Submitted:
03 June 2025
Posted:
04 June 2025
Read the latest preprint version here
Abstract
Keywords:
1. Introduction
- Introduction of Network Shape Automata (NSA): We propose NSA, a novel memory-based collaborative filtering method that, for the first time, integrates CH theory into similarity computation by leveraging local topological features of the network.
- Comprehensive Hyperparameter Learning and Evaluation Across Domains: NSA was evaluated on 13 datasets across both recommendation and link prediction domains, consistently demonstrating stable and often superior performance compared to state-of-the-art models. Rather than relying on a single perspective, we innovatively examined both sides of the bipartite network structure. By incorporating a broader range of datasets and application scenarios, as detailed in Table 1, our evaluation provides a more comprehensive and rigorous assessment of model performance. Notably, we conducted extensive and systematic hyperparameter learning, involving over 105300 model assessments. This ensured unbiased and automated hyperparameter selection, enabling fair and reproducible comparisons across all evaluated methods.
- Revealing the Power of Structural Information: This work underscores the often-overlooked potential of structural information in recommendation tasks and shows how NSA effectively bridges the gap between interpretability and accuracy. NSA demonstrates that tools from network science can be effectively used to uncover intrinsic patterns in user-item interactions, providing a principled way to model real-world information. By exploiting network structural properties, NSA can capture meaningful relationships even when user-item interactions are extremely limited.
- Unified Perspective on Recommendation and Link Prediction: This work is among the most recent efforts to systematically bridge the tasks of link prediction and recommendation through the lens of network science. By viewing recommendation as a dynamic network evolution process, we provide a unified framework that captures the underlying mechanisms of real-world information systems.
2. Related Work
2.1. Bipartite Network Projection
2.2. Collaborative Filtering
2.3. Cannistraci-Hebb Theory
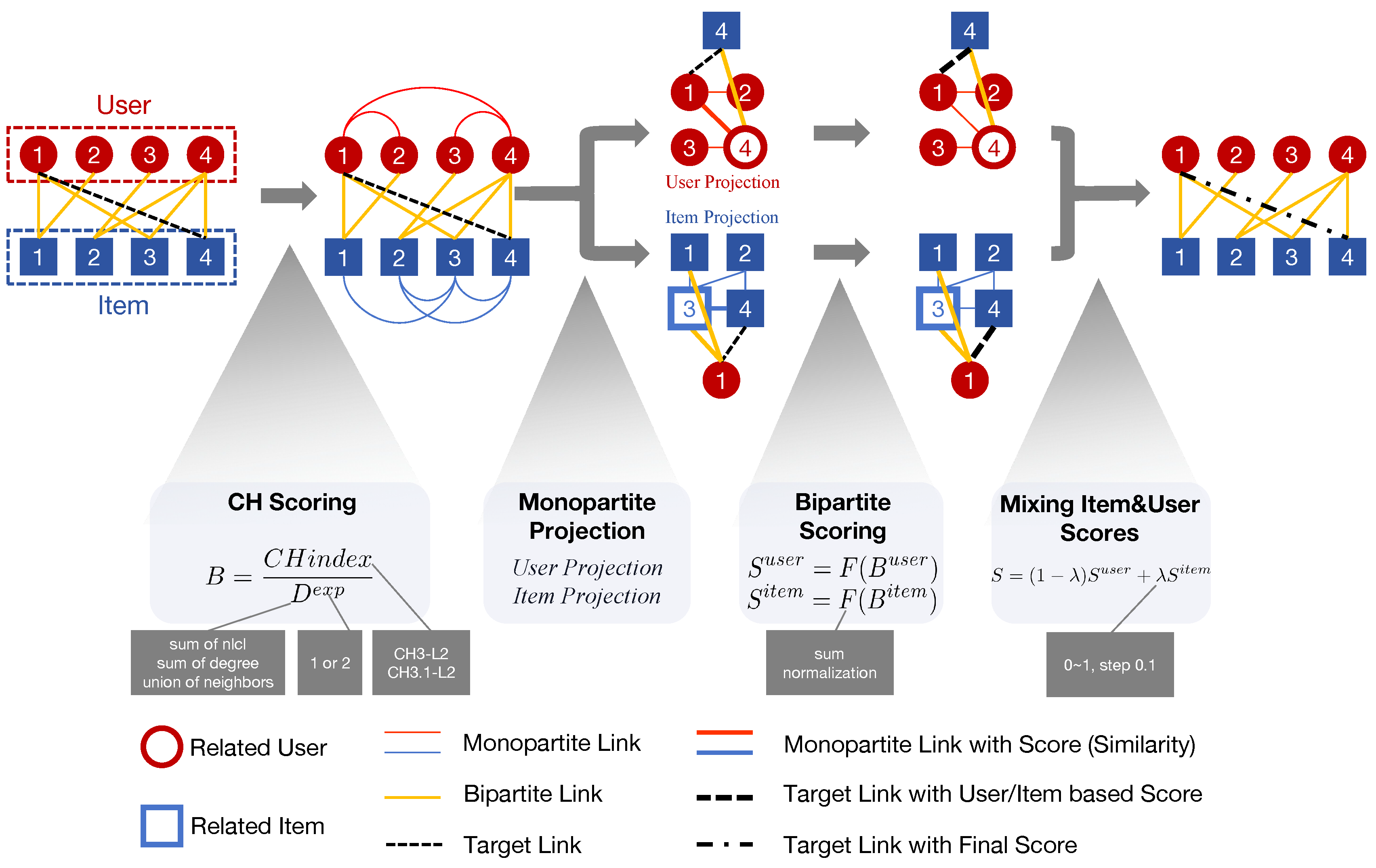
3. Network Shape Automata
3.1. CH Scoring
CH index
Denominator
- sum of degree: the sum of the degrees of the two seed nodes
- union of neighbors: the total number of neighbor nodes of the two seed nodes
- sum of nlcl: the number of non local community links (nlcl) of the two seed nodes (i.e., the number of neighbors of the two seed nodes that are not in the set)
Exponent
3.2. Monopartite Projection
3.3. Bipartite Scoring
- sum
- normalization
3.4. Mixing Item and User Scores
4. Experiments
4.1. Baselines
4.2. Datasets
4.3. Hyperparameter Learning and Evaluation
Metrics
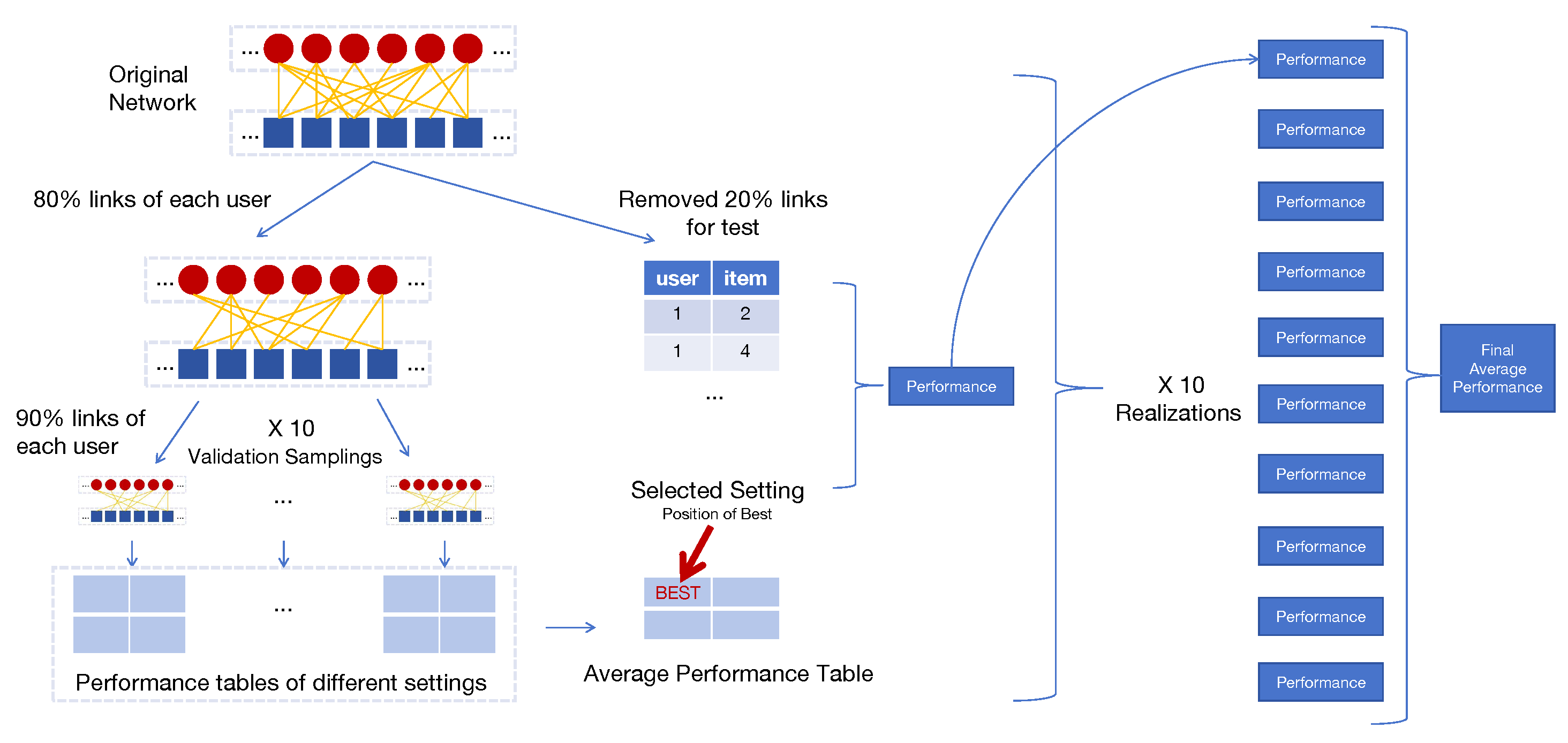
Train-Test Split
Hyperparameter Learning
Evaluation Process
5. Results
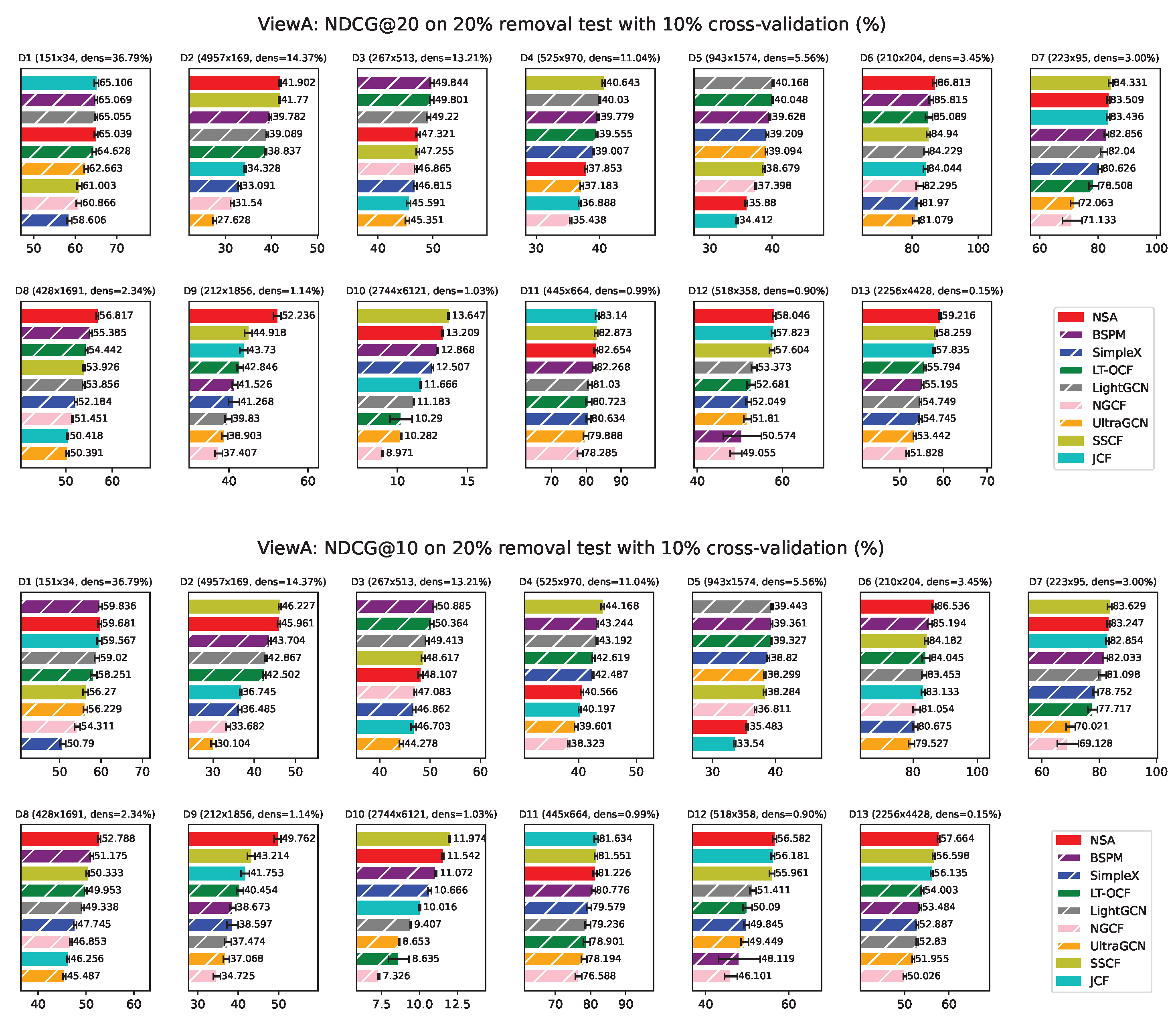
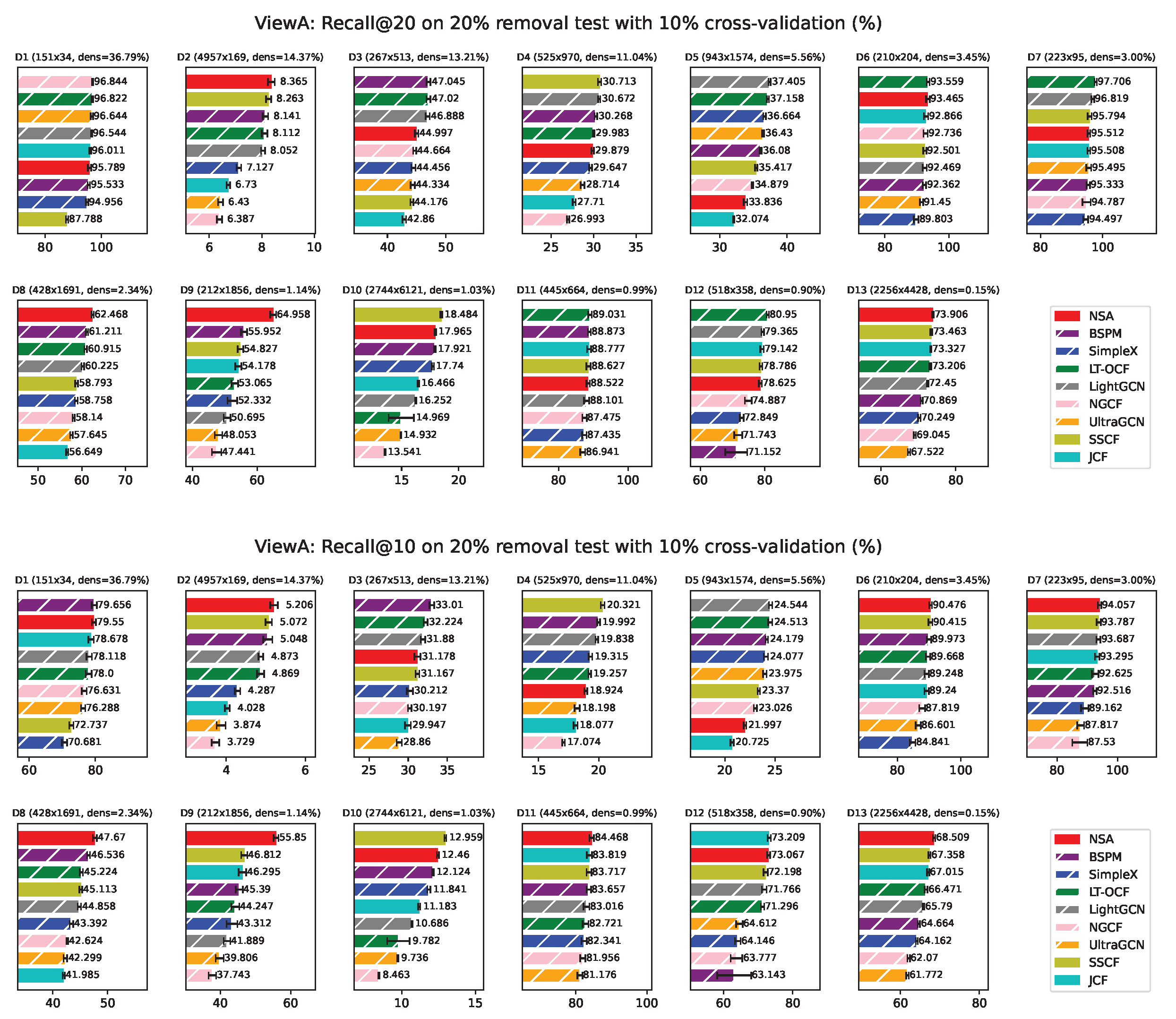
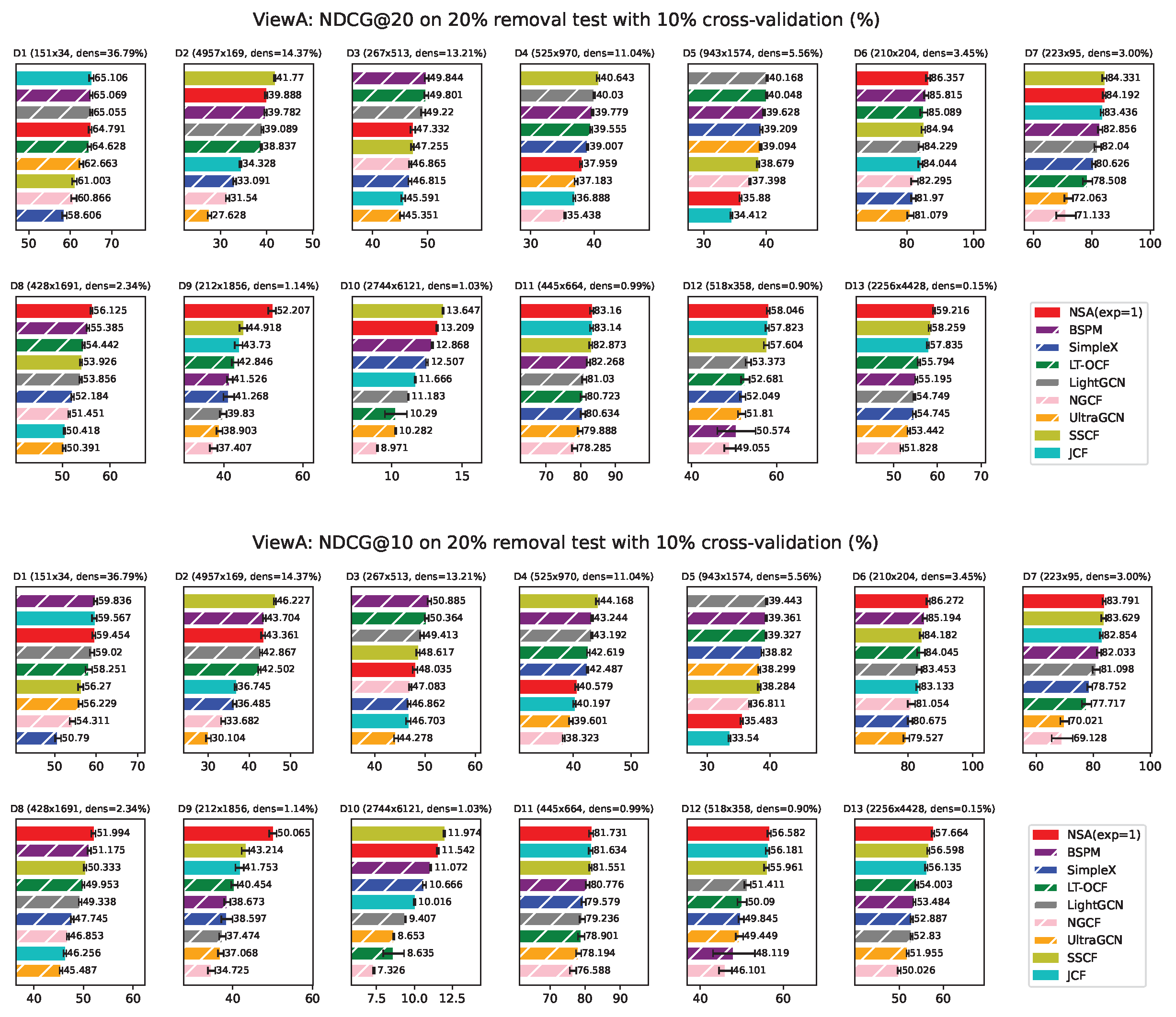
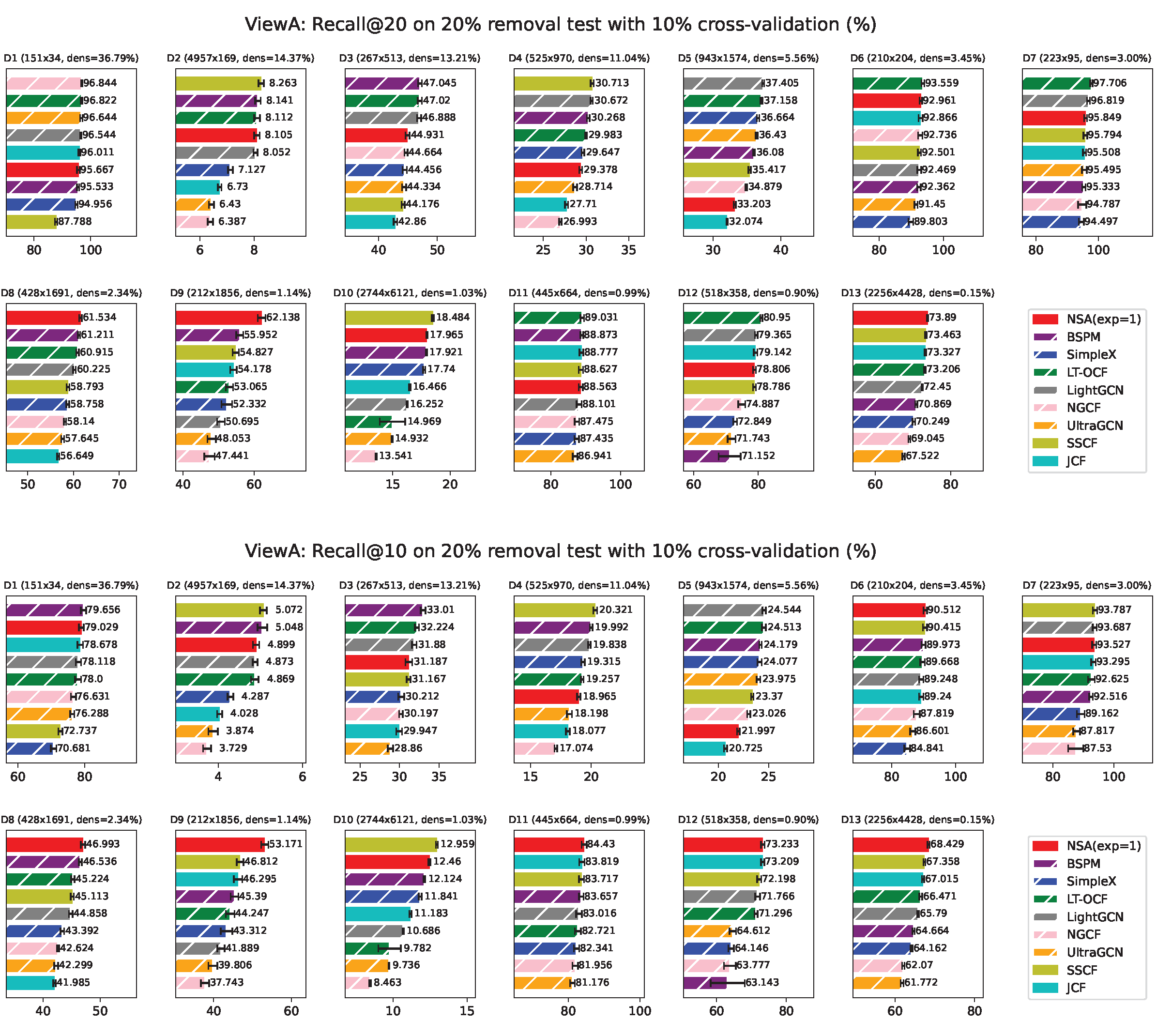
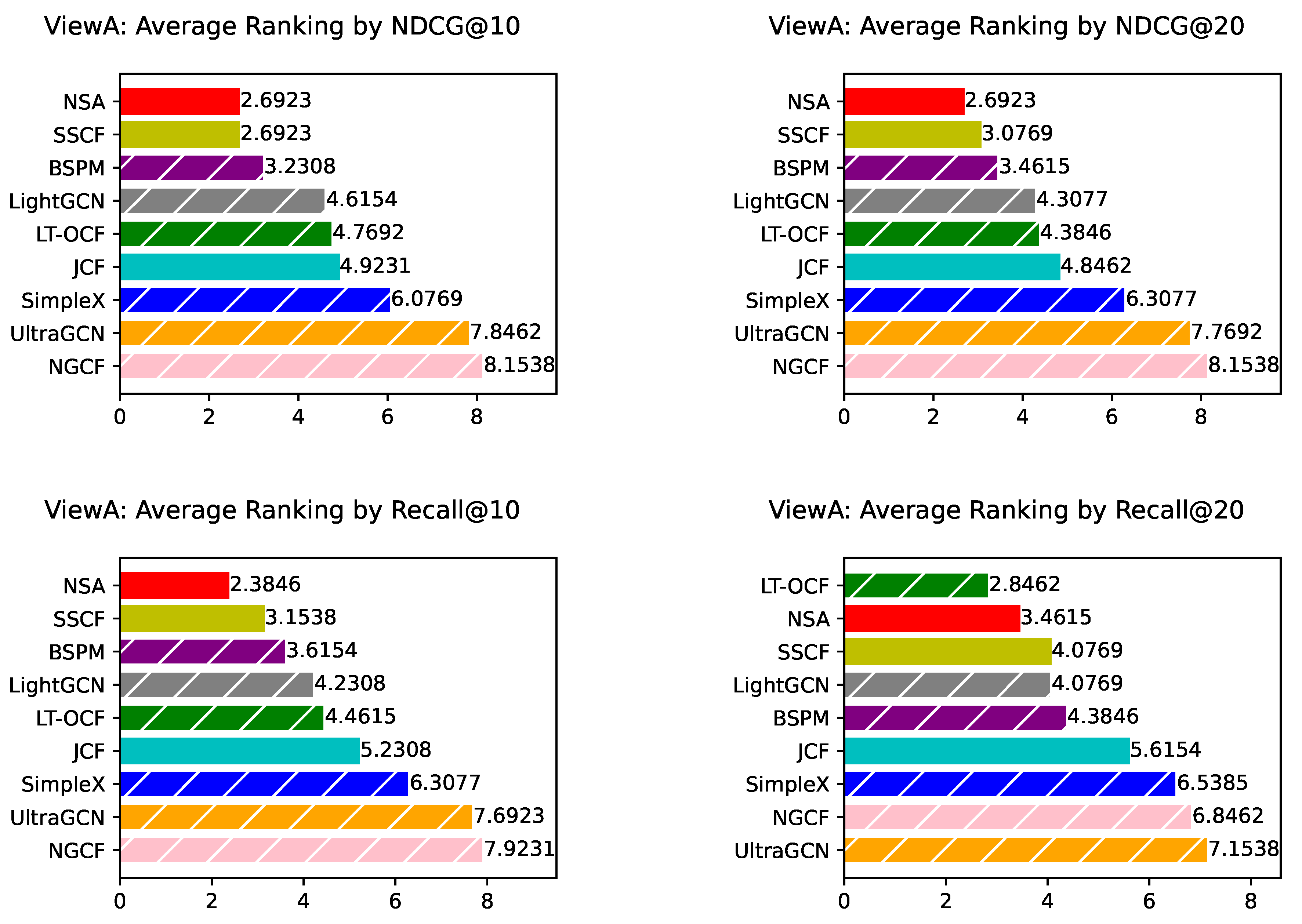
ViewA Results
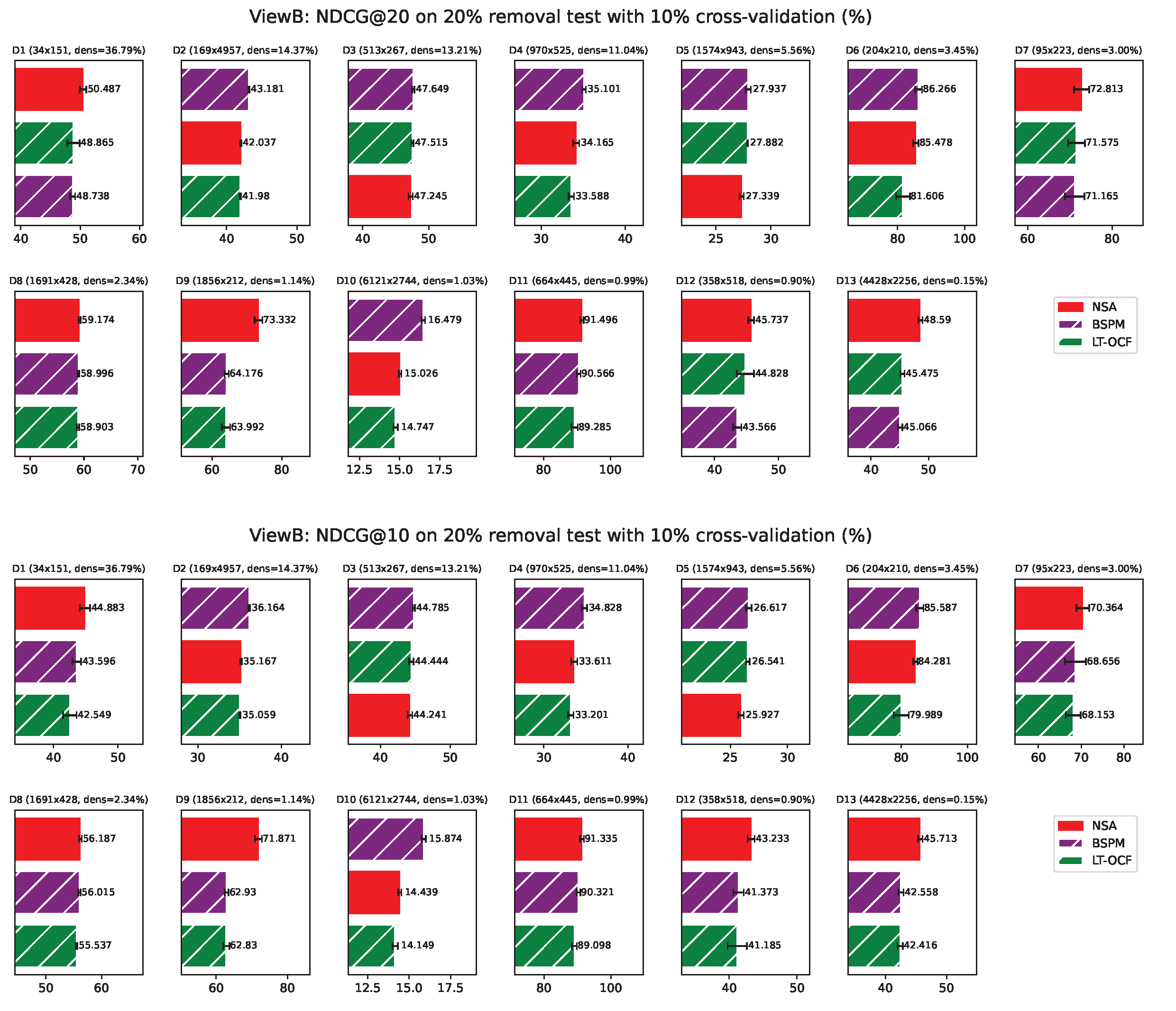
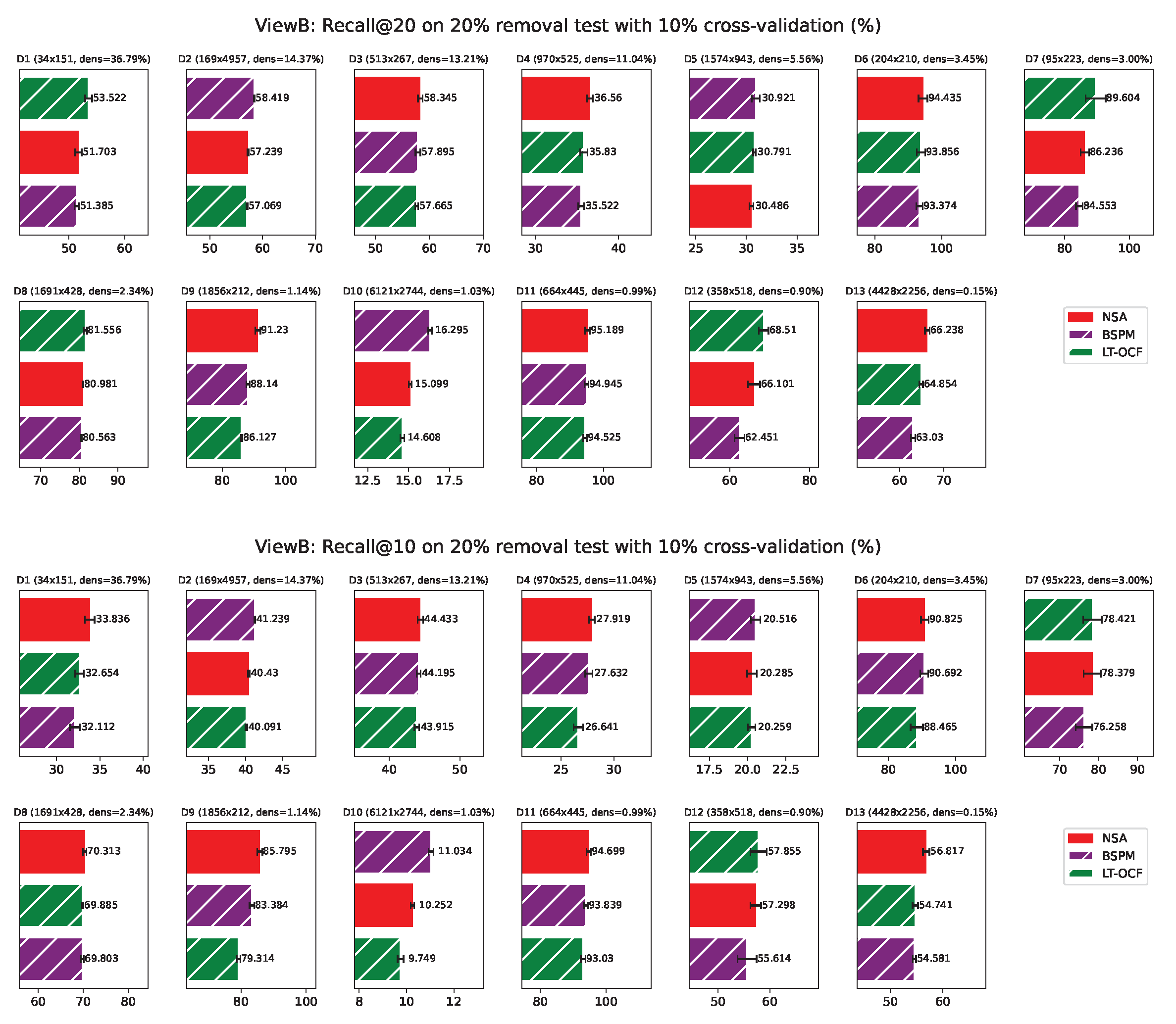
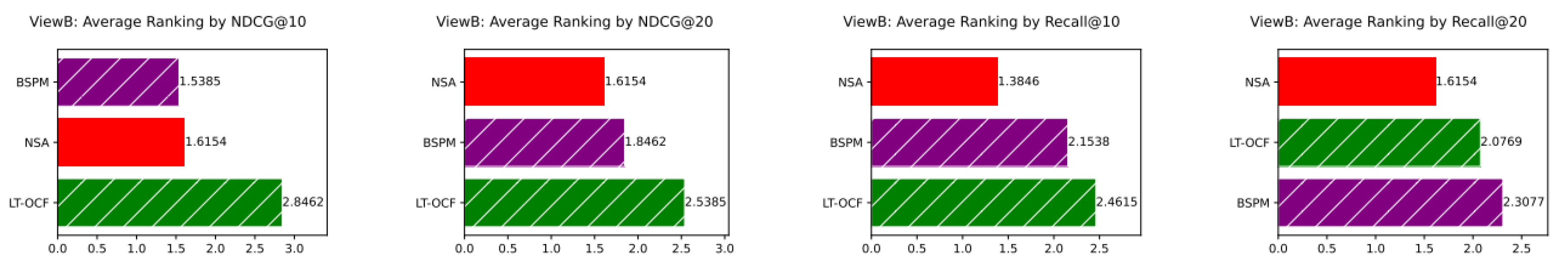
ViewB Results
Effectiveness of a Simplified NSA Variant
Training-Free Robustness of NSA
High Sparsity Robustness of NSA
6. Conclusion and Discussion
Acknowledgments
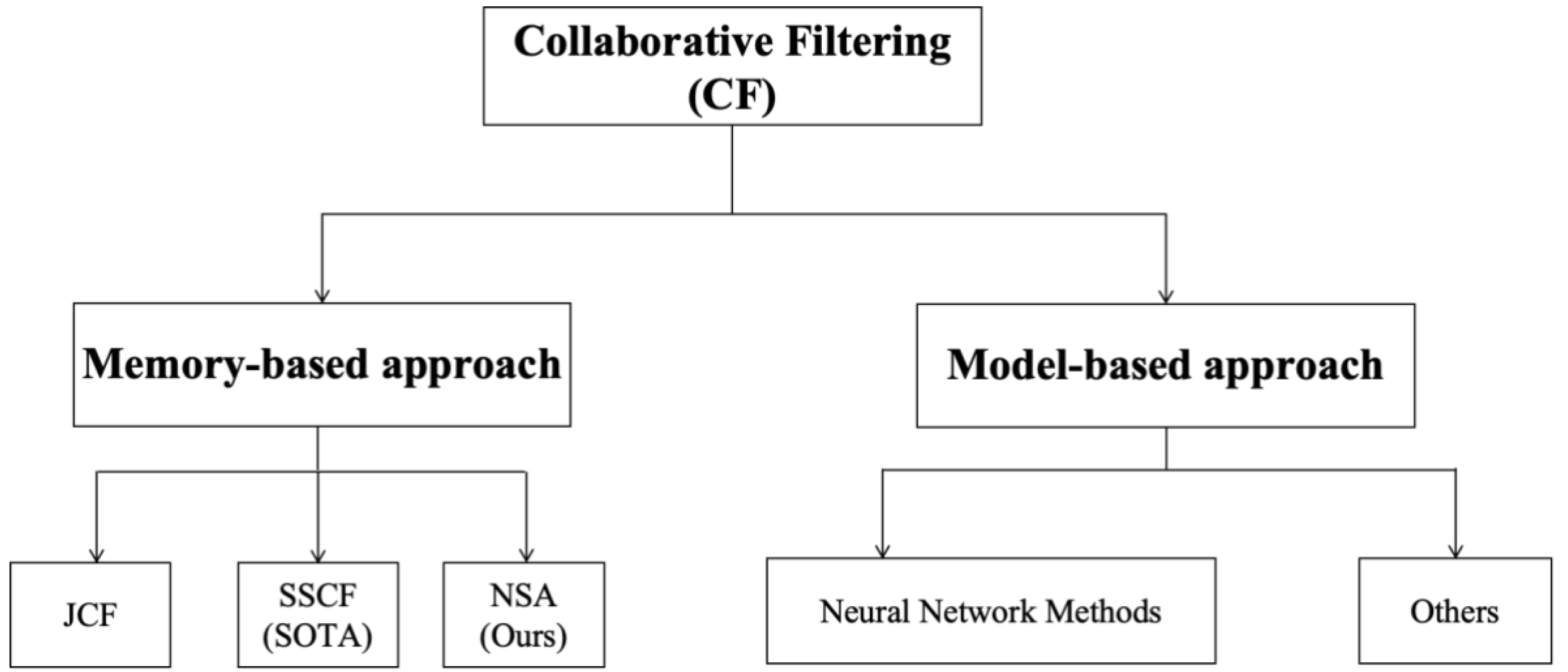
Appendix A. Classification of Collaborative Filtering
Appendix B. Statistics of Datasets
| Index | Name | Field | TypeA | #NodeA | TypeB | #NodeB | #Link | Density |
|---|---|---|---|---|---|---|---|---|
| D1 | aidorganizations_issues [38] | Social | orgnization | 151 | issue | 34 | 1889 | 36.79% |
| D2 | export [42] | Social | country | 169 | item | 4957 | 120377 | 14.37% |
| D3 | industries_educationfields_IPUMS [39] | Social | industry | 267 | education | 513 | 18088 | 13.21% |
| D4 | congressmen_topics_US [40] | Social | congressmen | 525 | topic | 970 | 56215 | 11.04% |
| D5 | users_movies_movielens100k | Social | user | 943 | movie | 1574 | 82520 | 5.56% |
| D6 | drug_target_ionchannel_2009 [41] | Biological | drug | 210 | target | 204 | 1476 | 3.45% |
| D7 | drug_target_GPCR_2009 [41] | Biological | drug | 223 | target | 95 | 635 | 3.00% |
| D8 | occupations_tasks_ONET [40] | Social | occupation | 428 | task | 1691 | 16936 | 2.34% |
| D9 | tfs_genes_regulation_ecoli | Biological | protein | 212 | gene | 1856 | 4496 | 1.14% |
| D10 | amazon-product [45,46] | Social | user | 6121 | item | 2744 | 172206 | 1.03% |
| D11 | drug_target_enzyme_2009 [41] | Biological | drug | 445 | target | 664 | 2926 | 0.99% |
| D12 | drug_target_HQ_2014 [43] | Biological | drug | 518 | target | 358 | 1666 | 0.90% |
| D13 | drug_target_moesm4_esm [44] | Biological | drug | 4428 | target | 2256 | 15051 | 0.15% |
Appendix C. Experimental Environment
Appendix D. Hyperparameter Setting
| Classification | Algorithm | Parameter | Tuning value |
|---|---|---|---|
| Memory-based | CH index | CH3-L2, CH3.1-L2 | |
| denominator | sum of degree, sum of nlcl, union of neighbours | ||
| exponent | 1, 2 | ||
| bipartite scoring | sum, normalization | ||
| NSA | mixing parameter | 0-1, interval 0.1 | |
| SSCF | mixing parameter | 0-1, interval 0.1 | |
| JCF | mixing parameter | 0-1, interval 0.1 | |
| Model-based | lr | 1e-3, 1e-4, 1e-5 | |
| reg | 1e-4, 1e-5, 1e-6 | ||
| embed_size | 64 | ||
| layer size | [64, 64, 64] | ||
| batch size | 1024 | ||
| node dropout | 0.1 | ||
| NGCF | mess dropout | [0.1, 0.1, 0.1] | |
| lr | 1e-2, 1e-3, 1e-4 | ||
| decay | 1e-3, 1e-4, 1e-5 | ||
| recdim | 64 | ||
| dropout | 0 | ||
| layer | 3 | ||
| LightGCN | bpr_batch | 2048 | |
| lr | 1e-3, 1e-4, 1e-5 | ||
| gamma | 0.8, 0.5 | ||
| negative weight | 250, 10 | ||
| embedding_dim | 64 | ||
| num neg | 1000 | ||
| margin | 0.9 | ||
| net_dropout | 0.1 | ||
| SimpleX | batch size | 1024 | |
| lr | 1e-2, 1e-1 | ||
| gamma | 1e-3, 1e-4, 1e-5 | ||
| lambda | 5e-4, 1e-5 | ||
| batch size | 512 | ||
| negative weight | 300 | ||
| UltraGCN | embedding dim | 64 | |
| lr | 1e-2, 1e-3, 1e-4 | ||
| k | 4, 2 | ||
| decay | 1e-4 | ||
| LT-OCF | lrt | 1e-5 | |
| lr | 1e-3, 1e-2 | ||
| idl_betas | 0.2, 0.3 | ||
| factor_dims | 12, 50 | ||
| decay | 1e-4 | ||
| dropout | 0 | ||
| BSPM | layer | 3 |
Appendix E. Hyperparameter Learning and Evaluation Process
Appendix F. ViewA Results on Individual Network
Appendix G. ViewB Results on Individual Network
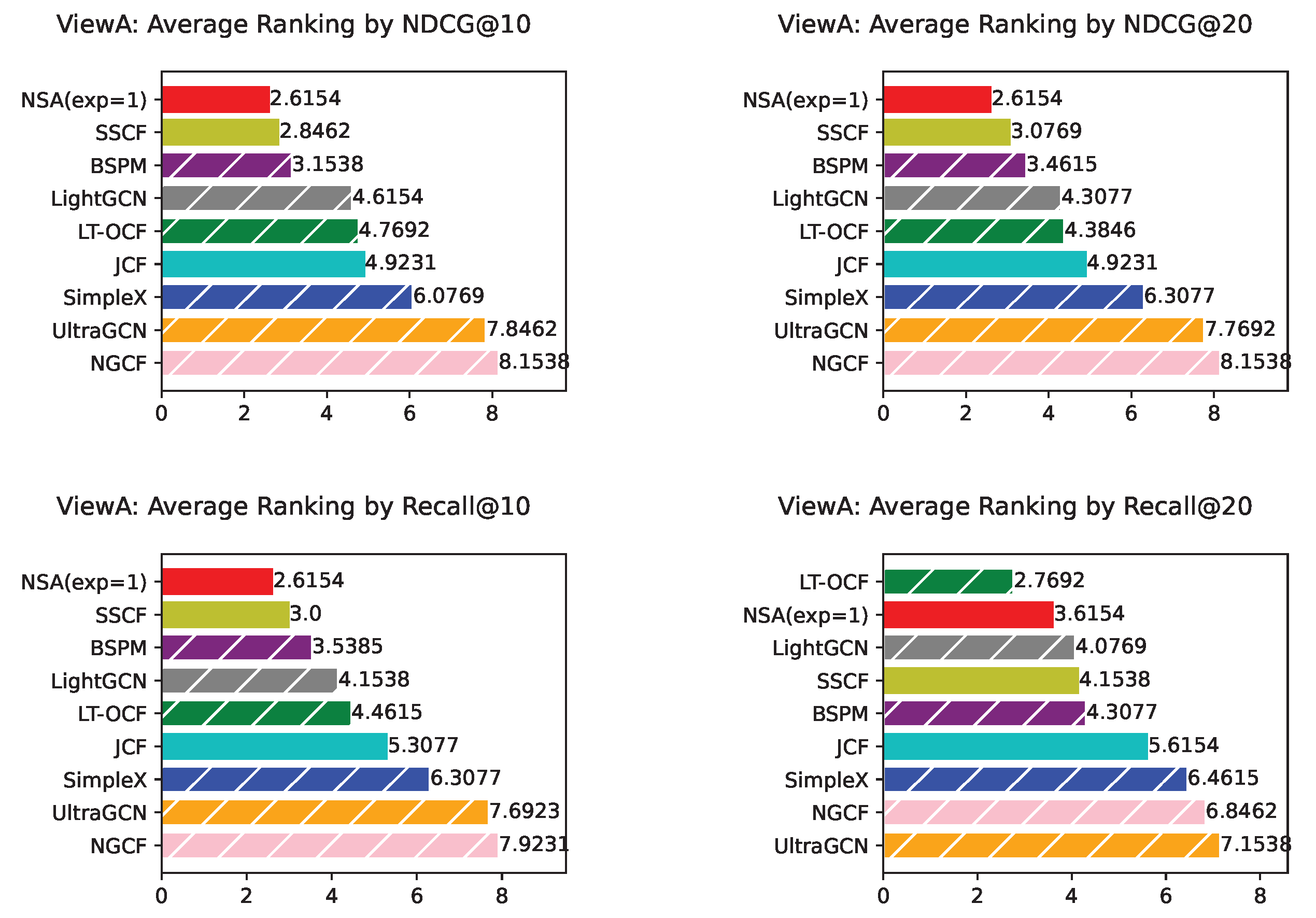
Appendix H. NSA with Fixed Exponent 1 Results from ViewA
Appendix I. Broader Impact and Future Work
Broader Impact
Future Work
Appendix J. Time Complexity of NSA
Appendix J.1. Basic Definition
- U: number of users
- I: number of items
Appendix J.2. CH Scoring and Monopartite Projection
Appendix J.2.2.15. CH index
- Path count. Each length-2 path is defined by an intermediate node z connected to both u and v. The total number of such paths is given by:where is the degree of node z. This represents the number of unique unordered two-hop paths in the network.
- Computation per path. For each length-2 path, CHA computes a score based on the iLCL and eLCL of the intermediate node z. This requires checking the neighbors of z against the local community associated with the pair , which takes time per path.
- Overall time complexity. Multiplying the path count and per-path cost gives the total time complexity:
-
Sparse, degree-homogeneous: If the graph is Sparse (i.e. ) with relatively uniform degrees (i.e., for all z), then:So the overall time complexity of .
-
Sparse, degree-heterogeneous: If the graph is sparse (i.e., ), but has a skewed degree distribution (e.g., power law), we can no longer assume for all nodes. To handle this case, we apply a relaxation via Hölder’s inequality to upper-bound the root-mean-cube degree in terms of the average degree:This relaxation allows us to express the cubic-degree term in the overall complexity as:Thus, the overall time complexity in this case is .
- Dense graphs: In the worst-case scenario of dense graphs, where for all nodes, we obtain:leading to an overall time complexity of .
Denominator
Appendix J.3. Bipartite Scoring
Appendix J.4. Mix Item and User Scores
Appendix J.5. Summary
- Sparse, degree-homogeneous: The dominant component of the time complexity is the collaborative filtering mechanism, result in overall complexity of .
- Sparse, degree-heterogeneous: The dominant component of the time complexity is CH score computation, result in overall complexity of .
- Dense graphs: The dominant component of the time complexity is CH score computation, result in overall complexity of which is rare for recommendation system tasks.
Appendix K. Experimental Time
| Dataset | NSA | SSCF | JCF | NGCF | LightGCN | UltraGCN | SimpleX | LT-OCF | BSPM |
|---|---|---|---|---|---|---|---|---|---|
| aidorganizations_issues | 0.06± 0.00 | 0.06± 0.00 | 0.06± 0.00 | 28.90± 1.52 | 11.30± 0.07 | 35.80± 0.74 | 22.39± 0.42 | 13.41± 0.09 | 17.31± 0.27 |
| export | 5.40± 0.01 | 1.55± 0.00 | 1.20± 0.00 | 1129.81± 8.55 | 435.42± 3.44 | 277.43± 2.16 | 267.52± 0.63 | 617.89± 31.94 | 22.75± 0.03 |
| industries_eductionfields_IPUMS | 0.35± 0.00 | 0.34± 0.00 | 0.40± 0.00 | 150.53± 2.49 | 67.50± 0.54 | 75.34± 0.37 | 34.17± 0.94 | 96.63± 0.85 | 17.99± 0.13 |
| congressmen_topics_US | 1.21± 0.01 | 1.24± 0.00 | 1.41± 0.02 | 340.44± 1.96 | 209.35± 2.30 | 157.58± 1.26 | 113.68± 3.61 | 269.45± 4.39 | 18.97± 0.20 |
| users_movies_movielens100k | 2.90± 0.00 | 3.15± 0.01 | 3.65± 0.01 | 477.04± 5.27 | 288.40± 0.68 | 205.45± 2.18 | 132.38± 2.65 | 402.87± 2.50 | 19.33± 0.07 |
| drug_target_ionchannel_2009 | 0.13± 0.00 | 0.13± 0.00 | 0.12± 0.00 | 70.05± 2.24 | 9.85± 0.37 | 35.58± 0.70 | 12.84± 0.41 | 11.10± 0.27 | 17.71± 0.04 |
| drug_target_GPCR_2009 | 0.09± 0.00 | 0.09± 0.00 | 0.09± 0.00 | 39.53± 1.36 | 6.98± 0.06 | 34.92± 1.14 | 13.85± 0.95 | 8.58± 0.13 | 17.87± 0.13 |
| occupations_tasks_ONET | 1.32± 0.00 | 1.51± 0.00 | 1.76± 0.01 | 141.58± 0.41 | 61.06± 0.37 | 76.30± 0.47 | 63.22± 2.62 | 83.36± 0.36 | 17.91± 0.12 |
| tfs_genes_regulation_ecoli | 0.65± 0.00 | 1.00± 0.00 | 0.57± 0.01 | 82.75± 0.23 | 19.40± 0.13 | 41.33± 0.54 | 24.49± 0.93 | 23.85± 0.11 | 18.04± 0.05 |
| amazon-product | 26.81± 0.12 | 25.67± 0.05 | 25.25± 0.14 | 924.45± 6.48 | 600.66± 1.88 | 385.63± 1.44 | 394.01± 2.27 | 836.37± 4.80 | 22.19± 0.05 |
| drug_target_enzyme_2009 | 0.37± 0.00 | 0.54± 0.00 | 0.30± 0.00 | 64.72± 0.05 | 14.46± 0.20 | 39.43± 1.12 | 12.96± 0.24 | 19.76± 0.14 | 17.70± 0.11 |
| drug_target_HQ_2014 | 0.32± 0.00 | 0.43± 0.00 | 0.30± 0.00 | 67.52± 0.36 | 10.33± 0.12 | 35.92± 1.32 | 18.43± 0.59 | 12.29± 0.27 | 17.83± 0.09 |
| drug_target_moesm4_esm | 12.30± 0.10 | 16.36± 0.04 | 11.66± 0.01 | 135.00± 2.90 | 60.33± 0.11 | 74.57± 0.37 | 58.64± 1.48 | 85.20± 0.63 | 18.85± 0.07 |
Appendix L. Time Complexity of Baselines
Appendix L.1. Definition
- U: number of users
- I: number of items
- E: number of edges in the network
- L: number of layers for neural-network based methods
- D: dimension of embedding in model-based methods
- N: number of negative samples
- K: number of sampling similar neighbors
- T: number of epochs for neural-network based methods
Appendix L.2. Time Complexity
- NGCF:
- LightGCN: Not declared
- UltraGCN:
- SimpleX: Not declared
- LT-OCF: Not declared
- BSPM: Not declared
- SSCF:
- JCF:
References
- Watts, D.J.; Strogatz, S.H. Collective dynamics of ‘small-world’networks. nature 1998, 393, 440–442. [Google Scholar] [CrossRef] [PubMed]
- Albora, G.; Zaccaria, A.; et al. Machine learning to assess relatedness: the advantage of using firm-level data. Complexity 2022, 2022. [Google Scholar] [CrossRef]
- Tacchella, A.; Cristelli, M.; Caldarelli, G.; Gabrielli, A.; Pietronero, L. A new metrics for countries’ fitness and products’ complexity. Scientific reports 2012, 2, 723. [Google Scholar] [CrossRef]
- Straccamore, M.; Pietronero, L.; Zaccaria, A. Which will be your firm’s next technology? Comparison between machine learning and network-based algorithms. Journal of Physics: Complexity 2022, 3, 035002. [Google Scholar] [CrossRef]
- Ricci, F.; Rokach, L.; Shapira, B. Recommender systems: Techniques, applications, and challenges. Recommender systems handbook.
- Goldberg, D.; Nichols, D.; Oki, B.M.; Terry, D. Using collaborative filtering to weave an information tapestry. Communications of the ACM 1992, 35, 61–70. [Google Scholar] [CrossRef]
- Schafer, J.B.; Frankowski, D.; Herlocker, J.; Sen, S. Collaborative filtering recommender systems. In The adaptive web: methods and strategies of web personalization; Springer, 2007; pp. 291–324.
- Chen, R.; Hua, Q.; Chang, Y.S.; Wang, B.; Zhang, L.; Kong, X. A survey of collaborative filtering-based recommender systems: From traditional methods to hybrid methods based on social networks. IEEE access 2018, 6, 64301–64320. [Google Scholar] [CrossRef]
- Albora, G.; Mori, L.R.; Zaccaria, A. Sapling similarity: A performing and interpretable memory-based tool for recommendation. Knowledge-Based Systems 2023, 275, 110659. [Google Scholar] [CrossRef]
- Muscoloni, A.; Abdelhamid, I.; Cannistraci, C.V. Local-community network automata modelling based on length-three-paths for prediction of complex network structures in protein interactomes, food webs and more. bioRxiv, 2018. [Google Scholar] [CrossRef]
- Muscoloni, A.; Michieli, U.; Cannistraci, C.V. Adaptive network automata modelling of complex networks 2020.
- Cannistraci, C.V.; Alanis-Lobato, G.; Ravasi, T. From link-prediction in brain connectomes and protein interactomes to the local-community-paradigm in complex networks. Scientific reports 2013, 3, 1613. [Google Scholar] [CrossRef]
- Abdelhamid, I.; Muscoloni, A.; Rotscher, D.M.; Lieber, M.; Markwardt, U.; Cannistraci, C.V. Network shape intelligence outperforms AlphaFold2 intelligence in vanilla protein interaction prediction. bioRxiv, 2023. [Google Scholar]
- Zhang, Y.; Zhao, J.; Wu, W.; Muscoloni, A.; Cannistraci, C.V. Ultra-sparse network advantage in deep learning via Cannistraci-Hebb brain-inspired training with hyperbolic meta-deep community-layered epitopology. In Proceedings of the The Twelfth International Conference on Learning Representations; 2023. [Google Scholar]
- Wang, X.; He, X.; Wang, M.; Feng, F.; Chua, T.S. Neural graph collaborative filtering. In Proceedings of the Proceedings of the 42nd international ACM SIGIR conference on Research and development in Information Retrieval, 2019, pp. 165–174.
- He, X.; Deng, K.; Wang, X.; Li, Y.; Zhang, Y.; Wang, M. Lightgcn: Simplifying and powering graph convolution network for recommendation. In Proceedings of the Proceedings of the 43rd International ACM SIGIR conference on research and development in Information Retrieval, 2020, pp. 639–648.
- Mao, K.; Zhu, J.; Xiao, X.; Lu, B.; Wang, Z.; He, X. UltraGCN: ultra simplification of graph convolutional networks for recommendation. In Proceedings of the Proceedings of the 30th ACM international conference on information & knowledge management, 2021, pp. 1253–1262.
- Mao, K.; Zhu, J.; Wang, J.; Dai, Q.; Dong, Z.; Xiao, X.; He, X. SimpleX: A simple and strong baseline for collaborative filtering. In Proceedings of the Proceedings of the 30th ACM International Conference on Information & Knowledge Management, 2021, pp. 1243–1252.
- Choi, J.; Jeon, J.; Park, N. Lt-ocf: learnable-time ode-based collaborative filtering. In Proceedings of the Proceedings of the 30th ACM International Conference on Information & Knowledge Management, 2021, pp. 251–260.
- Choi, J.; Hong, S.; Park, N.; Cho, S.B. Blurring-sharpening process models for collaborative filtering. In Proceedings of the Proceedings of the 46th international ACM SIGIR conference on research and development in information retrieval, 2023, pp. 1096–1106.
- Zhou, T.; Ren, J.; Medo, M.; Zhang, Y.C. Bipartite network projection and personal recommendation. Physical review E 2007, 76, 046115. [Google Scholar] [CrossRef]
- Fan, Y.; Li, M.; Zhang, P.; Wu, J.; Di, Z. The effect of weight on community structure of networks. Physica A: Statistical Mechanics and its applications 2007, 378, 583–590. [Google Scholar] [CrossRef]
- Newman, M.E. Scientific collaboration networks. II. Shortest paths, weighted networks, and centrality. Physical review E 2001, 64, 016132. [Google Scholar] [CrossRef] [PubMed]
- Brusilovski, P.; Kobsa, A.; Nejdl, W. The adaptive web: methods and strategies of web personalization; Vol. 4321, Springer Science & Business Media, 2007.
- Breese, J.S.; Heckerman, D.; Kadie, C. Empirical analysis of predictive algorithms for collaborative filtering. arXiv preprint arXiv:1301.7363, arXiv:1301.7363 2013.
- Su, X.; Khoshgoftaar, T.M. A survey of collaborative filtering techniques. Advances in artificial intelligence 2009, 2009. [Google Scholar] [CrossRef]
- Liben-Nowell, D.; Kleinberg, J. Proceedings of the Proceedings of the twelfth international conference on Information and knowledge management, 2003, pp. 556–559.
- Jaccard, P. Étude comparative de la distribution florale dans une portion des Alpes et des Jura. Bull Soc Vaudoise Sci Nat 1901, 37, 547–579. [Google Scholar]
- Zhou, T.; Lü, L.; Zhang, Y.C. Predicting missing links via local information. The European Physical Journal B 2009, 71, 623–630. [Google Scholar] [CrossRef]
- Shardanand, U.; Maes, P. Social information filtering: Algorithms for automating “word of mouth”. In Proceedings of the Proceedings of the SIGCHI conference on Human factors in computing systems, 1995, pp. 210–217.
- Chen, L.; Wu, L.; Hong, R.; Zhang, K.; Wang, M. Revisiting graph based collaborative filtering: A linear residual graph convolutional network approach. In Proceedings of the Proceedings of the AAAI conference on artificial intelligence, 2020, Vol. 34, pp. 27–34.
- Barabási, A.L.; Albert, R. Emergence of Scaling in Random Networks. Science 1999, 286, 509–512. [Google Scholar] [CrossRef]
- Papadopoulos, F.; Kitsak, M.; Serrano, M.Á.; Boguñá, M.; Krioukov, D. Popularity versus similarity in growing networks. Nature 2012, 489, 537–540. [Google Scholar] [CrossRef]
- Muscoloni, A.; Cannistraci, C.V. A nonuniform popularity-similarity optimization (nPSO) model to efficiently generate realistic complex networks with communities. New Journal of Physics 2018, 20, 052002. [Google Scholar] [CrossRef]
- Wolfram, S.; Gad-el Hak, M. A new kind of science. Appl. Mech. Rev. 2003, 56, B18–B19. [Google Scholar] [CrossRef]
- Smith, D.M.; Onnela, J.P.; Lee, C.F.; Fricker, M.D.; Johnson, N.F. Network automata: Coupling structure and function in dynamic networks. Advances in Complex Systems 2011, 14, 317–339. [Google Scholar] [CrossRef]
- Cannistraci, C.V. Modelling self-organization in complex networks via a brain-inspired network automata theory improves link reliability in protein interactomes. Scientific reports 2018, 8, 15760. [Google Scholar] [CrossRef]
- Coscia, M.; Hausmann, R.; Hidalgo, C.A. The structure and dynamics of international development assistance. Journal of Globalization and Development 2013, 3, 1–42. [Google Scholar] [CrossRef]
- Ruggles, S.; Hacker, J.D.; Sobek, M. General design of the integrated public use microdata series. Historical Methods: A Journal of Quantitative and Interdisciplinary History 1995, 28, 33–39. [Google Scholar] [CrossRef]
- Yildirim, M.A.; Coscia, M. Using random walks to generate associations between objects. PloS one 2014, 9, e104813. [Google Scholar] [CrossRef] [PubMed]
- Yamanishi, Y.; Araki, M.; Gutteridge, A.; Honda, W.; Kanehisa, M. Prediction of drug–target interaction networks from the integration of chemical and genomic spaces. Bioinformatics 2008, 24, i232–i240. [Google Scholar] [CrossRef]
- Balassa, B. Trade liberalisation and “revealed” comparative advantage 1. The manchester school 1965, 33, 99–123. [Google Scholar] [CrossRef]
- Hu, Y.; Bajorath, J. Monitoring drug promiscuity over time. F1000Research 2014, 3. [Google Scholar] [CrossRef]
- Cheng, F.; Kovács, I.A.; Barabási, A.L. Network-based prediction of drug combinations. Nature communications 2019, 10, 1197. [Google Scholar] [CrossRef]
- McAuley, J.; Targett, C.; Shi, Q.; Van Den Hengel, A. Image-based recommendations on styles and substitutes. In Proceedings of the Proceedings of the 38th international ACM SIGIR conference on research and development in information retrieval, 2015, pp. 43–52.
- Pasricha, R.; McAuley, J. Translation-based factorization machines for sequential recommendation. In Proceedings of the Proceedings of the 12th ACM Conference on Recommender Systems, 2018, pp. 63–71.
| Algorithm | Year | Networks | Ref |
|---|---|---|---|
| NGCF | 2019 | 3 | [15] |
| LightGCN | 2020 | 3 | [16] |
| UltraGCN | 2021 | 4 | [17] |
| SimpleX | 2021 | 11 | [18] |
| LT-OCF | 2021 | 3 | [19] |
| BSPM | 2022 | 3 | [20] |
| SSCF | 2023 | 5 | [9] |
| NSA | 2025 | 13 x 2 | Ours |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




