovic, S.; Gavrilov, D.; Oglesbee, D.; Matern, D.; Rinaldo, P.; Tortorelli, S.; Morava, E.; et al. A New D-Galactose Treatment Monitoring Index for PGM1-CDG. J Inherit Metab Dis 2021, 44. [Google Scholar] [CrossRef] [PubMed]
- Conte, F.; Morava, E.; Bakar, N.A.; Wortmann, S.B.; Poerink, A.J.; Grunewald, S.; Crushell, E.; Al-Gazali, L.; de Vries, M.C.; Mørkrid, L.; et al. Phosphoglucomutase-1 Deficiency: Early Presentation, Metabolic Management and Detection in Neonatal Blood Spots. Mol Genet Metab 2020, 131. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.; James, P.M.; Ng, B.G.; Li, X.; Xia, B.; Rong, J.; Asif, G.; Raymond, K.; Jones, M.A.; Hegde, M.; et al. A Novel N-Tetrasaccharide in Patients with Congenital Disorders of Glycosylation, Including Asparagine-Linked Glycosylation Protein 1, Phosphomannomutase 2, and Mannose Phosphate Isomerase Deficiencies. Clin Chem 2016, 62. [Google Scholar] [CrossRef]
- Bengtson, P.; Ng, B.G.; Jaeken, J.; Matthijs, G.; Freeze, H.H.; Eklund, E.A. Serum Transferrin Carrying the Xeno-Tetrasaccharide NeuAc-Gal-GlcNAc2 Is a Biomarker of ALG1-CDG. J Inherit Metab Dis 2016, 39. [Google Scholar] [CrossRef]
- Kitata, R.B.; Yang, J.C.; Chen, Y.J. Advances in Data-Independent Acquisition Mass Spectrometry towards Comprehensive Digital Proteome Landscape. Mass Spectrom Rev 2023, 42. [Google Scholar] [CrossRef]
- Hertzog, A.; Selvanathan, A.; Devanapalli, B.; Ho, G.; Bhattacharya, K.; Tolun, A.A. A Narrative Review of Metabolomics in the Era of “-Omics”: Integration into Clinical Practice for Inborn Errors of Metabolism. Transl Pediatr 2022, 11, 1704–1716. [Google Scholar] [CrossRef]
- Campbell, M.P.; Peterson, R.; Mariethoz, J.; Gasteiger, E.; Akune, Y.; Aoki-Kinoshita, K.F.; Lisacek, F.; Packer, N.H. UniCarbKB: Building a Knowledge Platform for Glycoproteomics. Nucleic Acids Res 2014, 42. [Google Scholar] [CrossRef]
- Alocci, D.; Mariethoz, J.; Gastaldello, A.; Gasteiger, E.; Karlsson, N.G.; Kolarich, D.; Packer, N.H.; Lisacek, F. GlyConnect: Glycoproteomics Goes Visual, Interactive, and Analytical. J Proteome Res 2019, 18. [Google Scholar] [CrossRef]
- Aoki-Kinoshita, K.; Agravat, S.; Aoki, N.P.; Arpinar, S.; Cummings, R.D.; Fujita, A.; Fujita, N.; Hart, G.M.; Haslam, S.M.; Kawasaki, T.; et al. GlyTouCan 1.0 - The International Glycan Structure Repository. Nucleic Acids Res 2016, 44. [Google Scholar] [CrossRef]
- Ceroni, A.; Maass, K.; Geyer, H.; Geyer, R.; Dell, A.; Haslam, S.M. GlycoWorkbench: A Tool for the Computer-Assisted Annotation of Mass Spectra of Glycans. J Proteome Res 2008, 7. [Google Scholar] [CrossRef] [PubMed]
- Aoki-Kinoshita, K.F.; Lisacek, F.; Karlsson, N.; Kolarich, D.; Packer, N.H. GlycoBioinformatics. Beilstein Journal of Organic Chemistry 2021, 17, 2726–2728. [Google Scholar] [CrossRef] [PubMed]
- Packer, N.H.; Von Der Lieth, C.W.; Aoki-Kinoshita, K.F.; Lebrilla, C.B.; Paulson, J.C.; Raman, R.; Rudd, P.; Sasisekharan, R.; Taniguchi, N.; York, W.S. Frontiers in Glycomics: Bioinformatics and Biomarkers in Disease: An NIH White Paper Prepared from Discussions by the Focus Groups at a Workshop on the NIH Campus, Bethesda MD (September 11-13, 2006). In Proceedings of the Proteomics; 2008; Vol. 8. [Google Scholar] [CrossRef]
- Bojar, D.; Lisacek, F. Glycoinformatics in the Artificial Intelligence Era. Chem Rev 2022, 122. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Chiang, A.W.T.; Lewis, N.E. Artificial Intelligence in the Analysis of Glycosylation Data. Biotechnol Adv 2022, 60. [Google Scholar] [CrossRef]
|
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).