Submitted:
06 May 2025
Posted:
07 May 2025
Read the latest preprint version here
Abstract
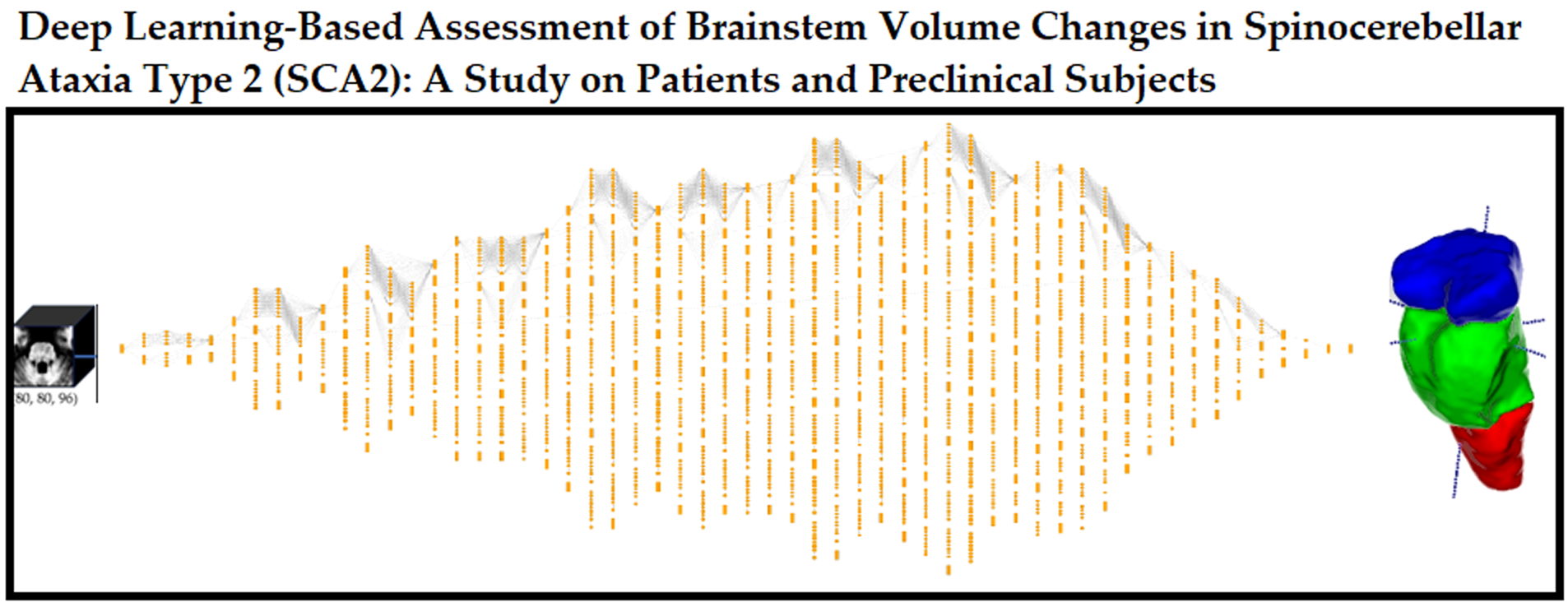
Keywords:
- Highlights
- What are the main findings?
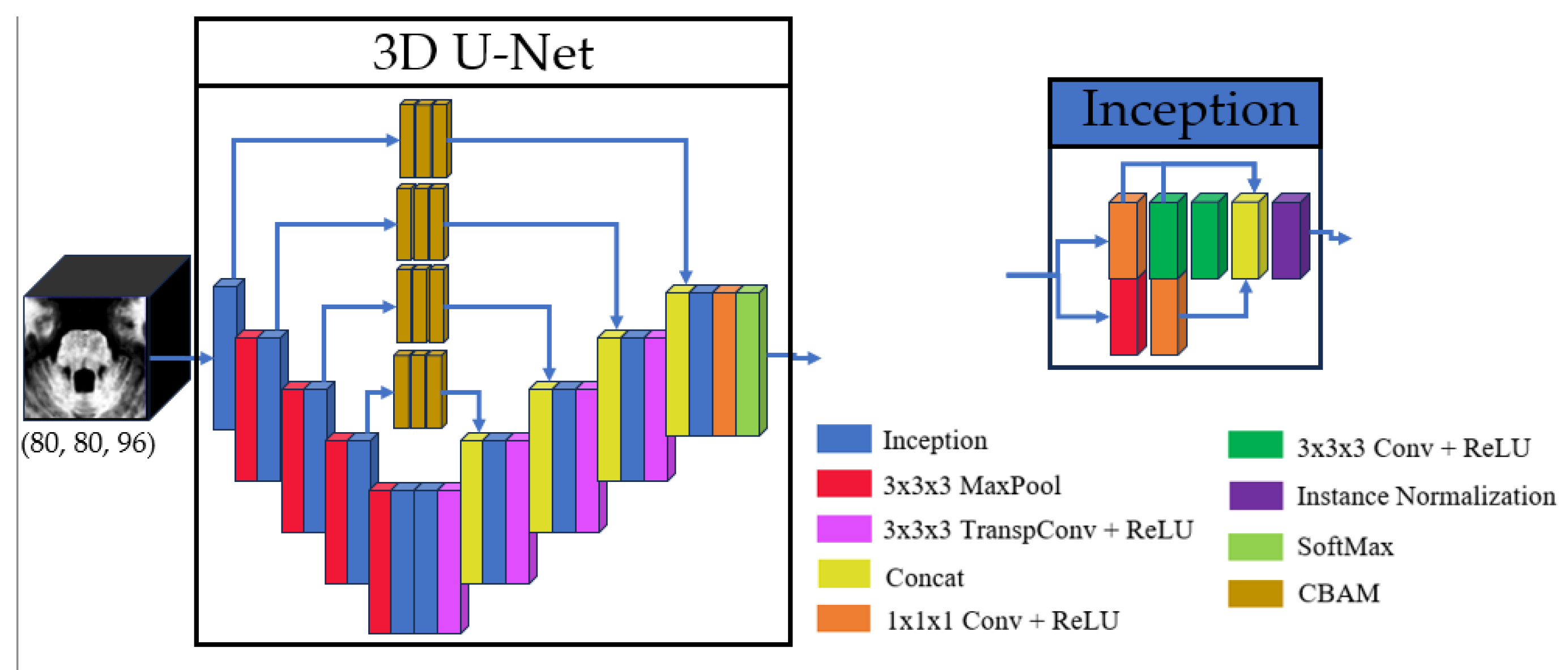
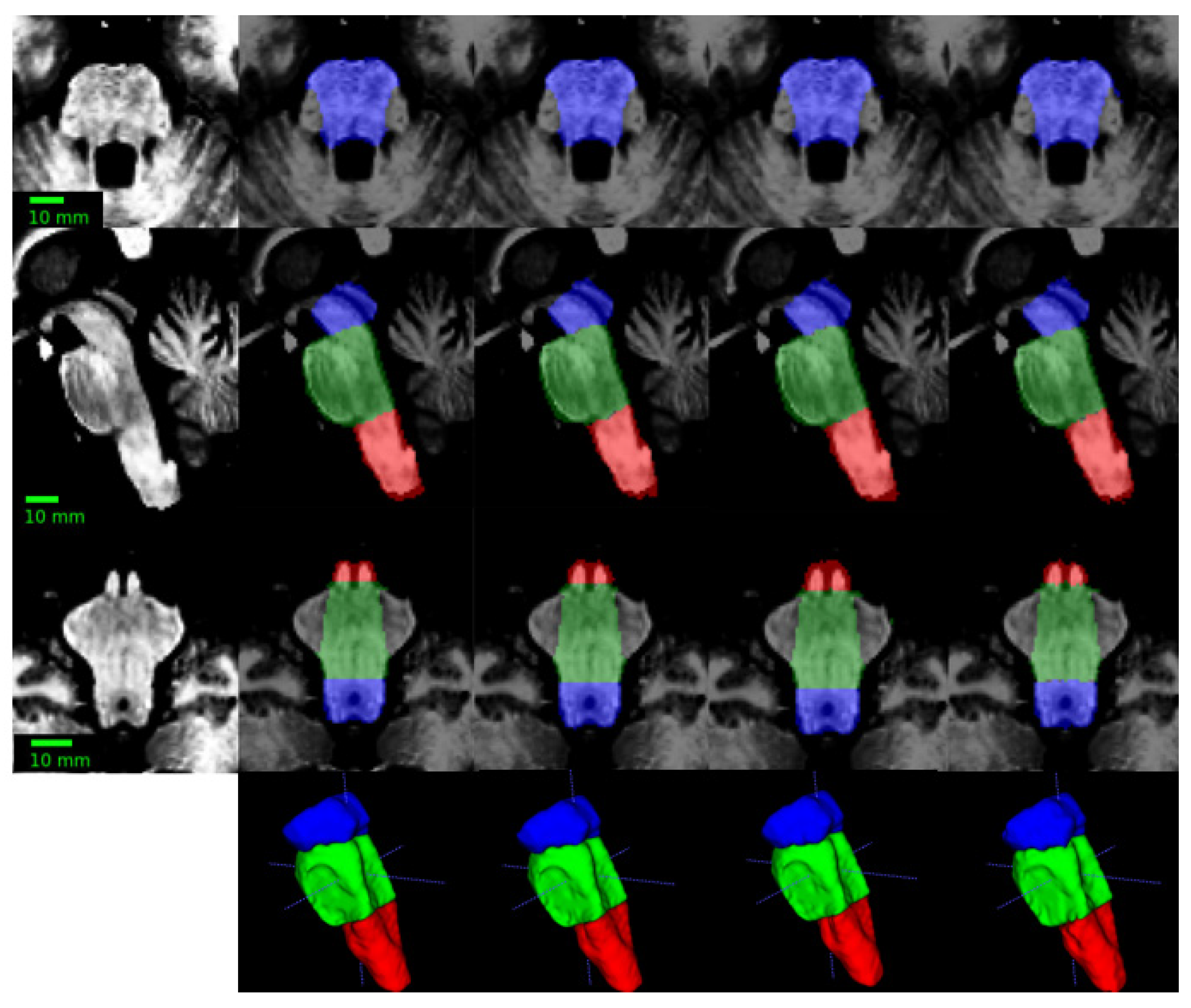
- Superior Segmentation Performance: The proposed modified U-Net architecture (with attention-enhanced skip connections and inception modules) significantly outperforms three comparative approaches in brainstem parcellation, achieving higher Dice scores across all substructures (medulla, pons, mesencephalon) and the whole brainstem.
- Volume Differences Across Groups: Automated segmentation reveals distinct volumetric patterns, with controls exhibiting larger volumes (whole brainstem: 1.62) compared to preclinical (1.49) and patient groups (1.12), suggesting potential atrophy linked to disease progression.
- What is the implication of the main finding?
- Clinical Utility: The method’s accuracy and robustness support its potential for precise brainstem assessment in neurodegenerative disorders, enabling earlier detection of structural changes (e.g., reduced medulla volume in patients: 0.26 vs. 0.31 in controls).
- Technical Advancements: The success of attention mechanisms and inception modules highlights their value for complex anatomical segmentation, paving the way for similar adaptations in other small-structure parcellation tasks.
1. Introduction
2. Materials and Methods
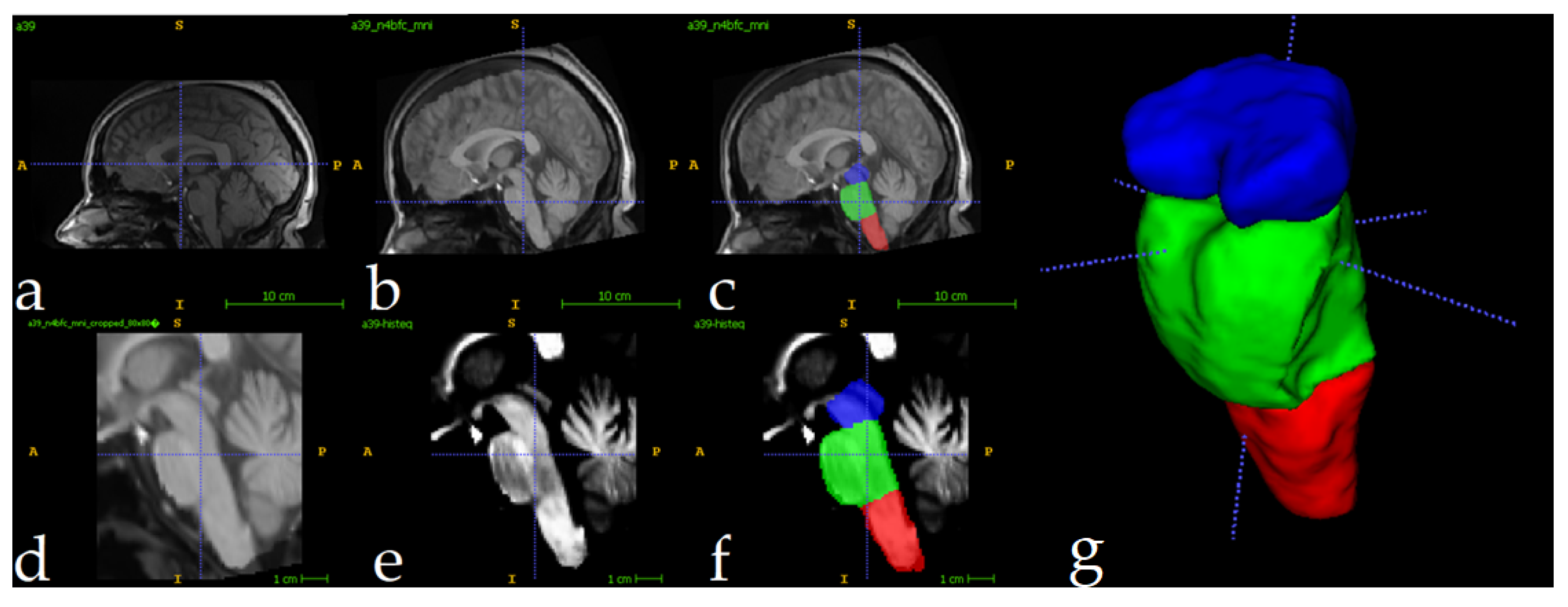
2.1. Image Preparation
2.2. Analysis Description
3. Results
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- K. Seidel, S. Siswanto, E.R.P. Brunt, W. Den Dunnen, H.W. Korf, U. Rüb, Brain pathology of spinocerebellar ataxias, Acta Neuropathol. 124 (2012) 1–21. [CrossRef]
- M. Mascalchi, S. Diciotti, M. Giannelli, A. Ginestroni, A. Soricelli, E. Nicolai, M. Aiello, C. Tessa, L. Galli, M.T. Dotti, S. Piacentini, Progression of Brain Atrophy in Spinocerebellar Ataxia Type 2 : A Longitudinal Tensor-Based Morphometry Study, PLoS One. 9 (2014). [CrossRef]
- C. Marzi, S. Ciulli, M. Giannelli, A. Ginestroni, C. Tessa, M. Mascalchi, S. Diciotti, Structural Complexity of the Cerebellum and Cerebral Cortex is Reduced in Spinocerebellar Ataxia Type 2, J. Neuroimaging. 28 (2018) 1–6. [CrossRef]
- A. Antenora, C. Rinaldi, A. Roca, C. Pane, M. Lieto, F. Saccà, S. Peluso, G. De Michele, A. Filla, The Multiple Faces of Spinocerebellar Ataxia type 2, Ann. Clin. Transl. Neurol. 4 (2017) 687–695. [CrossRef]
- L. Velázquez-Pérez, J. Medrano-Montero, R. Rodríguez-Labrada, N. Canales-Ochoa, J.C. Alí, F.J.C. Rodes, T.R. Graña, M.O.H. Oliver, R.A. Rodríguez, Y.D. Barrios, R.T. Vega, L.F. Angulo, Hereditary Ataxias in Cuba : A Nationwide Epidemiological and Clinical Study in 1001 Patients, (2020). [CrossRef]
- M. Mascalchi, Spinocerebellar ataxias, Neurol Sci. 29 (2008) 311–313. [CrossRef]
- S.D. Kumar, R.P. Chand, A.K. Gururaj, W.D. Jeans, CT features of olivopontocerebellar atrophy in children, Acta Radiol. 36 (1995) 593–596. [CrossRef]
- A.T. Meira, W.O. Arruda, S.E. Ono, A.D.C. Neto, S. Raskin, C.H. Camargo, H.A.G. Teive, Neuroradiological Findings in the Spinocerebellar Ataxias, Tremor and Other Hyperkinetic Movements. (2019) 1–8. [CrossRef]
- M. Mascalchi, A. Vella, Neuroimaging Applications in Chronic Ataxias, Int. Rev. Neurobiol. 143 (2018) 109–162. [CrossRef]
- X.A. Klaes, X.E. Reckziegel, X.M.C.F. Jr, X.T.J.R. Rezende, X.L.M. Vedolin, X.L.B. Jardim, X.J.A. Saute, MR Imaging in Spinocerebellar Ataxias : A Systematic Review, Am. J. f Neuroradiol. 37 (2016) 1405–1412. [CrossRef]
- C.R. Hernandez-Castillo, V. Galvez, R. Mercadillo, R. Diaz, A. Campos-Romo, J. Fernandez-Ruiz, Extensive white matter alterations and its correlations with ataxia severity in SCA 2 patients, PLoS One. 10 (2015) 1–10. [CrossRef]
- S. Döhlinger, T.K. Hauser, J. Borkert, A.R. Luft, J.B. Schulz, Magnetic resonance imaging in spinocerebellar ataxias, Cerebellum. 7 (2008) 204–214. [CrossRef]
- D. Tamuli, M. Kaur, T. Sethi, A. Singh, M. Faruq, A.K. Jaryal, A.K. Srivastava, S. Senthil, Cortical and Subcortical Brain Area Atrophy in SCA1 and SCA2 Patients in India : The Structural MRI Underpinnings and Correlative Insight Among the Atrophy and Disease Attributes, Neurol. India. 69 (2021) 1318–1325. [CrossRef]
- L. Peipei, L. Yang, G. Weihong, S. Xiaonan, Clinical Manifestation, Imaging, and Genotype Analysis of Two Pedigrees with Spinocerebellar Ataxia, Cell Biochem. Biophys. 61 (2011) 691–698. [CrossRef]
- H. Jacobi, T. Hauser, P. Giunti, C. Globas, P. Bauer, Spinocerebellar Ataxia Types 1 , 2 , 3 and 6 : the Clinical Spectrum of Ataxia and Morphometric Brainstem and Cerebellar Findings, Cerebellum. 11 (2012) 155–166. [CrossRef]
- T. Van Dijk, P. Barth, L. Reneman, B. Appelhof, F. Baas, B.T. Poll-the, A De Novo Missense Mutation in the Inositol 1 , 4 , 5-Triphosphate Receptor Type 1 Gene Causing Severe Pontine and Cerebellar Hypoplasia : Expanding the Phenotype of ITPR1 -Related Spinocerebellar Ataxia ’ s, Am. J. Med. Genet. (2016). [CrossRef]
- R. Della Nave, A. Ginestroni, C. Tessa, E. Salvatore, D. De Grandis, R. Plasmati, F. Salvi, G. De Michele, M.T. Dotti, S. Piacentini, M. Mascalchi, Brain white matter damage in SCA1 and SCA2. An in vivo study using voxel-based morphometry, histogram analysis of mean diffusivity and tract-based spatial statistics, Neuroimage. 43 (2008) 10–19. [CrossRef]
- L.S. Politi, S. Bianchi Marzoli, C. Godi, M. Panzeri, P. Ciasca, G. Brugnara, A. Castaldo, D. Di Bella, F. Taroni, L. Nanetti, C. Mariotti, MRI evidence of cerebellar and extraocular muscle atrophy differently contributing to eye movement abnormalities in SCA2 and SCA28 diseases, Investig. Ophthalmol. Vis. Sci. 57 (2016) 2714–2720. [CrossRef]
- M. Mascalchi, A. Vella, Neuroimaging biomarkers in SCA2 gene carriers, Int. J. Mol. Sci. 21 (2020) 1–14. [CrossRef]
- Y. LeCun, B. Boser, J.S. Denker, D. Henderson, R.E. Howard, W. Hubbard, L.D. Jackel, Backpropagation applied to digit recognition, Neural Comput. 1 (1989) 541–551. https://www.ics.uci.edu/~welling/teaching/273ASpring09/lecun-89e.pdf.
- M.D. Zeiler, R. Fergus, Visualizing and Understanding Convolutional Networks, Anal. Chem. Res. 12 (2014) 818–833. [CrossRef]
- B. Hariharan, P. Arbeláez, L. Bourdev, S. Maji, J. Malik, Semantic Contours from Inverse Detectors, in: Int. Conf. Comput. Vis., 2011: pp. 991–998. http://home.bharathh.info/pubs/pdfs/BharathICCV2011.pdf.
- R. Jaroensri, A. Zhao, G. Balakrishnan, D. Lo, J.D. Schmahmann, F. Durand, J. Guttag, A Video-Based Method for Automatically Rating Ataxia, in: Proc. Mach. Learn., 2017: pp. 1–13.
- C. Kawahara, C.J. Brown, S.P. Miller, B.G. Booth, V. Chau, R.E. Grunau, J.G. Zwicker, G. Hamarneh, BrainNetCNN : Convolutional Neural Networks for Brain Networks ; Towards Predicting Neurodevelopment, Neuroimage. 146 (2017) 1038–1049. [CrossRef]
- C. Stoean, R. Stoean, M. Atencia, M. Abdar, L. Velázquez-Pérez, A. Khosrabi, S. Nahavandi, U.R. Acharya, G. Joya, Automated Detection of Presymptomatic Conditions in Spinocerebellar Ataxia Type 2 Using Monte Carlo Dropout and Deep Neural Network Techniques with Electrooculogram Signals, Sensors. 20 (2020) 3032. [CrossRef]
- J. Dolz, C. Desrosiers, L. Wang, J. Yuan, D. Shen, I. Ben Ayed, Deep CNN ensembles and suggestive annotations for infant brain MRI segmentation, Comput. Med. Imaging Graph. 79 (2020) 101660. [CrossRef]
- K. Kamnitsas, C. Ledig, V.F.J. Newcombe, J.P. Simpson, A.D. Kane, D.K. Menon, D. Rueckert, B. Glocker, Efficient multi-scale 3D CNN with fully connected CRF for accurate brain lesion segmentation, Med. Image Anal. 36 (2016) 61–78. [CrossRef]
- B. Erden, N. Gamboa, S. Wood, 3D Convolutional Neural Network for Brain Tumor Segmentation, Stanford, 2017.
- O. Ronneberger, P. Fischer, T. Brox, U-net: Convolutional networks for biomedical image segmentation, Lect. Notes Comput. Sci. (Including Subser. Lect. Notes Artif. Intell. Lect. Notes Bioinformatics). 9351 (2015) 234–241. [CrossRef]
- B. Mansencal, B.D. De Senneville, V. Ta, V. Lepetit, AssemblyNet : A large ensemble of CNNs for 3D whole brain MRI segmentation, Neuroimage. 219 (2020) 117026. [CrossRef]
- I. Mecheter, M. Abbod, A. Amira, H. Zaidi, Deep Learning with Multiresolution Handcrafted Features for Brain MRI Segmentation, Artif. Intell. Med. 131 (2022) 102365. [CrossRef]
- J. Faber, D. Kügler, E. Bahrami, L.S. Heinz, D. Timmann, T.M. Ernst, K. Deike-Hofmann, T. Klockgether, B. van de Warrenburg, J. van Gaalen, K. Reetz, S. Romanzetti, G. Oz, J.M. Joers, J. Diedrichsen, P. Giunti, H. Garcia-Moreno, H. Jacobi, J. Jende, J. de Vries, M. Povazan, P.B. Barker, K.M. Steiner, J. Krahe, M. Reuter, CerebNet: A fast and reliable deep-learning pipeline for detailed cerebellum sub-segmentation, Neuroimage. 264 (2022) 119703. [CrossRef]
- P. Moeskops, M. Veta, M.W. Lafarge, K.A.J. Eppenhof, J.P.W. Pluim, Adversarial training and dilated convolutions for brain MRI segmentation, in: Deep Learn. Med. Image Anal. Multimodal Learn. Clin. Decis. Support, Springer, 2017: pp. 56–64. [CrossRef]
- R. Mehta, J. Sivaswamy, M-NET : A Convolutional Neural Network for Deep Brain Structure Segmentation, in: 2017 IEEE Int. Symp. Biomed. Imaging, 2017: pp. 437–440. [CrossRef]
- M. Havaei, A. Davy, D. Warde-farley, A. Biard, A. Courville, Y. Bengio, C. Pal, P. Jodoin, H. Larochelle, Brain Tumor Segmentation with Deep Neural Networks, Med. Image Anal. 35 (2017) 18–31. [CrossRef]
- L. Chen, P. Bentley, K. Mori, K. Misawa, M. Fujiwara, D. Rueckert, DRINet for Medical Image Segmentation, IEEE Trans. Med. Imaging. 37 (2018) 1–11. [CrossRef]
- D.E. Cahall, G. Rasool, N.C. Bouaynaya, H.M. Fathallah-Shaykh, Inception Modules Enhance Brain Tumor Segmentation, Front. Comput. Neurosci. 13 (2019) 1–8. [CrossRef]
- R. Mehta, A. Majumdar, J. Sivaswamy, BrainSegNet : a convolutional neural network architecture for automated segmentation of human brain structures BrainSegNet : a convolutional neural network architecture for automated segmentation of, J. Med. Imaging. 4 (2017). [CrossRef]
- H. Chen, Q. Dou, L. Yu, J. Qin, P. Heng, VoxResNet : Deep voxelwise residual networks for brain segmentation from 3D MR images, Neuroimage. 170 (2018) 446–455. [CrossRef]
- S. Han, Y. He, A. Carass, S.H. Ying, J.L. Prince, Cerebellum Parcellation with Convolutional Neural Networks, Proc SPIE Int Soc Opt Eng. 10949 (2019). [CrossRef]
- S. Han, A. Carass, Y. He, J.L. Prince, Automatic Cerebellum Anatomical Parcellation using U-Net with Locally Constrained Optimization, Neuroimage. (2020) 116819. [CrossRef]
- R. Cabeza-Ruiz, L. Velázquez-Pérez, A. Linares-Barranco, R. Pérez-Rodríguez, Convolutional Neural Networks for Segmenting Cerebellar Fissures from Magnetic Resonance Imaging, Sensors. 22 (2022) 1345. [CrossRef]
- S. Woo, J. Park, J. Lee, I.S. Kweon, CBAM : Convolutional Block Attention Module, in: Proc. Eur. Conf. Comput. Vis., 2018: pp. 3–19.
- N.J. Tustison, B.B. Avants, P.A. Cook, Y. Zheng, A. Egan, P.A. Yushkevich, J.C. Gee, N4ITK : Improved N3 Bias Correction, in: IEEE Trans. Med. Imaging, 2010: pp. 1310–1320. [CrossRef]
- V.S. Fonov, A.C. Evans, R.C. Mckinstry, C.R. Almli, D.L. Collins, Unbiased nonlinear average age-appropriate brain templates from birth to adulthood, Neuroimage. 47 (2009). [CrossRef]
- B.B. Avants, N. Tustison, H. Johnson, Advanced Normalization Tools ( ANTS ), Insight J. 2 (2009).
- A. Agarwal, P. Barham, E. Brevdo, Z. Chen, C. Citro, G.S. Corrado, A. Davis, J. Dean, M. Devin, S. Ghemawat, I. Goodfellow, A. Harp, G. Irving, M. Isard, Y. Jia, R. Jozefowicz, L. Kaiser, M. Kudlur, J. Levenberg, D. Man, R. Monga, S. Moore, D. Murray, C. Olah, M. Schuster, J. Shlens, B. Steiner, I. Sutskever, K. Talwar, P. Tucker, V. Vanhoucke, V. Vasudevan, F. Vi, O. Vinyals, P. Warden, M. Wattenberg, M. Wicke, Y. Yu, X. Zheng, TensorFlow: Large-Scale Machine Learning on Heterogeneous Distributed Systems, ArXiv Prepr. ArXiv1603.04467. (2016).
- F. Chollet, Keras: The Python deep learning library, Astrophys. Source Code Libr. ascl-1806 (2018).
- D.P. Kingma, J.L. Ba, Adam: a method for stochastic optimization, in: 3rd Int. Conf. Learn. Represent., 2015.
- M. Magnusson, A. Love, L.M. Ellingsen, Automated brainstem parcellation using multi-atlas segmentation and deep neural network, in: Med. Imaging 2021 Image Process., 2021: pp. 645–650. [CrossRef]
- J.E. Iglesias, C.Y. Liu, P.M. Thompson, Z. Tu, Robust brain extraction across datasets and comparison with publicly available methods, IEEE Trans. Med. Imaging. 30 (2011) 1617–1634. [CrossRef]
- K. Reetz, R. Rodríguez, I. Dogan, S. Mirzazade, S. Romanzetti, J.B. Schulz, E.M. Cruz-Rivas, J.A. Alvarez-Cuesta, R. Aguilera Rodríguez, Y. Gonzalez Zaldivar, G. Auburger, L. Velázquez-Pérez, Brain atrophy measures in preclinical and manifest spinocerebellar ataxia type 2, Ann. Clin. Transl. Neurol. 5 (2018) 128–137. [CrossRef]
- Ö. Çiçek, A. Abdulkadir, S.S. Lienkamp, T. Brox, O. Ronneberger, 3D U-net: Learning dense volumetric segmentation from sparse annotation, in: Med. Image Comput. Comput. Interv. 2016 19th Int. Conf., Springer International Publishing, Athens, Greece, 2016: pp. 424–432. [CrossRef]

| Label | Mean DSC±stdev | |||
|---|---|---|---|---|
| This research | [42] | Han et al.[41] | Magnusson et al.[50] | |
| Mesencephalon | 0.96±0.022 | 0.91±0.023 | 0.93±0.019 | 0.89±0.031 |
| Pons | 0.96±0.015 | 0.93±0.016 | 0.94±0.013 | 0.91±0.029 |
| Medulla | 0.95±0.021 | 0.91±0.022 | 0.92±0.021 | 0.91±0.023 |
| Full brainstem | 0.96±0.008 | 0.94±0.008 | 0.95±0.007 | 0.93±0.013 |
| Brainstem section | Mean volumes (% TICV) | P | ||
|---|---|---|---|---|
| Patients | Preclinical | Controls | ||
| Mesencephalon | 0.4 | 0.44 | 0.48 | 0.007 |
| Pons | 0.47 | 0.76 | 0.82 | < 0.0001 |
| Medulla | 0.26 | 0.29 | 0.31 | 0.00012 |
| Whole brainstem | 1.12 | 1.49 | 1.62 | < 0.0001 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




