Submitted:
20 May 2025
Posted:
20 May 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1 Study Design
2.2. Population and Case Selection
2.2.1. Inclusion Criteria
- Histopathologically confirmed diagnosis of sporadic colorectal cancer.
- Availability of complete sequencing data for KRAS, NRAS, BRAF, and MSI.
- Sufficient tumor tissue for molecular analysis.
2.2.2. Exclusion Criteria
- Cases of hereditary non-polyposis colorectal cancer (HNPCC) or Lynch syndrome at the time of diagnosis.
- Patients lacking complete information regarding the genes studied or MSI status.
- Inadequate or degraded samples unsuitable for molecular testing.
2.3. Molecular Analysis Procedures
2.3.1. DNA Extraction
2.3.2 Next-Generation Sequencing (NGS) with Action OncoKitDx® Panel
- The Action OncoKitDx panel allows full exon sequencing of KRAS, NRAS, and BRAF genes. MSI is assessed through a panel of 110 microsatellite regions. A minimum of 99 evaluable markers is required, and classification is based on the percentage of unstable markers as follows: high MSI (31–100%, MSI-H), low MSI (21–30%, MSI-L), microsatellite stable (0–17%, MSS), and inconclusive results (18–20%)
- The panel is prepared using an automated workflow with the Magnis Dx NGS Prep System (Agilent). After DNA extraction, samples undergo enzymatic fragmentation and enrichment of target regions via hybridization with capture probes using SureSelectXT HS technology, according to the manufacturer’s instructions. High-throughput NGS is performed on the NextSeq 550 platform (Illumina, San Diego, CA, US) using paired-end sequencing (2 × 75 bp) by cyclic reversible termination chemistry. This allows detection of point mutations and larger sequence alterations in the targeted genes.
- Bioinformatic analysis is performed using a dedicated pipeline through the Data Genomics platform (www.datagenomics.es). It includes alignment of reads to the reference genome (GRCh37/hg19), quality filtering, and variant calling. Variant nomenclature follows the guidelines of the Human Genome Variation Society (HGVS; www.hgvs.org).
- Analytical validation and clinical utility of the Action OncoKitDx panel have been established (Martínez-Fernández et al. J Pers Med 2021; 11:360. doi:10.3390/jpm11050360). Both the panel and the Data Genomics software are CE-IVD certified.
- According to the manufacturer, the panel can detect alterations present at a minimum allele frequency of 5%. Detection may be compromised if sequencing depth is below 200×. To achieve this detection threshold, a minimum tumor cellularity of 30% is recommended, along with input DNA quantity between 50–200 ng and a DNA Integrity Number (DIN) >3. For MSI interpretation, the recommended tumor cellularity is also ≥30%. Samples with lower cellularity or suboptimal DNA quality may still be analyzed, though with reduced sensitivity and specificity.
2.3.3. Determination of Microsatellite Instability (MSI)
- NGS, as described above, using the Action OncoKitDx panel.
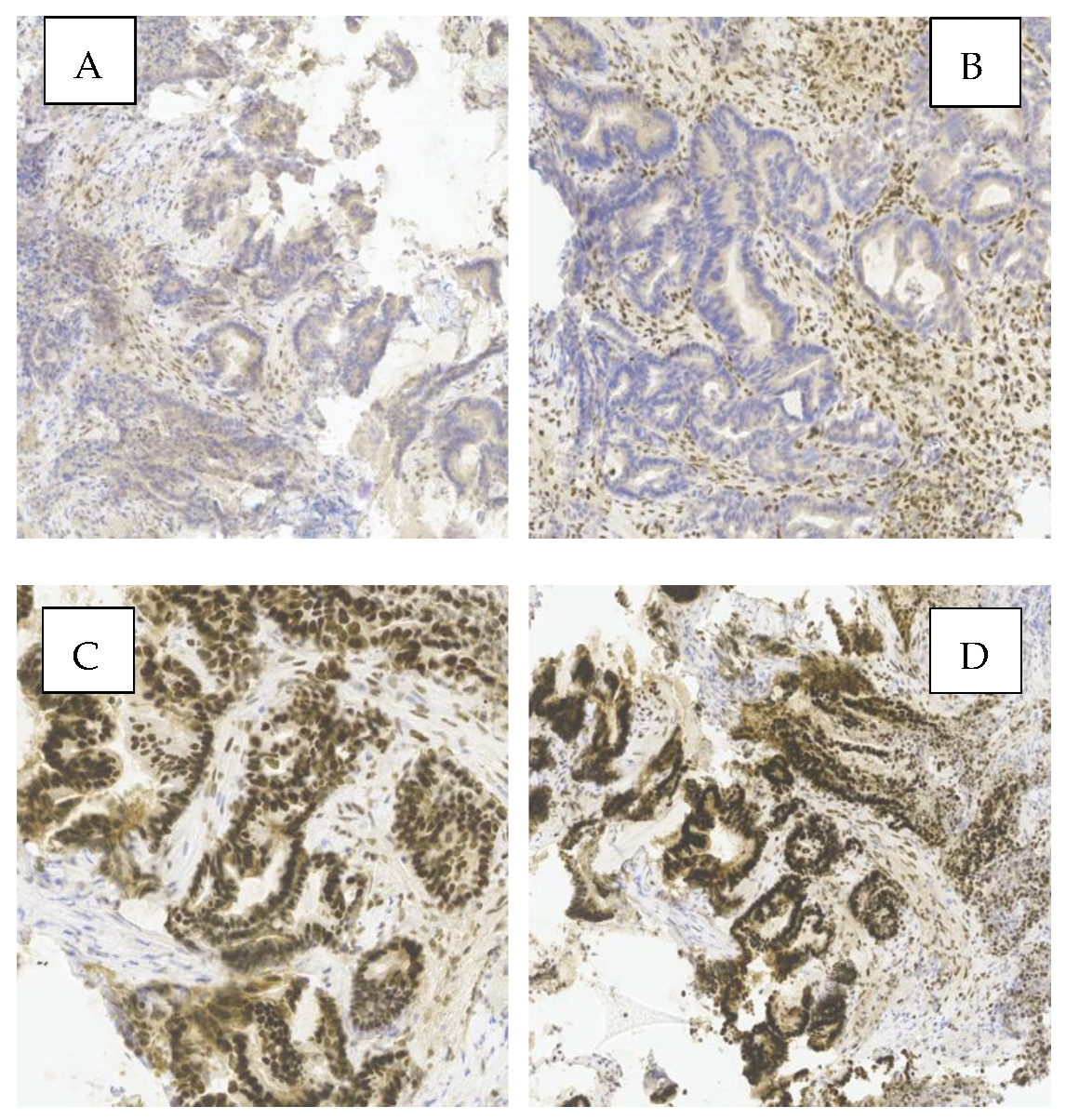
- Immunohistochemistry (IHC), by evaluating the expression of DNA mismatch repair (MMR) proteins MLH1, MSH2, MSH6, and PMS2.
2.3.4. IHC Interpretation Criteria for MMR Protein Expression
2.3.5. Real-Time PCR: Idylla KRAS Mutation Test and Idylla NRAS-BRAF Mutation Test
2.4 Statistical Analysis
- Chi-square test (χ²) and Fisher’s exact test, depending on data distribution.
- Calculation of odds ratios (ORs) with 95% confidence intervals (95% CI).
- A p-value < 0.05 was considered statistically significant.
2.5. Artificial Inteligence
3. Results
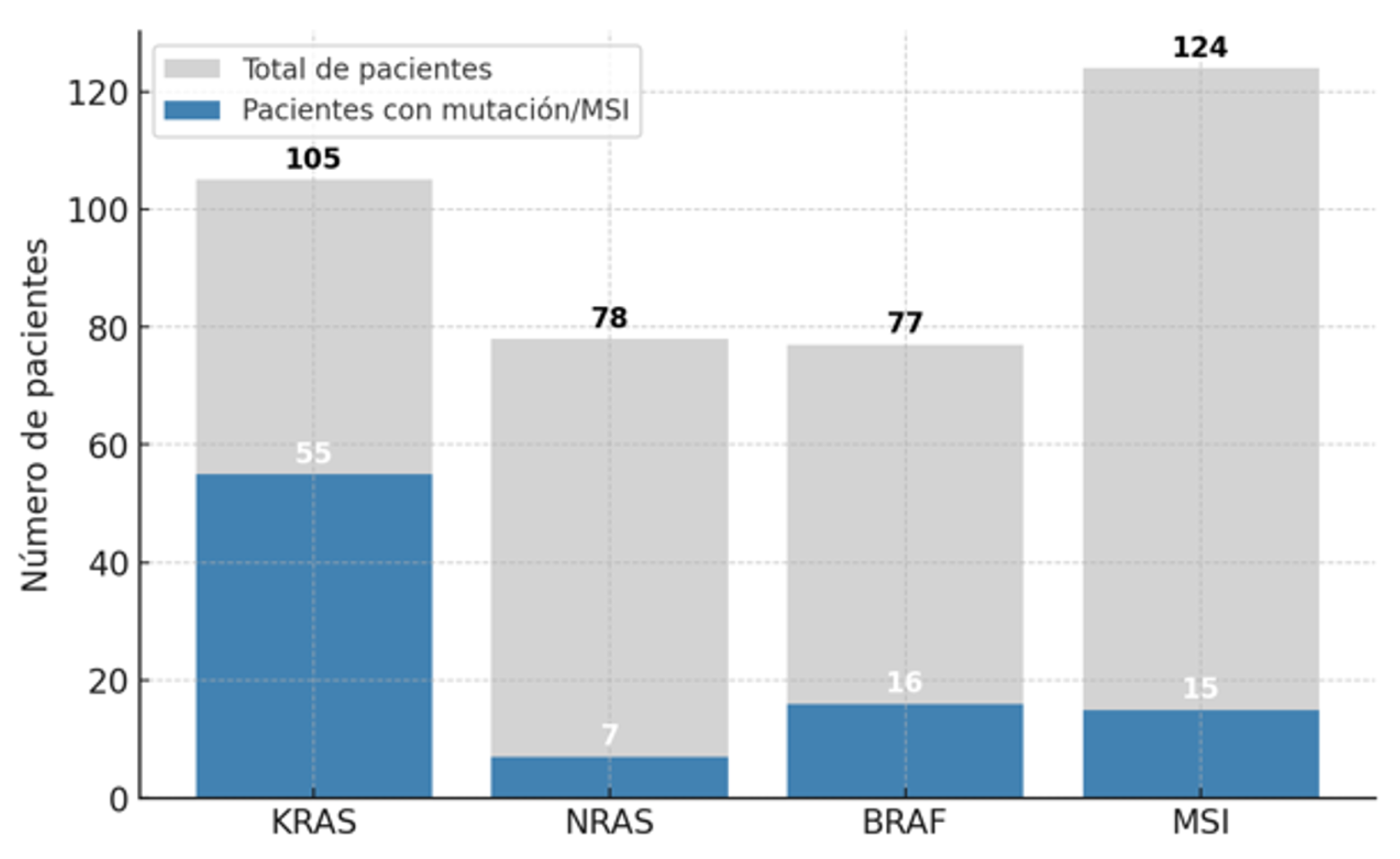
3.1. Descriptive Study
3.2 Comparative Study
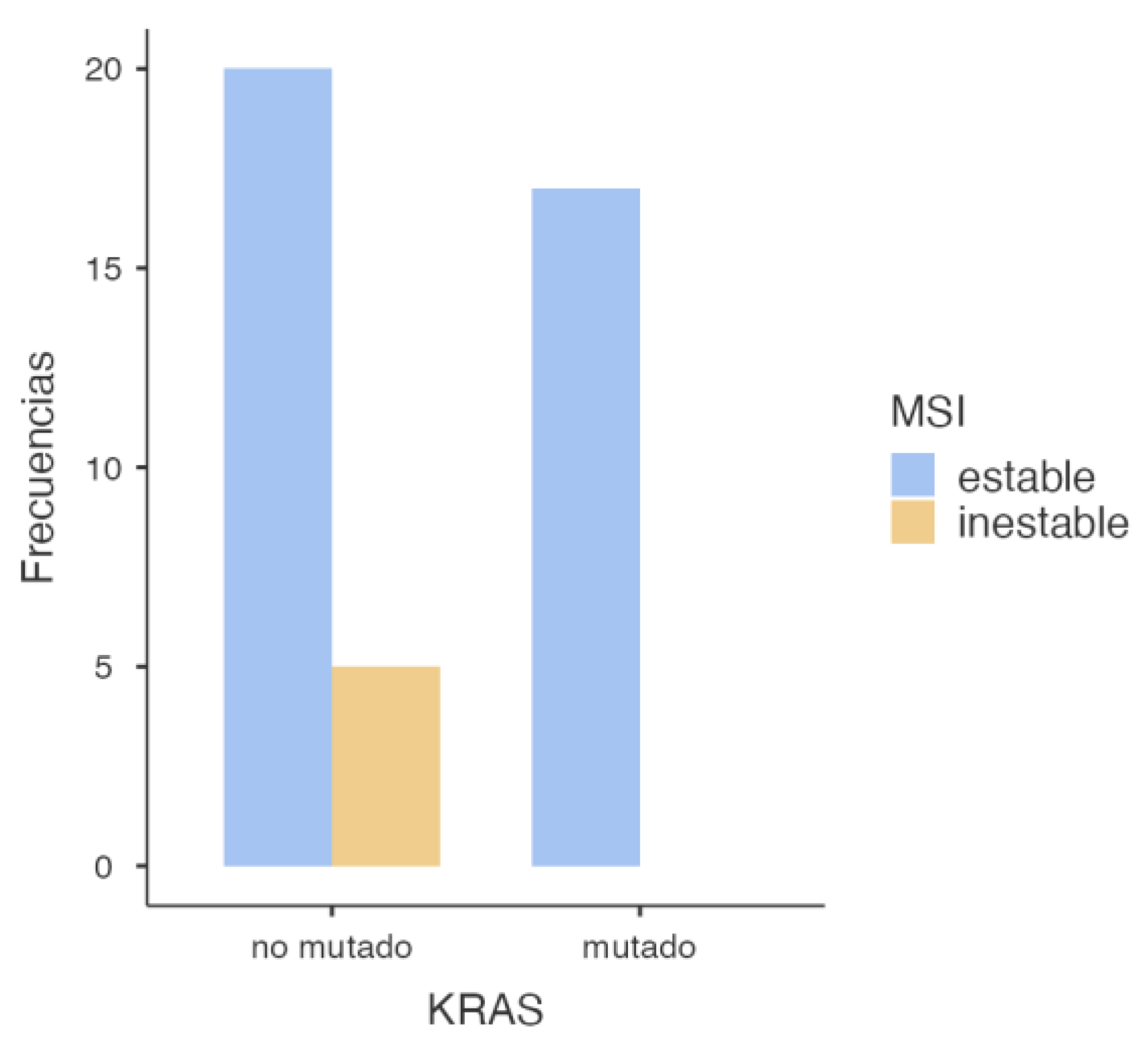
3.2.1. KRAS and MSI Association
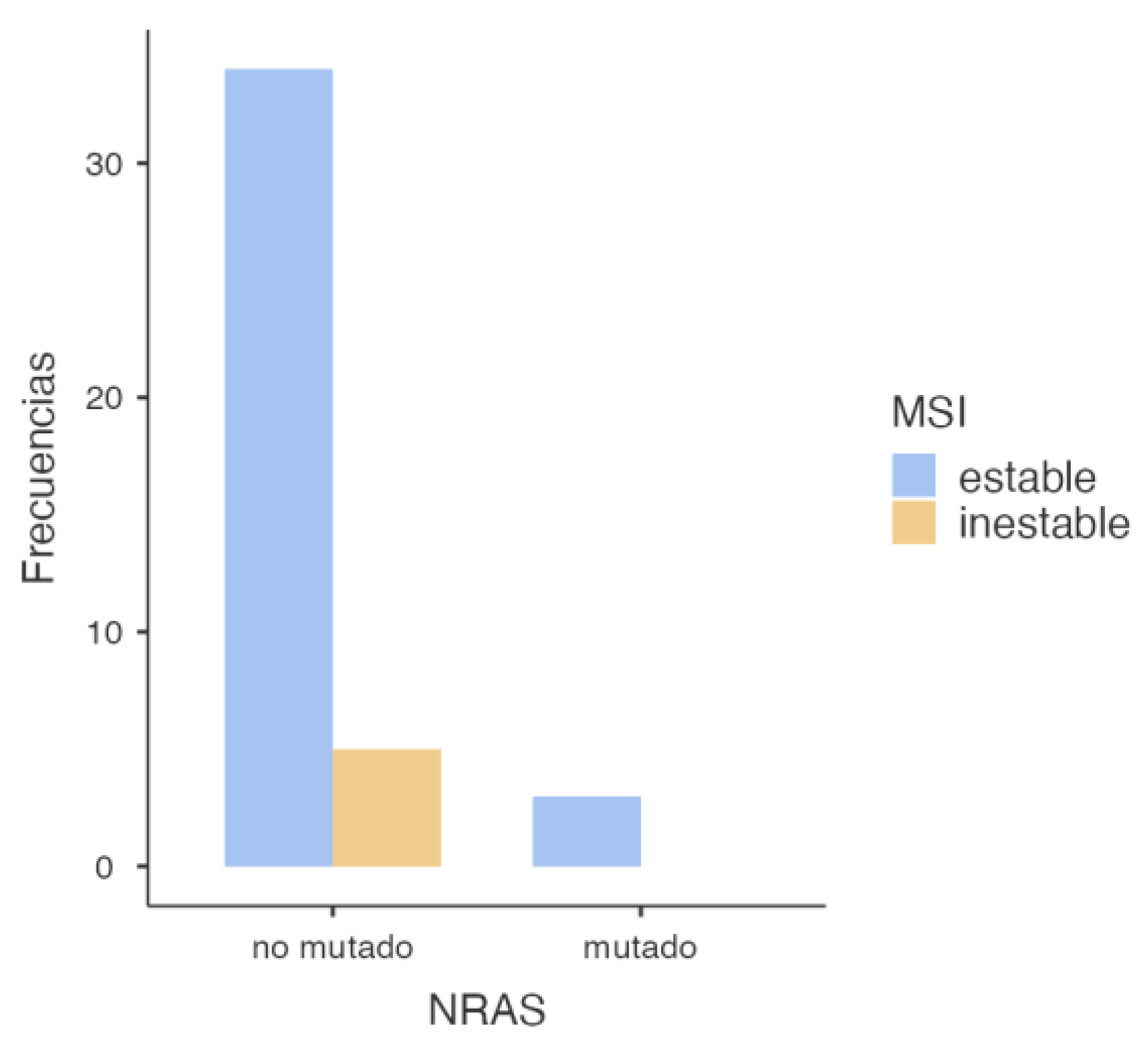
3.2.2. NRAS and MSI Association
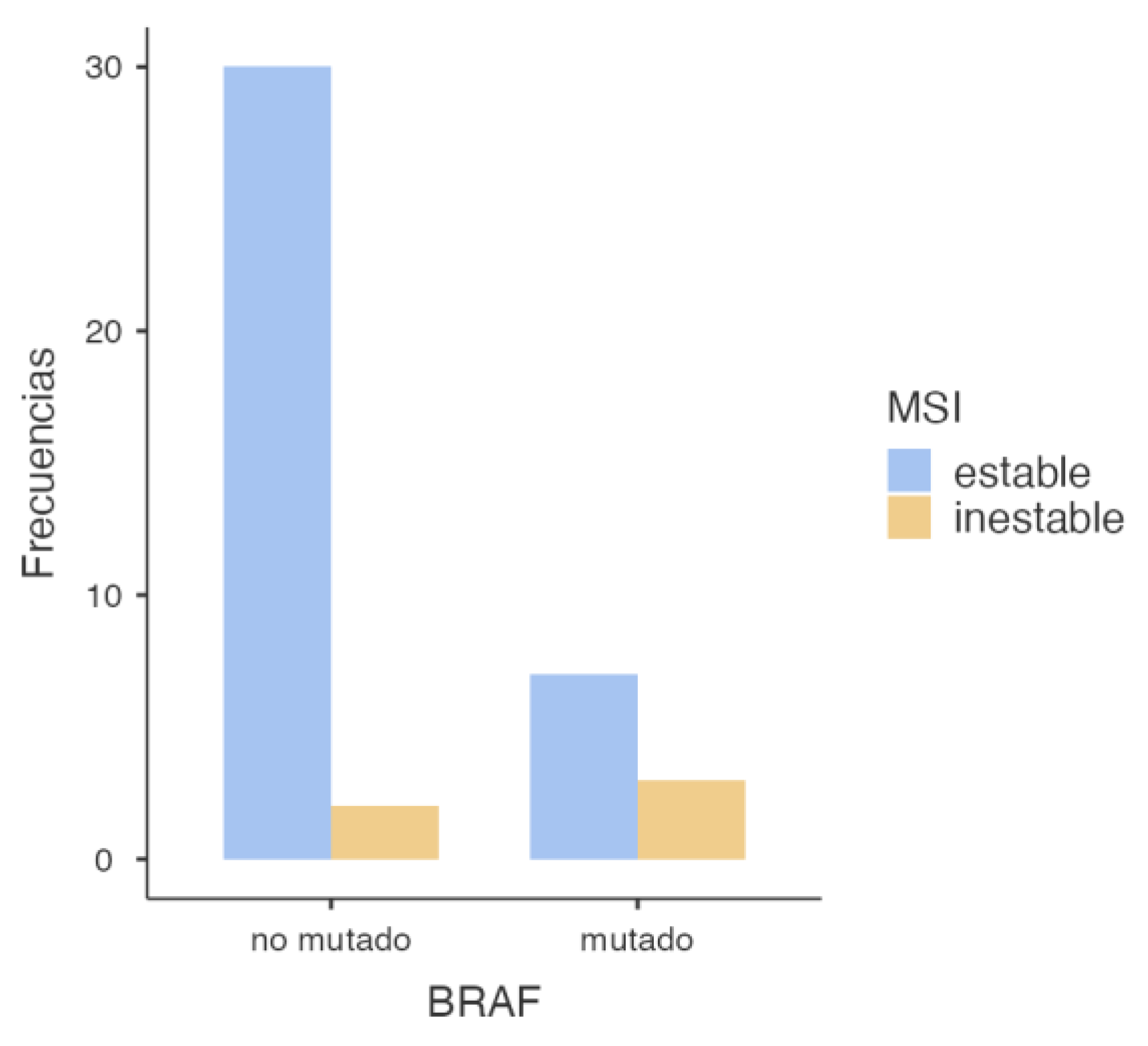
3.2.3. BRAF and MSI Association
| Value | Gl | p | |
|---|---|---|---|
| X2 | 4.10 | 1 | 0.043 |
| Fisher’s Exact Test p-value | 0.078 | ||
| N | 42 |
| 95% Confidence Intervals | |||
|---|---|---|---|
| Value | Lower | Upper | |
| Odds Ratio | 6.43 | 0.897 | 46.1 |
| Relative Risk | 1.341 | 0.884 | 2.03 |
4. Discussion
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
| CRC | Colorectal Cancer |
| SCRC | Sporadic Colorectal Cancer |
| MSI | Microsatellite Inestability |
| MSS | Microsatellite Stability |
| MMR | Mismatch Repair |
| NGS | Next Generation Sequencing |
| IHC | Inmunohistochemical |
References
- Keshinro A, Ganesh K, Vanderbilt C, Firat C, Kim JK, Chen CT, et al. Characteristics of Mismatch Repair-Deficient Colon Cancer in Relation to Mismatch Repair Protein Loss, Hypermethylation Silencing, and Constitutional and Biallelic Somatic Mismatch Repair Gene Pathogenic Variants. Dis Colon Rectum. 2023;66(4):549–58.
- Orlandi E, Giuffrida M, Trubini S, Luzietti E, Ambroggi M, Anselmi E, et al. Unraveling the Interplay of KRAS, NRAS, BRAF, and Micro-Satellite Instability in Non-Metastatic Colon Cancer: A Systematic Review. Diagnostics. 2024;14(10). [CrossRef]
- Lochhead P, Kuchiba A, Imamura Y, Liao X, Yamauchi M, Nishihara R, et al. Microsatellite instability and braf mutation testing in colorectal cancer prognostication. J Natl Cancer Inst. 2013;105(15):1151–6. [CrossRef]
- Birgisson H, Edlund K, Wallin U, Påhlman L, Kultima HG, Mayrhofer M, et al. Microsatellite instability and mutations in BRAF and KRAS are significant predictors of disseminated disease in colon cancer. BMC Cancer. 2015;15(1):1–11. [CrossRef]
- Sánchez M, Monteagudo F, Rodríguez M, García G. Marcadores inmunohistoquímicos de inestabilidad microsatelital para latipificación del cáncer colorrectal. Aspectos claves para la interpretaciónpor el patólogo. 2022;23(4). Available from: https://orcid.org/0000-0002-5149-2450.
- Fan JZ, Wang GF, Cheng X Bin, Dong ZH, Chen X, Deng YJ, et al. Relationship between mismatch repair protein, RAS, BRAF, PIK3CA gene expression and clinicopathological characteristics in elderly colorectal cancer patients. World J Clin Cases. 2021;9(11):2458–68.
- Remonatto G, Ferreira Salles Pilar E, De-Paris F, Schaefer PG, Kliemann LM. Integrated molecular profiling of RAS, BRAF mutations, and mismatch repair status in advanced colorectal carcinoma: insights from gender and tumor laterality. J Gastrointest Oncol [Internet]. 2024 Aug 31;15(4):1580–91. Available from: http://www.ncbi.nlm.nih.gov/pubmed/39279928. [CrossRef]
- Sinicrope FA. Evaluating the Combination of Microsatellite Instability and Mutation in BRAF as Prognostic Factors for Patients With Colorectal Cancer. Clin Gastroenterol Hepatol. 2019;17(3):391–4. [CrossRef]
- Salem ME, Bodor JN, Puccini A, Xiu J, Goldberg RM, Grothey A, et al. Relationship between MLH1, PMS2, MSH2 and MSH6 gene-specific alterations and tumor mutational burden in 1057 microsatellite instability-high solid tumors. Int J Cancer. 2020;147(10):2948–56.
- Parsons MT, Buchanan DD, Thompson B, Young JP, Spurdle AB. Correlation of tumour BRAF mutations and MLH1 methylation with germline mismatch repair (MMR) gene mutation status: a literature review assessing utility of tumour features for MMR variant classification. J Med Genet [Internet]. 2012 Mar;49(3):151–7. Available from: http://www.ncbi.nlm.nih.gov/pubmed/22368298. [CrossRef]
- Bläker H, Alwers E, Arnold A, Herpel E, Tagscherer KE, Roth W, et al. The Association Between Mutations in BRAF and Colorectal Cancer-Specific Survival Depends on Microsatellite Status and Tumor Stage. Clin Gastroenterol Hepatol [Internet]. 2019 Feb;17(3):455-462.e6. Available from: http://www.ncbi.nlm.nih.gov/pubmed/29660527. [CrossRef]
- Kopetz S, Grothey A, Yaeger R, Van Cutsem E, Desai J, Yoshino T, et al. Encorafenib, Binimetinib, and Cetuximab in BRAF V600E-Mutated Colorectal Cancer. N Engl J Med [Internet]. 2019 Oct 24;381(17):1632–43. Available from: http://www.ncbi.nlm.nih.gov/pubmed/31566309.
- Ye JX, Liu Y, Qin Y, Zhong HH, Yi WN, Shi XY. KRAS and BRAF gene mutations and DNA mismatch repair status in Chinese colorectal carcinoma patients. World J Gastroenterol. 2015;21(5):1595–605.
- Rencuzogullari A, Bisgin A, Erdogan KE, Gumus S, Yalav O, Boga I, et al. Site specific genetic differences in colorectal cancer via Next-Generation-Sequencing using a multigene panel. Ann Ital Chir [Internet]. 2023;94:605–11. Available from: http://www.ncbi.nlm.nih.gov/pubmed/38131395.
- Ashish S, Raj M. Importance of Early Next-Generation Sequencing in Microsatellite Unstable Colon Cancer With a High Tumor Mutation Burden. Cureus. 2022;2022(3):4–8. [CrossRef]
- Hernando-Calvo A, Nguyen P, Bedard PL, Chan KKW, Saleh RR, Weymann D, et al. Impact on costs and outcomes of multi-gene panel testing for advanced solid malignancies: a cost-consequence analysis using linked administrative data. eClinicalMedicine. 2024;69:1–12. [CrossRef]

| MSI | |||
|---|---|---|---|
| KRAS | Stable | Inestable | Total |
| Wild-Type | 20 | 5 | 25 |
| Mutated | 17 | 0 | 17 |
| Total | 37 | 5 | 42 |
| Value | Gl | p | |
|---|---|---|---|
| X2 | 3.86 | 1 | 0.049 |
| Fisher’s Exact Test p-value | 0.070 | ||
| N | 42 |
| 95% Confidence Intervals | |||
|---|---|---|---|
| Value | Lower | Upper | |
| Odds Ratio | 0.1061 | 0.00549 | 2.06 |
| Relative Risk | 0.8002 | 0.658 | 0.973 |
| MSI | |||
|---|---|---|---|
| NRAS | Stable | Inestable | Total |
| Wild-Type | 34 | 5 | 39 |
| Mutated | 3 | 0 | 3 |
| Total | 37 | 5 | 42 |
| Value | Gl | p | |
|---|---|---|---|
| X2 | 0.437 | 1 | 0.509 |
| Fisher’s Exact Test p-value | 1.000 | ||
| N | 42 |
| 95% Confidence Intervals | |||
|---|---|---|---|
| Value | Lower | Upper | |
| Odds Ratio | 0.8961 | 0.0405 | 19.8 |
| Relative Risk | 0.8722 | 0.773 | 0.983 |
| MSI | |||
|---|---|---|---|
| BRAF | Stable | Inestable | Total |
| Wild-Type | 30 | 2 | 32 |
| Mutated | 7 | 3 | 10 |
| Total | 37 | 5 | 42 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




