Submitted:
09 May 2025
Posted:
12 May 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Overview of Small Non-Coding RNA in Bacteria
2.1. Types and Structures of sRNA
2.2. RNA Chaperones
2.3. 6S RNA
2.4. Orthogonality of sRNA
3. Regulatory Mechanisms of sRNAs in Bacteria
3.1. Anti-Sense or Cis-Encoded sRNA
3.2. Trans-Encoded sRNA
4. Role of sRNA in Regulation of Antimicrobial Resistant Bacteria
4.1. Methicillin-Resistant Staphylococcus aureus
4.2. Multidrug-Resistant Pseudomonas aeruginosa
4.3. Extended-Spectrum β-Lactamase (ESBL)-Producing Enterobacterales
4.4. Carbapenem-Resistant Acinetobacter baumannii
4.5. Vancomycin Resistance Enterococcus faecium and E. faecalis
5. High-Throughput Technologies for Identification of sRNA
6. Therapeutic Strategies of sRNA
6.1. Stability of sRNA
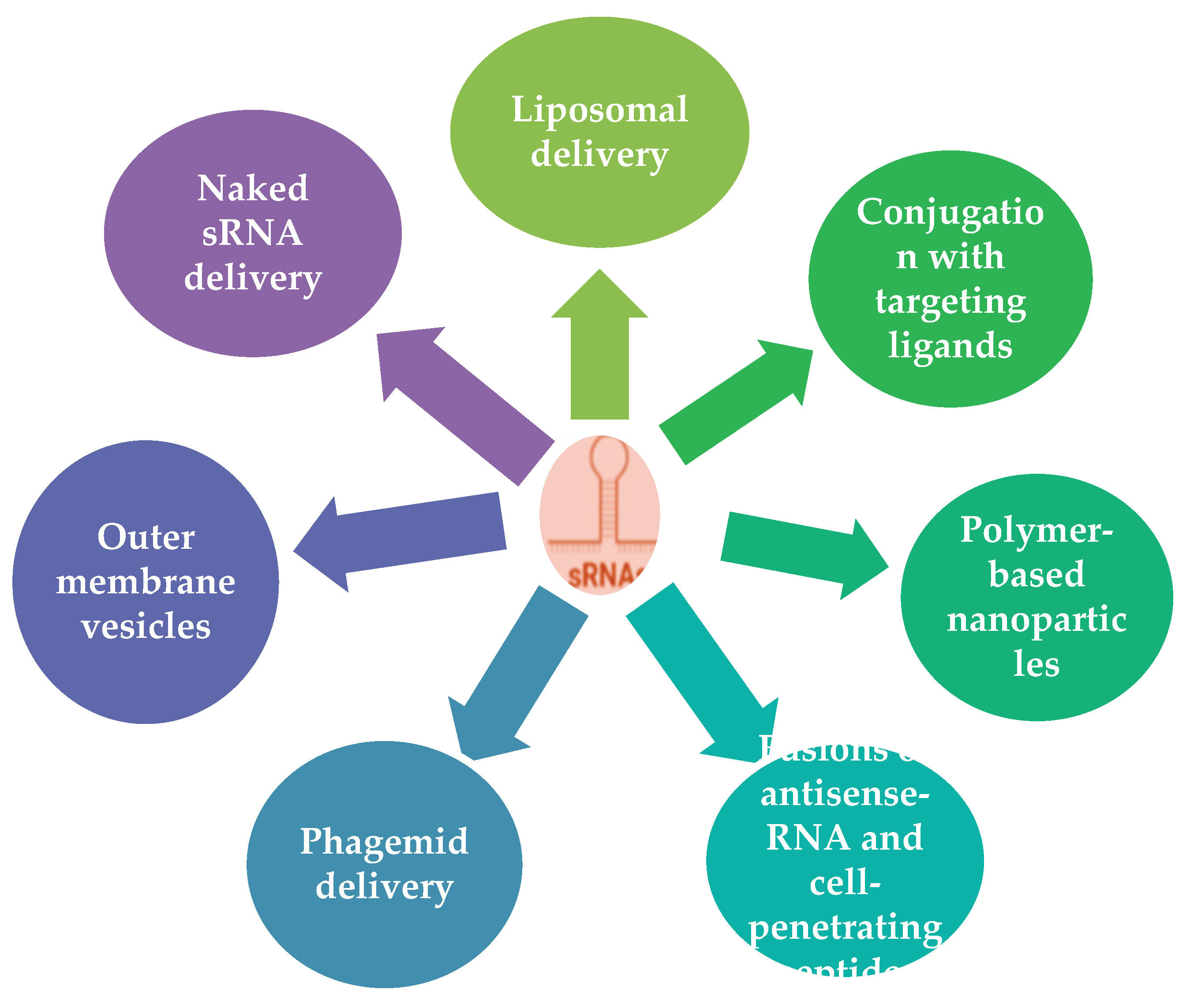
6.2. sRNA Delivery to Target Cell
7. Advantage and Disadvantage of sRNAs Drug
8. Conclusion and Future Perspective
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Conflicts of Interest
References
- 1. Parmeciano Di Noto, Gisela, María Carolina Molina, and Cecilia Quiroga. Insights into non-coding RNAs as novel antimicrobial drugs. Front Genet. 2019, 10, 57. [Google Scholar] [CrossRef] [PubMed]
- Murray, C.J.; Ikuta, K.S.; Sharara, F.; Swetschinski, L.; Aguilar, G.R.; Gray, A.; Han, C.; Bisignano, C.; Rao, P.; Wool, E.; Johnson, S.C. Global burden of bacterial antimicrobial resistance in 2019: A systematic analysis. The Lancet. 2022, 399, 629–655. [Google Scholar] [CrossRef]
- Ahmed, S.K.; Hussein, S.; Qurbani, K.; Ibrahim, R.H.; Fareeq, A.; Mahmood, K.A.; Mohamed, M.G. Antimicrobial resistance: Impacts, challenges, and future prospects. Journal of Medicine, Surgery, and Public Health. 2024, 2, 100081. [Google Scholar] [CrossRef]
- Cattoir, V.; Felden, B. Future antibacterial strategies: From basic concepts to clinical challenges. J. Infect. Dis. 2019, 220, 350–360. [Google Scholar] [CrossRef]
- Maxwell, A.; Bush, N.G.; Germe, T.; McKie, S.J. Non-quinolone topoisomerase inhibitors. Antimicrobial Resistance in the 21st Century. 2018, 593–618. [Google Scholar]
- Felden, B.; Cattoir, V. Bacterial adaptation to antibiotics through regulatory RNAs. Antimicrob. Agents Chemother. 2018, 62, 10–128. [Google Scholar] [CrossRef]
- Henderson, C.A.; Vincent, H.A.; Callaghan, A.J. Reprogramming gene expression by targeting RNA-based interactions: A novel pipeline utilizing RNA array technology. ACS Synthetic Biology. 2021, 10, 1847–1858. [Google Scholar] [CrossRef] [PubMed]
- Borgmann, J.; Schäkermann, S.; Bandow, J.E.; Narberhaus, F. A small regulatory RNA controls cell wall biosynthesis and antibiotic resistance. MBio 2018, 9, 10–128. [Google Scholar] [CrossRef]
- Hoyos, M.; Huber, M.; Förstner, K.U.; Papenfort, K. Gene autoregulation by 3’UTR-derived bacterial small RNAs. eLife 2020, 9, e58836. [Google Scholar] [CrossRef]
- Djapgne, L.; Oglesby, A.G. Impacts of small RNAs and their chaperones on bacterial pathogenicity. Front. Cell. Infect. Microbiol. 2021, 11. [Google Scholar] [CrossRef]
- Chan, H.; Ho, J.; Liu, X.; Zhang, L.; Wong, S.H.; Chan, M.T.; Wu, W.K. Potential and use of bacterial small RNAs to combat drug resistance: A systematic review. Infect Drug Resist. 2017, 521–532. [Google Scholar] [CrossRef]
- Fu, H.; Elena, R.C.; Marquez, P.H. The roles of small RNAs: Insights from bacterial quorum sensing. ExRNA 2019, 1, 1–8. [Google Scholar] [CrossRef]
- Lalaouna, D.; Eyraud, A.; Chabelskaya, S.; Felden, B.; Masse, E. Regulatory RNAs involved in bacterial antibiotic resistance. PLoS Pathog. 2014, 10, e1004299. [Google Scholar] [CrossRef] [PubMed]
- Svensson, S.L.; Sharma, C.M. Small RNAs in bacterial virulence and communication. Virulence Mechanisms of Bacterial Pathogens. 5th edition. 2016, 169–212. [Google Scholar]
- Harris, J.F.; Micheva-Viteva, S.; Li, N.; Hong-Geller, E. Small RNA-mediated regulation of host–pathogen interactions. Virulence. 2013, 4, 785–795. [Google Scholar] [CrossRef]
- Soltani-Fard, E.; Taghvimi, S.; Abedi Kichi, Z.; Weber, C.; Shabaninejad, Z.; Taheri-Anganeh, M.; et al. Insights into the Function of Regulatory RNAs in Bacteria and Archaea. J Transl Med. 2021, 1, 403–423. [Google Scholar] [CrossRef]
- Dutta, T.; Srivastava, S. Small RNA-mediated regulation in bacteria: A growing palette of diverse mechanisms. Gene. 2018, 656, 60–72. [Google Scholar] [CrossRef] [PubMed]
- Kavita, K.; de Mets, F.; Gottesman, S. New aspects of RNA-based regulation by Hfq and its partner sRNAs. Current opinion in microbiology. 2018, 42, 53–61. [Google Scholar] [CrossRef]
- De Lay, N.; Schu, D.J.; Gottesman, S. Bacterial small RNA-based negative regulation: Hfq and its accomplices. J Biol Chem. 2013, 288, 7996–8003. [Google Scholar] [CrossRef]
- Watkins, D.; Arya, D.P. Regulatory roles of small RNAs in prokaryotes: Parallels and contrast with eukaryotic mirna. Non-coding RNA Investig. 2019, 3, 21037. [Google Scholar] [CrossRef]
- Dendooven, T.; Sinha, D.; Roeselová, A.; Cameron, T.A.; De Lay, N.R.; Luisi, B.F.; Bandyra, K.J. A cooperative PNPase-Hfq-RNA carrier complex facilitates bacterial riboregulation. Molecular Cell. 2021, 81, 2901–2913. [Google Scholar] [CrossRef]
- Westermann, A.J.; Venturini, E.; Sellin, M.E.; Förstner, K.U.; Hardt, W.D.; Vogel, J. The major RNA-binding protein ProQ impacts virulence gene expression in Salmonella enterica serovar Typhimurium. MBio. 2019, 10, 10–128. [Google Scholar] [CrossRef]
- Gottesman, S.; Storz, G. Bacterial small RNA regulators: Versatile roles and rapidly evolving variations. Cold Spring Harbor perspectives in biology. 2011, 3, a003798. [Google Scholar] [CrossRef] [PubMed]
- Christopoulou, N.; Granneman, S. The role of RNA-binding proteins in mediating adaptive responses in Gram-positive bacteria. The FEBS journal. 2022, 289, 1746–1764. [Google Scholar] [CrossRef] [PubMed]
- Matera, G.; Altuvia, Y.; Gerovac, M.; El Mouali, Y.; Margalit, H.; Vogel, J. Global RNA interactome of Salmonella discovers a 5′ UTR sponge for the MicF small RNA that connects membrane permeability to transport capacity. Mol. Cell. 2022, 82, 629–644. [Google Scholar] [CrossRef] [PubMed]
- Warrier, I.; Hicks, L.D.; Battisti, J.M.; Raghavan, R.; Minnick, M.F. Identification of novel small RNAs and characterization of the 6S RNA of Coxiella burnetii. PLoS ONE. 2014, 9, 1–13. [Google Scholar] [CrossRef]
- Klein, G.; Raina, S. Small regulatory bacterial regulating the envelope stress response. Biochem. Soc. Trans. 2017, 45, 417–425. [Google Scholar] [CrossRef]
- Sakai, Y.; Abe, K.; Nakashima, S.; Yoshida, W.; Ferri, S.; Sode, K.; Ikebukuro, K. Improving the gene-regulation ability of small RNAs by scaffold engineering in Escherichia coli. ACS Synth. Biol. 2014, 3, 152–162. [Google Scholar] [CrossRef]
- Kobayashi, Y.; Tian, S.; Ui-Tei, K. The siRNA off-target effect is determined by base-pairing stabilities of two different regions with opposite effects. Genes. 2022, 13, 319. [Google Scholar] [CrossRef]
- Balasubramanian, D.; Vanderpool, C.K. New developments in post-transcriptional regulation of operons by small RNAs. RNA biology. 2013, 10, 337–341. [Google Scholar] [CrossRef]
- Hoekzema, M.; Romilly, C.; Holmqvist, E.; Wagner, E.G.H. Hfq-dependent mRNA unfolding promotes sRNA-based inhibition of translation. EMBO J. 2019, 38, e101199. [Google Scholar] [CrossRef] [PubMed]
- Villa, J.K.; Su, Y.; Contreras, L.M.; Hammond, M.C. Synthetic Biology of Small RNAs and Riboswitches. Microbiol Spectr. 2018, 6, 10. [Google Scholar]
- Ignatov, D.; Vaitkevicius, K.; Durand, S.; Cahoon, L.; Sandberg, S.S.; Liu, X.; Kallipolitis, B.H.; Rydén, P.; Freitag, N.; Condon, C.; Johansson, J. An mRNA-mRNA interaction couples expression of a virulence factor and its chaperone in Listeria monocytogenes. Cell reports. 2020, 30, 4027–4040. [Google Scholar] [CrossRef]
- Sousa, J.P.; Silva, A.F.; Arraiano, C.M.; Andrade, J.M. Bacterial Small RNAs: Diversity of Structure and Function. InRNA Structure and Function. Cham: Springer International Publishing, 2023, 259-277.
- Storz, G.; Vogel, J.; Wassermann, K.M. Regulation by small RNAs in bacteria: Expanding frontiers. Molecular cell. 2011, 43, 880–891. [Google Scholar] [CrossRef]
- Waters, L.S.; Storz, G. Regulatory RNAs in bacteria. Cell. 2009, 136, 615–628. [Google Scholar] [CrossRef] [PubMed]
- González Plaza, J.J. Small RNAs as fundamental players in the transference of Information during Bacterial Infectious Diseases. Front. Mol. Biosci. 2020, 7. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Ying, X.; Lu, Q.; Chen, L. Predicting sRNAs and their targets in bacteria. Genomics Proteomics and Bioinformatics 2012, 10, 276–284. [Google Scholar] [CrossRef]
- Mulani, M.S.; Kamble, E.E.; Kumkar, S.N.; Tawre, M.S.; Pardesi, K.R. Emerging strategies to combat ESKAPE pathogens in the era of antimicrobial resistance: A review. Front. Microbiol. 2019, 10. [Google Scholar] [CrossRef]
- Dersch, P.; Khan, M.A.; Mühlen, S.; Görke, B. Roles of regulatory RNAs for antibiotic resistance in bacteria and their potential value as novel drug targets. Front. Microbiol. 2017, 8, 803. [Google Scholar] [CrossRef]
- Dar, D.; Sorek, R. Regulation antibiotics resist distance by non-coding RNAs in bacteria. Curr. Opin. Microbiol. 2017, 36, 111–117. [Google Scholar] [CrossRef]
- Mishra, B.M.A.K.; Easwaran, N.; Gothandam, K.M. Staphylococcus aureus and Virulence-Related Small RNA. Insights into Drug Resistance in Staphylococcus aureus. IntechOpen. 2021, 41. [Google Scholar]
- Mediati, D.G.; Wu, S.; Wu, W.; Tree, J.J. Networks of resistance: Small RNA control of antibiotic resistance. Trends Genet. 2021, 37, 35–45. [Google Scholar] [CrossRef] [PubMed]
- Eyraud, A.; Tattevin, P.; Chabelskaya, S.; Felden, B. A small RNA controls a protein regulator involved in antibiotic resistance in Staphylococcus aureus. Nucleic Acids Res. 2014, 42, 4892–4905. [Google Scholar] [CrossRef]
- Pusic, P.; Sonnleitner, E.; Bläsi, U. Specific and global RNA regulators in Pseudomonas aeruginosa. Int. J. Mol. Sci. 2021, 22. [Google Scholar] [CrossRef]
- Coleman, S.R.; Smith, M.L.; Spicer, V.; Lao, Y.; Mookherjee, N.; Hancock, R.E. Overexpression of the small RNA PA0805. 1 in Pseudomonas aeruginosa modulates the expression of a large set of genes and proteins, resulting in altered motility, cytotoxicity, and tobramycin resistance. Msystems. 2020, 5, 10–128. [Google Scholar] [CrossRef]
- Law, C.O.K.; Huang, C.; Pan, Q.; Lee, J.; Hao, Q.; Chan, T.F.; Lo, N.W.S.; et al. A Small RNA transforms the multidrug resistance of Pseudomonas aeruginosa to drug susceptibility. Molecular Therapy Nucleic Acids 2019, 16, 218–228. [Google Scholar] [CrossRef]
- Kröger, C.; MacKenzie, K.D.; Alshabib, E.Y.; Kirzinger, M.W.B.; Suchan, D.M.; Chao, T.; et al. The primary transcriptome, small RNAs, and regulation of antimicrobial resistance in Acinetobacter baumannii ATCC 17978. Nucleic Acids Res. 2018, 46, 9684–9698. [Google Scholar] [CrossRef]
- Klein, G.; Raina, S. Small regulatory bacterial regulating the envelope stress response. Biochem. Soc. Trans. 2017, 45, 417–425. [Google Scholar] [CrossRef] [PubMed]
- Montzka Wassarman, K.; Zhang, A.; Storz, G. Small RNAs in Escherichia coli. Trends.Microbiol 1999, 7, 37–45. [Google Scholar] [CrossRef]
- Andreassen, P.R.; Pettersen, J.S.; Szczerba, M.; Valentin-hansen, P.; Møller-jensen, J.; Jørgensen, M.G. sRNA-dependent control of curli biosynthesis in Escherichia coli: McaS directs endonucleolytic cleavage of csgD, m.R.N.A. Nucleic Acids Res. 2018, 46, 6746–6760. [Google Scholar] [CrossRef]
- Huang, S.H.; Wang, C.K.; Peng, H.L.; Wu, C.C.; Chen, Y.T.; Hong, Y.M.; Lin, C.T. Role of the small RNA RyhB in the fur regulon in mediating the capsular polysaccharide biosynthesis and iron acquisition systems in Klebsiella pneumoniae. BMC Microbiol. 2012, 12, 148. [Google Scholar] [CrossRef] [PubMed]
- Kyriakidis, I.; Vasileiou, E.; Pana, Z.D.; Tragiannidis, A. Acinetobacter baumannii antibiotic resistance mechanisms. Pathogens. 2021, 10, 1–31. [Google Scholar] [CrossRef] [PubMed]
- Allen, J.L.; Tomlinson, B.R.; Casella, L.G.; Shaw, L.N. Regulatory networkswork important for the survival of Acinetobacter baumannii within the host. Curr. Opin. Microbiol. 2020, 55, 74–80. [Google Scholar] [CrossRef]
- Sharma, R.; Arya, S.; Patil, S.D.; Sharma, A.; Jain, P.K.; Navani, N.K.; Pathania, R. Identification of novel regulatory small RNAs in Acinetobacter baumannii. PLoS ONE 2014, 9. [Google Scholar] [CrossRef] [PubMed]
- Ahmed, M.O.; Baptiste, K.E. Vancomycin-Resistant Enterococci: A Review of Antimicrobial Resistance Mechanisms and Perspectives of Human and Animal Health. Microb. Drug Resist. 2018, 24, 590–606. [Google Scholar] [CrossRef]
- Sinel, C.; Augagneur, Y.; Sassi, M.; Bronsard, J.; Cacaci, M.; Guérin, F.; et al. Small RNAs in vancomycin-resistant Enterococcus faecium involved in daptomycin response and resistance. Scientific Reports 2017, 7, 11067. [Google Scholar] [CrossRef]
- Diallo, I. Provost PRNA-sequencing analyses of small bacterial RNAs their emergence host pathogen- factors in host pathogen interactions. Int J. Mol. Sci. 2020, 21. [Google Scholar] [CrossRef]
- Barman, R.K.; Mukhopadhyay, A.; Das, S. An improved method for identification of small non-coding RNAs in bacteria using support vector machine. Sci. Rep. 2017, 7, 1–8. [Google Scholar] [CrossRef]
- Sridhar, J.; Narmada, S.R.; Sabarinathan, R.; Ou, H.Y.; Deng, Z.; Sekar, K.; et al. sRNAscanner: A computational tool for intergenic small RNA detection in bacterial genomes. PLoS ONE 2010, 5. [Google Scholar] [CrossRef]
- Lott, S.C.; Schäfer, R.A.; Mann, M.; Backofen, R.; Hess, W.R.; Voß, B.; Georg, J. GLASSgo - Automated and reliable detection of sRNA homologs from a single input sequence. Front. Genet. 2018, 9, 1–12. [Google Scholar] [CrossRef]
- Hu, B.; Zhong, L.; Weng, Y.; Peng, L.; Huang, Y.; Zhao, Y.; Liang, X.J. Therapeutic siRNA: State of the art. Signal Transduct Target Ther. 2020, 5, 101. [Google Scholar] [CrossRef] [PubMed]
- Pandey, R.P.; Mukherjee, R.; Chang, C.M. Emerging concern with imminent therapeutic strategies for treating resistance in biofilm. Antibiotics 2022, 11, 476. [Google Scholar] [CrossRef]
- Wang, Y.; Yin, G.; Weng, H.; Zhang, L.; Du, G.; Chen, J.; Kang, Z. Gene knockdown by structure defined single-stem loop small non-coding RNAs with programmable regulatory activities. Synth. Syst. Biotechnol. 2023, 8, 86–96. [Google Scholar] [CrossRef]
- Wang, Y.; Zhang, Z.; Luo, J.; Han, X.; Wei, Y.; Wei, X. mRNA vaccine: A potential therapeutic strategy. Mol. Cancer. 2021, 20, 33. [Google Scholar] [CrossRef] [PubMed]
- Yu, A.M.; Tu, M.J. Deliver the promise: RNAs as a new class of molecular entities for therapy and vaccination. Pharmacol Ther. 2022, 230, 107967. [Google Scholar] [CrossRef]
- Williford, J.M.; Wu, J.; Ren, Y.; Archang, M.M.; Leong, K.W.; Mao, H.Q. Recent advances in nanoparticle-mediated siRNA delivery. Annu. Rev. Biomed. Eng. 2014, 16, 347–370. [Google Scholar] [CrossRef] [PubMed]
- Guo, S.; Fu, D.; Utupova, A.; Sun, D.; Zhou, M.; Jin, Z.; Zhao, K. Applications of polymer-based nanoparticles in vaccine field. Nanotechnol. Rev. 2019, 8, 143–155. [Google Scholar] [CrossRef]
- Wang, L.; Wang, N.; Zhang, W.; Cheng, X.; Yan, Z.; Shao, G.; Wang, X.; Wang, R.; Fu, C. Therapeutic peptides: Current applications and future directions. Signal Transduct Target Ther. 2022, 7, 48. [Google Scholar] [CrossRef] [PubMed]
- Koeppen, K.; Hampton, T.H.; Jarek, M.; Scharfe, M.; Gerber, S.A.; Mielcarz, D.W.; Demers, E.G.; Dolben, E.L.; Hammond, J.H.; Hogan, D.A.; Stanton, B.A. A novel mechanism of host-pathogen interaction through sRNA in bacterial outer membrane vesicles. PLoS Pathog. 2016, 12, e1005672. [Google Scholar] [CrossRef]
- Suzuki, Y.; Ishimoto, T.; Fujita, S.; Kiryu, S.; Wada, M.; Akatsuka, T.; et al. Antimicrobial antisense RNA delivery to F-pili producing multidrug-resistant bacteria via a genetically engineered bacteriophage. Biochem. Biophys. Res. Commun. 2020, 530, 533–540. [Google Scholar] [CrossRef]
- Libis, V.K.; Bernheim, A.G.; Basier, C.; Jaramillo-Riveri, S.; Deyell, M.; Aghoghogbe, I.; Atanaskovic, I.; Bencherif, A.C.; Benony, M.; Koutsoubelis, N.; Löchner, A.C. Silencing of antibiotic resistance in E. coli with engineered phage bearing small regulatory RNAs. ACS Synth. Biol. 2014, 3, 1003–1006. [Google Scholar] [CrossRef] [PubMed]
- Sadeghian, I.; Heidari, R.; Sadeghian, S.; Raee, M.J.; Negahdaripour, M. Potential of cell-penetrating peptides (CPPs) in delivery of antiviral therapeutics and vaccines. European Journal of Pharmaceutical Sciences. 2022, 169, 106094. [Google Scholar] [CrossRef] [PubMed]
- Eriksson, M.; Nielsen, P.E.; Good, L. Cell Permeabilization and Uptake of Antisense Peptide-Peptide Nucleic Acid (PNA) into Escherichia Coli. J. Biol. Chem. 2002, 277, 7144–7147. [Google Scholar] [CrossRef] [PubMed]
- Gooding, M.; Malhotra, M.; Evans, J.C.; Darcy, R.; O’Driscoll, C.M. Oligonucleotide conjugates–Candidates for gene silencing therapeutics. Eur J Pharm Biopharm. 2016, 107, 321–340. [Google Scholar] [CrossRef]
- Nielsen, P.E.; Shiraishi, T. Peptide nucleic acid (PNA) cell penetrating peptide (CPP) conjugates as carriers for cellular delivery of antisense oligomers. Artificial Dna Pna Xna 2011, 2, 90–99. [Google Scholar] [CrossRef]
- Yu, A.M.; Choi, Y.H.; Tu, M.J. RNA drugs and RNA targets for small molecules: Principles, progress, and challenges. Pharmacol Res. 2020, 72, 862–898. [Google Scholar] [CrossRef]
- Cattoir, V.; Felden, B. Future Antibacterial Strategies: From Basic Concepts to Clinical Challenges. J Infect Dis. 2019, 220, 350–360. [Google Scholar] [CrossRef]
- Wang, L.; Wang, N.; Zhang, W.; Cheng, X.; Yan, Z.; Shao, G.; Wang, X.; Wang, R.; Fu, C. Therapeutic peptides: Current applications and future directions. Signal Transduct Target Ther. 2022, 7, 48. [Google Scholar] [CrossRef]
| challenges of maintaining the orthogonally of sRNAs | Mechanism | Bacterial strategy |
|---|---|---|
| Competing Interactions | sRNAs can bind to multiple target mRNAs, potentially leading to unintended regulation of native genes. This can disrupt essential cellular processes and affect overall cell function. | To mitigate this, synthetic biologists can design sRNAs with unique binding sites that do not match any native mRNA sequences. Bacteria like E. coli have been engineered with orthogonal sRNA systems that utilize modified or synthetic RNA components to ensure specificity. |
| Cellular Context and Concentration | The effectiveness of sRNA can vary based on their concentration and the presence of other competing RNAs within the cell, leading to unpredictable outcomes. | Bacteria can be engineered to express sRNAs at controlled levels, using tightly regulated promoters to ensure that they remain at optimal concentrations. Synthetic systems can also incorporate feedback mechanisms to modulate sRNA levels in response to cellular conditions. |
| Degradation and Stability | sRNAs are subject to degradation by RNases, which can limit their availability and functionality. This instability can lead to fluctuations in their activity, compromising orthogonality. | To enhance stability, synthetic sRNAs can be engineered with modified nucleotides or specific secondary structures that resist degradation. Additionally, incorporating sequences that prevent rapid degradation can prolong sRNA half-lives. |
| Context-Dependent Regulation | The regulatory effects of sRNAs can depend on the specific cellular context, including the presence of ribosomal binding sites and other regulatory elements, which may lead to unwanted interactions. | Engineers can use context-specific regulatory elements, such as synthetic riboswitches, that only activate under certain conditions. This ensures that sRNAs only exert their effects in predefined scenarios, maintaining orthogonality. |
| Evolutionary Pressure and Adaptation: | Bacterial populations can rapidly evolve, leading to mutations that may allow for cross-reactivity between engineered sRNAs and native targets. | To counter this, researchers can use directed evolution techniques to screen for and select for sRNAs that maintain their function without interacting with native targets. Incorporating redundancy into the design, such as multiple orthogonal sRNAs targeting the same mRNA, can also enhance robustness. |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




