1. Introduction
The hallmark of diabetes mellitus (DM), a chronic metabolic disease, is persistent hyperglycemia brought on by either decreased insulin action, insulin secretion, or both. Diabetes mellitus has become a pandemic in prevalence, impacting millions of people globally and dramatically raising morbidity, mortality, and medical costs of patients. To effectively manage diabetes mellitus, it is essential to avoid major complications such as retinopathy, neuropathy, and cardiovascular diseases, while also significantly reducing healthcare costs. Accurate prediction and early diagnosis of diabetes and its related health outcomes are crucial [
1,
2]. Machine learning (ML) and deep learning (DL) techniques are now essential for delivering predictive insights, facilitating individualized patient care, and supporting clinical decision-making processes with high precision due to improvements in processing power and data availability [
3,
4,
5]. Obesity, lifestyle changes, and genetic factors have all contributed to the significant increase in diabetes incidence. Diabetes can cause serious consequences, such as renal failure, neuropathy, and CVD, if it is not treated or is not adequately controlled [
6,
7].
The International Diabetes Foundation (IDF) has reported the rapid rise of people with diabetes aged 18 to 79 years from 4.7% to 8.5% within three decades from 1980 to 2015. The prevalence in 2019 increased to an estimated percentage of 9.3% (463 million) and is projected to rise to 10.2% (578 million) by 2030 and 10.9% (700 million) by 2045, respectively [
2,
8]. This indicates a serious problem for both developed and developing countries. China, India, and the United States of America are the most impacted nations, although this rise is unevenly spread, with estimates of 143% in Africa (undiagnosed cases) and 15% in Europe [
8].
Early identification and precise diabetes prediction are essential for prompt management and better patient outcomes, given the disease's increasing cost on healthcare systems [
9,
10,
11]. Wearable technology combined with powerful ML and DL algorithms has enabled real-time glucose monitoring and insulin adjustment, significantly enhancing patients' freedom and lifestyle [
12]. Recent research has proven that ML and DL techniques have evolved in this area. These case studies demonstrate industry improvements while laying the groundwork for future advancements [
13]. DL-based prediction models have also revealed remarkable accuracy in detecting early signs and progressions of DM-related issues, such as retinopathy, neuropathy, and nephropathy.
On the other hand, healthcare systems are designed to improve sickness detection and diagnosis while simultaneously providing patients with the essentials for optimum health [
13,
14]. Concerns over the quality of care offered by the healthcare system and the availability of treatment resources are common among patients [
15]. Most people who would immediately benefit from better healthcare systems are those who have serious illnesses, including diabetes, hypertension, and irregular blood sugar levels [
16]. A healthy society must prioritize health and healthcare. Hence, it is imperative to use state-of-the-art techniques to track the development of diabetes. Encouraging a healthy population and reducing the risk of illnesses like diabetes in future generations enables the development of novel techniques or hybrids that may be used in healthcare systems to improve the quality of life [
17,
18,
19,
20].
With their automated, data-driven insights that can improve clinical decision-making, ML and DL models have become potent medical diagnosis and prediction technologies [
21,
22]. While DL models like convolutional neural networks (CNNs) and recurrent neural networks (RNNs) offer sophisticated feature extraction capabilities, a variety of ML models, such as decision trees (DT), random forest (RF), logistic regression (LR), and support vector machines (SVM), have been extensively utilized for diabetes prediction. Research is ongoing to determine how well these models perform in comparison regarding accuracy, dependability, and computing economy.
This study focuses on two main research topics. The first centres around the differences in accuracy and reliability of ML and DL models and their hybrids in predicting diabetic patient outcomes across various healthcare settings. The second one compares ML, DL, their hybrid models, and ensemble strategies regarding processing time and computational efficiency when applied to selected datasets for DM personalized medicine. This demonstrates the effectiveness of various ML, DL models and ensemble strategies in diagnosing diabetes, tracking its progression, and evaluating performance indicators by analyzing multiple datasets and comparing different predictive models. This is true because the architectural complexity and internal mechanisms of ML and DL models significantly influence differences in processing speed, RAM usage, and overall computing efficiency.
The rest of the paper is organized into sections as follows:
Section 2 presents the review of previous related literature addressing diabetes prediction,
Section 3 provides an overview of the methodology, a report on the datasets used, including data preprocessing performance metrics and the models employed in this study;
Section 4 presents the methodology flow diagram of the study;
Section 5 presents the results of each model, highlighting their respective metrics and time efficiency; Section 6 presents a detailed discussion of the results and the comparative analysis;
Section 7 provides the conclusion to the study and future direction.
2. Related Works
2.1. Synopsis of Diabetes Mellitus
“Diabetes” refers to a group of metabolic disorders that are characterised by elevated blood glucose levels resulting from insufficient insulin production, impaired insulin utilisation, or a combination of both [
23]. Chronic hyperglycemia is linked to long-term damage and dysfunction of organs such as the heart, blood vessels, kidneys, eyes, and nerves [
23,
24]. Individuals with diabetes have varying effects based on their age, income, race, and ethnicity. Environmental and genetic factors are catalysts for diabetes, resulting in insulin resistance and beta-cell death [
25,
26,
27].
To prevent comorbidities such as CVD, neuropathy, and retinopathy, diabetes care entails initial identification and aggressive control. Diabetes is a complicated condition with a tendency to develop silently due to lifestyle, environmental, and hereditary factors [
9]. Early indicators of prediabetic diseases are often misrepresented by traditional diagnostic and treatment techniques, which can increase healthcare expenses and delay interventions. Thus, new methods for controlling diabetes are crucial for reducing its impact on people and enhancing positive world health outcomes [
24,
28]. Type 1 diabetes mellitus (T1DM), type 2 diabetes mellitus (T2DM), and gestational diabetes mellitus (GDM) are the three general forms of diabetes mellitus [
29]. The characteristic feature of T1DM, also referred to as insulin-dependent diabetes, is the autoimmune destruction of pancreatic beta cells, resulting in insufficient insulin production. T1DM affects 5–10% of people with diabetes. Ketoacidosis, or high blood acidity due to ketones, is often the initial sign of T1DM, which can develop slowly in adults or swiftly in children. It is one of the irreversible types. T1DM is becoming more common worldwide at a rate of 3% every year, affecting both sexes equally and leading to a sharp decline in life expectancy [
29,
30].
Non-insulin-dependent diabetes is another name for T2DM. It is characterized by beta-cell malfunction and insulin resistance [
29,
30]. T2DM accounts for 90 to 95 percent of all diabetes cases. The body creates more insulin to compensate for the deficiency; nevertheless, beta-cell activity progressively decreases, leading to insulin insufficiency [
31]. T2DM is associated with aging, obesity, sedentary lifestyles, high blood pressure, impaired lipid metabolism, and genetic factors. Ethnicity, which is more prevalent in some racial groups, is another aspect of T2DM prevalence [
31,
32,
33].
Pregnancy-related hyperglycemia is a common side effect of gestational diabetes mellitus (GDM) [
30,
34]. Despite impacting the mother and the foetus, it is frequently controllable with medicine, food, and exercise. GDM risk factors include obesity, advanced maternal age, and a history of glucose intolerance. Women with GDM have a greater lifetime risk of developing T2DM diabetes. Although there are differences in international diagnostic methods for GDM, early detection is crucial for therapy and issue prevention [
35,
36].
2.2. Existing Comparative Analysis of ML, DL, and ensemble models for DM prediction
Recent studies have investigated various ML and DL techniques for predicting chronic illnesses, offering valuable insights into the effectiveness and application of these models. Mahajan et al. [
37] assessed 15 ensemble ML models across 16 datasets, concluding that stacking methods yielded the best performance in chronic illness prediction. Similarly, Flores et al. [
38] employed feature selection techniques to evaluate SVM, RF, and neural networks (NN), revealing that RF achieved the highest accuracy of 98.5% for early-stage diabetes prediction.
In another study, Gupta et al. [
39] compared DL and quantum machine learning (QML) using the PIMA dataset, finding that a DL-based Multi-Layer Perceptron (MLP) outperformed QML approaches. Aggarwal et al. [
40] investigated eight classifiers, identifying Naïve Bayes (NB) as the most accurate model, while Refat et al. [
3] established that XGBoost surpassed both DL and traditional ML models, achieving an impressive 99% accuracy.
Swathy and Saruladha [
41] reviewed CVD prediction models, advocating for hybrid approaches to enhance predictive accuracy. Fregoso-Aparicio et al. [
42] and Butt et al. [
5] highlighted the effectiveness of tree-based models combined with Internet of Things (IoT) integration for real-time glucose monitoring. Additionally, Uddin et al. [
43] identified RF and SVM as consistently high-performing ML algorithms in disease prediction tasks. Zarkogianni et al. [
9] validated the benefits of ensemble learning in assessing CVD risk associated with T2DM, showing that hybrid models like HWNN and Self-Organizing Maps (SOM) improved predictive capabilities.
Further notable contributions include Hasan et al., [
44] who achieved a 95% area under the curve (AUC) using an ensemble framework; Ayon and Islam [
4], as well as Naz and Ahuja [
45], whose DL models reached accuracy levels exceeding 98%; Lai et al. [
46], who optimized Gradient Boosting Machine (GBM) techniques for Canadian demographics; Dagliati et al. [
25], who predicted diabetic complications with an accuracy of 83.8% using LR; and Sahoo et al. [
47], who emphasized the superiority of CNN in managing high-dimensional healthcare data.
Building upon these findings, the current research utilizes five publicly available datasets and implements essential preprocessing steps such as outlier removal and imputation. A comparative analysis of various models, including LR, NB, Decision Trees (DT), RF, SVM, K-Nearest Neighbours (KNN), XGBoost, AdaBoost, as well as several neural networks like CNN, Deep Neural Networks (DNN), Recurrent Neural Networks (RNN), Long Short-Term Memory networks (LSTM), Autoencoders, and Gated Recurrent Units (GRU), is conducted. Furthermore, hybrids of these models and ensemble strategies, such as systematic bagging and stacking, are evaluated. The performance of these models is measured using a comprehensive set of metrics, including accuracy, precision, recall, F1-score, area under the receiver operating characteristic curve (AU-ROC), coefficient of determination (R²), mean squared error (MSE), mean absolute error (MAE), root mean square deviation (RMSD), number of parameters, optimal parameters, memory usage, and computation time.
.
3. Materials and Methods
This section provides a summary of the techniques and algorithms used in the study, outlining the methods and how they work. It is organized into different sections: (i.) sampling techniques for dataset imbalance, (ii.) ML and DL techniques used, where each model offers an overview of the fundamental concepts behind the techniques, ensuring their role in the research is understood, (iii) Performance metrics used (iv.) Datasets, and finally (v.) Preprocessing.
3.1. Sampling Techniques for Datasets Imbalance
3.1.1. Oversampling Techniques
- a)
Synthetic Minority Oversampling Techniques (SMOTE): SMOTE balances class distribution by creating artificial samples for the minority class. Instead of duplicating existing samples, it generates new instances by interpolating between them, selecting
k nearest neighbours, and using a random interpolation factor to promote diversity. [
48]. SMOTE is represented as:
where = ith minority instances, n = No. of features (dimensions) and N = number of minority class instances.
The
k nearest neighbours of
based on a distance metric (usually Euclidean distance), denoting the set of these neighbours as:
where
= k-nearest neighbours of
. Finally, it creates a new synthetic sample
by randomly choosing a neighbour
and then generate the
through interpolation between
and
where
is the random scalar randomly drawn from the uniform distribution between
0 and
1 i.e.
U(0,1). These steps continue until the desired number of synthetic minority samples has been created.
- b)
Adaptive Synthetic Sampling (ADASYN): ADASYN, an adaptive extension of SMOTE, emphasizes complex minority class samples by assigning greater weights to those near the decision boundary or surrounded by majority class samples. It generates synthetic data in these difficult areas, improving model robustness and refining the decision boundary in imbalanced datasets. Mathematically, it is represented in this regard:
and
nearest neighbours’ computation for the majority class for each minority sample
is given as:
where if
,
is easy to classify, but if
,
is difficult to classify and hence requires more synthetic samples. Normalized density distribution for each minority sample (difficult scores)
where the distribution
represents the importance of each minority sample in oversampling. The method then computes how many synthetics to generate from each minority sample as:
where
can be rounded to the nearest integer. Therefore, for each minority sample
, it then generates
synthetic samples by randomly selecting a minority-class neighbour
from the
K-nearest neighbours of
belonging to the minority class and then generates the synthetic samples
This process continues times for each minority sample
- c)
SMOTE-ENN and Random Oversampling are other techniques used to
address class imbalance in datasets. SMOTE-ENN enhances decision boundaries by generating synthetic samples for the minority class and removing ambiguous instances using Edited Nearest Neighbours [
49,
50]. Random Oversampling, on the contrary, increases the minority class size by duplicating existing samples, which is simple and efficient but carries a risk of overfitting. This risk can be mitigated by resampling with replacement to maintain a more diverse and balanced dataset [
51].
3.1.2. Undersampling Techniques
Several undersampling techniques have been developed to address class imbalance in datasets. Among these, clustering-based undersampling methods are specifically utilized to manage such imbalances effectively. One effective method involves using clustering centroids, particularly through the
K-means algorithm. This method consolidates clusters of majority class instances into singular representative points, effectively diminishing data volume while maintaining critical patterns [
52]. In contrast, random undersampling, although straightforward and computationally efficient, may discard valuable samples and increase variance. To improve upon this, random undersampling can be enhanced with Tomek Links, which removes borderline samples that blur the class boundaries, ultimately improving clarity and classifier performance [
53]. NearMiss-3 selects the majority class instances that are farthest from minority samples. This strategy enhances separability and reduces class overlap. One-Sided Selection (OSS), an alternative approach, refines the dataset further by combining Tomek Link removal with the Condensed Nearest Neighbour algorithm, retaining only a compact and representative subset of the majority class. Additionally, Neighbourhood Cleaning (NCR) employs
k-NN classification to identify and eliminate noisy or misclassified samples from the majority class. This process helps maintain the integrity of the dataset while minimizing overlapping [
52,
54]. Among these techniques, clustering is highlighted as a structured, data-preserving method for our study. It offers a strategic advantage by retaining meaningful patterns while significantly reducing the majority class, ultimately improving the model’s efficiency and classification performance [
52,
54].
3.2. Machine Learning and Deep Learning Techniques employed.
3.2.1. Machine learning (ML)
ML is a subfield of artificial intelligence (AI) that allows computers to recognize patterns in data and learn from them with minimal human intervention. ML techniques fall into three main categories: supervised learning (classification and regression with labelled datasets), unsupervised learning (clustering and dimensionality reduction with unlabelled datasets), and reinforcement learning.
- a)
Logistic Regression (LR): A binary classification algorithm that uses the sigmoid function to map inputs to a 0-1 range, indicating class likelihood. It optimizes the log-likelihood function through Gradient Descent, assuming a linear relationship between variables [
55,
56].
- b)
Naïve Bayes (NB): A probabilistic classifier that applies Bayes' theorem, relying on the assumption of conditional independence among features. It's effective in spam filtering and text categorization by calculating posterior probabilities [
56,
57,
58].
- c)
Decision Trees (DT): This supervised learning method splits data into subsets based on features to make predictions. It consists of nodes (decisions), branches (outcomes), and leaves (predictions), using criteria like MSE or Gini Index to determine splits [
58].
- d)
Random Forest (RF): An ensemble method that trains multiple decision trees and combines their outputs. It reduces overfitting by bagging (training on random data samples) and selecting random feature subsets. Predictions are made through majority voting or averaging [
10,
16,
56,
59].
- e)
Support Vector Machine (SVM): A technique that identifies the optimal hyperplane to separate classes by maximizing the margin between them, utilizing support vectors. Kernel functions transform non-linearly separable data into higher dimensions for separation [
10,
16,
56,
60].
- f)
K-Nearest Neighbours (KNN): A classification method that assigns data points based on the majority class of their k-nearest neighbors using distance metrics like Euclidean. It has a low training cost but high inference cost, with performance influenced by the choice of k [
16,
61,
62,
63].
- g)
Extreme Gradient Boosting (XGBoost): An efficient gradient boosting method for accuracy, using a second-order Taylor expansion for loss function approximation. It enhances performance with cache-aware access and regularization techniques to mitigate overfitting [
10,
16,
59,
60].
- h)
Adaptive Boosting (AdaBoost): An ensemble method that combines weak learners, usually decision stumps, into a strong classifier. It dynamically adjusts sample weights to focus on misclassified instances, improving performance [
16,
56].
3.2.2. Deep Learning models
DL models, built on complex artificial neural networks (ANN), excel at extracting nonlinear patterns from large datasets. They develop hierarchical feature representations automatically, reducing the need for manual engineering. This enhances their effectiveness in image recognition, natural language processing (NLP), speech recognition, and healthcare diagnostics. However, they require significant data and processing power to perform optimally.
- a)
Convolutional Neural Networks (CNN): CNNs are deep learning models for grid-like data (e.g., images). They utilize convolutional layers for spatial feature extraction, pooling layers for dimensionality reduction, and fully connected layers for classification or regression, leveraging weight sharing and local connectivity [
16,
64,
65].
- b)
Deep Neural Networks (DNN): DNNs consist of hidden layers between input and output, enabling the learning of complex patterns through interconnected neurons and nonlinear activation functions [
5,
14,
66].
- c)
Recurrent Neural Networks (RNN): RNNs retain memory of previous inputs using hidden states, making them suitable for interpreting sequential data and capturing temporal dependencies [
16].
- d)
Long Short-Term Memory (LSTM): LSTMs enhance RNNs by addressing the vanishing gradient problem with gates that manage information flow. This allows them to effectively capture long-term relationships in data, useful in tasks like time-series forecasting and speech recognition [
14,
16,
68].
- e)
Gated Recurrent Unit (GRU): GRUs are a type of RNN that uses gating techniques to manage information flow, helping retain important historical data while discarding irrelevant details [
16].
3.2.3. Hybrids and Ensemble strategies
These ML and DL models combine predictions from individual models to enhance overall generalization, accuracy, and resilience. By leveraging the diversity among individual classifiers or regressors, these techniques reduce variance, bias, and sensitivity to noisy data [
67,
68]. General ensemble methods, including stacking and bagging, were routinely implemented using the best-performing base learners discovered for each dataset. By integrating the advantages of several separate models, these ensemble approaches seek to improve generalization, mitigate overfitting, and reduce variation, to improve prediction performance. Using bootstrap sampling, several instances of the same learning algorithm were trained on various data subsets in the bagging technique. The predictions of these instances would then be combined, usually by majority vote or averaging. This approach was particularly effective for stabilizing models such as decision trees, which often experience significant variation.
In contrast, stacking involves training a meta-learner to aggregate the results of multiple base models. The complementary strengths of heterogeneous models enhance the effectiveness of stacked ensembles. The effectiveness of these ensemble approaches compared to their base models, that is, the un-stacked and un-bagged counterparts, was consistently observed across all datasets examined. This improvement in performance highlights the advantage of ensemble learning in leveraging several hypotheses to create a more reliable and accurate predictive model, particularly in varied healthcare data contexts like diabetes progression prediction and categorization
3.3. Performance Metrics Tools
3.3.1. Hyperparameter Tuning
Through methodical adjustment of configuration parameters that govern the learning process, hyperparameter tuning is crucial for optimizing model performance. While more sophisticated approaches like Bayesian optimization provide more effective substitutes, conventional methods like grid search and random search are frequently computationally costly. To intelligently explore the hyperparameter space, this study uses Optuna, a sophisticated optimization system that uses Tree-structured Parzen Estimators (TPE). Optuna is especially well-suited for intricate ML and DL models because of its adaptive sampling and early pruning features, drastically lowering computing expenses while guaranteeing ideal parameter selection [
69,
70]. Utilizing Optuna leads to faster convergence on high-performing configurations, seamless interaction with various ML frameworks, and enhanced reproducibility through detailed logging and visualization. Optuna is more efficient than traditional methods since it dynamically prioritizes promising trials and discards underperforming ones. This makes it the perfect option for creating reliable models with enhanced generalization powers, especially when computing resources are limited. The framework has shown itself to be a helpful tool for contemporary ML pipelines due to its efficacy in various applications.
3.3.2. Evaluation Metrics
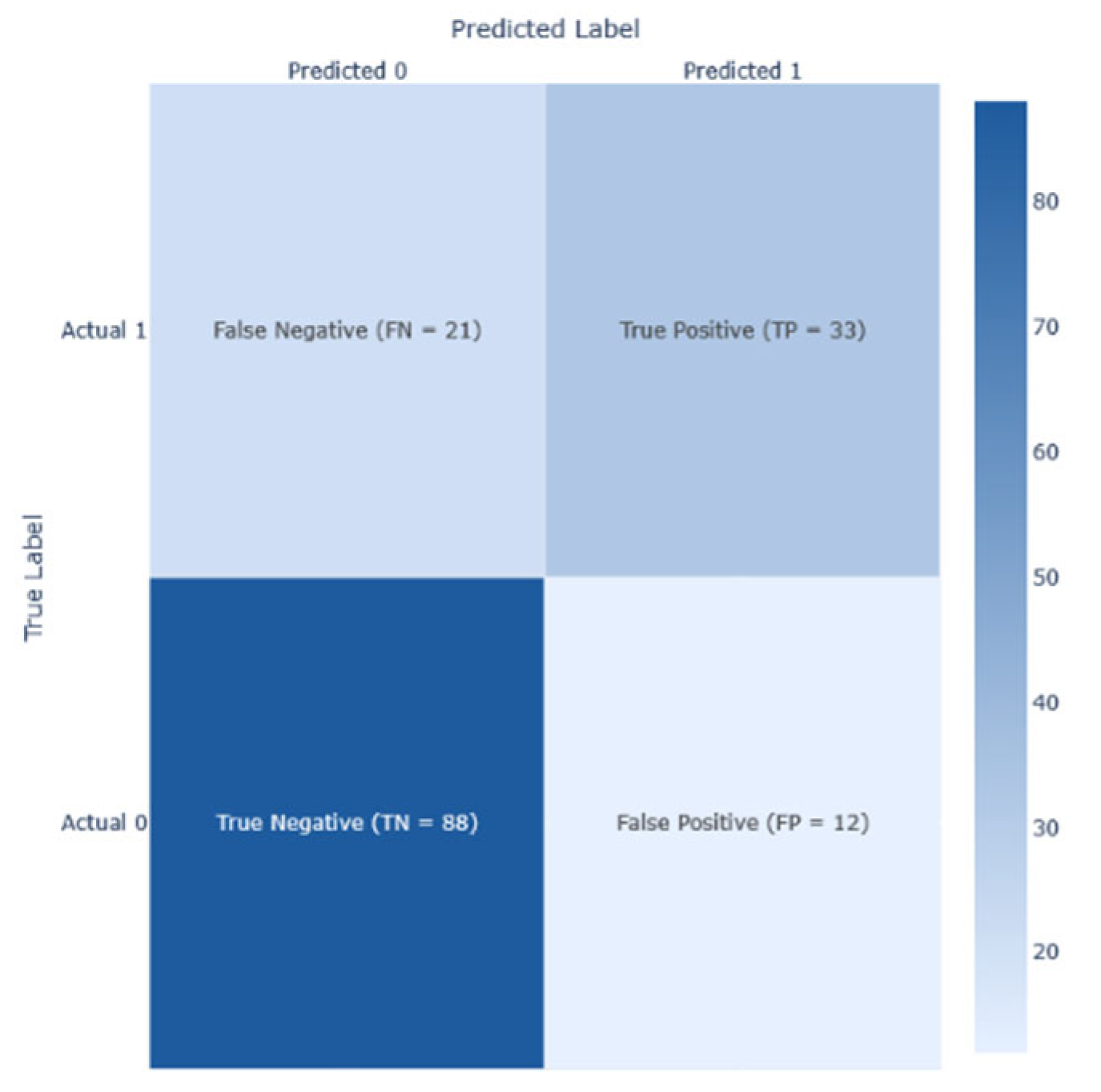
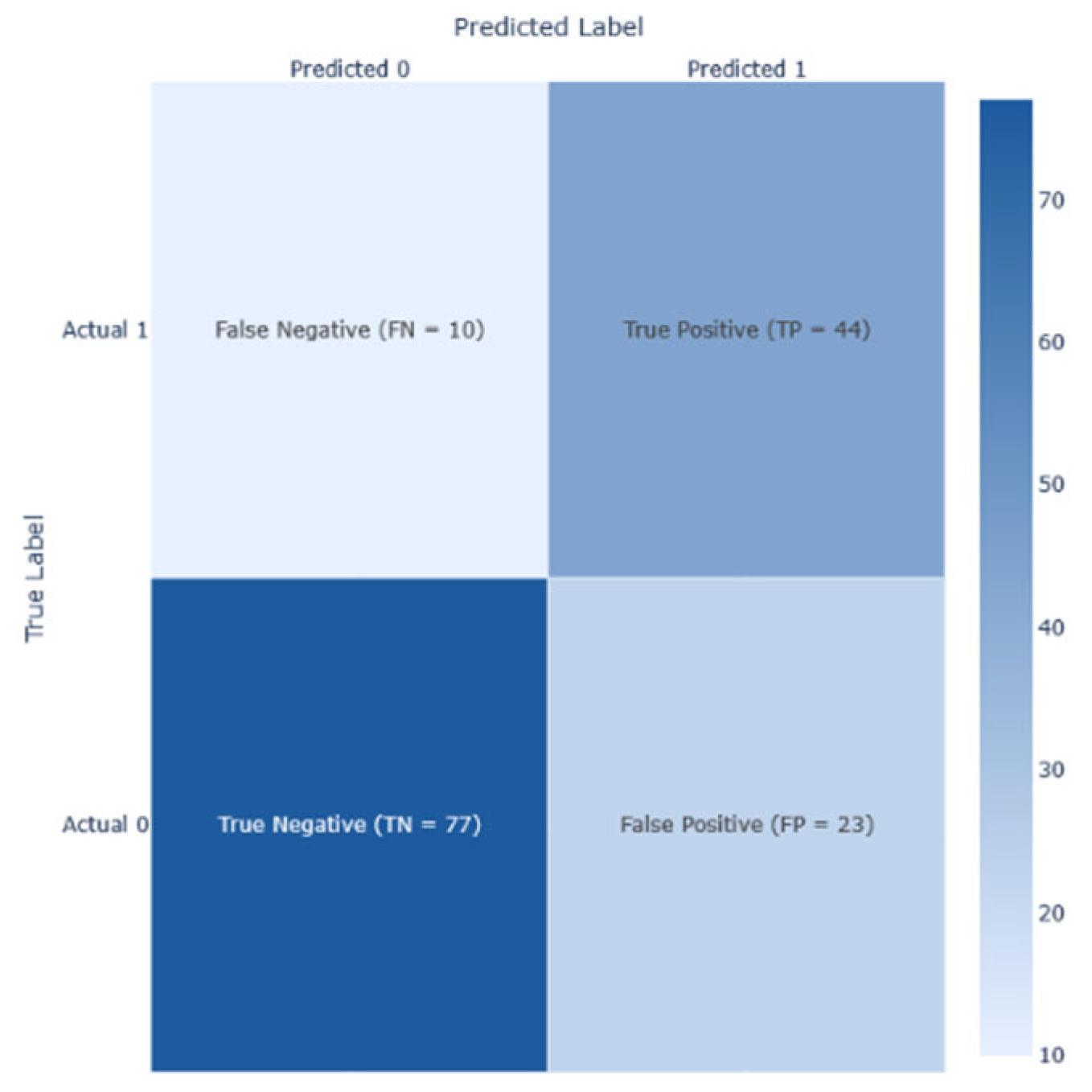
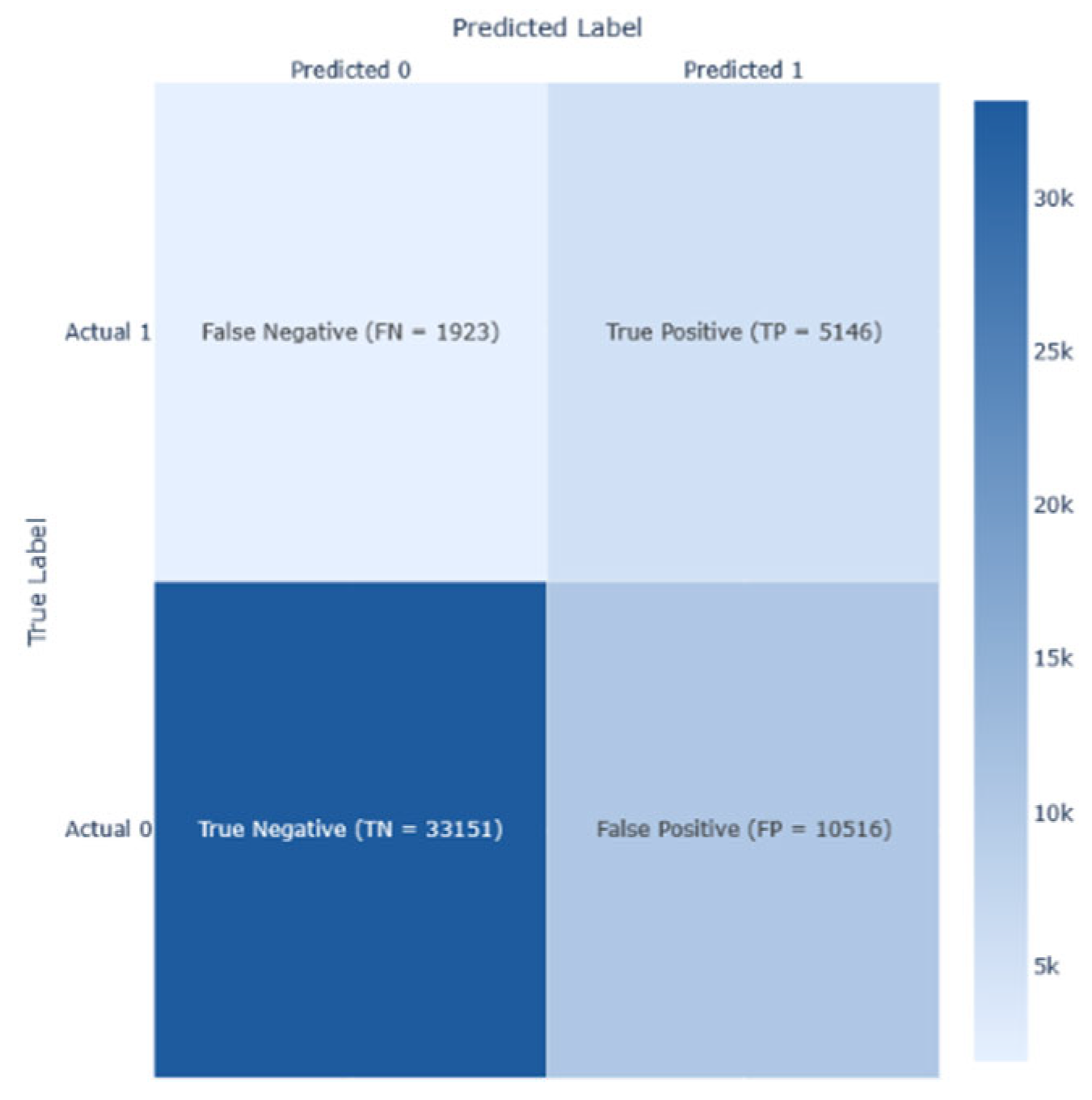
To guarantee a thorough model evaluation, performance metrics were used. True positive (TP) indicated that the model predicted diabetes I present or has progressed; true negative (TN) signifies that the model predicts the absence of diabetes and its progression; false positive (FP) indicated that the model predicted incorrectly the presence of diabetes; and false negative (FN) signifies the failure of the model predicting the presence of diabetes while it exists.
Accuracy measures the proportion of correct predictions, both positive and negative, against the total number of predictions made, resulting in the overall percentage of accurate predictions. While accuracy appears simple, it may be misleading for imbalanced datasets as it does not account for different types of errors.
Precision calculates the percentage of accuracy by which diabetes is correctly identified by the model. This measure is critical when FP can lead to high costs, such as unnecessary medical procedures or false fraud alerts.
Recall (Sensitivity): calculates the percentage of
TP that are successfully detected, which indicates how well the model detects positive cases.
F1-score combines precision and recall using their harmonic means to assess model performance fairly. This is our primary assessment statistic since it evenly weights
FP and
FN, effectively managing class imbalance.
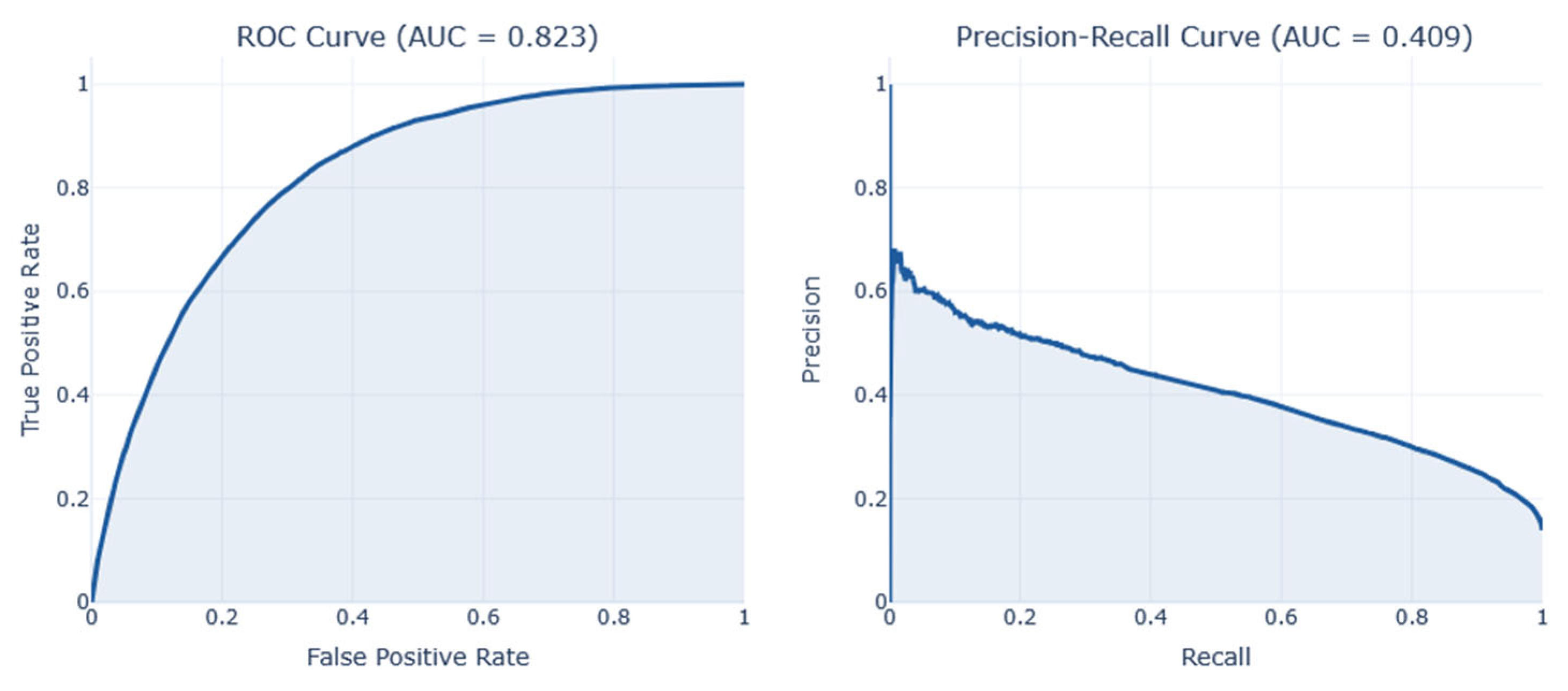
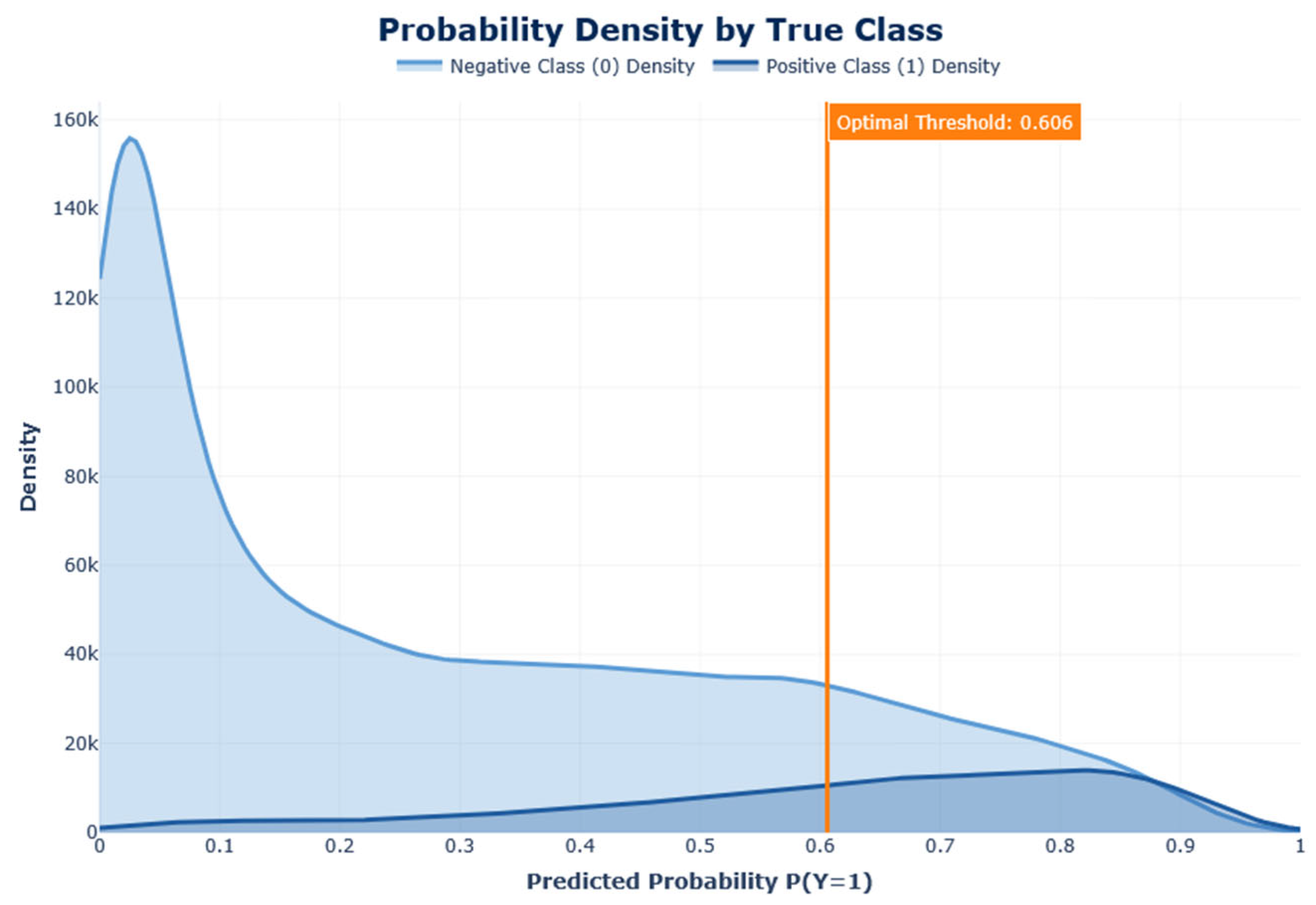
AUC-ROC justifies the model's capacity to differentiate between classes across all potential classification thresholds. A perfect classifier obtains an AUC of 1, whereas 0.5 is obtained by random guessing.
where
represents the decision threshold
Mean Squared Error (MSE) measures the average squared difference between predicted and actual values penalizing large errors more heavily.
Mean Absolute Error (MAE) measures the average of absolute difference between predicted and actual values treating all errors equally.
Root Mean Square Deviation or Error (RMSD/RMSE) performs the square root of MSE keeping the same units as the predicted value and more interpretable than MSE.
Number of Parameters (NoP) signifies the total number of learnable elements (such as weights and biases) with respect to the selected model. It is evident that more parameters signify higher complexity and capacity, but higher risks of overfitting.
Inference Time, or Time taken (TT) as noted in the results tables, logs the time needed to produce predictions to assess the model's computational efficiency. Although it has no bearing on the statistical performance of the model, this parameter is essential for real-time applications and deployment in contexts with limited resources.
Since the F1-score provides the most balanced evaluation for medical diagnostics by equally weighing false positives and false negatives, the results in
Section 4 are organized according to F1-score.
3.4. Datasets
This study analyzes five diabetes-related datasets from the UCI Machine Learning Repository, CDC, and Kaggle, summarized in
Table 1, which outlines their sources, characteristics, total instances, and positive/negative counts. Data preprocessing included normalization to ensure consistency and enhance result precision. Recursive Feature Elimination (RFE) was applied for feature selection, and hyperparameter tuning using Optuna was conducted for each classifier during model construction.
3.4.1. Dataset 1
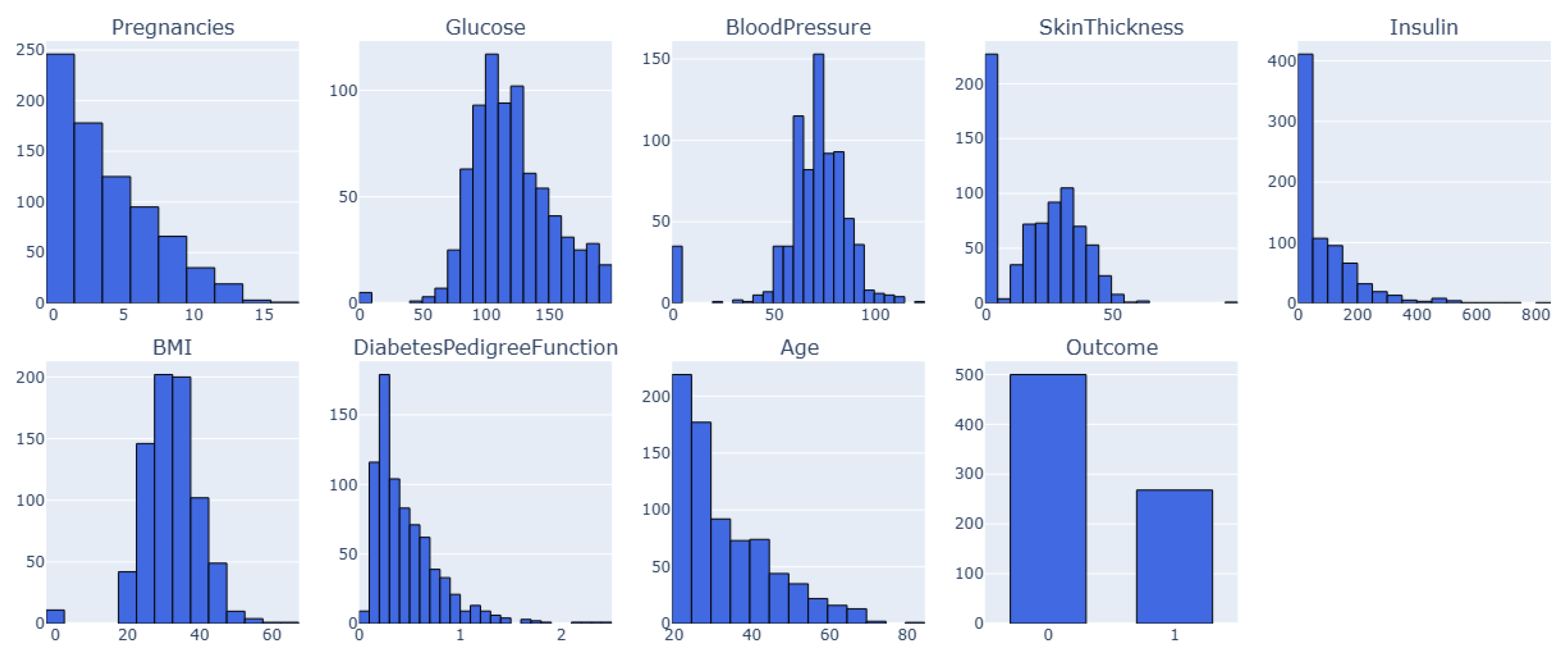
This is the PIMA Indian Diabetes dataset called Dataset 1. It has 768 samples and nine features, including clinical measures and patient characteristics as visualized in
Figure 1. The dataset features are Pregnancy, Glucose, Blood Pressure, Insulin, Skin Thickness, BMI, Diabetes Pedigree-Function, Age, and Outcome. The dataset contains no duplicate entries or missing values (NaNs); all characteristics are numerical. However, several features, especially those related to blood pressure, skin thickness, insulin, glucose, and BMI, contain sundry zero values, which is biologically impossible.
Section 3.3 will discuss these discrepancies and their ramifications [
71,
72,
73,
74,
75].
3.4.2. Dataset 2
This is also PIMA Indian Diabetes dataset, henceforth referred to as Dataset 2. It also has numerical characteristics about clinical measures and patient demographics and is structured similarly to Dataset 1. However, it is much larger with 2000 samples rather than 768, but 9 features.
3.4.3. Dataset 3
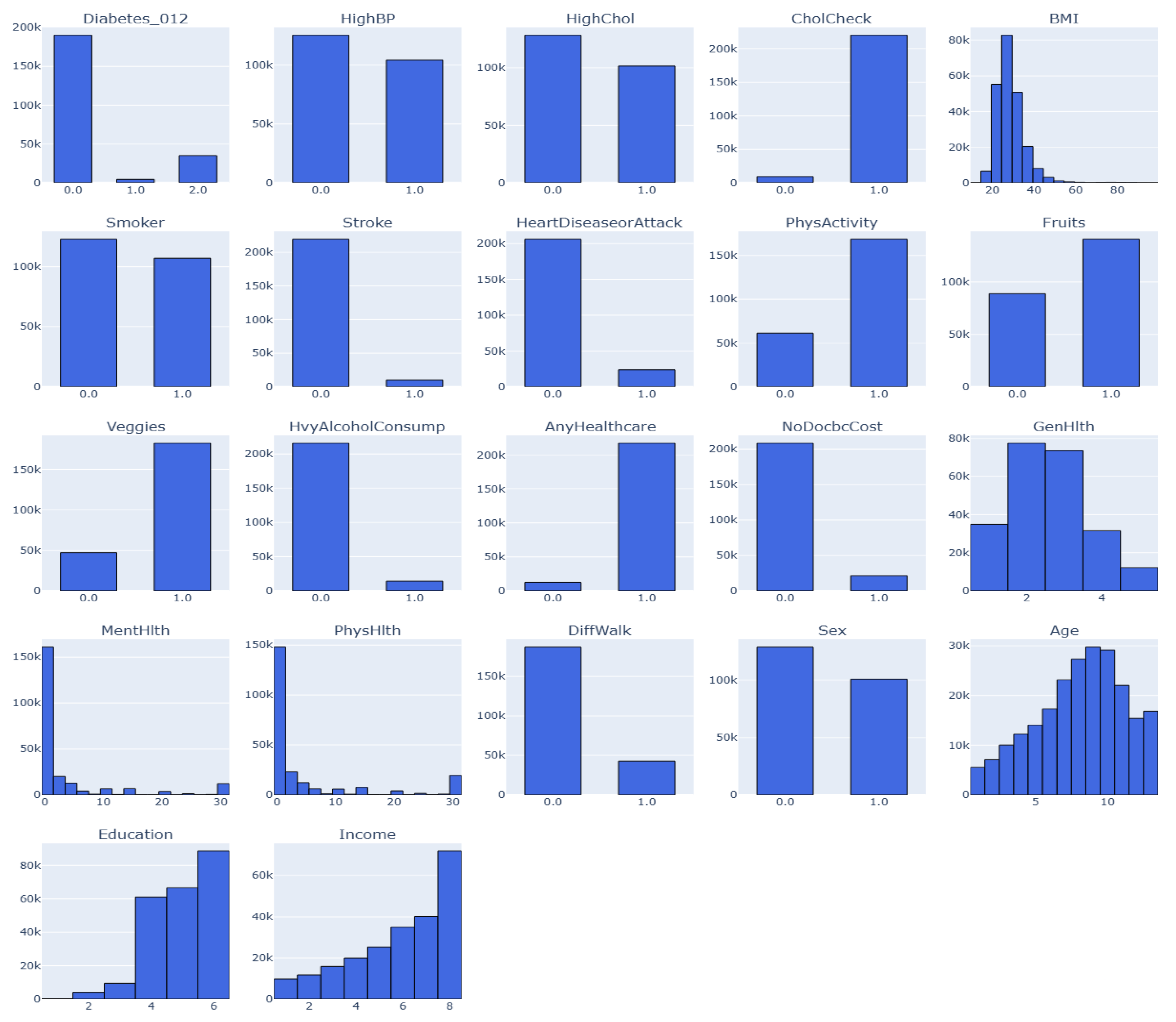
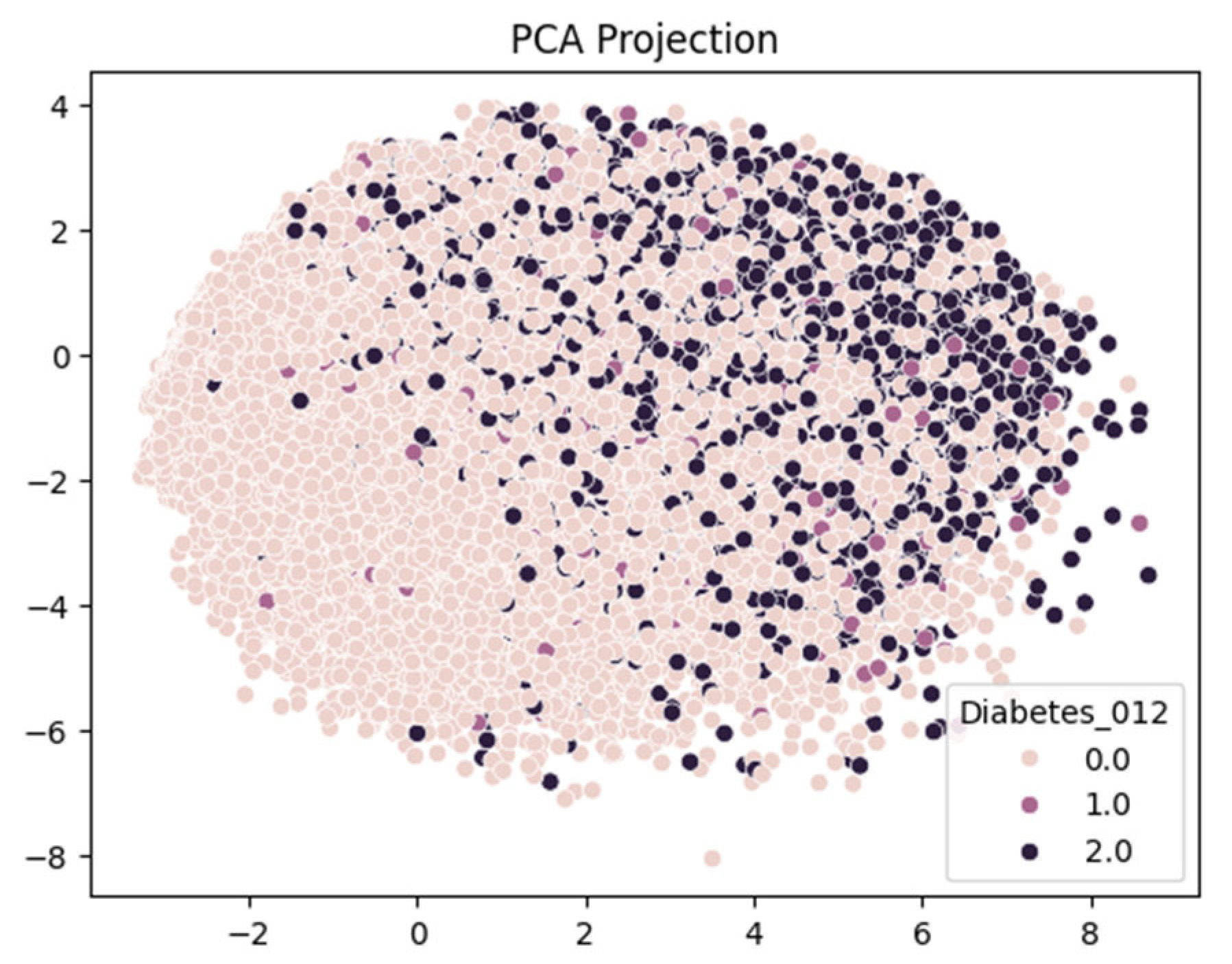
This is an annual Behavioral Risk Factor Surveillance System (BRFSS) dataset captured by the Centre for Disease Control (CDC). This dataset is for the year 2015. Henceforth, the dataset would be known as Dataset 3. The target variable has three classes (0, 1, 2). 0 is for no diabetes or only during pregnancy, 1 is for prediabetes, and 2 is for diabetes, as depicted as feature Diabeter_012 in
Figure 2. There is a class imbalance in the dataset, but it has 21 features and 253,680 samples [
76]
3.4.4. Dataset 4
This variant of Dataset 3 consists of 70,692 samples and 21 features of the BRFSS dataset captured by CDC for 2015. Here, the target consists of two classes (0, 1). 0 is for no diabetes, and 1 is for prediabetes or diabetes. It also contains class imbalance and would be known as Dataset 4 in this study.
3.4.5. Dataset 5
The early-stage diabetes risk prediction of patients from Sylhet Diabetes Hospital, Bangladesh, were captured in this dataset. Direct surveys from the patients were used in the study [
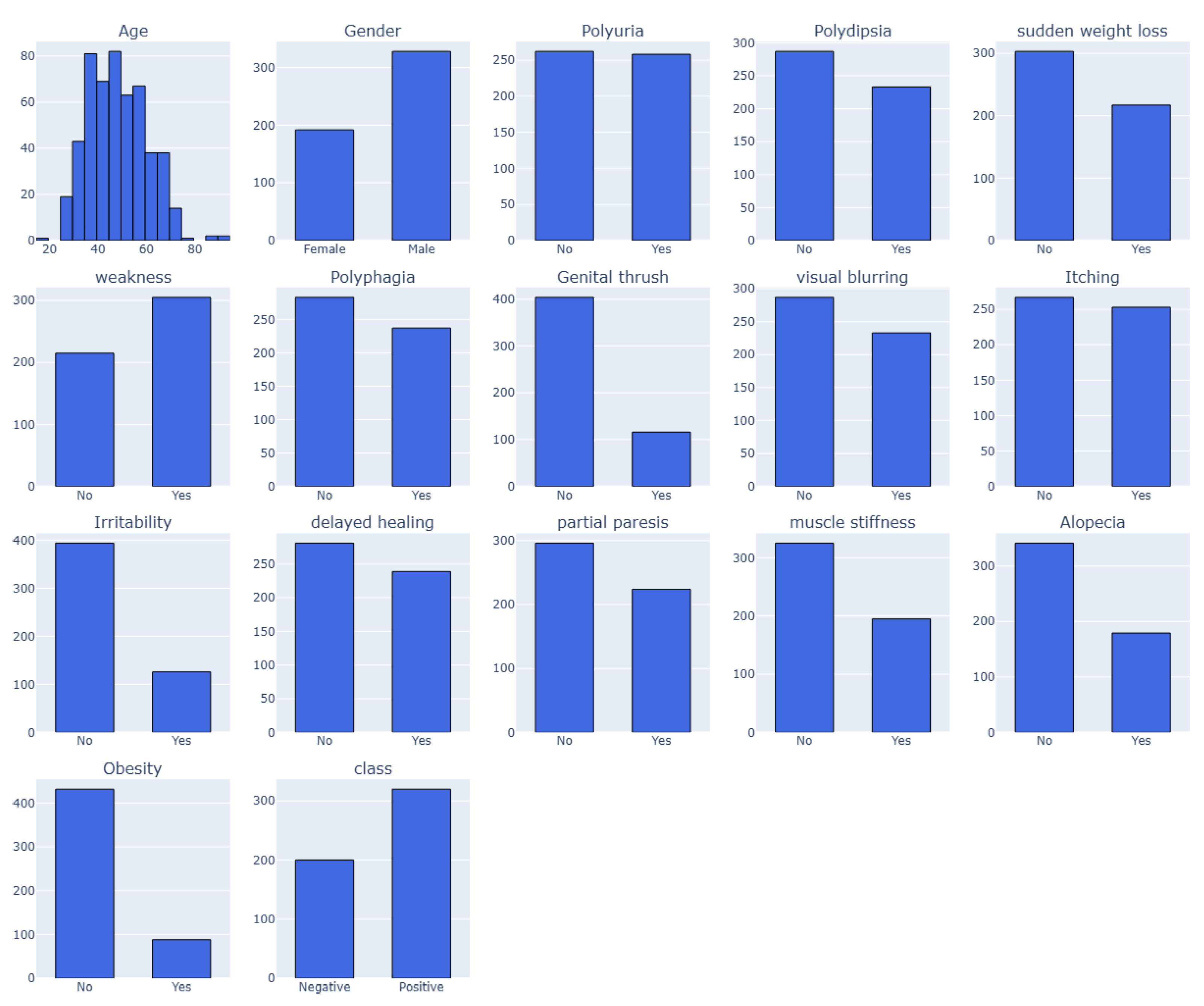
77]. This dataset report includes 520 people with diabetes-related symptoms and information on people who may have diabetes-related symptoms. The dataset has 520 cases and 17 features, including the target class. The dataset, collected in 2020, was verified by a certified physician from Sylhet Diabetes Hospital. The dataset, which includes several categorical (Yes/No) variables associated with diabetes diagnosis, is displayed in
Table 1. The "Class" property indicates the patient's diabetes status as either positive (1) or negative (2). The values of 1 (yes) or 2 (no) for each feature indicate whether the associated symptom or condition is present. However, there are four categories for the "Age" attribute: 1 for those aged 20–35, 2 for those aged 36–45, 3 for those aged 46–55, 4 for those aged 56–65 and 5 for those aged above 65 as visualized in
Figure 3. These characteristics and values serve as the foundation for developing a classification algorithm that uses patient data to forecast the diagnosis of diabetes [
78,
79].
3.5. Preprocessing
Improving model accuracy and reliability through preprocessing datasets is essential for preparing raw data for ML procedures. This process often includes cleaning the data to address outliers and missing values, transforming the data through standardization or normalization, and converting categorical features using one-hot encoding. Various dimensionality reduction techniques help manage large sets of features. Additionally, sampling techniques like ADASYN and Clustering was employed to correct class imbalances.
To effectively evaluate the performance of the study's model, the five datasets are divided into subsets with an 80:20 ratio for training and testing/validation. Proper preprocessing not only reduces computational complexity, but also enhances the predictive ability of ML models, ensuring high dataset quality.
Performing the exploratory data analysis (EDA) of each dataset, it was observed that zero values exist in columns where they are not physiologically conceivable, which is a significant problem in both Datasets 1 and 2. Missing data may be entered as zeros instead of NaNs, resulting in inaccurate numbers.
Table 2 shows zero values concerning affected features under Datasets 1 and 2.
Two imputation techniques are employed to deal with the problem of zero values in columns such as BMI, Insulin, Glucose, Blood Pressure, and Skin Thickness) is biologically impossible:
Median Imputation: In each column, the median of non-zero values for zeros is substituted.
Minimum Imputation: Instead of actual measurement, the zeros may mean data was not collected. This might indicate that the physiological levels of the patients with missing results were normal. Consequently, we used each column's smallest non-zero value to impute missing data.
Remarkably, models trained using minimum imputation on the datasets consistently performed better than those trained with median imputation. This validates our prediction that missing data were likely connected with patients having normal measures rather than abnormal or severe results. Given that various imputation techniques can substantially influence model performance, this conclusion implies that comprehending the nature of missing data is essential in medical datasets.
The target variable exhibited class imbalance, complicating the study’s analysis. In Dataset 1, there were 400 entries for 0 (No) and 214 for 1 (Yes), while Dataset 2 had 1053 for 0 and 547 for 1. We focused on oversampling techniques, as undersampling was unfeasible due to the limited data. Various methods were tested, including ADASYN, SMOTE-ENN, random oversampling, and SMOTE, with ADASYN yielding the best results. This method generates synthetic samples near the decision boundary, highlighting the importance of selecting the right data balancing strategy for model performance.
Datasets 3 and 4 had considerable data points and were unbalanced, but Datasets 1 and 2 had fewer data points, as shown in
Table 3. We thus used undersampling techniques on the datasets to lessen this problem. Instead of random undersampling, we employed clustering-based undersampling on datasets 3 and 4, which maintains the underlying data distribution. Clustering-based undersampling chooses representative samples from each cluster, guaranteeing that important patterns and class features are preserved, in contrast to conventional techniques that randomly exclude data points. It keeps crucial information from being lost despite its high computational cost.
Simple binary encoding was used to transform (encode) categorical characteristics into numerical representations to guarantee consistency across all datasets. To normalize the data and guarantee that each feature had a similar range, feature scaling was also used. This step is essential for optimising ML models because it keeps characteristics with bigger magnitudes from overpowering those with smaller values.
Due to the considerable class imbalance, where the dominant class significantly outnumbered the minority class, the experimental assessment revealed that modelling datasets 3 and 4 presented significant obstacles. The models' total incapacity to detect any occurrences of the minority class demonstrates that this extreme imbalance ratio made it difficult to create useful prediction models on the original datasets. However, applying hyperparameter tuning, the model was able to present reasonable results. This is essentially based on the size of the datasets and the corresponding features.
4. Methodology Flow Diagram
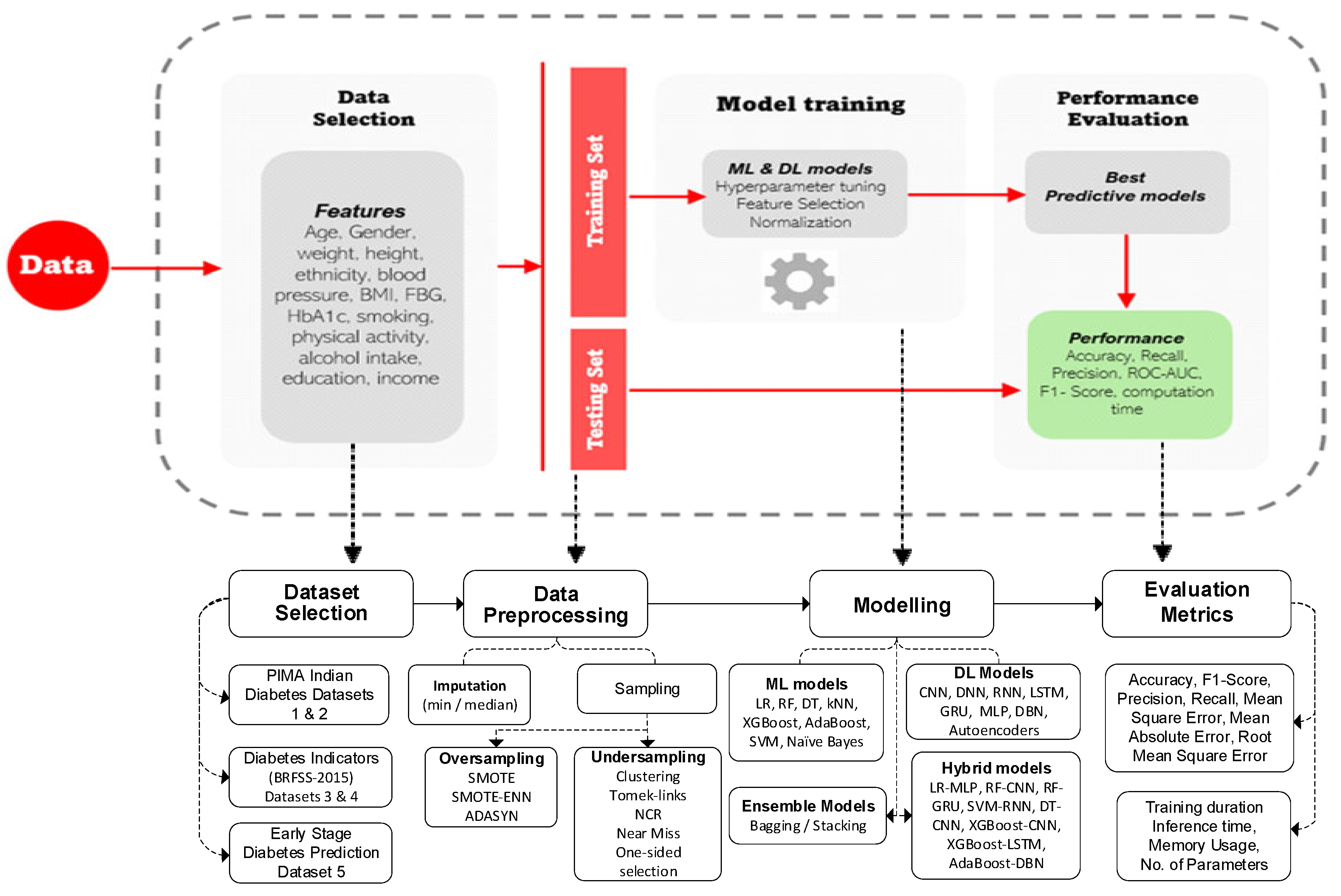
The flow diagram in
Figure 4 illustrates a comprehensive pipeline for predicting diabetes outcomes using ML and DL models. It begins with data selection, which incorporates diverse features and lifestyle factors. The next step involves dividing the data into training and testing sets. During model training, several preprocessing steps were conducted, including imputation, normalization, feature selection, and hyperparameter tuning. Different ML, DL, and ensemble strategies models are then applied to the data. Finally, the performance of the models is evaluated using metrics such as accuracy, precision, recall, F1-score, ROC-AUC, MSE, MAE, R
2, RMSE, and computation time, ensuring both predictive accuracy and efficiency.
The 80:20 train-test split ratio employed in this study is a commonly accepted standard in ML applications, as it strikes a balance between model training and evaluation. By allocating 80% of the data for training, the model has access to a large and representative subset of the dataset, enabling it to effectively learn the underlying patterns, relationships, and distributions. The remaining 20% is set aside for testing, serving as an independent evaluation set. This allows this study to assess the model's ability to generalize to new, unseen data, which is essential for understanding how well the model may perform in real-world scenarios.
Choosing lower split ratios, such as 70:30 or 60:40, can lead to a smaller training set. This limitation can significantly hinder the model’s ability to learn, especially when the overall size of the dataset is limited. This issue is particularly evident in Datasets 1, 2, and 5, which have few samples. Reducing the training data in these cases can worsen problems like underfitting, unstable model behavior, and poor predictive performance. Therefore, maintaining an 80:20 split in this study is not only methodologically sound but also strategically important, especially for small or sensitive healthcare datasets where maximizing training information is crucial for the model's success.
5. Results Analysis
The results demonstrate the outcomes of a comprehensive investigation, utilizing comparison tables, confusion matrices, density graphs, and informative bar charts across all models employed. The Python programming language platform, version 3.11.5 packaged by Anaconda3, was used to implement all these processes. The model training procedure was systematically conducted for each model, following an encoded sequence of features. The datasets were split into training and testing groups. The training process was managed using the X_train and y_train values. The performance of the models was recorded by generating the predictions on the test datasets (X_test). In contrast, the efficiency of the models was assessed by evaluating their performance through metrics such as accuracy, precision, recall, F1-score, AUC-ROC, among others.
Confusion matrix and AUC-ROC visualization were also used in this study to gain detailed information on the performance of each model. This allowed for TP, TN, FP, and FN identification, while heatmap visualization was presented to enhance the perception of performance complexities in these matrices. Graphs were used to visualize the outputs and comparisons, while the tables illustrate the values assigned to each model’s performance.
The study also employs Principal Component Analysis (PCA), t-distributed Stochastic Neighbor Embedding (t-SNE), and Linear Discriminant Analysis (LDA) to facilitate feature extraction, noise filtering, and the visualization of high-dimensional data. These methods are particularly useful for handling multi-class outputs, such as in Dataset 3, by transforming high-dimensional data into a lower-dimensional space.
5.1. Result Analysis on Dataset 1
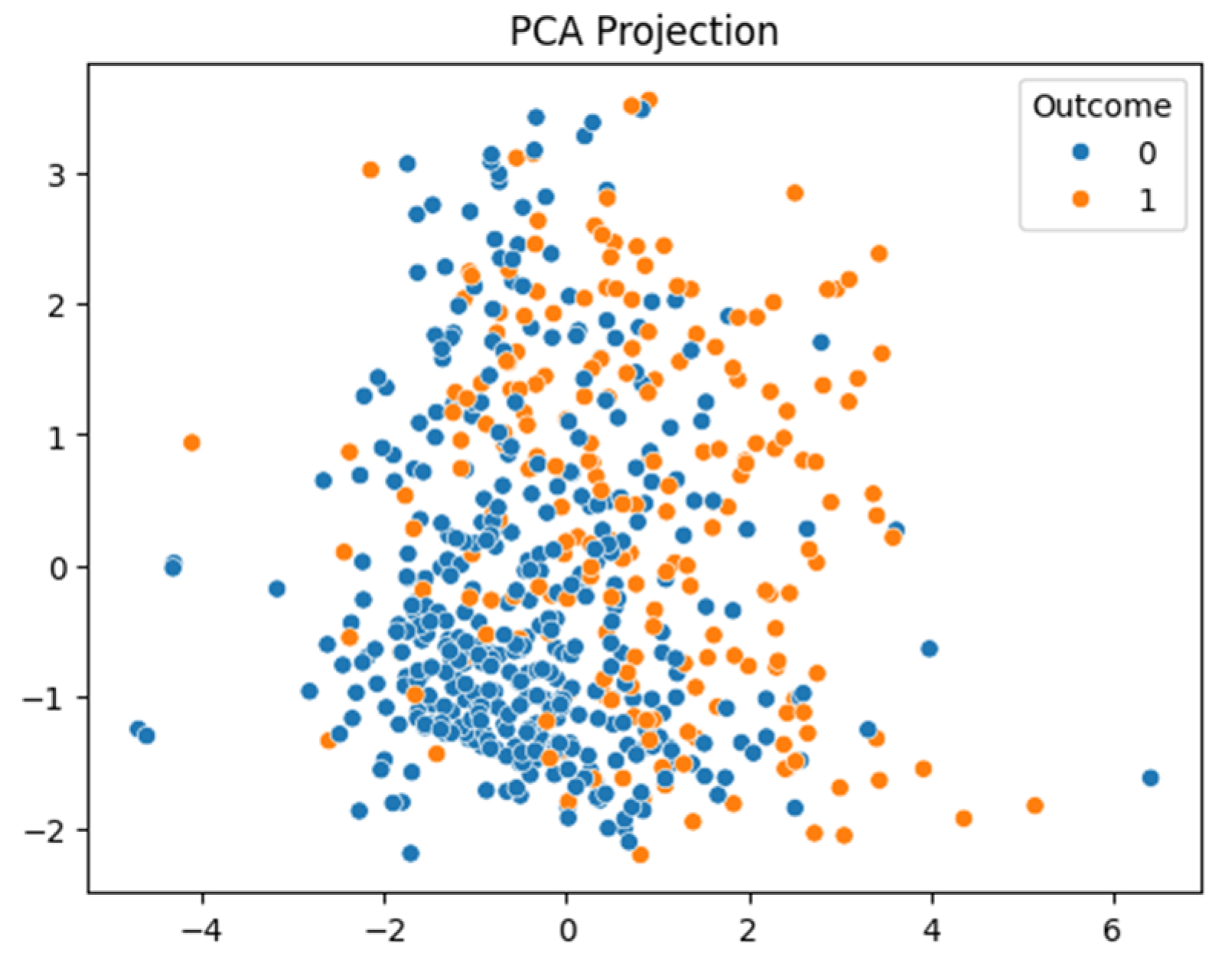
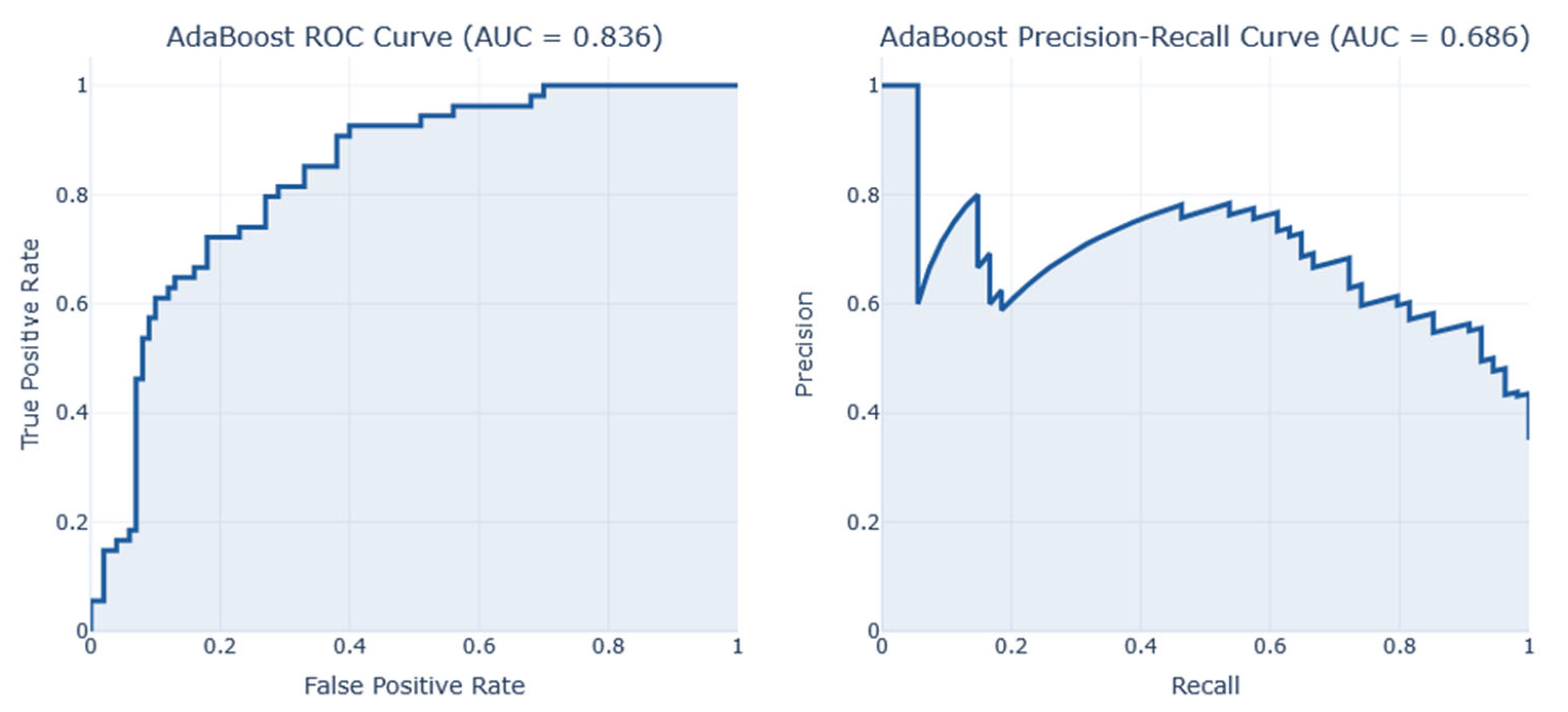
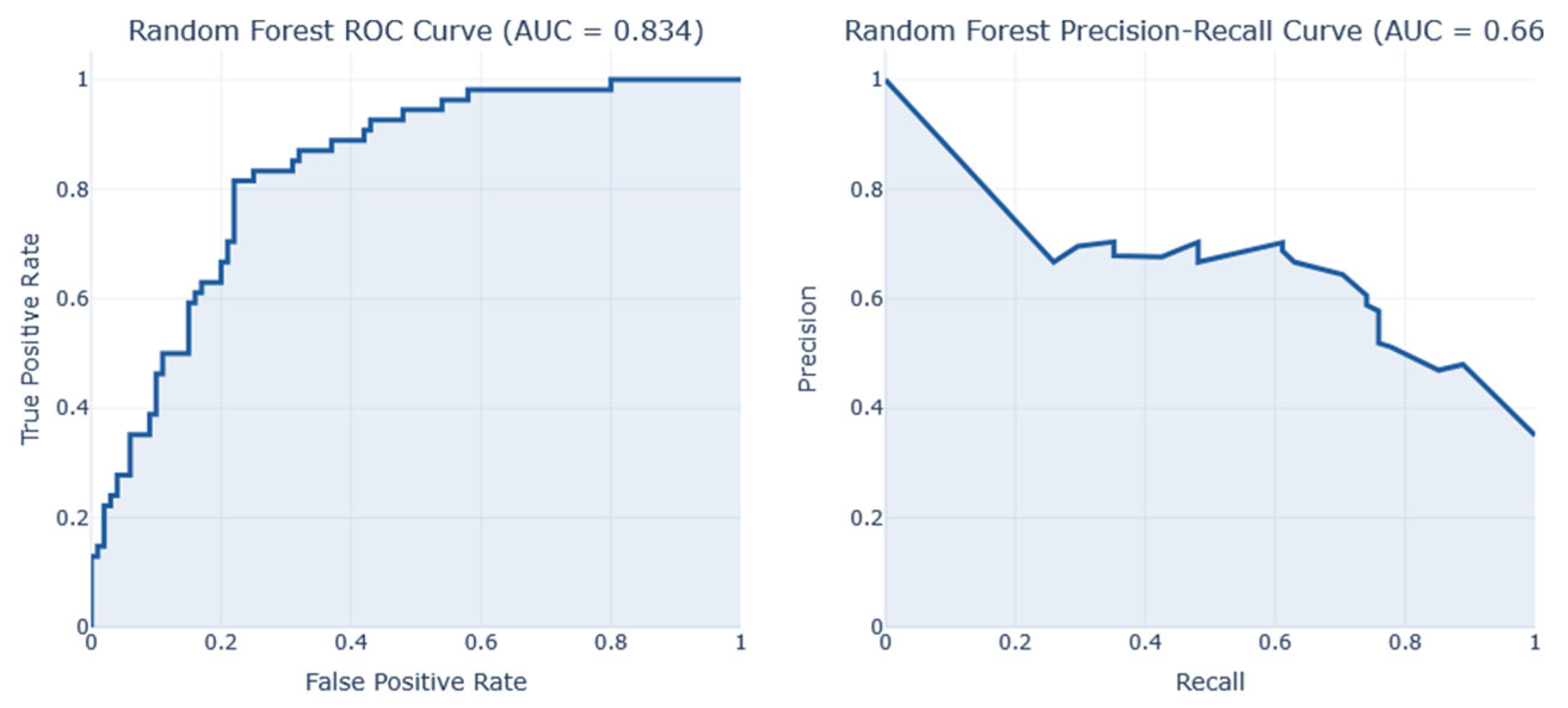
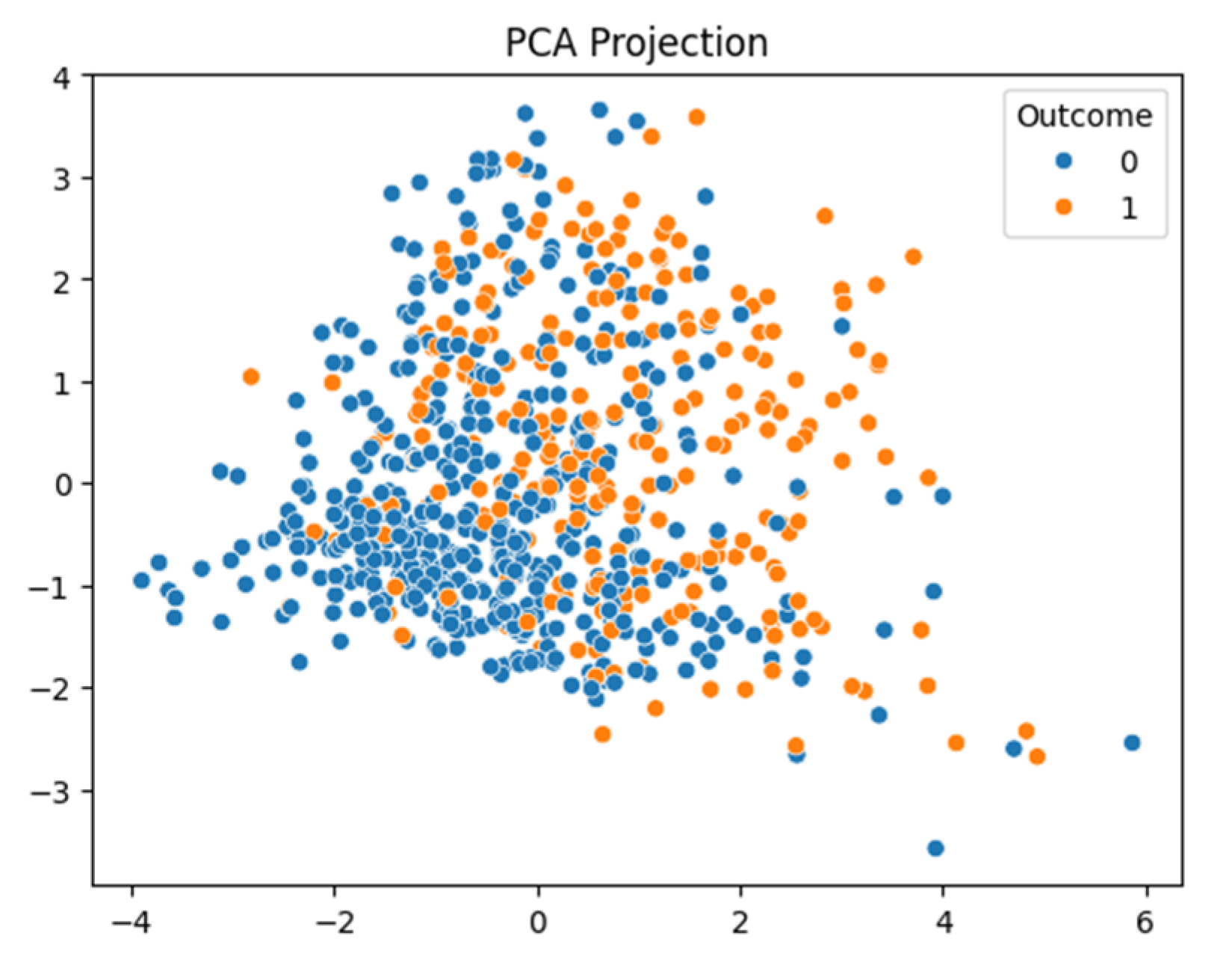
After conducting a series of analyses on Dataset 1 (PIMA—768/9), results are presented as illustrated in
Table 4,
Figure 5,
Figure 6,
Figure 7,
Figure 8. These figures show the analysis outcomes, including the corresponding confusion matrix, precision and recall metrics, the AUC-ROC representation, heatmaps, and the PCA projections of the results. The AdaBoost model performed the best on this dataset, achieving an F1-score of 0.74.
5.2. Result Analysis on Dataset 2
The performance analysis of Dataset 2 (PIMA – 2000/9), presented in
Table 5 and
Figure 9,
Figure 10,
Figure 11, illustrates the results of the analysis, including the confusion matrix, Precision/Recall metrics, AUC-ROC, and PCA projection of the class outcome representation. The RF model demonstrated the highest performance on this dataset, achieving an F1-score of ~0.73.
5.3. Result Analysis on Dataset 3
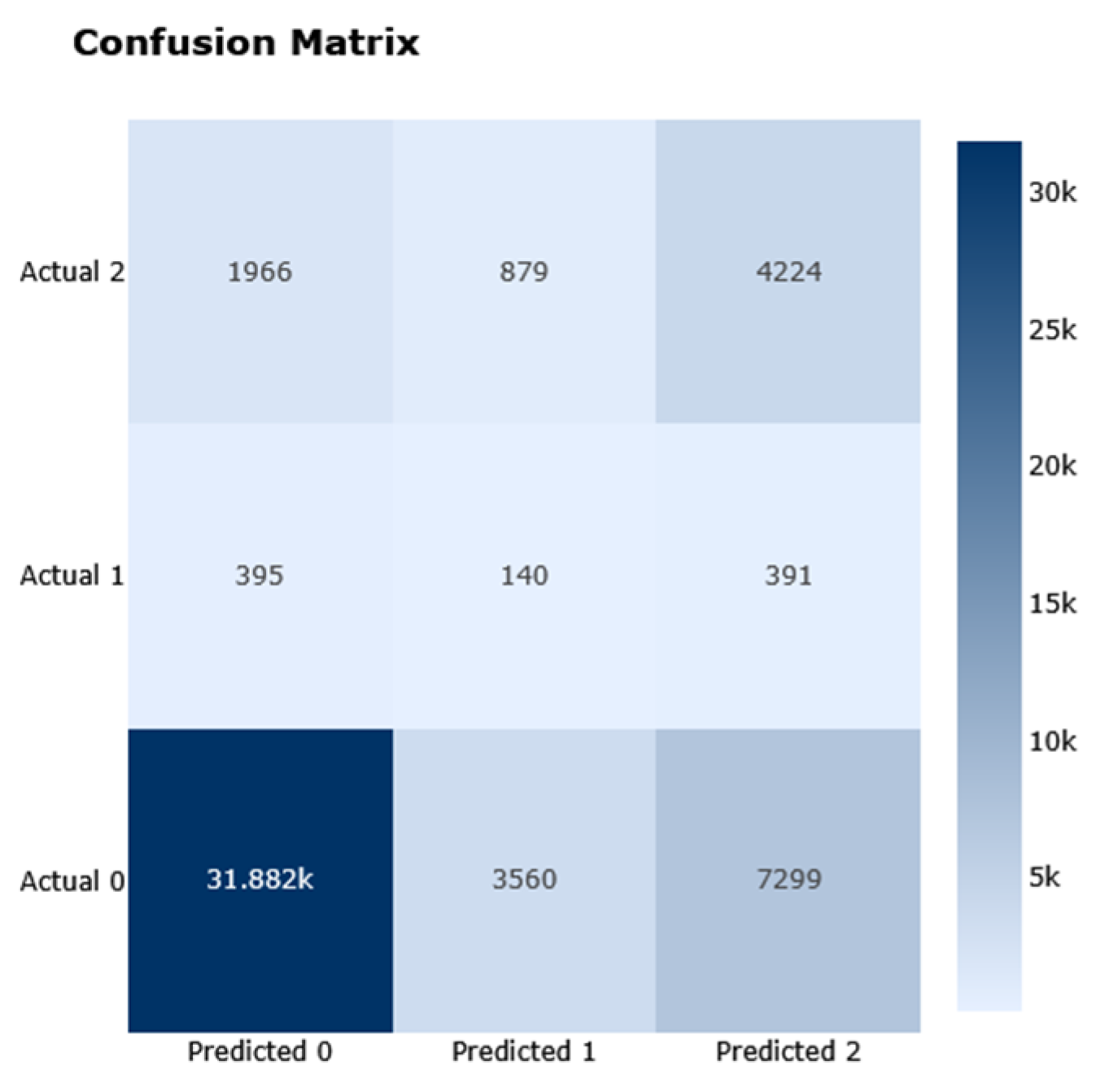
The performance analysis of Dataset 3 (BRFSS), which includes 253,680 samples and 21 features across three outcome classes, is presented in
Table 6 and
Figure 12,
Figure 13,
Figure 14,
Figure 15,
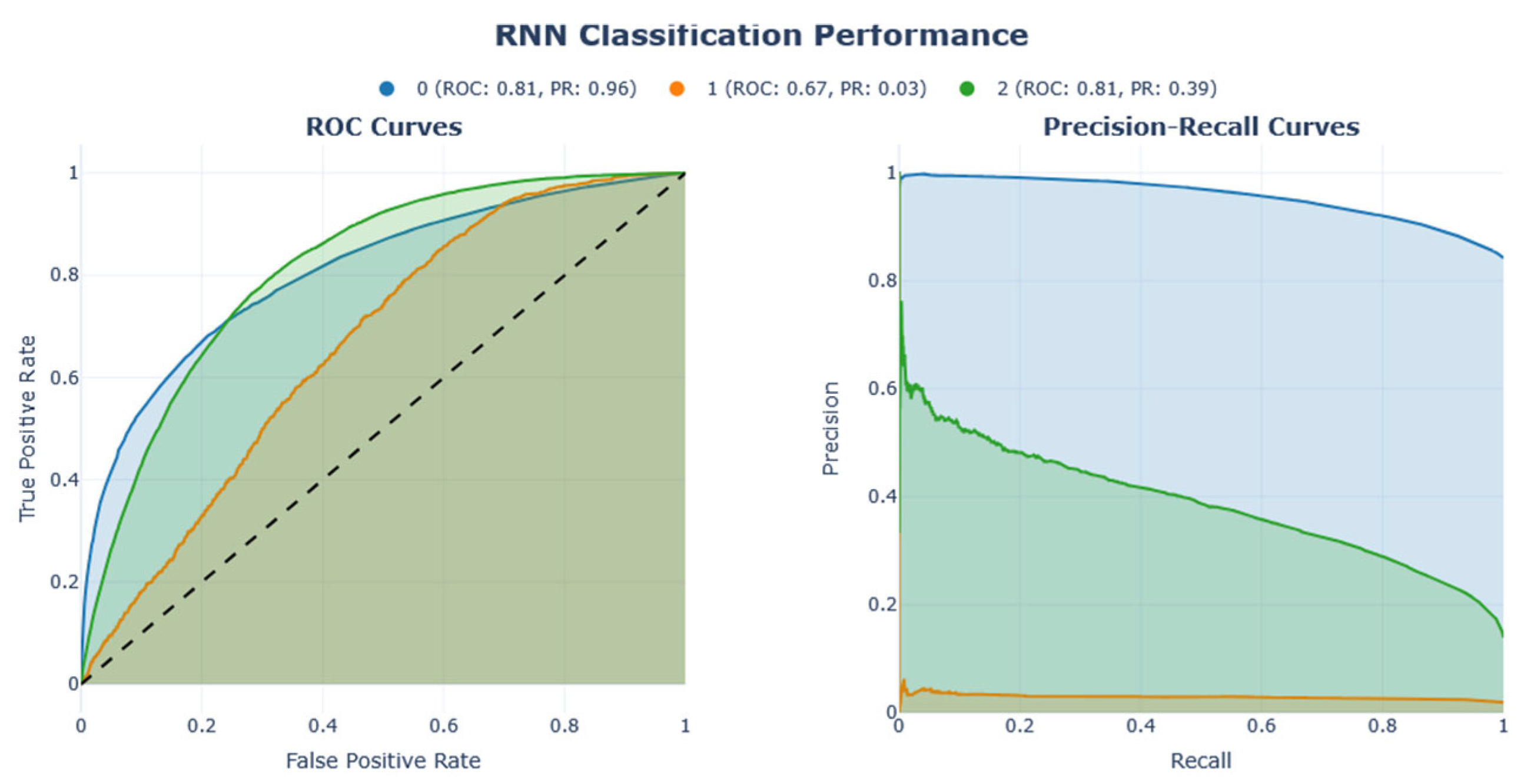
Figure 16. These illustrations demonstrate the results of the analysis, including the corresponding confusion matrix, Precision/Recall metrics, AUC-ROC representation, and projections using LDA, PCA, and t-SNE. The RNN model performed better than other models on this dataset, achieving an F1-score of 0.44.
The analysis of Dataset 3 provides several crucial insights into the structure and complexity of the data, particularly in predicting diabetes status with multiclass outcomes: class 0 (no diabetes or diabetes only during pregnancy), class 1 (pre-diabetes), and class 2 (diabetes).
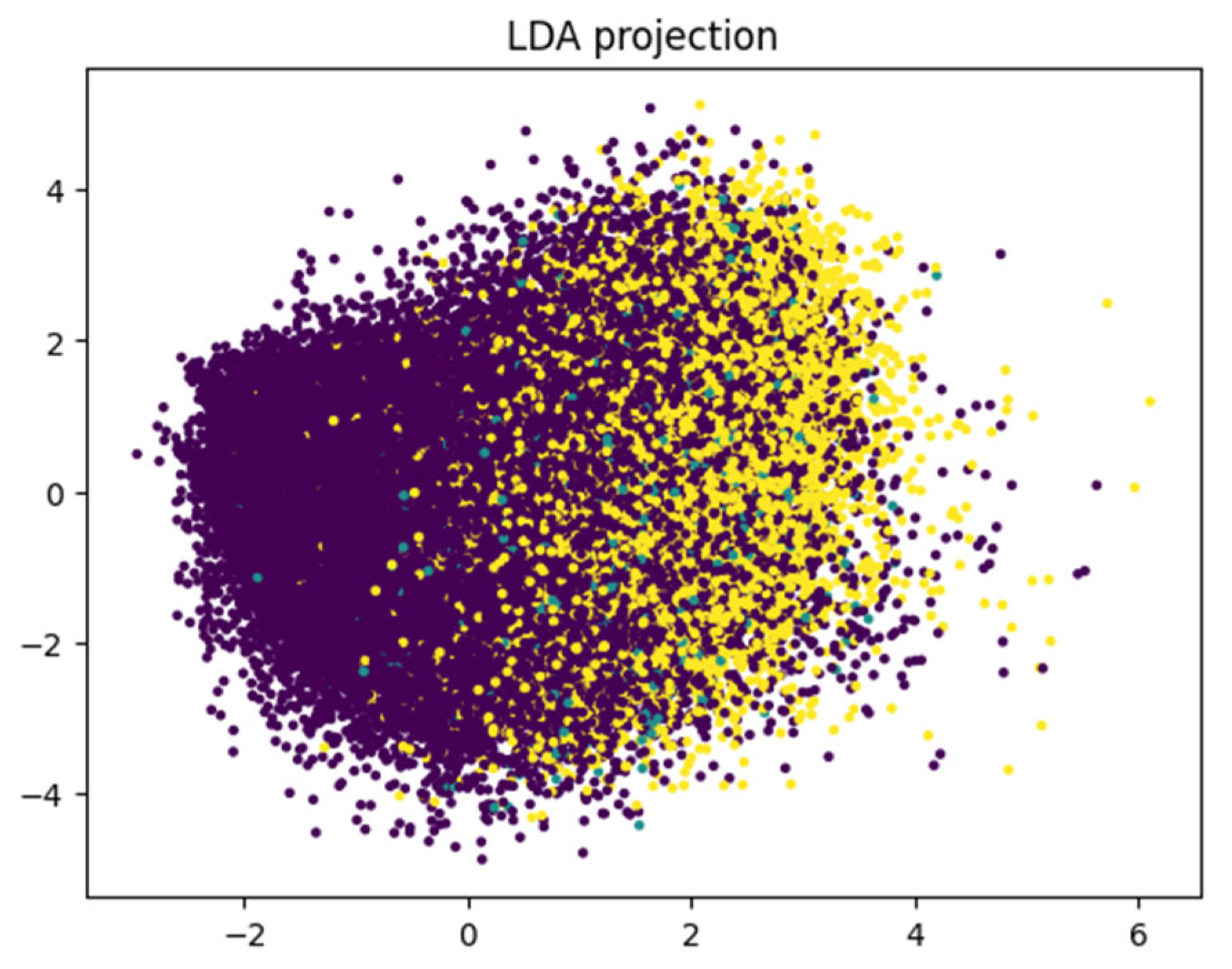
Although the dataset includes medically relevant features such as BMI, blood pressure, cholesterol levels, physical activity, and age, the boundaries between diabetes stages are unclear. This is evident from the projections of LDA and PCA (
Figure 14 and
Figure 15), which show significant overlap, especially between the pre-diabetic and diabetic categories. This suggests that while the features are informative, they may not be sufficient in their linear form to fully distinguish between the classes.
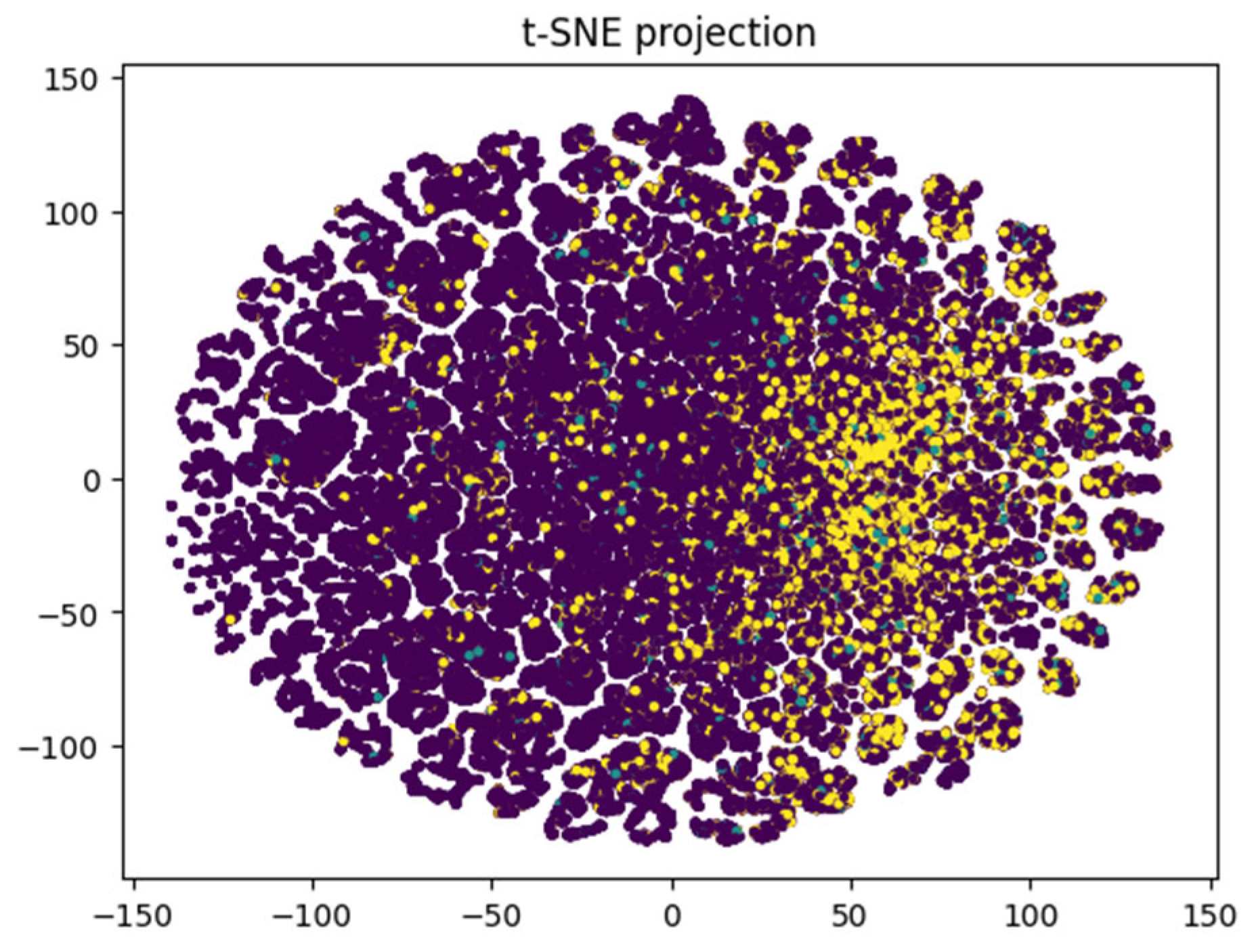
The t-SNE projection reveals more distinct clustering patterns (
Figure 16) compared to linear dimensionality reduction techniques such as PCA or LDA. This suggests the presence of non-linear relationships within the data that linear methods fail to capture. Consequently, this supports the use of more sophisticated ML or DL models capable of modelling such non-linearities. The RNN model achieved an impressive F1-score of 0.44 and accuracy of 0.71, highlighting its ability to effectively utilize complex patterns. Initial insights from the confusion matrix and class distribution analysis confirmed a significant class imbalance, with class 0 (no diabetes) being overrepresented. This imbalance underscores the necessity of employing resampling techniques such as Clustering undersampling to synthetically balance the dataset. Additionally, it highlights the importance of using evaluation metrics like the F1-score, which provide a more balanced assessment of model performance in the presence of skewed class distributions.
Furthermore, all models produced negative R² scores, indicating that none outperformed a naive mean predictor in explaining the variance of the target variable. This suggests a fundamental misalignment between the models' assumptions and the underlying data complexity or target structure. Despite this, evaluation using error-based metrics (MSE, MAE, and RMSE) revealed that RNN and Logistic Regression models achieved the lowest error values (MSE: ~0.79–0.83, MAE: ~0.46–0.51, RMSE: ~0.89–0.91), suggesting relatively better performance in minimizing prediction errors. In contrast, models such as XGBoost-LSTM, Stacking Classifier, and kNN variants exhibited higher error rates and greater variability, indicating less stable predictive behavior. The consistently high error metrics and negative R² values across models highlight challenges in generalization, likely due to overlapping class structures and persistent data imbalance.
5.4. Result Analysis on Dataset 4
Performance analysis on Dataset 4 (BRFSS – 253,680 samples/21 features with two classes outcomes) shown in
Table 7,
Figure 17,
Figure 18,
Figure 19 demonstrate the results of the analysis, its corresponding confusion matrix, Precision/Recall, and the AUC-ROC representation. The DNN model performed better than other models on this dataset, achieving an F1-score of 0.45.
Dataset 4, a binary variant of Dataset 3 (0: no diabetes or pre-diabetes; 1: diabetes), with a 50:50 split (i.e.,
Table 1), also yielded negative R² values across all models, ranging from approximately -1.04 (DNN, GRU) to -1.87 (RNN). These results indicate that none of the models outperformed a naive mean predictor, reinforcing the notion that regression framing may be ill-suited for this classification-oriented task. The persistent data imbalance contributes to the models' inability to capture variance effectively. Despite this, models such as DNN, GRU, and CNN achieved the lowest error rates (MSE and MAE in the range of 0.245–0.265, and RMSE around 0.49–0.51), suggesting better error minimization. These models also demonstrated stronger classification performance, with accuracies around 75% and notably high recall scores. Dataset 4, a binary variant of Dataset 3 (0: no diabetes or pre-diabetes; 1: diabetes), with a 50:50 split (i.e.,
Table 1), also yielded negative R² values across all models, ranging from approximately -1.04 (DNN, GRU) to -1.87 (RNN). These results indicate that none of the models outperformed a naive mean predictor, reinforcing the notion that regression framing may be ill-suited for this classification-oriented task. The persistent data imbalance contributes to the models' inability to capture variance effectively. Despite this, models such as DNN, GRU, and CNN achieved the lowest error rates (MSE and MAE in the range of 0.245–0.265, and RMSE around 0.49–0.51), suggesting better error minimization. These models also demonstrated stronger classification performance, with accuracies around 75% and notably high recall scores.
5.5. Result Analysis on Dataset 5
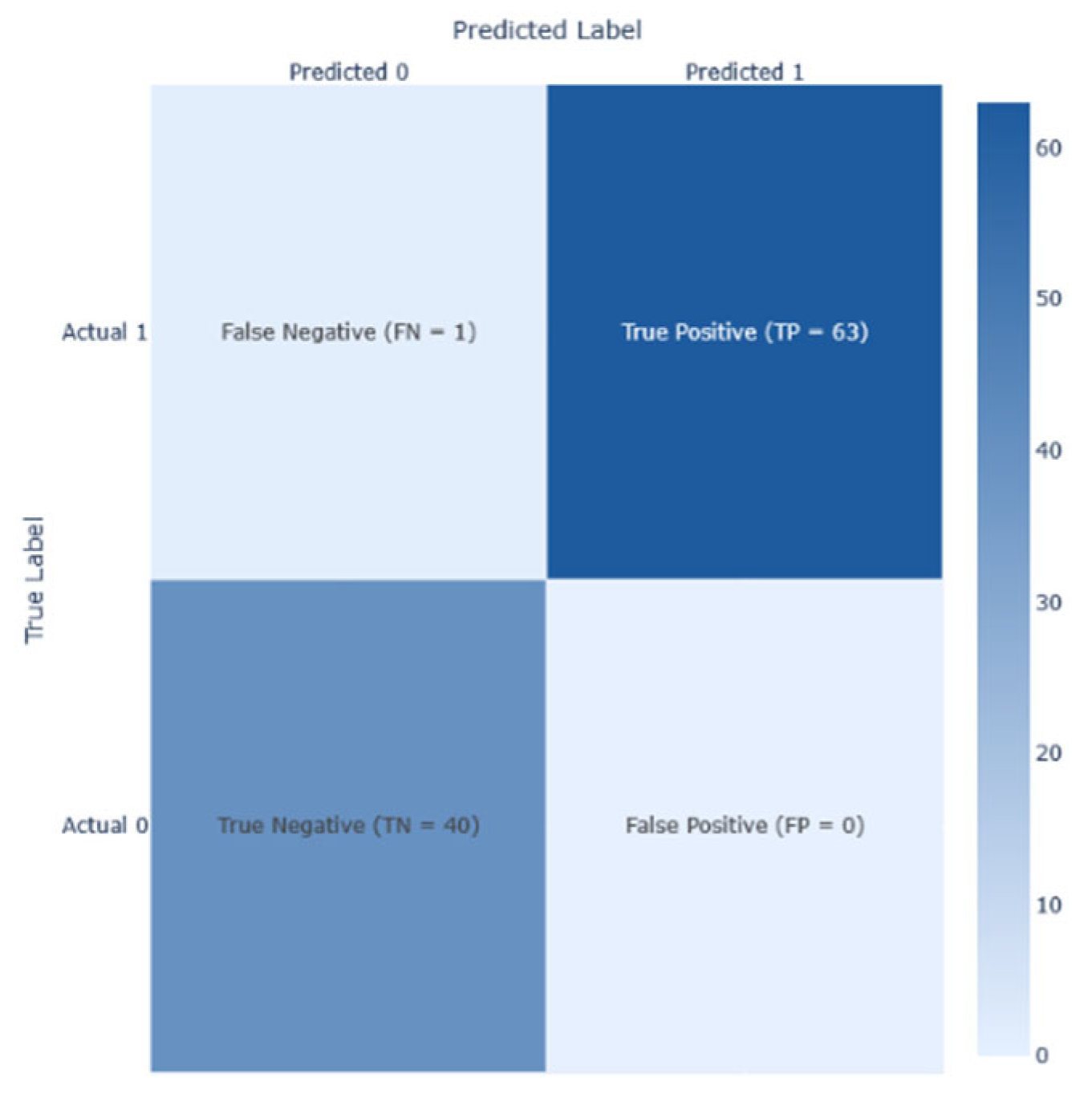
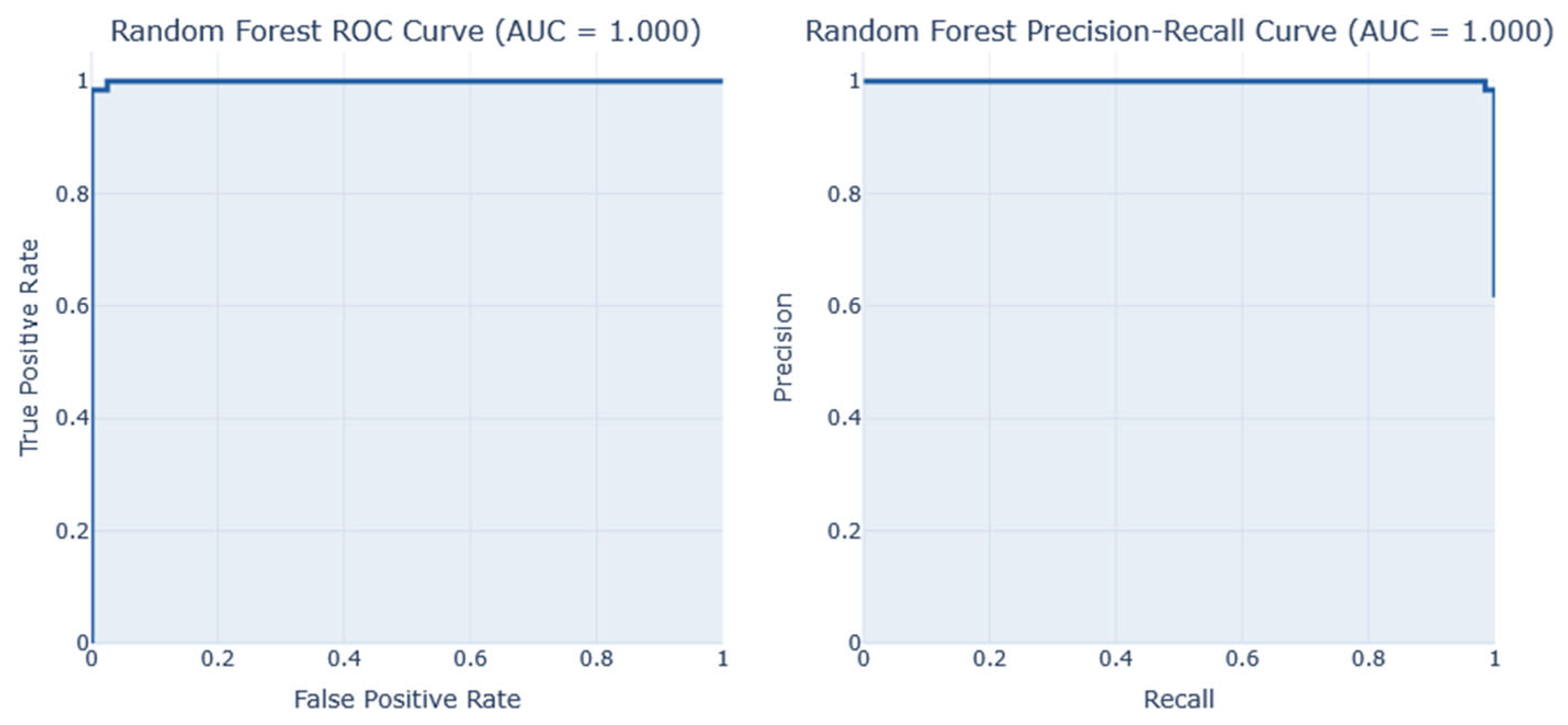
Performance analysis on Dataset 5 (early-stage diabetes risk prediction of patients of 520 samples and 17 features from Sylhet Diabetes Hospital, Bangladesh, shown in
Table 8,
Figure 20 and
Figure 21demonstrates the results of the analysis, its corresponding confusion matrix, Precision/Recall, and the AUC-ROC representation. The RF and Stacking Classifier models performed the best on this dataset, achieving an F1-score of 1.00 and a reasonable accuracy of 1.0 each. However. The Random Forest (RF) is selected as the best due to its lower computation time in predicting diabetes at 0.58s, compared to the Stacking classifier, which took 37.05s.
Regarding both computational efficiency and predictive effectiveness, this study performs a comparative analysis of ML, DL, hybrid models, and ensemble strategies applied to five publicly available datasets, highlighting considerable variations in performance, influenced by model architecture, complexity, and the inherent characteristics of the data. The evaluation utilized critical metrics to identify optimal predictive tools relevant to healthcare settings, with the F1-score serving as a baseline measure.
Ensemble models, particularly Random Forest (RF), AdaBoost, Bagging, and Stacking Classifier, consistently achieved high F1-scores and accuracies across most datasets. Among these, RF and its variants stood out as top performers. AdaBoost achieved an impressive F1-score of 0.7438, using minimal memory (0.0 B) and completing computations in just 1.18 seconds on Dataset 1. This performance significantly surpassed that of deeper models such as LSTM and GRU, which, while consuming more resources (up to 2052 kB and over 20 seconds of computation time), yielded lower F1-scores in the vicinity of 0.56.
In the analysis summarized in
Table 5 on Dataset 2, RF achieved a commendable F1-score of 0.7273 alongside minimal memory usage (32 kB) and a computation time of 0.65 seconds. Similarly, models like Bagging, AdaBoost, and XGBoost demonstrated high precision with reasonable memory requirements, indicating the scalability of ensemble strategies. On the other hand, DL models, particularly GRU and RNN, although exhibiting moderate accuracy, were identified as computationally intensive, with memory usage reaching up to 154790 kB and training times exceeding 1000 seconds.
While
Table 6 illustrates some overall degraded performance attributed to Dataset 3 due to dataset challenges, neural network variants such as RNN, DNN, and CNN showed strong results, with RNN maintaining the highest rank in this context with an F1-score of 0.44.
Table 7 highlighted the performance of DNN and GRU, with both achieving F1-scores between approximately 0.45 and 0.44 on Dataset 4, a variant of Dataset 3., but with two classes (0 and 1). However, their computational costs were high; DNN outperformed GRU with an F1-score of 0.45 while also demonstrating lower computation time and memory usage. In addition, Xie et al. [
76] also proved that NN produces a better accuracy of 0.8240 but a lower recall of 0.3781. This is evident because the dataset size is inadequate for DL models.
Finally,
Table 8 showcased exemplary performance by RF and the Stacking Classifier on Dataset 5, both attaining an F1-score and accuracy of 1.000, which could suggest either overfitting or optimal conditions within the dataset. Random Forest remained the preferred choice due to its reasonable memory consumption of 24 kB. Xie et al. [
78] demonstrated that RF outperformed other classical ML models. However, their analysis reported a score of 0.9740 across all metrics. In contrast, our study achieved a score of 1.0000 using the same model. Overall, the Random Forest model emerged as the most robust and resource-efficient option, delivering consistent high performance while ensuring low memory usage and rapid computation time, making it particularly suited for practical applications in diabetes prediction systems.
There are several key insights to be gained from this study. The quantity, complexity, and structure of the dataset that ML and DL models are trained on affect their performances. Empirical findings from our experiments indicate that conventional ML models are generally most effective on small to moderately sized structured datasets, particularly when the patterns exhibit linear or significantly non-linear separability. When the feature space is small and well-defined, these models benefit from simplicity, reduced computing cost, and strong generalization. DL models like CNNs, DNNs, and RNNs, on the other hand, excel with complex, high-dimensional data such as text, images, or time-series inputs. They require large datasets to avoid overfitting and ensure generalization, but are computationally demanding, frequently requiring large amounts of memory, processing power, and extended training periods. This might provide real-world challenges in settings with limited resources. Aligning dataset properties with model selection is essential for optimal prediction performance, especially in resource-limited environments.
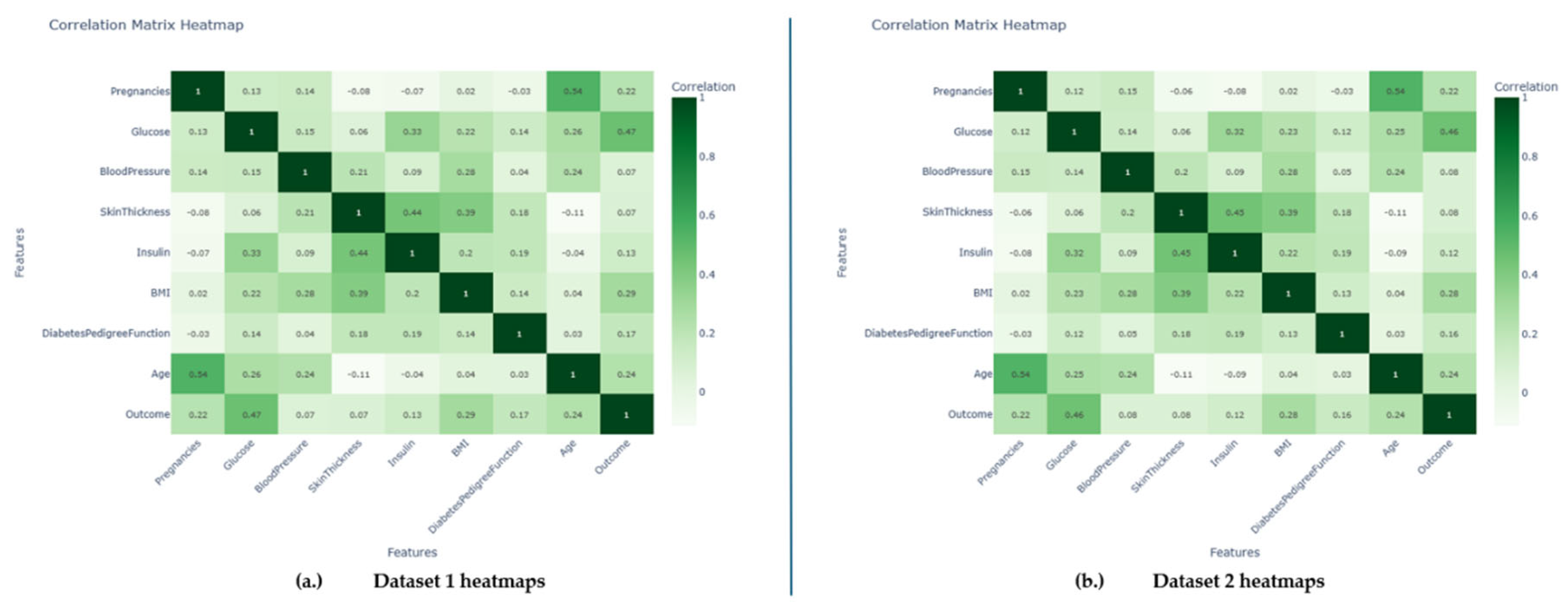
The quality, applicability, and predictive power of the features found in each dataset are primarily responsible for the variation in model performances shown across the various datasets. Specifically, the correlation values of 0.47 and 0.46 in Datasets 1 and 2 (
Figure 8a and 8b) indicate that the characteristic "Glucose" has a comparatively substantial positive link with the diabetic mellitus result. This strong correlation suggests that changes in blood sugar levels are significantly linked to the existence or non-existence of diabetes, which gives predictive algorithms a reliable signal to work with. Therefore, models trained on these datasets perform better because they have high-value features related to the target variable. On the contrary, Dataset 3 shows moderate predictive performance across all evaluated ML and DL models. This result is mainly due to the quality and informativeness of its features, which do not show a strong correlation with the DM outcome. The variables lack discriminative power, reducing model efficacy due to limited signals differentiating diabetic from non-diabetic cases. This highlights the importance of feature selection and dataset quality for achieving accurate predictions in healthcare-related AI applications.
Additionally, the architectural complexity and internal mechanisms of ML and DL models significantly influence differences in processing speed, RAM usage, and overall computing efficiency. Deep learning architecture can differ significantly in the number of parameters, layer depth, and internal processes, all of which directly affect resource usage. For example, LSTM networks are commonly used for sequence modelling due to their strong ability to capture long-range temporal relationships. However, this capability comes at a computational cost. LSTMs require increased model size, higher memory demands, and longer training times because they incorporate multiple gating mechanisms, including input, output, and forget gates, each with its own set of parameters [
16,
68].
GRU is a more lightweight alternative that simplifies the gating process by combining the input and forget gates into a single update gate. This results in a more straightforward architecture with fewer parameters, which accelerates training and reduces memory usage, often with only minor changes in performance. These differences emphasize the importance of aligning model choices with computational constraints, particularly in scenarios requiring real-time processing or when working with limited hardware resources [
16].
6.1. Top-performing Models and Their Implications
The analysis of the study examines the complex relationship between the observed F1-scores and the inference times (TT) of the highest-performing models within the selected datasets. It examines how the unique mechanics of each algorithm align with critical factors such as data size, feature topology, and class structure. By doing so, it uncovers the underlying principles that contribute to model performance. For instance, larger datasets usually necessitate more sophisticated algorithms to manage complexity, while feature topology could influence the model's ability to capture relevant patterns. Additionally, understanding class structure is essential, as imbalanced classes require specialized techniques to ensure accurate predictions, as demonstrated by the ADASYN and Clustering techniques in our study. This comprehensive examination offers valuable insights for selecting and optimizing ML and DL algorithms tailored to specific data characteristics.
High F1-score arises when a model’s bias–variance profile and feature handling align with the dataset’s intrinsic complexity. In contrast, run-time reflects algorithmic depth and feature dimensionality; that is, shallow boosted or bagged trees provide quick, accurate results on small tabular data, while recurrent or fully connected nets sacrifice speed for the representational power needed to model high-dimensional, progression-laden surveys.
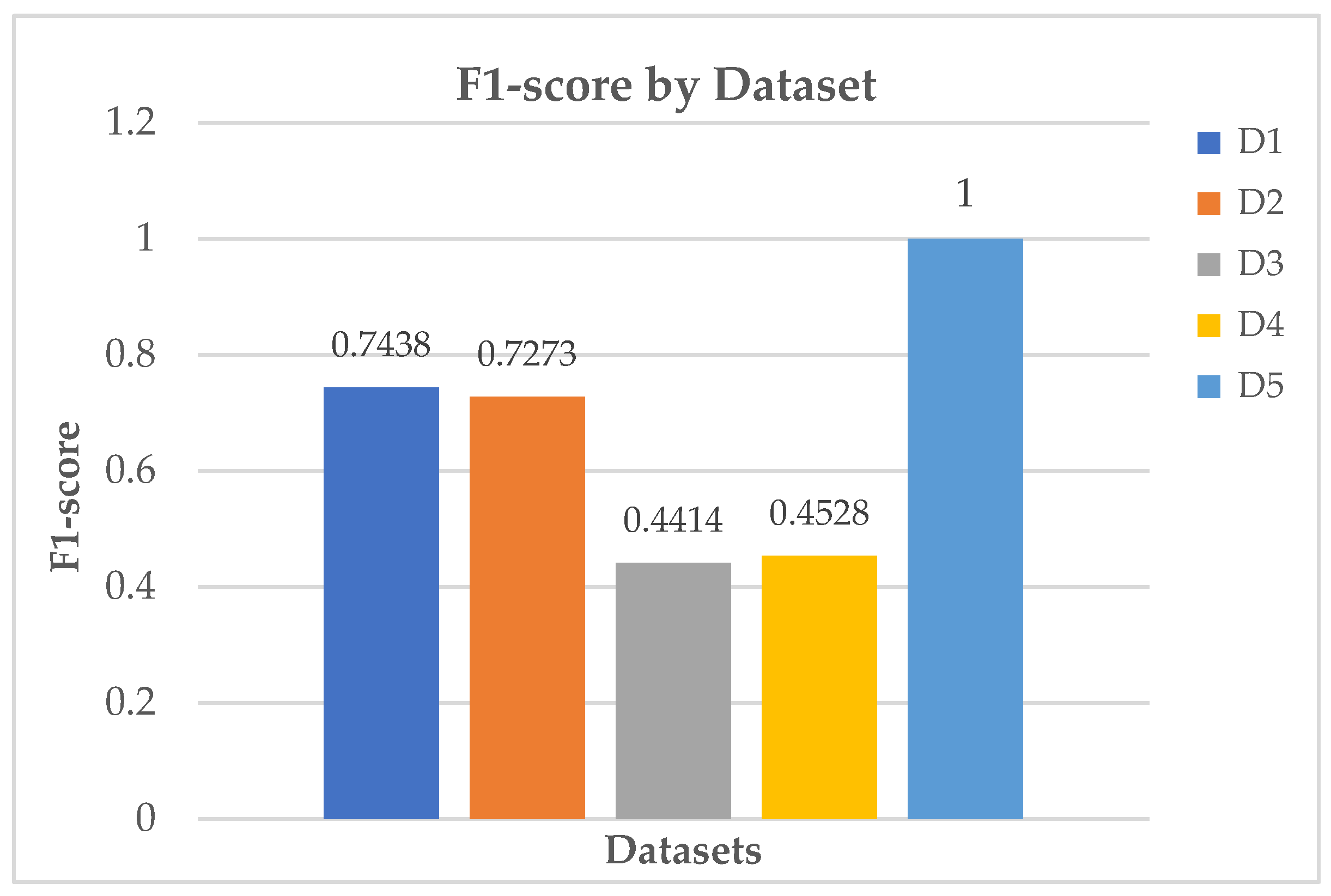
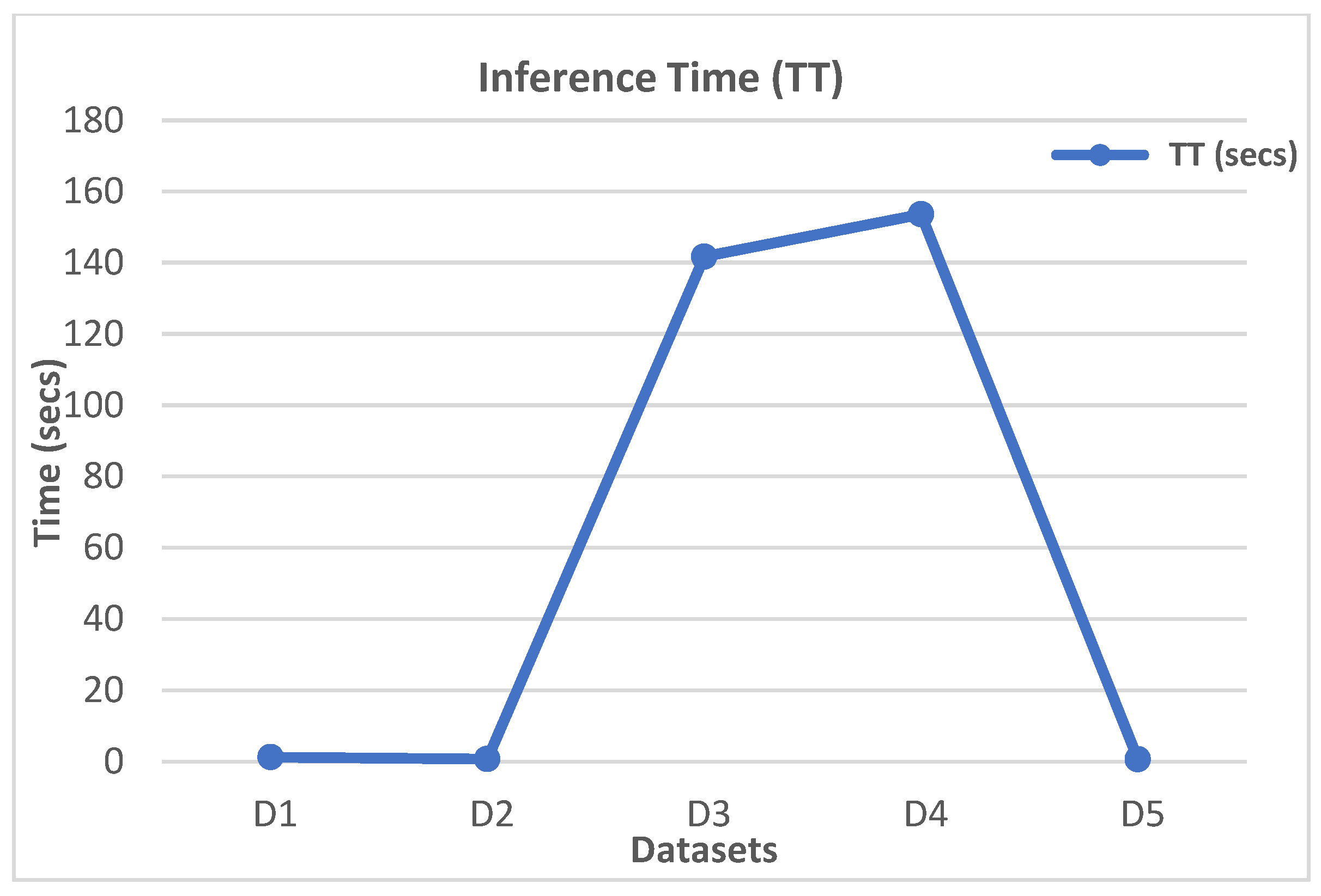
Table 9,
Figure 22, and
Figure 23 depict the extracted top-performing models and their respective computation times.
The first variant of the PIMA dataset (Dataset 1/D1) is small by today’s ML standards, with only 614 training samples after the 80:20 train-test split, and 9 mostly straightforward numeric features, making it a manageable challenge for analysis. Simple models like decision stumps can capture some patterns, but they often struggle with hard-to-classify cases, especially around borderline pregnancies and rare insulin levels. AdaBoost works well in this situation by focusing on the misclassified data points for improvement. The algorithm changes the weight of these difficult cases, creating a series of weak classifiers that better identify and understand these minority areas, while keeping the model simple. Given the low-dimensional nature of the data, AdaBoost demonstrates a reduced likelihood of overfitting. It reduces bias effectively while only slightly increasing variance. Using oversampling techniques like ADASYN boosts AdaBoost’s performance even more. This method creates a denser group of hard-to-classify cases, giving AdaBoost an edge over other methods like bagged DTs and NNs. This combination leads to a stronger model for classifying challenging data
With 2,000 observations, the second version of the PIMA dataset (Dataset 2/D2) provides sufficient samples for high-capacity models, while still maintaining the same features. In this context, the RF algorithm performs best because the dominant source of error is variance rather than bias, as in Dataset 1. The additional data points help reduce bias naturally, but the dataset still includes noisy measurements, such as imputed zeros, which can mislead individual trees or boosted models. Using bagging to create hundreds of decorrelated trees stabilizes predictions and captures non-linear interactions, such as the thresholds between glucose and BMI. Additionally, RF incorporates built-in resilience to class imbalance through balanced subsampling at each split. Inference remains fast (less than 0.7 seconds) because only a few dozen features are evaluated per tree, giving RF the best speed-to-accuracy ratio in this scenario.
Dataset 3 (D3) presents the full BRFSS survey categorizes diabetes status on an ordinal scale: 0 = No diabetes, 1 = prediabetes, and 2 = diabetes, emphasizing progression in conditions. Many of the 21 features in the survey represent behavioural patterns, such as weekly exercise, daily sugar intake, and smoking frequency, which are often autocorrelated and recorded as ordered categorical bands. After applying Clustering-based undersampling balancing, a RNN model can interpret each respondent’s feature vector as a short "time-axis," where neighbouring fields demonstrate interdependence (e.g., age band à blood pressure band à medication usage). The gated recurrent mechanism of the RNN integrates these conditional patterns more effectively than feed-forward networks or tree ensembles, leading to the highest macro-F1 score despite longer inference times. In summary, the RNN effectively utilizes the quasi-sequential, progression-based structure that tabular models treat as independent columns
Dataset 4 (D4) presents the multi-class labels being collapsed into a binary outcome, although it still contains over 56,000 training rows and a heterogeneous mix of ordinal, binary, and scaled numeric features. The class boundary now resides in a densely populated area where subtle high-order interactions, such as age × BMI × physical activity or diet score × sex, become crucial. A deep, fully connected network with multiple hidden layers can automatically learn these hierarchical combinations, especially after applying feature scaling and clustering-based undersampling to improve the representation of minority classes. Compared to tree ensembles, DNNs benefit from weight sharing and batch optimization, making them less sensitive to redundant variables and more tolerant of noise. Thus, their slightly superior F1-score reflects an architecture that is adequately expressive for the high-dimensional, highly non-linear boundary while remaining computationally efficient.
Dataset 5 (D5) presents an EMR dataset from the early-stage Sylhet survey, containing 520 records with 17 binary symptoms and a 5-band age code, validated by a physician. This clean, categorical data is ideal for decision-tree splits, and with RF emerging as the top-performing model, offers three advantages: (1) Low variance via bootstrapping prevents overfitting common in single trees with limited data. (2) It efficiently processes binary inputs, resulting in clear leaf nodes without complicated weighting. (3) It discovers non-linear symptom interactions (e.g., polyuria ^ polydipsia ^ age > 45) that linear models miss while achieving perfect class separation. The result is an F1 score of 1.00 in under 0.6 seconds, outperforming stacking classifiers and NNs.
6.2. Comparative Analysis of Results with already developed diabetes prediction models.
The analysis presented evaluates various approaches, including ML, DL, hybrid models, and ensemble strategies, in predicting health outcomes for diabetic patients. The outcomes generated from these methods were compared against other existing predictive models utilizing multiple datasets (specifically Datasets 1 – 5). The Random Forest (RF) model demonstrated exceptional performance, achieving high F1-scores, accuracy, and efficient computation times.. In contrast, other ML models also delivered commendable results in terms of accuracy, speed, F1-scores, and AUC-ROC, all within a reasonable timeframe for computation. Additionally, some DL models and ensemble strategies showed promising results based on the same dataset samples and features. A comprehensive comparative analysis of the performance of the models in this study, relative to existing predictive model research, can be found in
Table 10.
7. Conclusions
People of all ages are becoming more susceptible to diabetes. The current study showed that early diabetes identification might be crucial for treatment and enhanced health outcomes for individuals with the disease. Obesity may be prevented by taking easy awareness-raising steps like eating a low-sugar diet, exercising frequently, and leading a healthy lifestyle. Its relevance in healthcare is apparent since models and ensemble strategies show increasing promise in predicting diabetes and eventually lowering treatment costs and increasing computing efficiency. Finding the optimal model for predicting datasets created for diabetes progression and risk prediction is the primary contribution of this work.
Choosing the best ML or DL model to predict clinical outcomes in diabetes patients relies heavily on the characteristics of the dataset used; there is no universally optimal model. Key factors that can significantly influence model performance include sample size, feature richness (the variety and significance of input variables), and data distribution across classes. A model may perform poorly on a smaller or more diverse dataset that has missing values or imbalanced classes, even if it excels on a larger, balanced, and feature-rich dataset. Furthermore, how well models generalize can be affected by slight variations in clinical recording procedures, population characteristics, and measurement standards across different institutions.
In this study, traditional ML models, including Random Forest (RF) and AdaBoost demonstrated superior predictive performance on Datasets 1, 2, and 5. These datasets were characterized by relatively small sample sizes and structured data formats. The ML models are less data-intensive by nature and perform effectively in low-data environments, particularly when the datasets contain high-quality and well-engineered features. Their ensemble-based architecture helps reduce variance and improve robustness, making them well-suited for medical datasets where data may be limited but well-defined.
Deep learning models, especially RNNs and DNNs, demonstrated superior performance compared to traditional ML models on Datasets 3 and 4. These datasets were significantly larger and more complex, featuring high-dimensional feature spaces and potentially nonlinear patterns, conditions where deep learning models excel. DL architectures are designed to learn hierarchical and abstract representations of features, enabling them to capture intricate, non-linear relationships that traditional ML algorithms might struggle to detect. However, the enhanced performance of DL models relies heavily on the availability of large, diverse datasets and adequate computational resources for training. These results underscore the established differences in the suitability of ML versus DL models across various data scenarios. Nonetheless, our prediction algorithms could be more effective in forecasting the health outcomes of diabetes patients now that clinical data and biomarkers are available.
We strongly recommend clinical researchers, data scientists, and healthcare practitioners against relying solely on benchmark performances reported in the literature. It is advised that before implementing any prediction tool for practical use, it is essential to conduct a thorough assessment and validation of the model using their institution's datasets. This approach enhances accuracy, security, and confidence in AI-assisted healthcare decision-making while also improving alignment with regional patient characteristics and clinical workflows.
Author Contributions
Conceptualization, methodology, software, validation, formal analysis, investigation, resources, data curation, writing—original draft preparation, O.B.A.; writing—review and editing, visualization, supervision, project administration, S.S., C.R.; funding acquisition, S.S., O.B.A. All authors have read and agreed to the published version of the manuscript
Funding
The APC charge was funded by a waiver from the Journal’s Guest Editor to my Principal Supervisor. The PhD research work was funded by the Australian Research Training Program (RTP) Postgraduate Research Scholarship Award, under the Australian Government.
Institutional Review Board Statement
Not applicable
Informed Consent Statement
Not Applicable
Data Availability Statement
All datasets used in this research are publicly available in Kaggle, CDC, and UCI Machine Learning Repository.
Acknowledgments
This work is part of the doctorate research under the Research Training Program (RTP) scholarship opportunity. The authors thank the anonymous reviewers for their valuable suggestions and comments on this paper.
Conflicts of Interest
The authors declare no conflicts of interest.
Abbreviations
DM Diabetes Mellitus
ML Machine Learning
DL Deep Learning
AU-ROC Area under the ROC
KPI Key Performance Indicators
IDF International Diabetes Federation
T1DM Type 1 DM
T2DM Type 2 DM
GDM Gestational DM
RF Random Forest
LR Logistic Regression
XGBoost Extreme Gradient Boosting
NB Naive Bayes
SVM Support Vector Machine
NN Neural Networks
RNN Recurrent NN
CNN Convolutional NN
DNN Deep NN
QML Quantum ML
KNN k-Nearest Neighbour
CVD Cardiovascular diseases
DT Decision Trees
LSTM Long Short-Term Memory
AdaBoost Adaptive Boosting
GRU Gated Recurrent Unit
ANN Artificial Neural Networks
MU Memory Usage
TT Inference time |
References
- Kavakiotis, I. O. Tsave, A. Salifoglou, N. Maglaveras, I. Vlahavas, and I. Chouvarda, "Machine Learning and Data Mining Methods in Diabetes Research," Computational and Structural Biotechnology Journal, vol. 15, pp. 104-116, 2017. [CrossRef]
- IDF. International Diabetes Federation (IDF) Diabetes Atlas 2021 (IDF Atlas 2021); International Diabetes Federation: Brussels, Belgium, 2021; pp. 1–141.
- Refat, M.A.R.; Amin, M.A.; Kaushal, C.; Yeasmin, M.N.; Islam, M.K. A Comparative Analysis of Early Stage Diabetes Prediction using Machine Learning and Deep Learning Approach. In Proceedings of the 6th IEEE International Conference on Signal Processing, Computing and Control (ISPCC), Solan, India, 7–9 October 2021; IEEE: Piscataway, NJ, USA, 2021; pp. 654–659. [CrossRef]
- Ayon, I.S.; Islam, M.M. Diabetes Prediction: A Deep Learning Approach. Int. J. Inf. Eng. Electron. Bus. 2019, 11, 21–27. [CrossRef]
- Butt, U.M.; Letchmunan, S.; Ali, M.; Hassan, F.H.; Baqir, A.; Sherazi, H.H.R.; Espino, D. Machine Learning Based Diabetes Classification and Prediction for Healthcare Applications. J. Healthc. Eng. 2021, 2021, 9930985. [CrossRef]
- David, S.A.; Varsha, V.; Ravali, Y.; Naga Amrutha Saranya, N. Comparative Analysis of Diabetes Prediction Using Machine Learning. In Soft Computing for Security Applications; Ranganathan, G., Fernando, X., Piramuthu, S., Eds.; Advances in Intelligent Systems and Computing; Springer: Singapore, 2022; Volume 1428, pp. 155–163, Chapter 13.
- Longato, E.; Fadini, G.P.; Sparacino, G.; Avogaro, A.; Tramontan, L.; Di Camillo, B. A Deep Learning Approach to Predict Diabetes’ Cardiovascular Complications From Administrative Claims. IEEE J. Biomed. Health Inform. 2021, 25, 3608–3617. [CrossRef]
- Saeedi, P.; Petersohn, I.; Salpea, P.; Malanda, B.; Karuranga, S.; Unwin, N.; Colagiuri, S.; Guariguata, L.; Motala, A.A.; Ogurtsova, K.; et al. Global and regional diabetes prevalence estimates for 2019 and projections for 2030 and 2045: Results from the International Diabetes Federation Diabetes Atlas, 9(th) editions. Diabetes Res. Clin. Pract. 2019, 157, 107843. [CrossRef]
- Zarkogianni, K.; Athanasiou, M.; Thanopoulou, A.C.; Nikita, K.S. Comparison of Machine Learning Approaches Toward Assessing the Risk of Developing Cardiovascular Disease as a Long-Term Diabetes Complication. IEEE J. Biomed. Health Inform. 2018, 22, 1637–1647. [CrossRef]
- Dinh, A.; Miertschin, S.; Young, A.; Mohanty, S.D. A data-driven approach to predicting diabetes and cardiovascular disease with machine learning. BMC Med. Inf. Decis. Mak. 2019, 19, 211. [CrossRef]
- Hasan, M.M.; Ahmad, S.; Ahmed, A.H.; Sayed, A.; Mia, T.; Ayon, E.H.; Koli, T.; Thakur, H.N. Cardiovascular Disease Prediction Through Comparative Analysis of Machine Learning Models. In Proceedings of the 2023 International Conference on Modelling & E-Information Research, Artificial Learning and Digital Applications (ICMERALDA), Karawang, Indonesia, 24 November 2023.
- Lin, X.; Xu, Y.; Pan, X.; Xu, J.; Ding, Y.; Sun, X.; Song, X.; Ren, Y.; Shan, P.F. Global, regional, and national burden and trend of diabetes in 195 countries and territories—An analysis from 1990 to 2025. Sci. Rep. 2020, 10, 14790. [CrossRef]
- Kodama, S.; Fujihara, K.; Horikawa, C.; Kitazawa, M.; Iwanaga, M.; Kato, K.; Watanabe, K.; Nakagawa, Y.; Matsuzaka, T.; Shimano, H.; et al. Predictive ability of current machine learning algorithms for type 2 diabetes mellitus: A meta-analysis. J. Diabetes Investig. 2022, 13, 900–908. [CrossRef]
- Larabi-Marie-Sainte, S.; Aburahmah, L.; Almohaini, R.; Saba, T. Current Techniques for Diabetes Prediction: Review and Case Study. Appl. Sci. 2019, 9, 4604. [CrossRef]
- Islam, S.; Tariq, F. Machine Learning-Enabled Detection and Management of Diabetes Mellitus. In Artificial Intelligence for Disease Diagnosis and Prognosis in Smart Healthcare; Ghita Kouadri Mostefaoui, S. M. Riazul Islam, and Tariq F.; Eds.; CRC Press: Boca Raton, New York, USA; 2020, Chapter 12, pp. 113–125. 2023; pp. 203–218. [CrossRef]
- Afsaneh, E.; Sharifdini, A.; Ghazzaghi, H.; Ghobadi, M.Z. Recent applications of machine learning and deep learning models in the prediction, diagnosis, and management of diabetes: A comprehensive review. Diabetol. Metab. Syndr. 2022, 14, 196. [CrossRef]
- Giacomo, C.; Martina, V.; Giovanni, S.; Andrea, F. Continuous Glucose Monitoring Sensors for Diabetes Management—A Review of Technologies and Applications. Diabetes Metab. J. 2019, 43, 383–397. [CrossRef]
- Nomura, A.; Noguchi, M.; Kometani, M.; Furukawa, K.; Yoneda, T. Artificial Intelligence in Current Diabetes Management and Prediction. Curr. Diab Rep. 2021, 21, 61. [CrossRef]
- Guan, Z.; Li, H.; Liu, R.; Cai, C.; Liu, Y.; Li, J.; Wang, X.; Huang, S.; Wu, L.; Liu, D.; et al. Artificial intelligence in diabetes management: Advancements, opportunities, and challenges. Cell Rep. Med. 2023, 4, 101213. [CrossRef]
- Lu, H.Y.; Ding, X.; Hirst, J.E.; Yang, Y.; Yang, J.; Mackillop, L.; Clifton, D.A. Digital Health and Machine Learning Technologies for Blood Glucose Monitoring and Management of Gestational Diabetes. IEEE Rev. Biomed. Eng. 2024, 17, 98–117. [CrossRef]
- Ba, T.; Li, S.; Wei, Y. A data-driven machine learning integrated wearable medical sensor framework for elderly care service. Measurement 2021, 167, 108383. [CrossRef]
- Kakoly, I.J.; Hoque, M.R.; Hasan, N. Data-Driven Diabetes Risk Factor Prediction Using Machine Learning Algorithms with Feature Selection Technique. Sustainability 2023, 15, 4930. [CrossRef]
- Mora, T.; Roche, D.; Rodriguez-Sanchez, B. Predicting the onset of diabetes-related complications after a diabetes diagnosis with machine learning algorithms. Diabetes Res. Clin. Pract. 2023, 204, 110910. [CrossRef]
- Han, B.C.; Kim, J.; Choi, J. Prediction of complications in diabetes mellitus using machine learning models with transplanted topic model features. Biomed. Eng. Lett. 2024, 14, 163–171. [CrossRef]
- Dagliati, A.; Marini, S.; Sacchi, L.; Cogni, G.; Teliti, M.; Tibollo, V.; De Cata, P.; Chiovato, L.; Bellazzi, R. Machine Learning Methods to Predict Diabetes Complications. J. Diabetes Sci. Technol. 2018, 12, 295–302. [CrossRef]
- Ochocinski, D.; Dalal, M.; Black, L.V.; Carr, S.; Lew, J.; Sullivan, K.; Kissoon, N. Life-Threatening Infectious Complications in Sickle Cell Disease: A Concise Narrative Review. Front. Pediatr. 2020, 8, 38. [CrossRef]
- Tan, K.R.; Seng, J.J.B.; Kwan, Y.H.; Chen, Y.J.; Zainudin, S.B.; Loh, D.H.F.; Liu, N.; Low, L.L. Evaluation of Machine Learning Methods Developed for Prediction of Diabetes Complications: A Systematic Review. J. Diabetes Sci. Technol. 2023, 17, 474–489. [CrossRef]
- Chauhan, A.S.; Varre, M.S.; Izuora, K.; Trabia, M.B.; Dufek, J.S. Prediction of Diabetes Mellitus Progression Using Supervised Machine Learning. Sensors 2023, 23, 4658. [CrossRef]
- Skyler, J.S.; Bakris, G.L.; Bonifacio, E.; Darsow, T.; Eckel, R.H.; Groop, L.; Groop, P.-H.; Handelsman, Y.; Insel, R.A.; Mathieu, C.; et al. Differentiation of Diabetes by Pathophysiology, Natural History, and Prognosis. Diabetes 2017, 66, 241–255. [CrossRef]
- Banday, M.Z.; Sameer, A.S.; Nissar, S. Pathophysiology of diabetes—An overview. Avicenna J. Med. 2020, 10, 174–188. [CrossRef]
- Fujimoto, W.Y. The Importance of Insulin Resistance in the Pathogenesis of Type 2 Diabetes Mellitus. Am. J. Med. 2000, 108, 9S–14S. [CrossRef]
- Galicia-Garcia, U.; Benito-Vicente, A.; Jebari, S.; Larrea-Sebal, A.; Siddiqi, H.; Uribe, K.B.; Ostolaza, H.; Martín, C. Pathophysiology of Type 2 Diabetes Mellitus. Int. J. Mol. Sci. 2020, 21, 6275. [CrossRef]
- Agliata, A.; Giordano, D.; Bardozzo, F.; Bottiglieri, S.; Facchiano, A.; Tagliaferri, R. Machine Learning as a Support for the Diagnosis of Type 2 Diabetes. Int. J. Mol. Sci. 2023, 24, 6775. [CrossRef]
- McIntyre, H.D.; Catalano, P.; Zhang, C.; Desoye, G.; Mathiesen, E.R.; Damm, P.; Primers, N.R.D. Gestational diabetes mellitus. Nat. Reviews. Dis. Primers 2019, 5, 47. [CrossRef]
- Plows, J.F.; Stanley, J.L.; Baker, P.N.; Reynolds, C.M.; Vickers, M.H. The Pathophysiology of Gestational Diabetes Mellitus. Int. J. Mol. Sci. 2018, 19, 3342. [CrossRef]
- Ahmad, R.; Narwaria, M.; Haque, M. Gestational diabetes mellitus prevalence and progression to type 2 diabetes mellitus: A matter of global concern. Adv. Hum. Biol. 2023, 13, 232–237. [CrossRef]
- Mahajan, P.; Uddin, S.; Hajati, F.; Moni, M.A.; Gide, E. A comparative evaluation of machine learning ensemble approaches for disease prediction using multiple datasets. Health Technol. 2024, 14, 597–613. [CrossRef]
- Flores, L.; Hernandez, R.M.; Macatangay, L.H.; Garcia, S.M.G.; Melo, J.R. Comparative analysis in the prediction of early-stage diabetes using multiple machine learning techniques. Indones. J. Electr. Eng. Comput. Sci. 2023, 32, 887. [CrossRef]
- Gupta, H.; Varshney, H.; Sharma, T.K.; Pachauri, N.; Verma, O.P. Comparative performance analysis of quantum machine learning with deep learning for diabetes prediction. Complex. Intell. Syst. 2022, 8, 3073–3087. [CrossRef]
- Aggarwal, N.; Basha, C.B.; Arya, A.; Gupta, N. A Comparative Analysis of Machine Leaming-Based Classifiers for Predicting Diabetes. In Proceedings of the 2023 International Conference on Advanced Computing & Communication Technologies (ICACCTech), Banur, India, 23–24 December 2023.
- Swathy, M.; Saruladha, K. A comparative study of classification and prediction of Cardio-vascular diseases (CVD) using Machine Learning and Deep Learning techniques. ICT Express 2022, 8, 109–116. [CrossRef]
- Fregoso-Aparicio, L.; Noguez, J.; Montesinos, L.; Garcia-Garcia, J.A. Machine learning and deep learning predictive models for type 2 diabetes: A systematic review. Diabetol. Metab. Syndr. 2021, 13, 148. [CrossRef]
- Uddin, S.; Khan, A.; Hossain, M.E.; Moni, M.A. Comparing different supervised machine learning algorithms for disease prediction. BMC Med. Inform. Decis. Mak. 2019, 19, 281. [CrossRef]
- Naz, H.; Ahuja, S. Deep learning approach for diabetes prediction using PIMA Indian dataset. J. Diabetes Metab. Disord. 2020, 19, 391–403. [CrossRef]
- Hasan, M.K.; Alam, M.A.; Das, D.; Hossain, E.; Hasan, M. Diabetes Prediction Using Ensembling of Different Machine Learning Classifiers. IEEE Access 2020, 8, 76516–76531. [CrossRef]
- Sahoo, A.K.; Pradhan, C.; Das, H.; Rout, M.; Das, H.; Rout, J.K. Performance Evaluation of Different Machine Learning Methods and Deep-Learning Based Convolutional Neural Network for Health Decision Making. In Nature Inspired Computing for Data Science; Rout, M., Rout, J.K., Das, H., Eds.; Studies in Computational Intelligence; Springer International Publishing AG: Cham, Switzerland, 2020; Volume 871, pp. 201–212, Chapter 8.
- Lai, H.; Huang, H.; Keshavjee, K.; Guergachi, A.; Gao, X. Predictive models for diabetes mellitus using machine learning techniques. BMC Endocr. Disord. 2019, 19, 101. [CrossRef]
- Elreedy, D.; Atiya, A.F. A Comprehensive Analysis of Synthetic Minority Oversampling Technique (SMOTE) for handling class imbalance. Inf. Sci. 2019, 505, 32–64. [CrossRef]
- Wongvorachan, T.; He, S.; Bulut, O. A Comparison of Undersampling, Oversampling, and SMOTE Methods for Dealing with Imbalanced Classification in Educational Data Mining. Information 2023, 14, 54. [CrossRef]
- Kaur, R.; Sharma, R.; Dhaliwal, M.K. Evaluating Performance of SMOTE and ADASYN to Classify Falls and Activities of Daily Living. In Proceedings of the 12th International Conference on Soft Computing for Problem Solving. SocProS 2023; Pant, M., Deep, K., Nagar, A., Eds.; Lecture Notes in Networks and Systems; Springer: Singapore, 2024; Volume 995. [CrossRef]
- Panigrahi, R.; Kumar, L.; Kuanar, S.K. An Empirical Study to Investigate Different SMOTE Data Sampling Techniques for Improving Software Refactoring Prediction. In Neural Information Processing. ICONIP 2020. Communications in Computer and Information Science; Yang, H., Pasupa, K., Leung, A.C., Kwok, J.T., Chan, J.H., King, I.e., Eds.; Springer: Cham, Switzerland, 2020; Volume 1332, pp. 23–31.
- Sahlaoui, H.; Alaoui, E.A.A.; Agoujil, S.; Nayyar, A. An empirical assessment of smote variants techniques and interpretation methods in improving the accuracy and the interpretability of student performance models. Educ. Inf. Technol. 2023, 29, 5447–5483. [CrossRef]
- Haibo, H.; Yang, B.; Garcia, E.A.; Shutao, L. ADASYN: Adaptive synthetic sampling approach for imbalanced learning. In Proceedings of the 2008 IEEE International Joint Conference on Neural Networks (IEEE World Congress on Computational Intelligence), Hong Kong, China, 1–8 June 2008.
- Elsoud, E.A.; Hassan, M.; Alidmat, O.; Al Henawi, E.; Alshdaifat, N.; Igtait, M.; Ghaben, A.; Katrawi, A.; Dmour, M. Under Sampling Techniques for Handling Unbalanced Data with Various Imbalance Rates—A Comparative Study. Int. J. Adv. Comput. Sci. Appl. (IJACSA) 2024, 15, 1274–1284.
- Bach, M.; Werner, A. Improvement of Random Undersampling to Avoid Excessive Removal of Points from a Given Area of the Majority Class. In Computational Science—ICCS 2021, Proceedings of the 21st International Conference, Krakow, Poland, 16–18 June 2021; Part III; Paszynski, M., Kranzlmüller, D., Krzhizhanovskaya, V.V., Dongarra, J.J., Sloot, P.M., Eds.; Lecture Notes in Computer Science; Springer: Cham, Switzerland, 2021; Volume 12744, pp. 172–186. [CrossRef]
- Rekha, G.; Tyagi, A.K.; Reddy, V.K. Performance Analysis of Under-Sampling and Over-Sampling Techniques for Solving Class Imbalance Problem. In Proceedings of the International Conference on Sustainable Computing in Science, Technology & Management (SUSCOM-2019), Jaipur, India, 26–28 February 2019; pp. 1305–1315.
- Joshi, R.D.; Dhakal, C.K. Predicting Type 2 Diabetes Using Logistic Regression and Machine Learning Approaches. Int. J. Environ. Res. Public. Health 2021, 18, 7346. [CrossRef]
- Maniruzzaman, M.; Rahman, J.; Hasan, A.M.; Suri, H.S.; Abedin, M.; El-Baz, A.; Suri, J.S. Accurate Diabetes Risk Stratification Using Machine Learning: Role of Missing Value and Outliers. J. Med. Syst. 2018, 42, 92. [CrossRef]
- Mittal, S.; Hasija, Y. Applications of Deep Learning in Healthcare and Biomedicine. In Deep Learning Techniques for Biomedical and Health Informatics; Dash, S., Acharya, B.R., Mittal, M., Abraham, A., Kelemen, A., Eds.; Springer International Publishing AG: Cham, Switzerland, 2019: Volume 68, pp. 57–78, Chapter 4.
- Iyer, A.; Jeyalatha, S.; Sumbaly, R. Diagnosis of Diabetes Using Classification Mining Techniques. Int. J. Data Min. Knowl. Manag. Process 2015, 5, 1–14. [CrossRef]
- Barik, S.; Mohanty, S.; Mohanty, S.; Singh, D. Analysis of Prediction Accuracy of Diabetes Using Classifier and Hybrid Machine Learning Techniques. In Intelligent and Cloud Computing; Mishra, D., Buyya, R., Mohapatra, P., Patnaik, S., Eds.; Smart Innovation, Systems and Technologies; Springer: Singapore, 2020; pp. 399–409.
- Ganie, S.M.; Malik, M.B.; Arif, T. Performance analysis and prediction of type 2 diabetes mellitus based on lifestyle data using machine learning approaches. J. Diabetes Metab. Disord. 2022, 21, 339–352. [CrossRef]
- Iparraguirre-Villanueva, O.; Espinola-Linares, K.; Castaneda, R.O.F.; Cabanillas-Carbonell, M. Application of Machine Learning Models for Early Detection and Accurate Classification of Type 2 Diabetes. Diagnostics 2023, 13, 2383. [CrossRef]
- Altamimi, A.; Alarfaj, A.A.; Umer, M.; Alabdulqader, E.A.; Alsubai, S.; Kim, T.-H.; Ashraf, I. An automated approach to predict diabetic patients using KNN imputation and effective data mining techniques. BMC Med. Res. Methodol. 2024, 24, 221. [CrossRef]
- Suriya, S.; Muthu, J.J. Type 2 Diabetes Prediction using K-Nearest Neighbor Algorithm. J. Trends Comput. Sci. Smart Technol. 2023, 5, 190–205. [CrossRef]
- Salam, S.S.; Rafi, R. Deep Learning Approach for Sleep Apnea Detection Using Single Lead ECG: Comparative Analysis Between CNN and SNN. In Proceedings of the 2023 26th International Conference on Computer and Information Technology (ICCIT), Cox’s Bazar, Bangladesh, 13–15 December 2023.
- Rahman, M.; Islam, D.; Mukti, R.J.; Saha, I. A deep learning approach based on convolutional LSTM for detecting diabetes. Comput. Biol. Chem. 2020, 88, 107329. [CrossRef]
- Nadesh, R.K.; Arivuselvan, K. Type 2: Diabetes mellitus prediction using Deep Neural Networks classifier. Int. J. Cogn. Comput. Eng. 2020, 1, 55–61. [CrossRef]
- Wadghiri, M.Z.; Idri, A.; Idrissi, T.E.; Hakkoum, H. Ensemble blood glucose prediction in diabetes mellitus—A review. Comput. Struct. Biotechnol. J. 2022, 147, 105674. [CrossRef]
- Guan, Y.; Plotz, T. Ensembles of Deep LSTM Learners for Activity Recognition using Wearables. In Proceedings of the ACM on Interactive, Mobile, Wearable and Ubiquitous Technologies, ACM: New York, USA; 2017; pp. 1–28.
- Shams, M.Y.; Tarek, Z.; Elshewey, A.M. A novel RFE-GRU model for diabetes classification using PIMA Indian dataset. Sci. Rep. 2025, 15, 982. [CrossRef]
- Hossain, M.R.; Hossain, M.J.; Rahman, M.M.; Alam, M.M. Machine Learning Based Prediction and Insights of Diabetes Disease: Pima Indian and Frankfurt Datasets. J. Mech. Contin. Math. Sci. 2025, pp. 99–114. 20. [CrossRef]
- Mousa, A.; Mustafa, W.; Marqas, R.B. A Comparative Study of Diabetes Detection Using the Pima Indian Diabetes Database. J. Univ. Duhok 2023, 26, 277–288. [CrossRef]
- Zargar, O.S.; Bhagat, A.; Teli, T.A.; Sheikh, S. Early Prediction of Diabetes Mellitus on Pima Dataset Using ML And DL Techniques. J. Army Eng. Univ. PLA 2023, 23, 230–249.
- Chang, V.; Bailey, J.; Xu, Q.A.; Sun, Z. Pima Indians diabetes mellitus classification based on machine learning (ML) algorithms. Neural Comput. Appl. 2022, 35, 16157–16173. [CrossRef]
- Xie, Z.; Nikolayeva, O.; Luo, J.; Li, D. Building Risk Prediction Models for Type 2 Diabetes Using Machine Learning Techniques. Prev. Chronic Dis. 2019, 16, E130. [CrossRef]
- Islam, M.M.F.; Ferdousi, R.; Rahman, S.; Bushra, H.Y. Likelihood Prediction of Diabetes at Early Stage Using Data Mining Techniques. In Computer Vision and Machine Intelligence in Medical Image Analysis; Advances in Intelligent Systems and Computing; Gupta, M., Konar, D., Bhattacharyya, S., Biswas, S.; Springer, Singapore; 2020, Chapter 12, pp. 113–125. [CrossRef]
- Sadhu, A.; Jadli, A. Early-Stage Diabetes Risk Prediction—A Comparative Analysis of Classification Algorithms. Int. Adv. Res. J. Sci. Eng. Technol. 2021, 8, 193–201. [CrossRef]
- Al-Haija, Q.A.; Smadi, M.; Al-Bataineh, O.M. Early Stage Diabetes Risk Prediction via Machine Learning. In Proceedings of the 13th International Conference on Soft Computing and Pattern Recognition (SoCPaR 2021); Springer: Cham, Switzerland, 2022; Volume 417, pp. 451–461. [CrossRef]
- Chatrati, S.P.; Hossain, G.; Goyal, A.; Bhan, A.; Bhattacharya, S.; Gaurav, D.; Tiwari, S.M. Smart home health monitoring system for predicting type 2 diabetes and hypertension. J. King Saud. Univ.—Comput. Inf. Sci. 2022, 34, 862–870. [CrossRef]
- Bozkurt, M.R.; Yurtay, N.; Yilmaz, Z.; Sertkaya, C. Comparison of different methods for determining diabetes. Turk. J. Electr. Eng. Comput. Sci. 2014, 22, 1044–1055. [CrossRef]
- Bashir, S.; Qamar, U.; Khan, F.H. IntelliHealth: A medical decision support application using a novel weighted multi-layer classifier ensemble framework. J. Biomed. Inf. 2016, 59, 185–200. [CrossRef]
- Wang, Q.; Cao, W.; Guo, J.; Ren, J.; Cheng, Y.; Davis, D.N. DMP_MI: An Effective Diabetes Mellitus Classification Algorithm on Imbalanced Data with Missing Values. IEEE Access 2019, 7, 102232–102238. [CrossRef]
- Kaur, H.; Kumari, V. Predictive modelling and analytics for diabetes using a machine learning approach. Appl. Comput. Inform. 2020, 18, 90–100. [CrossRef]
- Yuvaraj, N.; SriPreethaa, K.R. Diabetes prediction in healthcare systems using machine learning algorithms on Hadoop cluster. Clust. Comput. 2017, 22, 1–9. [CrossRef]
Figure 1.
Feature Distribution for Datasets 1 and 2 (PIMA dataset).
Figure 1.
Feature Distribution for Datasets 1 and 2 (PIMA dataset).
Figure 2.
Feature Distribution for Datasets 3 (BRFSS_2015 dataset). .
Figure 2.
Feature Distribution for Datasets 3 (BRFSS_2015 dataset). .
Figure 3.
Feature Distribution for Dataset 5 (BRFSS_2015 dataset).
Figure 3.
Feature Distribution for Dataset 5 (BRFSS_2015 dataset).
Figure 4.
Methodology Flow Diagram .
Figure 4.
Methodology Flow Diagram .
Figure 5.
Confusion matrix for the AdaBoost model.
Figure 5.
Confusion matrix for the AdaBoost model.
Figure 6.
PCA Projection for Dataset 1 class outcomes. .
Figure 6.
PCA Projection for Dataset 1 class outcomes. .
Figure 7.
AUC Curves for the AdaBoost model.
Figure 7.
AUC Curves for the AdaBoost model.
Figure 8.
Heatmaps for Datasets 1 and 2.
Figure 8.
Heatmaps for Datasets 1 and 2.
Figure 9.
Confusion matrix for the Random Forest model.
Figure 9.
Confusion matrix for the Random Forest model.
Figure 10.
AUC and Precision-Recall Curves for the Random Forest model.
Figure 10.
AUC and Precision-Recall Curves for the Random Forest model.
Figure 11.
PCA Projection for Dataset 2 class outcomes.
Figure 11.
PCA Projection for Dataset 2 class outcomes.
Figure 12.
Confusion matrix for the RNN model.
Figure 12.
Confusion matrix for the RNN model.
Figure 13.
AUC and Precision-Recall curves of the RNN model demonstrate better performance, as indicated by the ROC curve being above the 45-degree dotted line. The blue line (Class 0) and green line (Class 2) above the dotted line show good performance, while the orange line (Class 1) shows moderate performance.
Figure 13.
AUC and Precision-Recall curves of the RNN model demonstrate better performance, as indicated by the ROC curve being above the 45-degree dotted line. The blue line (Class 0) and green line (Class 2) above the dotted line show good performance, while the orange line (Class 1) shows moderate performance.
Figure 14.
LDA Projection for Dataset 3 class outcomes.
Figure 14.
LDA Projection for Dataset 3 class outcomes.
Figure 15.
PCA Projection for Dataset 3 class outcomes .
Figure 15.
PCA Projection for Dataset 3 class outcomes .
Figure 16.
t-SNE Projection for Dataset 3 class outcomes .
Figure 16.
t-SNE Projection for Dataset 3 class outcomes .
Figure 17.
Confusion matrix for the DNN model.
Figure 17.
Confusion matrix for the DNN model.
Figure 18.
AUC Curves of DNN Model.
Figure 18.
AUC Curves of DNN Model.
Figure 19.
Threshold-dependent metrics for DNN. The vertical line denotes the chosen threshold.
Figure 19.
Threshold-dependent metrics for DNN. The vertical line denotes the chosen threshold.
Figure 20.
Confusion matrix for the Random Forest model.
Figure 20.
Confusion matrix for the Random Forest model.
Figure 21.
AUC Curves for the Random Forest model.
Figure 21.
AUC Curves for the Random Forest model.
Figure 22.
F1-score distribution by Dataset.
Figure 22.
F1-score distribution by Dataset.
Figure 23.
Inference Time by Dataset.
Figure 23.
Inference Time by Dataset.
Table 1.
Datasets Statistics.
Table 1.
Datasets Statistics.
| Description |
Dataset 1 |
Dataset 2 |
Dataset 3 |
Dataset 4 |
Dataset 5 |
| Source |
UCL Machine Learning Repository, Kaggle and CDC websites |
|
| Samples |
768 |
2000 |
253,680 |
70692 |
520 |
| Features |
9 |
9 |
21 |
21 |
17 |
| Positive instances |
268 |
684 |
35346 |
35346 |
320 |
| Negative instances |
500 |
1316 |
218334 |
35346 |
200 |
Table 2.
Number of dataset features labelled as zero values.
Table 2.
Number of dataset features labelled as zero values.
| Feature |
Dataset 1 |
Dataset 2 |
| Pregnancies |
111 |
301 |
| Glucose |
5 |
13 |
| BloodPressure |
35 |
90 |
| SkinThickness |
227 |
573 |
| Insulin |
374 |
956 |
| BMI |
11 |
28 |
| DiabetesPedigreeFunction |
0 |
0 |
| Age |
0 |
0 |
Table 3.
Imbalanced values in the Outcome (Target) variable.
Table 3.
Imbalanced values in the Outcome (Target) variable.
| Outcome (Target class) |
Dataset 1 |
Dataset 2 |
Dataset 3 |
Dataset 4 |
Dataset 5 |
| 0 |
400 |
1053 |
213,703 |
218,334 |
200 |
| 1 |
214 |
547 |
4631 |
35,346 |
320 |
| 2 |
- |
- |
35,346 |
- |
- |
Table 4.
Model Performance Comparison for Dataset 1 using F1-score as reference.
Table 4.
Model Performance Comparison for Dataset 1 using F1-score as reference.
| Model |
Accuracy |
Precision |
Recall |
F1 Score |
AUC-ROC |
R2 |
MSE |
MAE |
RMSE |
TT |
MU |
NOP |
| AdaBoost |
0.7987 |
0.6716 |
0.8333 |
0.7438 |
0.8386 |
0.1159 |
0.2013 |
0.2013 |
0.4487 |
1.1812 |
0.0 B |
987 |
| Bagging AdaBoost |
0.7857 |
0.6615 |
0.7963 |
0.7227 |
0.8439 |
0.0589 |
0.2143 |
0.2143 |
0.4629 |
2.1976 |
80.0 kB |
1392 |
| RNN |
0.7792 |
0.6471 |
0.8148 |
0.7213 |
0.8202 |
0.0304 |
0.2208 |
0.2208 |
0.4699 |
7.9969 |
76.0 kB |
7831 |
| Bagging DNN |
0.7727 |
0.6338 |
0.8333 |
0.72 |
0.8198 |
0.0019 |
0.2273 |
0.2273 |
0.4767 |
60.4006 |
1228.0 kB |
110652 |
| RF |
0.7792 |
0.6563 |
0.7778 |
0.7119 |
0.8304 |
0.0304 |
0.2208 |
0.2208 |
0.4699 |
0.4222 |
36.0 kB |
6269 |
| Bagging XGBoost |
0.7727 |
0.6418 |
0.7963 |
0.7107 |
0.828 |
0.0019 |
0.2273 |
0.2273 |
0.4767 |
1.4211 |
0.0 B |
19164 |
| XGBoost |
0.7662 |
0.6286 |
0.8148 |
0.7097 |
0.8381 |
-0.0267 |
0.2338 |
0.2338 |
0.4835 |
0.395 |
0.0 B |
1618 |
| Stacking Classifier |
0.7727 |
0.6462 |
0.7778 |
0.7059 |
0.8302 |
0.0019 |
0.2273 |
0.2273 |
0.4767 |
60.8967 |
428.0 kB |
26603 |
| Bagging RF |
0.7662 |
0.6324 |
0.7963 |
0.7049 |
0.832 |
-0.0267 |
0.2338 |
0.2338 |
0.4835 |
9.6091 |
24.0 kB |
243870 |
| LR-MLP |
0.7662 |
0.6364 |
0.7778 |
0.7000 |
0.8248 |
-0.0267 |
0.2338 |
0.2338 |
0.4835 |
4.4264 |
0.0 B |
10 |
| LR |
0.7597 |
0.6269 |
0.7778 |
0.6942 |
0.8244 |
-0.0552 |
0.2403 |
0.2403 |
0.4902 |
0.2405 |
0.0 B |
9 |
| XGBoost-CNN |
0.7662 |
0.6452 |
0.7407 |
0.6897 |
0.8126 |
-0.0267 |
0.2338 |
0.2338 |
0.4835 |
6.3776 |
0.0 B |
20809 |
| SVM |
0.7597 |
0.6308 |
0.7593 |
0.6891 |
0.8271 |
-0.0552 |
0.2403 |
0.2403 |
0.4902 |
0.9922 |
0.0 B |
8 |
| Bagging RNN |
0.7468 |
0.6056 |
0.7963 |
0.688 |
0.8128 |
-0.1122 |
0.2532 |
0.2532 |
0.5032 |
42.0664 |
492.0 kB |
113785 |
| XGBoost-LSTM |
0.7338 |
0.5867 |
0.8148 |
0.6822 |
0.8145 |
-0.1693 |
0.2662 |
0.2662 |
0.516 |
9.9012 |
0.0 B |
3168 |
| RF-GRU |
0.7403 |
0.6000 |
0.7778 |
0.6774 |
0.8189 |
-0.1407 |
0.2597 |
0.2597 |
0.5096 |
7.5671 |
108.0 kB |
28314 |
| KNN |
0.7338 |
0.589 |
0.7963 |
0.6772 |
0.806 |
-0.1693 |
0.2662 |
0.2662 |
0.516 |
0.1758 |
16.0 kB |
6288 |
| RF-CNN |
0.7338 |
0.5915 |
0.7778 |
0.672 |
0.8174 |
-0.1693 |
0.2662 |
0.2662 |
0.516 |
4.1751 |
40.0 kB |
5572 |
| SVM-RNN |
0.7338 |
0.5915 |
0.7778 |
0.672 |
0.8152 |
-0.1693 |
0.2662 |
0.2662 |
0.516 |
6.396 |
0.0 B |
9 |
| AdaBoost-DBN |
0.7208 |
0.5714 |
0.8148 |
0.6718 |
0.7947 |
-0.2263 |
0.2792 |
0.2792 |
0.5284 |
24.0248 |
4.0 kB |
1491 |
| KNN-Autoencoders |
0.6948 |
0.5432 |
0.8148 |
0.6519 |
0.7839 |
-0.3404 |
0.3052 |
0.3052 |
0.5524 |
10.054 |
0.0 B |
24366 |
| NB |
0.7208 |
0.5797 |
0.7407 |
0.6504 |
0.7804 |
-0.2263 |
0.2792 |
0.2792 |
0.5284 |
0.2365 |
0.0 B |
34 |
| DNN |
0.7338 |
0.6032 |
0.7037 |
0.6496 |
0.807 |
-0.1693 |
0.2662 |
0.2662 |
0.516 |
5.0588 |
92.0 kB |
8067 |
| DT-CNN |
0.7013 |
0.5526 |
0.7778 |
0.6462 |
0.712 |
-0.3119 |
0.2987 |
0.2987 |
0.5465 |
5.1779 |
0.0 B |
81 |
| CNN |
0.7208 |
0.5846 |
0.7037 |
0.6387 |
0.807 |
-0.2263 |
0.2792 |
0.2792 |
0.5284 |
4.9617 |
68.0 kB |
1579 |
| DT |
0.7208 |
0.5873 |
0.6852 |
0.6325 |
0.7915 |
-0.2263 |
0.2792 |
0.2792 |
0.5284 |
0.2345 |
0.0 B |
31 |
| LSTM |
0.6818 |
0.5424 |
0.5926 |
0.5664 |
0.7085 |
-0.3974 |
0.3182 |
0.3182 |
0.5641 |
18.418 |
2052.0 kB |
64639 |
| GRU |
0.6753 |
0.5333 |
0.5926 |
0.5614 |
0.7081 |
-0.4259 |
0.3247 |
0.3247 |
0.5698 |
20.1196 |
1404.0 kB |
34126 |
Table 5.
Model Performance Comparison for Dataset 2 using F1-score as reference.
Table 5.
Model Performance Comparison for Dataset 2 using F1-score as reference.
| Model |
Accuracy |
Precision |
Recall |
F1 Score |
AUC-ROC |
R2 |
MSE |
MAE |
RMSE |
TT |
MU |
NOP |
| RF |
0.7857 |
0.6567 |
0.8148 |
0.7273 |
0.8341 |
0.0589 |
0.2143 |
0.2143 |
0.4629 |
0.6486 |
32.0 kB |
13697 |
| Bagging DNN |
0.7792 |
0.6429 |
0.8333 |
0.7258 |
0.8283 |
0.0304 |
0.2208 |
0.2208 |
0.4699 |
69.5682 |
1256.0 kB |
154790 |
| Bagging RNN |
0.7792 |
0.6429 |
0.8333 |
0.7258 |
0.8207 |
0.0304 |
0.2208 |
0.2208 |
0.4699 |
56.3799 |
1452.0 kB |
31885 |
| Bagging AdaBoost |
0.7792 |
0.6471 |
0.8148 |
0.7213 |
0.8446 |
0.0304 |
0.2208 |
0.2208 |
0.4699 |
5.1711 |
8.0 kB |
3036 |
| AdaBoost |
0.7727 |
0.6338 |
0.8333 |
0.7200 |
0.8401 |
0.0019 |
0.2273 |
0.2273 |
0.4767 |
1.8813 |
0.0 B |
1500 |
| XGBoost |
0.7727 |
0.6338 |
0.8333 |
0.7200 |
0.8376 |
0.0019 |
0.2273 |
0.2273 |
0.4767 |
0.4041 |
0.0 B |
1272 |
| Stacking Classifier |
0.7662 |
0.6286 |
0.8148 |
0.7097 |
0.8344 |
-0.0267 |
0.2338 |
0.2338 |
0.4835 |
52.1551 |
320.0 kB |
57376 |
| Bagging XGBoost |
0.7857 |
0.6780 |
0.7407 |
0.7080 |
0.8274 |
0.0589 |
0.2143 |
0.2143 |
0.4629 |
3.1946 |
4.0 kB |
118269 |
| DNN |
0.7662 |
0.6324 |
0.7963 |
0.7049 |
0.8170 |
-0.0267 |
0.2338 |
0.2338 |
0.4835 |
7.7426 |
48.0 kB |
11339 |
| RF-GRU |
0.7597 |
0.6197 |
0.8148 |
0.7040 |
0.8161 |
-0.0552 |
0.2403 |
0.2403 |
0.4902 |
8.3945 |
60.0 kB |
27302 |
| RF-CNN |
0.7468 |
0.5974 |
0.8519 |
0.7023 |
0.8159 |
-0.1122 |
0.2532 |
0.2532 |
0.5032 |
7.3780 |
40.0 kB |
4184 |
| LR-MLP |
0.7532 |
0.6143 |
0.7963 |
0.6935 |
0.8222 |
-0.0837 |
0.2468 |
0.2468 |
0.4967 |
14.3566 |
28.0 kB |
10 |
| Bagging RF |
0.7662 |
0.6452 |
0.7407 |
0.6897 |
0.8298 |
-0.0267 |
0.2338 |
0.2338 |
0.4835 |
11.5716 |
12024.0 kB |
342125 |
| SVM |
0.7532 |
0.6176 |
0.7778 |
0.6885 |
0.8219 |
-0.0837 |
0.2468 |
0.2468 |
0.4967 |
0.2106 |
0.0 B |
8 |
| XGBoost-LSTM |
0.7338 |
0.5844 |
0.8333 |
0.6870 |
0.8228 |
-0.1693 |
0.2662 |
0.2662 |
0.5160 |
9.3506 |
40.0 kB |
3214 |
| SVM-RNN |
0.7468 |
0.6087 |
0.7778 |
0.6829 |
0.8148 |
-0.1122 |
0.2532 |
0.2532 |
0.5032 |
5.1516 |
0.0 B |
9 |
| LR |
0.7468 |
0.6119 |
0.7593 |
0.6777 |
0.8215 |
-0.1122 |
0.2532 |
0.2532 |
0.5032 |
0.4155 |
0.0 B |
9 |
| KNN |
0.7273 |
0.5789 |
0.8148 |
0.6769 |
0.7935 |
-0.1978 |
0.2727 |
0.2727 |
0.5222 |
0.1915 |
36.0 kB |
6304 |
| AdaBoost-DBN |
0.7273 |
0.5833 |
0.7778 |
0.6667 |
0.8135 |
-0.1978 |
0.2727 |
0.2727 |
0.5222 |
24.3952 |
0.0 B |
1314 |
| XGBoost-CNN |
0.7532 |
0.6333 |
0.7037 |
0.6667 |
0.7983 |
-0.0837 |
0.2468 |
0.2468 |
0.4967 |
8.3154 |
0.0 B |
10900 |
| KNN-Autoencoders |
0.7273 |
0.5857 |
0.7593 |
0.6613 |
0.7693 |
-0.1978 |
0.2727 |
0.2727 |
0.5222 |
10.7923 |
84.0 kB |
10244 |
| RNN |
0.7338 |
0.5970 |
0.7407 |
0.6612 |
0.8087 |
-0.1693 |
0.2662 |
0.2662 |
0.5160 |
9.4642 |
436.0 kB |
3539 |
| DT |
0.7597 |
0.6667 |
0.6296 |
0.6476 |
0.7770 |
-0.0552 |
0.2403 |
0.2403 |
0.4902 |
0.2572 |
0.0 B |
129 |
| DT-CNN |
0.7208 |
0.5902 |
0.6667 |
0.6261 |
0.7525 |
-0.2263 |
0.2792 |
0.2792 |
0.5284 |
6.4722 |
0.0 B |
101 |
| NB |
0.6948 |
0.5522 |
0.6852 |
0.6116 |
0.7676 |
-0.3404 |
0.3052 |
0.3052 |
0.5524 |
0.1878 |
0.0 B |
34 |
| CNN |
0.6818 |
0.5352 |
0.7037 |
0.6080 |
0.7665 |
-0.3974 |
0.3182 |
0.3182 |
0.5641 |
4.4649 |
16.0 kB |
50587 |
| LSTM |
0.6688 |
0.5231 |
0.6296 |
0.5714 |
0.7059 |
-0.4544 |
0.3312 |
0.3312 |
0.5755 |
12.0811 |
240.0 kB |
9649 |
| GRU |
0.6883 |
0.5517 |
0.5926 |
0.5714 |
0.7256 |
-0.3689 |
0.3117 |
0.3117 |
0.5583 |
13.9710 |
564.0 kB |
1401 |
Table 6.
Model Performance Comparison for Dataset 3 using F1-score as reference.
Table 6.
Model Performance Comparison for Dataset 3 using F1-score as reference.
| Model |
Accuracy |
Precision |
Recall |
F1 Score |
AUC-ROC |
R2 |
MSE |
MAE |
RMSE |
TT |
MU |
NOP |
| RNN |
0.7144 |
0.4387 |
0.4982 |
0.4414 |
0.7008 |
-0.7099 |
0.8334 |
0.4682 |
0.9129 |
141.7094 |
600.0 kB |
14277 |
| CNN |
0.6984 |
0.4403 |
0.5171 |
0.4411 |
0.7064 |
-0.8212 |
0.8877 |
0.4970 |
0.9422 |
77.7925 |
216.0 kB |
31823 |
| DNN |
0.6975 |
0.4397 |
0.5149 |
0.4401 |
0.7055 |
-0.8398 |
0.8967 |
0.5006 |
0.9470 |
959.0492 |
10316.0 kB |
19371 |
| AdaBoost |
0.6898 |
0.4337 |
0.5155 |
0.4330 |
0.7128 |
-1.0948 |
1.0210 |
0.5471 |
1.0105 |
38.0504 |
0.0 B |
11696 |
| XGBoost |
0.6834 |
0.4301 |
0.5109 |
0.4270 |
0.7143 |
-1.1936 |
1.0692 |
0.5674 |
1.0340 |
1.4653 |
4.0 kB |
1244 |
| XGBoost-LSTM |
0.7004 |
0.4301 |
0.5079 |
0.4252 |
0.7184 |
-1.2789 |
1.1108 |
0.5700 |
1.0539 |
385.1103 |
272.0 kB |
34830 |
| RF |
0.6755 |
0.4296 |
0.5119 |
0.4251 |
0.7091 |
-1.2227 |
1.0834 |
0.5775 |
1.0408 |
11.1173 |
68.0 kB |
738710 |
| RF-CNN |
0.6734 |
0.4302 |
0.5104 |
0.4245 |
0.7107 |
-1.1799 |
1.0625 |
0.5719 |
1.0308 |
44.9699 |
2220.0 kB |
281808 |
| RF-GRU |
0.6639 |
0.4307 |
0.5115 |
0.4229 |
0.7097 |
-1.1512 |
1.0485 |
0.5736 |
1.0240 |
136.2781 |
1240.0 kB |
269827 |
| DT-CNN |
0.6890 |
0.4227 |
0.4783 |
0.4218 |
0.6566 |
-0.9623 |
0.9564 |
0.5261 |
0.9780 |
22.8690 |
0.0 B |
15 |
| LR |
0.6260 |
0.4499 |
0.5147 |
0.4194 |
0.7077 |
-0.6358 |
0.7973 |
0.5151 |
0.8929 |
2.9852 |
336.0 kB |
64 |
| LR-MLP |
0.5930 |
0.4561 |
0.5197 |
0.4116 |
0.7118 |
-0.6116 |
0.7855 |
0.5332 |
0.8863 |
23.3801 |
20.0 kB |
67 |
| DT |
0.6384 |
0.4235 |
0.4935 |
0.4085 |
0.6876 |
-1.2310 |
1.0874 |
0.6036 |
1.0428 |
0.3724 |
0.0 B |
233 |
| NB |
0.6245 |
0.4364 |
0.4892 |
0.4083 |
0.6803 |
-0.7259 |
0.8412 |
0.5307 |
0.9172 |
0.2969 |
0.0 B |
129 |
| SVM |
0.5759 |
0.4591 |
0.5116 |
0.4044 |
0.7084 |
-0.5697 |
0.7651 |
0.5378 |
0.8747 |
277.1082 |
136.0 kB |
63 |
| KNN |
0.5626 |
0.4238 |
0.4778 |
0.3794 |
0.6617 |
-0.9820 |
0.9660 |
0.6136 |
0.9829 |
114.0517 |
0.0 B |
547344 |
| KNN-Autoencoders |
0.5279 |
0.4251 |
0.4760 |
0.3651 |
0.6665 |
-0.9109 |
0.9314 |
0.6252 |
0.9651 |
69.0643 |
1208.0 kB |
1537776 |
| Bagging XGBoost |
0.6899 |
0.4298 |
0.5098 |
0.4290 |
0.7029 |
-1.1983 |
1.0715 |
0.5639 |
1.0351 |
9.1290 |
72.0 kB |
15380 |
| Stacking Classifier |
0.6632 |
0.4266 |
0.5095 |
0.4169 |
0.7101 |
-1.3910 |
1.1654 |
0.6130 |
1.0795 |
552.4895 |
252.0 kB |
73996 |
Table 7.
Model Performance Comparison for Dataset 4 using F1-score as reference.
Table 7.
Model Performance Comparison for Dataset 4 using F1-score as reference.
| Model |
Accuracy |
Precision |
Recall |
F1 Score |
AUC-ROC |
R2 |
MSE |
MAE |
RMSD |
TT |
MU |
NOP |
| DNN |
0.7548 |
0.3286 |
0.7280 |
0.4528 |
0.8233 |
-1.0445 |
0.2452 |
0.2452 |
0.4951 |
153.6247 |
464.0 kB |
22892 |
| GRU |
0.7542 |
0.3243 |
0.7053 |
0.4443 |
0.8141 |
-1.0496 |
0.2458 |
0.2458 |
0.4958 |
1005.0657 |
1292.0 kB |
21498 |
| CNN |
0.7364 |
0.3145 |
0.7564 |
0.4443 |
0.8218 |
-1.1985 |
0.2636 |
0.2636 |
0.5135 |
89.3823 |
1096.0 kB |
55909 |
| Bagging AdaBoost |
0.7249 |
0.3080 |
0.7816 |
0.4419 |
0.8250 |
-1.2942 |
0.2751 |
0.2751 |
0.5245 |
172.3706 |
12.0 kB |
44017 |
| Bagging XGBoost |
0.7184 |
0.3050 |
0.7986 |
0.4414 |
0.8265 |
-1.3483 |
0.2816 |
0.2816 |
0.5307 |
13.8929 |
204.0 kB |
59953 |
| AdaBoost |
0.7206 |
0.3057 |
0.7908 |
0.4409 |
0.8250 |
-1.3300 |
0.2794 |
0.2794 |
0.5286 |
55.0732 |
0.0 B |
68311 |
| XGBoost |
0.7177 |
0.3044 |
0.7987 |
0.4408 |
0.8259 |
-1.3545 |
0.2823 |
0.2823 |
0.5314 |
2.1093 |
0.0 B |
851 |
| LR-MLP |
0.7259 |
0.3077 |
0.7739 |
0.4403 |
0.8206 |
-1.2858 |
0.2741 |
0.2741 |
0.5236 |
34.6740 |
0.0 B |
23 |
| LR |
0.7250 |
0.3069 |
0.7741 |
0.4396 |
0.8196 |
-1.2935 |
0.2750 |
0.2750 |
0.5244 |
1.5440 |
0.0 B |
22 |
| Stacking Classifier |
0.7168 |
0.3032 |
0.7953 |
0.4390 |
0.8248 |
-1.3612 |
0.2832 |
0.2832 |
0.5321 |
3029.2078 |
1148.0 kB |
173546 |
| XGBoost-LSTM |
0.7140 |
0.3016 |
0.8007 |
0.4382 |
0.8240 |
-1.3854 |
0.2860 |
0.2860 |
0.5348 |
227.6599 |
364.0 kB |
3377 |
| RF-CNN |
0.7097 |
0.2997 |
0.8109 |
0.4377 |
0.8252 |
-1.4207 |
0.2903 |
0.2903 |
0.5388 |
93.7112 |
1596.0 kB |
658218 |
| RF |
0.7124 |
0.3002 |
0.7994 |
0.4365 |
0.8226 |
-1.3986 |
0.2876 |
0.2876 |
0.5363 |
15.5934 |
28.0 kB |
1554750 |
| XGBoost-CNN |
0.7076 |
0.2983 |
0.8124 |
0.4363 |
0.8261 |
-1.4387 |
0.2924 |
0.2924 |
0.5408 |
70.9944 |
0.0 B |
11896 |
| RF-GRU |
0.7067 |
0.2974 |
0.8107 |
0.4351 |
0.8247 |
-1.4456 |
0.2933 |
0.2933 |
0.5415 |
553.4871 |
272.0 kB |
498831 |
| DT-CNN |
0.7121 |
0.2990 |
0.7928 |
0.4342 |
0.8178 |
-1.4005 |
0.2879 |
0.2879 |
0.5365 |
66.3589 |
24.0 kB |
31 |
| Bagging DNN |
0.7060 |
0.2960 |
0.8053 |
0.4329 |
0.8222 |
-1.4518 |
0.2940 |
0.2940 |
0.5422 |
447.2553 |
352.0 kB |
56725 |
| SVM |
0.7089 |
0.2967 |
0.7946 |
0.4321 |
0.8189 |
-1.4272 |
0.2911 |
0.2911 |
0.5395 |
882.2086 |
232.0 kB |
21 |
| Bagging GRU |
0.7206 |
0.3013 |
0.7622 |
0.4319 |
0.8120 |
-1.3297 |
0.2794 |
0.2794 |
0.5286 |
2105.2017 |
3892.0 kB |
139076 |
| SVM-RNN |
0.7023 |
0.2930 |
0.8046 |
0.4296 |
0.8183 |
-1.4827 |
0.2977 |
0.2977 |
0.5456 |
788.8983 |
28.0 kB |
674564 |
| LSTM |
0.7334 |
0.3049 |
0.7142 |
0.4274 |
0.8016 |
-1.2235 |
0.2666 |
0.2666 |
0.5164 |
1245.5357 |
404.0 kB |
61624 |
| AdaBoost-DBN |
0.7089 |
0.2942 |
0.7785 |
0.4270 |
0.8129 |
-1.4276 |
0.2911 |
0.2911 |
0.5396 |
592.0342 |
188.0 kB |
1317 |
| DT |
0.7124 |
0.2954 |
0.7687 |
0.4268 |
0.8077 |
-1.3987 |
0.2876 |
0.2876 |
0.5363 |
0.3540 |
0.0 B |
127 |
| Bagging CNN |
0.6939 |
0.2880 |
0.8127 |
0.4252 |
0.8191 |
-1.5526 |
0.3061 |
0.3061 |
0.5533 |
528.4152 |
992.0 kB |
87108 |
| NB |
0.7235 |
0.2941 |
0.7029 |
0.4147 |
0.7799 |
-1.3055 |
0.2765 |
0.2765 |
0.5258 |
0.2801 |
0.0 B |
86 |
| KNN-Autoencoders |
0.7156 |
0.2892 |
0.7141 |
0.4117 |
0.7808 |
-1.3713 |
0.2844 |
0.2844 |
0.5332 |
295.8879 |
16.0 kB |
1696620 |
| KNN |
0.7058 |
0.2848 |
0.7356 |
0.4107 |
0.7857 |
-1.4531 |
0.2942 |
0.2942 |
0.5424 |
66.8340 |
0.0 B |
1187634 |
| RNN |
0.6555 |
0.2602 |
0.7986 |
0.3925 |
0.7866 |
-1.8726 |
0.3445 |
0.3445 |
0.5869 |
98.3237 |
496.0 kB |
12431 |
Table 8.
Model Performance Comparison for Dataset 5 using F1-score as reference.
Table 8.
Model Performance Comparison for Dataset 5 using F1-score as reference.
| Model |
Accuracy |
Precision |
Recall |
F1 Score |
AUC-ROC |
R2 |
MSE |
MAE |
RMSE |
TT |
MU |
NOP |
| RF |
1.0000 |
1.0000 |
1.0000 |
1.0000 |
1.0000 |
1.0000 |
0.0000 |
0.0000 |
0.0000 |
0.58740 |
24.0 kB |
11455 |
| Stacking Classifier |
1.0000 |
1.0000 |
1.0000 |
1.0000 |
1.0000 |
1.0000 |
0.0000 |
0.0000 |
0.0000 |
37.0527 |
296.0 kB |
N/A |
| DT-CNN |
0.9904 |
0.9846 |
1.0000 |
0.9922 |
0.9992 |
0.9594 |
0.0096 |
0.0096 |
0.0981 |
5.7288 |
0.0 B |
27 |
| Bagging SVM |
0.9904 |
0.9846 |
1.0000 |
0.9922 |
0.9992 |
0.9594 |
0.0096 |
0.0096 |
0.0981 |
0.5832 |
0.0 B |
4360 |
| DT |
0.9904 |
1.0000 |
0.9844 |
0.9921 |
0.9922 |
0.9594 |
0.0096 |
0.0096 |
0.0981 |
0.2275 |
0.0 B |
67 |
| AdaBoost |
0.9904 |
1.0000 |
0.9844 |
0.9921 |
1.0000 |
0.9594 |
0.0096 |
0.0096 |
0.0981 |
0.7496 |
0.0 B |
9146 |
| Bagging DT |
0.9904 |
1.0000 |
0.9844 |
0.9921 |
1.0000 |
0.9594 |
0.0096 |
0.0096 |
0.0981 |
1.0822 |
16.0 kB |
10691 |
| SVM |
0.9808 |
0.9844 |
0.9844 |
0.9844 |
0.9977 |
0.9188 |
0.0192 |
0.0192 |
0.1387 |
0.3871 |
0.0 B |
1312 |
| DNN |
0.9808 |
0.9844 |
0.9844 |
0.9844 |
0.9988 |
0.9188 |
0.0192 |
0.0192 |
0.1387 |
7.3960 |
88.0 kB |
10911 |
| RF-CNN |
0.9808 |
0.9844 |
0.9844 |
0.9844 |
0.9980 |
0.9188 |
0.0192 |
0.0192 |
0.1387 |
5.2281 |
68.0 kB |
4230 |
| Bagging RF |
0.9808 |
0.9844 |
0.9844 |
0.9844 |
0.9965 |
0.9188 |
0.0192 |
0.0192 |
0.1387 |
1.6575 |
128.0 kB |
10245 |
| XGBoost |
0.9808 |
1.0000 |
0.9688 |
0.9841 |
0.9992 |
0.9188 |
0.0192 |
0.0192 |
0.1387 |
0.3725 |
0.0 B |
4488 |
| AdaBoost-DBN |
0.9808 |
1.0000 |
0.9688 |
0.9841 |
0.9984 |
0.9188 |
0.0192 |
0.0192 |
0.1387 |
15.2999 |
0.0 B |
1275 |
| RF-GRU |
0.9808 |
1.0000 |
0.9688 |
0.9841 |
1.0000 |
0.9188 |
0.0192 |
0.0192 |
0.1387 |
8.4001 |
56.0 kB |
6094 |
| CNN |
0.9712 |
0.9841 |
0.9688 |
0.9764 |
0.9980 |
0.8781 |
0.0288 |
0.0288 |
0.1698 |
6.8386 |
236.0 kB |
174659 |
| SVM-RNN |
0.9712 |
0.9841 |
0.9688 |
0.9764 |
0.9977 |
0.8781 |
0.0288 |
0.0288 |
0.1698 |
8.0761 |
0.0 B |
1394 |
| XGBoost-LSTM |
0.9712 |
1.0000 |
0.9531 |
0.9760 |
0.9973 |
0.8781 |
0.0288 |
0.0288 |
0.1698 |
14.8126 |
0.0 B |
1489 |
| Bagging AdaBoost |
0.9615 |
0.9688 |
0.9688 |
0.9688 |
0.9859 |
0.8375 |
0.0385 |
0.0385 |
0.1961 |
5.3577 |
0.0 B |
2970 |
| LR-MLP |
0.9615 |
0.9839 |
0.9531 |
0.9683 |
0.9984 |
0.8375 |
0.0385 |
0.0385 |
0.1961 |
11.7585 |
0.0 B |
18 |
| XGBoost-CNN |
0.9615 |
0.9839 |
0.9531 |
0.9683 |
0.9947 |
0.8375 |
0.0385 |
0.0385 |
0.1961 |
6.5654 |
0.0 B |
5082 |
| Bagging CNN-DT |
0.9615 |
0.9839 |
0.9531 |
0.9683 |
0.9969 |
0.8375 |
0.0385 |
0.0385 |
0.1961 |
38.0482 |
1072.0 kB |
N/A |
| KNN |
0.9519 |
0.9836 |
0.9375 |
0.9600 |
0.9820 |
0.7969 |
0.0481 |
0.0481 |
0.2193 |
0.1675 |
0.0 B |
6656 |
| LR |
0.9519 |
1.0000 |
0.9219 |
0.9593 |
0.9918 |
0.7969 |
0.0481 |
0.0481 |
0.2193 |
0.2580 |
0.0 B |
17 |
| KNN-Autoencoders |
0.9519 |
1.0000 |
0.9219 |
0.9593 |
0.9949 |
0.7969 |
0.0481 |
0.0481 |
0.2193 |
9.9981 |
0.0 B |
14560 |
| NB |
0.9423 |
0.9677 |
0.9375 |
0.9524 |
0.9863 |
0.7563 |
0.0577 |
0.0577 |
0.2402 |
0.2241 |
0.0 B |
66 |
| RNN |
0.9327 |
0.9831 |
0.9063 |
0.9431 |
0.9934 |
0.7156 |
0.0673 |
0.0673 |
0.2594 |
11.3673 |
408.0 kB |
15749 |
| LSTM |
0.8942 |
0.9206 |
0.9063 |
0.9134 |
0.9711 |
0.5531 |
0.1058 |
0.1058 |
0.3252 |
20.4970 |
1220.0 kB |
60635 |
| GRU |
0.8846 |
0.9643 |
0.8438 |
0.9000 |
0.9559 |
0.5125 |
0.1154 |
0.1154 |
0.3397 |
14.1540 |
712.0 kB |
38753 |
Table 9.
Top-performing models by Datasets.
Table 9.
Top-performing models by Datasets.
| Datasets |
Models |
Accuracy |
Precision |
Recall |
F1-score |
TT(s) |
MU |
| D1 |
AdaBoost |
0.798 |
0.671 |
0.833 |
0.743 |
1.181 |
0 B |
| D2 |
RF |
0.785 |
0.656 |
0.814 |
0.727 |
0.648 |
32 kB |
| D3 |
RNN |
0.714 |
0.438 |
0.498 |
0.441 |
141.709 |
600 kB |
| D4 |
DNN |
0.754 |
0.328 |
0.728 |
0.452 |
153.624 |
464 kB |
| D5 |
RF |
1.000 |
1.000 |
1.000 |
1.000 |
0.587 |
24 kB |
Table 10.
Comparative analysis of models used and existing diabetes prediction models using F1-score [
39].
Table 10.
Comparative analysis of models used and existing diabetes prediction models using F1-score [
39].
| Datasets |
Authors |
Outliers |
Missing Values |
Model |
Precision |
Accuracy |
Recall |
F1-score |
| |
[44] |
IQR |
Attribute Mean |
AB + XB |
– |
– |
0.7900 |
– |
| |
[46] |
– |
– |
GBM |
– |
– |
0.8700 |
– |
| |
[80] |
– |
– |
DA |
– |
0.7400 |
0.7200 |
– |
| |
[81] |
– |
– |
ANN |
– |
0.7600 |
0.5300 |
– |
| Dataset 1 |
[82] |
ESD |
k-NN |
HM-BagMoov |
– |
0.8600 |
0.8500 |
0.7900 |
| Dataset 2 |
[39] |
IQR |
CWM |
QML |
0.7400 |
0.8600 |
0.8500 |
0.7900 |
| |
[83] |
– |
NB |
RF |
0.8100 |
0.8700 |
0.8500 |
0.8300 |
| |
[84] |
– |
– |
k-NN |
0.8700 |
0.8800 |
0.9000 |
0.8800 |
| |
[56] |
GM |
Median |
RF |
– |
0.9300 |
0.7970 |
– |
| |
[85] |
– |
– |
RF |
0.9400 |
0.9400 |
0.8800 |
0.9100 |
| |
[39] |
IQR |
CWM |
DL |
0.9000 |
0.9500 |
0.9500 |
0.9300 |
| |
Our Study |
IQR |
ADASYN |
AdaBoost |
0.6716 |
0.7987 |
0.8333 |
0.7438 |
| |
Our Study |
IQR |
ADASYN |
RF |
0.6567 |
0.7857 |
0.8148 |
0.7273 |
| |
|
|
|
|
|
|
|
|
| Dataset 3 |
[76] |
– |
Excluded |
NN |
– |
0.8240 |
0.3781 |
– |
| Dataset 4 |
Our Study |
IQR |
Clustering |
RNN |
0.4387 |
0.7144 |
0.4982 |
0.4414 |
| |
Our Study |
IQR |
Clustering |
DNN |
0.3286 |
0.7548 |
0.7280 |
0.4526 |
| |
|
|
|
|
|
|
|
|
| Dataset 5 |
[77] |
– |
Ignoring Tuple |
RF |
0.9740 |
0.9740 |
0.9740 |
0.9740 |
| |
Our Study |
IQR |
– |
RF |
1.0000 |
1.0000 |
1.0000 |
1.0000 |
|
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).