Submitted:
30 April 2025
Posted:
30 April 2025
You are already at the latest version
Abstract
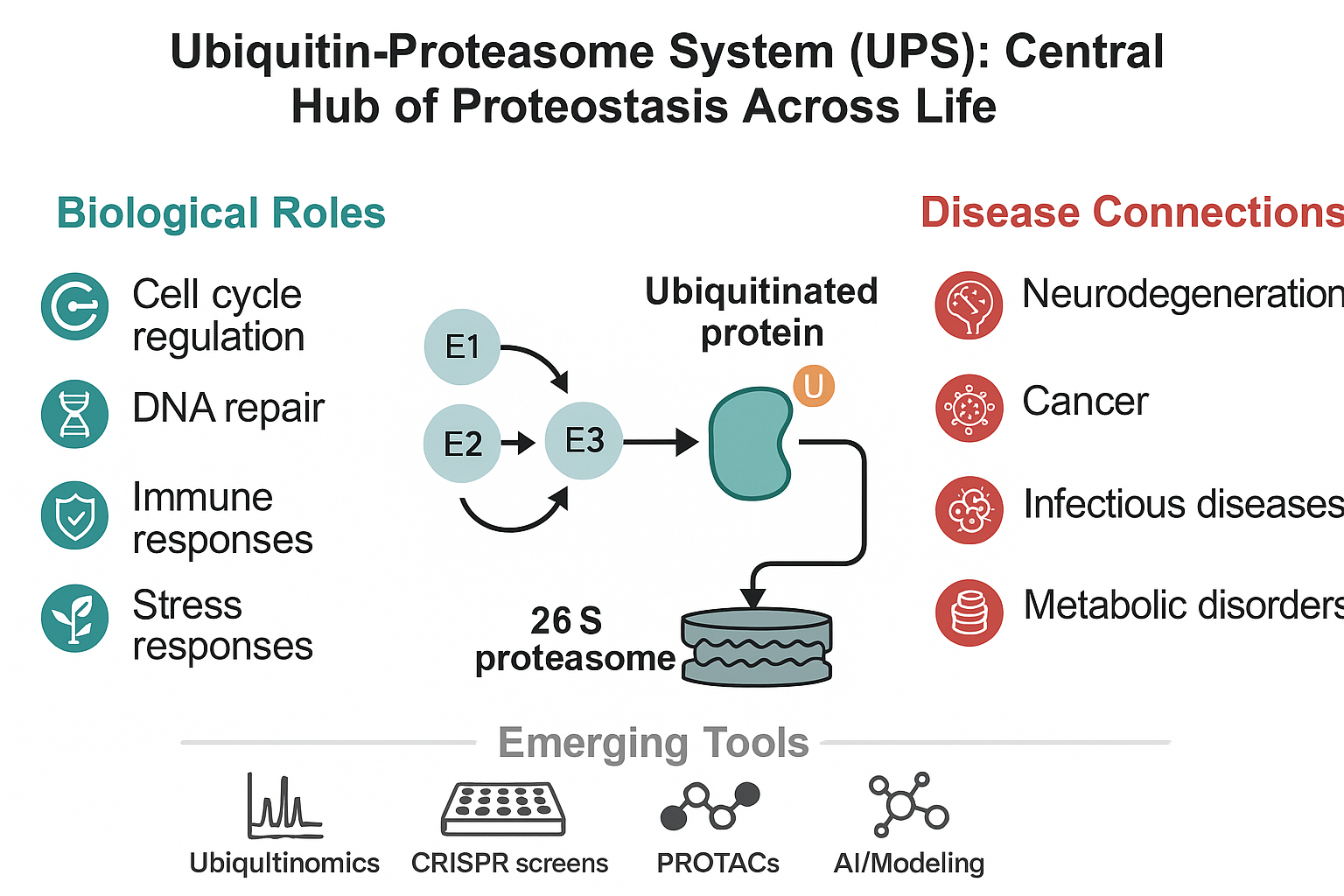
Keywords:
1. Introduction
1.1. Proteostasis and Cellular Protein Quality Control
1.2. The Ubiquitin-Proteasome System: A Central Regulator of Cellular Function
1.3. Rationale for a Cross-Disciplinary Exploration
2. The Ubiquitin-Proteasome System: Molecular Architecture and Function
2.1. Ubiquitination: The Enzymatic Cascade (E1, E2, E3)
2.2. Diversity of Ubiquitin Chains and Functional Outcomes
2.3. The 26S Proteasome: Structure and Regulation
2.4. Deubiquitinases (DUBs): Resetting the System
2.5. UPS vs Autophagy: Complementary or Redundant?
3. The UPS in Cellular and Systemic Homeostasis
3.1. Regulation of the Cell Cycle and DNA Damage Response
3.2. Adaptation to Proteotoxic Stress
3.3. Regulation of Cellular Signaling and Transcriptional Programs
3.4. Contribution of Systemic Homeostasis
4. Disease-Specific Insights into UPS Dysregulation
4.1. Neurodegenerative Disorders
UPS Dysfunction in Alzheimer’s, Parkinson’s, and Huntington’s Disease
Role of Protein Aggregates and Ubiquitin Tags
Therapeutic Strategies Targeting the UPS in Neurodegeneration
4.2. Cancer Biology
UPS Regulation of Oncoproteins and Tumor Suppressors
E3 Ligases as Tumor Drivers or Suppressors
Targeted Protein Degradation as a Novel Anti-Cancer Strategy
4.3. Infectious Diseases
Viral Manipulation of the Host UPS
Bacterial Subversion of Ubiquitination Pathways
Therapeutic Opportunities Targeting UPS-Pathogen Interactions
4.4. Immune Regulation and Inflammation
4.5. Metabolic Disorders and Aging
5. The UPS in Plant Biology and Agricultural Innovation
5.1. Regulation of Stress Responses and Hormone Signaling
5.2. Opportunities for Crop Improvement Through UPS Engineering
5.3. Cross-Kingdom Insights into Ubiquitination Mechanisms
6. Emerging Tools and Technologies to Study the UPS
6.1. Ubiquitin-omics and Mass Spectrometry
6.2. CRISPR-Based Screens for Functional Dissection of the UPS
6.3. Live-Cell Imaging of Ubiquitination and Proteasomal Activity
6.4. Artificial Intelligence and Computational Modeling of Ubiquitination
7. Challenges and Future Directions
7.1. Mapping E3 Ligase Specificity at Scale
7.2. Achieving Selectivity in Therapeutic Targeting
7.3. Expanding UPS Insights to Underexplored Diseases
7.4. Opportunities in Precision Medicine and Synthetic Biology
8. Conclusions
Author Contributions
Funding
Ethics declarations
Code availability
Data availability
Acknowledgment
Conflict of interest
References
- Aberger, F.; i Altaba, A. R. Context-dependent signal integration by the GLI code: the oncogenic load, pathways, modifiers and implications for cancer therapy. In Seminars in cell & developmental biology; Academic Press, September 2014; Vol. 33, pp. 93–104. [Google Scholar]
- Ajoolabady, A.; Lindholm, D.; Ren, J.; Pratico, D. ER stress and UPR in Alzheimer’s disease: mechanisms, pathogenesis, treatments. Cell death & disease 2022, 13(8), 706. [Google Scholar]
- Ajoolabady, A.; Lindholm, D.; Ren, J.; Pratico, D. ER stress and UPR in Alzheimer’s disease: mechanisms, pathogenesis, treatments. Cell death & disease 2022, 13(8), 706. [Google Scholar]
- Alam, I.; Batool, K.; Huang, Y.; Liu, J.; Ge, L. Developing genetic engineering techniques for control of seed size and yield. International journal of molecular sciences 2022, 23(21), 13256. [Google Scholar] [CrossRef]
- Alonso, A.; García del Portillo, F. Hijacking of eukaryotic functions by intracellular bacterial pathogens, 2004.
- An, Q.; Huang, L.; Wang, C.; Wang, D.; Tu, Y. New strategies to enhance the efficiency and precision of drug discovery. Frontiers in Pharmacology 2025, 16, 1550158. [Google Scholar] [CrossRef] [PubMed]
- Ashitomi, H.; Nakagawa, T.; Nakagawa, M.; Hosoi, T. Cullin-RING Ubiquitin Ligases in Neurodevelopment and Neurodevelopmental Disorders. Biomedicines 2025, 13(4), 810. [Google Scholar] [CrossRef] [PubMed]
- Banjare, R.; Nidhi, N.; Sood, A. Physiological Aspects of Flowering, Fruit Setting, Fruit Development and Fruit Drop, Regulation and their Manipulation: A Review. Int. J. Environ. Clim. Change 2023, 13(12), 205–224. [Google Scholar]
- Békés, M.; Langley, D. R.; Crews, C. M. PROTAC targeted protein degraders: the past is prologue. Nature reviews Drug discovery 2022, 21(3), 181–200. [Google Scholar] [CrossRef]
- Berkley, K.; Zalejski, J.; Sharma, N.; Sharma, A. Journey of PROTAC: From Bench to Clinical Trial and Beyond. Biochemistry 2025. [Google Scholar] [CrossRef] [PubMed]
- Biernacka, E. K.; Osadnik, T.; Bilińska, Z. T.; Krawczyński, M.; Latos-Bieleńska, A. I.; Gil, R. Genetic testing for inherited cardiovascular diseases. A position statement of the Polish Cardiac Society endorsed by Polish Society of Human Genetics and Cardiovascular Patient Communities. Polish Heart Journal (Kardiologia Polska) 2024, 82(5), 569–593. [Google Scholar] [CrossRef]
- Cabrera-Serrano, A. J.; Sánchez-Maldonado, J. M.; González-Olmedo, C.; Carretero-Fernández, M.; Díaz-Beltrán, L.; Gutiérrez-Bautista, J. F.; Sainz, J. Crosstalk Between Autophagy and Oxidative Stress in Hematological Malignancies: Mechanisms, Implications, and Therapeutic Potential. Antioxidants 2025, 14(3), 264. [Google Scholar] [CrossRef]
- Çetin, G.; Klafack, S.; Studencka-Turski, M.; Krüger, E.; Ebstein, F. The ubiquitin–proteasome system in immune cells. Biomolecules 2021, 11(1), 60. [Google Scholar] [CrossRef] [PubMed]
- Chaffai, R.; Ganesan, M.; Cherif, A. Gene Expression Regulation in Plant Abiotic Stress Response. In Plant Adaptation to Abiotic Stress: From Signaling Pathways and Microbiomes to Molecular Mechanisms; Springer Nature Singapore; Singapore, 2024; pp. 275–302. [Google Scholar]
- Chandra, P.; Grigsby, S. J.; Philips, J. A. Immune evasion and provocation by Mycobacterium tuberculosis. Nature Reviews Microbiology 2022, 20(12), 750–766. [Google Scholar] [CrossRef] [PubMed]
- Chao, C. C. K. Mechanisms of p53 degradation. Clinica Chimica Acta 2015, 438, 139–147. [Google Scholar] [CrossRef]
- Chopra, G.; Shabir, S.; Yousuf, S.; Kauts, S.; Bhat, S. A.; Mir, A. H.; Singh, M. P. Proteinopathies: deciphering physiology and mechanisms to develop effective therapies for neurodegenerative diseases. Molecular neurobiology 2022, 59(12), 7513–7540. [Google Scholar] [CrossRef]
- Chua, J. P.; De Calbiac, H.; Kabashi, E.; Barmada, S. J. Autophagy and ALS: mechanistic insights and therapeutic implications. Autophagy 2022, 18(2), 254–282. [Google Scholar] [CrossRef]
- Ciechanover, A.; Kwon, Y. T. Degradation of misfolded proteins in neurodegenerative diseases: therapeutic targets and strategies. Experimental & molecular medicine 2015, 47(3), e147–e147. [Google Scholar]
- Cui, X.; Wang, J.; Li, K.; Lv, B.; Hou, B.; Ding, Z. Protein post-translational modifications in auxin signaling. Journal of Genetics and Genomics 2024, 51(3), 279–291. [Google Scholar] [CrossRef] [PubMed]
- Dang, F.; Nie, L.; Wei, W. Ubiquitin signaling in cell cycle control and tumorigenesis. Cell Death & Differentiation 2021, 28(2), 427–438. [Google Scholar]
- Dhara, A.; Sinai, A. P. A cell cycle-regulated, 2016.
- Dhoundiyal, S.; Srivastava, S.; Kumar, S.; Singh, G.; Ashique, S.; Pal, R.; Taghizadeh-Hesary, F. Radiopharmaceuticals: navigating the frontier of precision medicine and therapeutic innovation. European Journal of Medical Research 2024, 29(1), 26. [Google Scholar] [CrossRef]
- Di Conza, G.; Ho, P. C.; Cubillos-Ruiz, J. R.; Huang, S. C. C. Control of immune cell function by the unfolded protein response. Nature Reviews Immunology 2023, 23(9), 546–562. [Google Scholar] [CrossRef]
- Dreher, K.; Callis, J. Ubiquitin, hormones and biotic stress in plants. Annals of botany 2007, 99(5), 787–822. [Google Scholar] [CrossRef]
- Enenkel, C.; Kang, R. W.; Wilfling, F.; Ernst, O. P. Intracellular localization of the proteasome in response to stress conditions. Journal of Biological Chemistry 2022, 298(7), 102083. [Google Scholar] [CrossRef]
- Fan, J.; Bellon, M.; Ju, M.; Zhao, L.; Wei, M.; Fu, L.; Nicot, C. Clinical significance of FBXW7 loss of function in human cancers. Molecular cancer 2022, 21(1), 87. [Google Scholar] [CrossRef] [PubMed]
- Farshi, P.; Deshmukh, R. R.; Nwankwo, J. O.; Arkwright, R. T.; Cvek, B.; Liu, J.; Dou, Q. P. Deubiquitinases (DUBs) and DUB inhibitors: a patent review. Expert opinion on therapeutic patents 2015. [Google Scholar]
- Fhu, C. W.; Ali, A. Dysregulation of the ubiquitin proteasome system in human malignancies: a window for therapeutic intervention. Cancers 2021, 13(7), 1513. [Google Scholar] [CrossRef]
- Fleming, A.; Bourdenx, M.; Fujimaki, M.; Karabiyik, C.; Krause, G. J.; Lopez, A.; Rubinsztein, D. C. The different autophagy degradation pathways and neurodegeneration. Neuron 2022, 110(6), 935–966. [Google Scholar] [CrossRef]
- Foster, B.; Attwood, M.; Gibbs-Seymour, I. Tools for decoding ubiquitin signaling in DNA repair. Frontiers in cell and developmental biology 2021, 9, 760226. [Google Scholar] [CrossRef]
- Frankowska, N.; Lisowska, K.; Witkowski, J. M. Proteolysis dysfunction in the process of aging and age-related diseases. Frontiers in Aging 2022, 3, 927630. [Google Scholar] [CrossRef] [PubMed]
- Galic, S.; Sachithanandan, N.; Kay, T. W.; Steinberg, G. R. Suppressor of cytokine signalling (SOCS) proteins as guardians of inflammatory responses critical for regulating insulin sensitivity. Biochemical Journal 2014, 461(2), 177–188. [Google Scholar] [CrossRef] [PubMed]
- Geddes-McAlister, J.; Uhrig, R. G. The plant proteome delivers from discovery to innovation. Trends in Plant Science 2025. [Google Scholar] [CrossRef]
- Gonçalves, M.; Vale, N.; Silva, P. Neuroprotective effects of olive oil: A comprehensive review of antioxidant properties. Antioxidants 2024, 13(7), 762. [Google Scholar] [CrossRef]
- Gonzalez-Santamarta, M.; Bouvier, C.; Rodriguez, M. S.; Xolalpa, W. Ubiquitin-chains dynamics and its role regulating crucial cellular processes. In Seminars in Cell & Developmental Biology; Academic Press, December 2022; Vol. 132, pp. 155–170. [Google Scholar]
- Greil, C.; Engelhardt, M.; Wäsch, R. The role of the APC/C and its coactivators Cdh1 and Cdc20 in cancer development and therapy. Frontiers in Genetics 2022, 13, 941565. [Google Scholar] [CrossRef]
- Groettrup, M.; Kirk, C. J.; Basler, M. Proteasomes in immune cells: more than peptide producers? Nature Reviews Immunology 2010, 10(1), 73–78. [Google Scholar] [CrossRef] [PubMed]
- Gu, W.; Wu, G.; Chen, G.; Meng, X.; Xie, Z.; Cai, S. Polyphenols alleviate metabolic disorders: the role of ubiquitin-proteasome system. Frontiers in Nutrition 2024, 11, 1445080. [Google Scholar] [CrossRef]
- Guo, Q.; Jin, Y.; Chen, X.; Ye, X.; Shen, X.; Lin, M.; Zhang, J. NF-κB in biology and targeted therapy: new insights and translational implications. Signal transduction and targeted therapy 2024, 9(1), 53. [Google Scholar] [CrossRef]
- Haberecht-Müller, S.; Krüger, E.; Fielitz, J. Out of control: the role of the ubiquitin proteasome system in skeletal muscle during inflammation. Biomolecules 2021, 11(9), 1327. [Google Scholar] [CrossRef]
- Hanna, J.; Guerra-Moreno, A.; Ang, J.; Micoogullari, Y. Protein degradation and the pathologic basis of disease. The American journal of pathology 2019, 189(1), 94–103. [Google Scholar] [CrossRef]
- Haq, E. U.; Yousaf, A.; Sardar, M.; Irshad, A.; Basharat, Z.; Ali, A.; Arsalan, M. Uncovering Genetic Interactions: CRISPR-Mediated Gene Knockouts and Activations in Understanding Complex Diseases. Pak-Euro Journal of Medical and Life Sciences 2024, 7(Special 2), S211–S220. [Google Scholar]
- Henning, R. H.; Brundel, B. J. Proteostasis in cardiac health and disease. Nature Reviews Cardiology 2017, 14(11), 637–653. [Google Scholar] [CrossRef] [PubMed]
- Hipp, M. S.; Park, S. H.; Hartl, F. U. Proteostasis impairment in protein-misfolding and-aggregation diseases. Trends in cell biology 2014, 24(9), 506–514. [Google Scholar] [CrossRef]
- Hoppe, T.; Cohen, E. Organismal protein homeostasis mechanisms. Genetics 2020, 215(4), 889–901. [Google Scholar] [CrossRef]
- Hou, C.; Li, Y.; Wang, M.; Wu, H.; Li, T. Systematic prediction of degrons and E3 ubiquitin ligase binding via deep learning. BMC biology 2022, 20(1), 162. [Google Scholar] [CrossRef] [PubMed]
- Javed, T.; I, I.; Singhal, R. K.; Shabbir, R.; Shah, A. N.; Kumar, P.; Siuta, D. Recent advances in agronomic and physio-molecular approaches for improving nitrogen use efficiency in crop plants. Frontiers in Plant Science 2022, 13, 877544. [Google Scholar] [CrossRef] [PubMed]
- Kacprzyk, J.; Burke, R.; Armengot, L.; Coppola, M.; Tattrie, S. B.; Vahldick, H.; McCabe, P. F. Roadmap for the next decade of plant programmed cell death research. New Phytologist 2024, 242(5), 1865–1875. [Google Scholar] [CrossRef] [PubMed]
- Kalies, S. M. K. Visualization and manipulation of repair and regeneration in biological systems using light, 2023.
- Kao, S. H.; Wu, H. T.; Wu, K. J. Ubiquitination by HUWE1 in tumorigenesis and beyond. Journal of biomedical science 2018, 25, 1–15. [Google Scholar] [CrossRef]
- Kaushik, S.; Cuervo, A. M. Proteostasis and aging. Nature medicine 2015, 21(12), 1406–1415. [Google Scholar] [CrossRef]
- Kim, J.; Byun, I.; Joh, H.; Kim, H. J.; Lee, M. J. Targeted protein degradation directly engaging lysosomes or proteasomes. In Chemical Society Reviews; 2024. [Google Scholar]
- Kloetzel, P. M. Generation of major histocompatibility complex class I antigens: functional interplay between proteasomes and TPPII. Nature immunology 2004, 5(7), 661–669. [Google Scholar] [CrossRef]
- Kocaturk, N. M.; Gozuacik, D. Crosstalk between mammalian autophagy and the ubiquitin-proteasome system. Frontiers in cell and developmental biology 2018, 6, 128. [Google Scholar] [CrossRef]
- Koliopoulos, M. G.; Alfieri, C. Cell cycle regulation by complex nanomachines. The FEBS Journal 2022, 289(17), 5100–5120. [Google Scholar] [CrossRef]
- Koszła, O.; Sołek, P. Misfolding and aggregation in neurodegenerative diseases: Protein quality control machinery as potential therapeutic clearance pathways. Cell Communication and Signaling 2024, 22(1), 421. [Google Scholar] [CrossRef]
- Kumar, A. V.; Mills, J.; Lapierre, L. R. Selective autophagy receptor p62/SQSTM1, a pivotal player in stress and aging. Frontiers in cell and developmental biology 2022, 10, 793328. [Google Scholar] [CrossRef]
- Kumar, P. Protein misfolding in nonneurological diseases. In Protein Misfolding in Neurodegenerative Diseases; Academic Press, 2025; pp. 493–523. [Google Scholar]
- Kumar, P.; Paul, D.; Jhajhriya, S.; Kumar, R.; Dutta, S.; Siwach, P.; Das, S. Understanding heat-shock proteins’ abundance and pivotal function under multiple abiotic stresses. Journal of Plant Biochemistry and Biotechnology 2024, 1–22. [Google Scholar] [CrossRef]
- Kurtishi, A.; Rosen, B.; Patil, K. S.; Alves, G. W.; Møller, S. G. Cellular proteostasis in neurodegeneration. Molecular neurobiology 2019, 56(5), 3676–3689. [Google Scholar] [CrossRef]
- Lang, B. J.; Guerrero, M. E.; Prince, T. L.; Okusha, Y.; Bonorino, C.; Calderwood, S. K. The functions and regulation of heat shock proteins; key orchestrators of proteostasis and the heat shock response. Archives of toxicology 2021, 95(6), 1943–1970. [Google Scholar] [CrossRef] [PubMed]
- Lee, B. H.; Lee, M. J.; Park, S.; Oh, D. C.; Elsasser, S.; Chen, P. C.; Finley, D. Enhancement of proteasome activity by a small-molecule inhibitor of USP14. Nature 2010, 467(7312), 179–184. [Google Scholar] [CrossRef] [PubMed]
- Lee, J. M.; Hammarén, H. M.; Savitski, M. M.; Baek, S. H. Control of protein stability by post-translational modifications. Nature Communications 2023, 14(1). [Google Scholar] [CrossRef]
- Lee, J. M.; Hammarén, H. M.; Savitski, M. M.; Baek, S. H. Control of protein stability by post-translational modifications. Nature Communications 2023, 14(1). [Google Scholar] [CrossRef]
- Li, L. Role of the metacaspase AtMC1 in stress-triggered protein aggregate formation in yeast and plants, 2021.
- Li, Y.; Li, S.; Wu, H. Ubiquitination-proteasome system (UPS) and autophagy two main protein degradation machineries in response to cell stress. Cells 2022, 11(5), 851. [Google Scholar] [CrossRef]
- Liu, F.; Xi, M.; Liu, T.; Wu, X.; Ju, L.; Wang, D. The central role of transcription factors in bridging biotic and abiotic stress responses for plants’ resilience. New crops 2024, 1, 100005. [Google Scholar] [CrossRef]
- Liu, J.; Liang, L.; Su, B.; Wu, D.; Zhang, Y.; Wu, J.; Fu, C. Transformative strategies in photocatalyst design: merging computational methods and deep learning. Journal of Materials Informatics 2024, 4(4), N–A. [Google Scholar] [CrossRef]
- Madsen, D. A.; Schmidt, S. I.; Blaabjerg, M.; Meyer, M. Interaction between Parkin and α-synuclein in PARK2-mediated Parkinson’s disease. Cells 2021, 10(2), 283. [Google Scholar] [CrossRef]
- Maiuolo, J.; Gliozzi, M.; Musolino, V.; Carresi, C.; Scarano, F.; Nucera, S.; Mollace, V. From metabolic syndrome to neurological diseases: role of autophagy. Frontiers in Cell and Developmental Biology 2021, 9, 651021. [Google Scholar] [CrossRef]
- Marine, J. C.; Lozano, G. Mdm2-mediated ubiquitylation: p53 and beyond. Cell Death & Differentiation 2010, 17(1), 93–102. [Google Scholar]
- Marshall, R. S.; Vierstra, R. D. Dynamic regulation of the 26S proteasome: from synthesis to degradation. Frontiers in Molecular Biosciences 2019, 6, 40. [Google Scholar] [CrossRef] [PubMed]
- Marshall, S. Role of insulin, adipocyte hormones, and nutrient-sensing pathways in regulating fuel metabolism and energy homeostasis: a nutritional perspective of diabetes, obesity, and cancer. Science’s STKE 2006, 2006(346), re7–re7. [Google Scholar] [CrossRef] [PubMed]
- Mevissen, T. E.; Komander, D. Mechanisms of deubiquitinase specificity and regulation. Annual review of biochemistry 2017, 86(1), 159–192. [Google Scholar] [CrossRef] [PubMed]
- Meyer-Schwesinger, C. The ubiquitin–proteasome system in kidney physiology and disease. Nature Reviews Nephrology 2019, 15(7), 393–411. [Google Scholar] [CrossRef]
- Millar, S. R. Evaluating Stress Granule Compositional Plasticity using a Systematic Proteomics Approach. Master’s thesis, University of Toronto (Canada)), 2023. [Google Scholar]
- Molinaro, C.; Martoriati, A.; Cailliau, K. Proteins from the DNA damage response: Regulation, dysfunction, and anticancer strategies. Cancers 2021, 13(15), 3819. [Google Scholar] [CrossRef]
- Müller-Rischart, A. K.; Pilsl, A.; Beaudette, P.; Patra, M.; Hadian, K.; Funke, M.; Winklhofer, K. F. The E3 ligase parkin maintains mitochondrial integrity by increasing linear ubiquitination of NEMO. Molecular cell 2013, 49(5), 908–921. [Google Scholar] [CrossRef]
- Nagel, Y. A.; Britschgi, A.; Ricci, A. From Degraders to Molecular Glues: New Ways of Breaking Down Disease-Associated Proteins. Successful Drug Discovery 2021, 47–85. [Google Scholar]
- Nagpal, I.; Yuan, Z. M. The basally expressed p53-mediated homeostatic function. Frontiers in cell and developmental biology 2021, 9, 775312. [Google Scholar] [CrossRef]
- Nerkar, G.; Devarumath, S.; Purankar, M.; Kumar, A.; Valarmathi, R.; Devarumath, R.; Appunu, C. Advances in crop breeding through precision genome editing. Frontiers in Genetics 2022, 13, 880195. [Google Scholar] [CrossRef] [PubMed]
- Ottens, F.; Franz, A.; Hoppe, T. Build-UPS and break-downs: metabolism impacts on proteostasis and aging. Cell Death & Differentiation 2021, 28(2), 505–521. [Google Scholar]
- Panwar, S.; Uniyal, P.; Kukreti, N.; Hashmi, A.; Verma, S.; Arya, A.; Joshi, G. Role of autophagy and proteostasis in neurodegenerative diseases: Exploring the therapeutic interventions. Chemical Biology & Drug Design 2024, 103(4), e14515. [Google Scholar]
- Papaevgeniou, N.; Chondrogianni, N. UPS activation in the battle against aging and aggregation-related diseases: an extended review. Proteostasis: Methods and Protocols 2016, 1–70. [Google Scholar]
- Papaioannou, A.; Chevet, E. Driving cancer tumorigenesis and metastasis through UPR signaling. Coordinating organismal physiology through the unfolded protein response 2017, 159–192. [Google Scholar]
- Parihar, N.; Bhatt, L. K. Deubiquitylating enzymes: potential target in autoimmune diseases. Inflammopharmacology 2021, 29, 1683–1699. [Google Scholar] [CrossRef]
- Pennington, K. L.; Chan, T. Y.; Torres, M. P.; Andersen, J. The dynamic and stress-adaptive signaling hub of 14-3-3: emerging mechanisms of regulation and context-dependent protein–protein interactions. Oncogene 2018, 37(42), 5587–5604. [Google Scholar] [CrossRef]
- Pettersson, M.; Crews, C. M. PROteolysis TArgeting Chimeras (PROTACs)—past, present and future. Drug Discovery Today: Technologies 2019, 31, 15–27. [Google Scholar] [CrossRef]
- Phanindhar, K.; Mishra, R. K. Auxin-inducible degron system: an efficient protein degradation tool to study protein function. Biotechniques 2023, 74(4), 186–198. [Google Scholar] [CrossRef]
- Pohl, C.; Dikic, I. Cellular quality control by the ubiquitin-proteasome system and autophagy. Science 2019, 366(6467), 818–822. [Google Scholar] [CrossRef]
- Potjewyd, F. M.; Axtman, A. D. Exploration of aberrant E3 ligases implicated in Alzheimer’s disease and development of chemical tools to modulate their function. Frontiers in Cellular Neuroscience 2021, 15, 768655. [Google Scholar] [CrossRef] [PubMed]
- Pożoga, M.; Armbruster, L.; Wirtz, M. From nucleus to membrane: a subcellular map of the N-acetylation machinery in plants. International Journal of Molecular Sciences 2022, 23(22), 14492. [Google Scholar] [CrossRef]
- PT, B.; Sahu, I. Decoding the ubiquitin landscape by cutting-edge ubiquitinomic approaches. Biochemical Society Transactions 2024, 52(2), 627–637. [Google Scholar]
- Qin, B.; Chen, X.; Wang, F.; Wang, Y. DUBs in Alzheimer’s disease: mechanisms and therapeutic implications. Cell Death Discovery 2024, 10(1), 475. [Google Scholar] [CrossRef] [PubMed]
- Rana, P. S.; Ignatz-Hoover, J. J.; Driscoll, J. J. Targeting proteasomes and the MHC class I antigen presentation machinery to treat cancer, infections and age-related diseases. Cancers 2023, 15(23), 5632. [Google Scholar] [CrossRef]
- Saha, B.; Nayak, J.; Srivastava, R.; Samal, S.; Kumar, D.; Chanwala, J.; Giri, M. K. Unraveling the involvement of WRKY TFs in regulating plant disease defense signaling. Planta 2024, 259(1), 7. [Google Scholar] [CrossRef]
- Samarasinghe, K. T.; Crews, C. M. Targeted protein degradation: A promise for undruggable proteins. Cell chemical biology 2021, 28(7), 934–951. [Google Scholar] [CrossRef] [PubMed]
- Sang, T.; Zhang, Z.; Liu, G.; Wang, P. Navigating the landscape of plant proteomics. Journal of Integrative Plant Biology 2025. [Google Scholar] [CrossRef]
- Sharma, V.; Verma, S.; Seranova, E.; Sarkar, S.; Kumar, D. Selective autophagy and xenophagy in infection and disease. Frontiers in cell and developmental biology 2018, 6, 147. [Google Scholar] [CrossRef]
- Sherman, M. Y.; Goldberg, A. L. Cellular defenses against unfolded proteins: a cell biologist thinks about neurodegenerative diseases. Neuron 2001, 29(1), 15–32. [Google Scholar] [CrossRef]
- Shi, D.; Gu, W. Dual roles of MDM2 in the regulation of p53: ubiquitination dependent and ubiquitination independent mechanisms of MDM2 repression of p53 activity. Genes & cancer 2012, 3(3-4), 240–248. [Google Scholar]
- Shin, J. Y.; Muniyappan, S.; Tran, N. N.; Park, H.; Lee, S. B.; Lee, B. H. Deubiquitination reactions on the proteasome for proteasome versatility. International Journal of Molecular Sciences 2020, 21(15), 5312. [Google Scholar] [CrossRef]
- Sikander, R.; Arif, M.; Ghulam, A.; Worachartcheewan, A.; Thafar, M. A.; Habib, S. Identification of the ubiquitin–proteasome pathway domain by hyperparameter optimization based on a 2D convolutional neural network. Frontiers in Genetics 2022, 13, 851688. [Google Scholar] [CrossRef] [PubMed]
- Singh, B. K.; Delgado-Baquerizo, M.; Egidi, E.; Guirado, E.; Leach, J. E.; Liu, H.; Trivedi, P. Climate change impacts on plant pathogens, food security and paths forward. Nature Reviews Microbiology 2023, 21(10), 640–656. [Google Scholar] [CrossRef]
- Sinha, R.; Bala, M.; Ranjan, A.; Lal, S. K.; Sharma, T. R.; Pattanayak, A.; Singh, A. K. Proteomic approaches to understand plant response to abiotic stresses. In Agricultural biotechnology: Latest research and trends; Springer Nature Singapore; Singapore, 2022; pp. 351–383. [Google Scholar]
- Snyder, N. A.; Silva, G. M. Deubiquitinating enzymes (DUBs): Regulation, homeostasis, and oxidative stress response. Journal of Biological Chemistry 2021, 297(3). [Google Scholar] [CrossRef]
- Snyder, N. A.; Silva, G. M. Deubiquitinating enzymes (DUBs): Regulation, homeostasis, and oxidative stress response. Journal of Biological Chemistry 2021, 297(3). [Google Scholar] [CrossRef]
- Song, Y.; Xu, Y.; Liu, Y.; Gao, J.; Feng, L.; Zhang, Y.; Zhang, M. Mitochondrial quality control in the maintenance of cardiovascular homeostasis: The roles and interregulation of UPS, mitochondrial dynamics and mitophagy. Oxidative medicine and cellular longevity 2021, 2021(1), 3960773. [Google Scholar] [CrossRef]
- Strebel, K. HIV accessory genes Vif and Vpu. Advances in pharmacology 2007, 55, 199–232. [Google Scholar] [PubMed]
- Su, M.; Zhang, Z.; Zhou, L.; Han, C.; Huang, C.; Nice, E. C. Proteomics, personalized medicine and cancer. Cancers 2021, 13(11), 2512. [Google Scholar] [CrossRef]
- Sun, R. Post-Translational Regulation of 4-1BB, an Emerging Target for Cancer Immunotherapy. Doctoral dissertation, Purdue University, 2022. [Google Scholar]
- Suranjika, S.; Barla, P.; Sharma, N.; Dey, N. A review on ubiquitin ligases: Orchestrators of plant resilience in adversity. Plant Science 2024, 112180. [Google Scholar] [CrossRef]
- Tabar, M. S.; Parsania, C.; Chen, H.; Su, X. D.; Bailey, C. G.; Rasko, J. E. Illuminating the dark protein-protein interactome. Cell reports methods 2022, 2(8). [Google Scholar]
- Teixeira, A. P.; Fussenegger, M. Synthetic Gene Circuits for Regulation of Next-Generation Cell-Based Therapeutics. Advanced Science 2024, 11(8), 2309088. [Google Scholar] [CrossRef] [PubMed]
- Teixeira, A. P.; Fussenegger, M. Synthetic Gene Circuits for Regulation of Next-Generation Cell-Based Therapeutics. Advanced Science 2024, 11(8), 2309088. [Google Scholar] [CrossRef] [PubMed]
- Thakur, A.; Mikkelsen, H.; Jungersen, G. Intracellular pathogens: host immunity and microbial persistence strategies. Journal of immunology research 2019, 2019(1), 1356540. [Google Scholar] [CrossRef]
- Tomaić, V. Functional roles of E6 and E7 oncoproteins in HPV-induced malignancies at diverse anatomical sites. Cancers 2016, 8(10), 95. [Google Scholar] [CrossRef]
- Tomaskovic, I.; Gonzalez, A.; Dikic, I. Ubiquitin and Legionella: From bench to bedside. In Seminars in Cell & Developmental Biology; Academic Press, December 2022; Vol. 132, pp. 230–241. [Google Scholar]
- Tracz, M.; Bialek, W. Beyond K48 and K63: non-canonical protein ubiquitination. Cellular & Molecular Biology Letters 2021, 26(1), 1. [Google Scholar]
- Tröster, V.; Wong, R. P.; Börgel, A.; Cakilkaya, B.; Renz, C.; Möckel, M. M.; Ulrich, H. D. Custom affinity probes reveal DNA-damage-induced, ssDNA-independent chromatin SUMOylation in budding yeast. Cell Reports 2025, 44(3). [Google Scholar] [CrossRef]
- Tu, Y.; Chen, C.; Pan, J.; Xu, J.; Zhou, Z. G.; Wang, C. Y. The Ubiquitin Proteasome Pathway (UPP) in the regulation of cell cycle control and DNA damage repair and its implication in tumorigenesis. International journal of clinical and experimental pathology 2012, 5(8), 726. [Google Scholar]
- Tubío-Santamaría, N.; Jayavelu, A. K.; Schnoeder, T. M.; Eifert, T.; Hsu, C. J.; Perner, F.; Heidel, F. H. Immunoproteasome function maintains oncogenic gene expression in KMT2A-complex driven leukemia. Molecular Cancer 2023, 22(1), 196. [Google Scholar] [CrossRef]
- Tyedmers, J.; Mogk, A.; Bukau, B. Cellular strategies for controlling protein aggregation. Nature reviews Molecular cell biology 2010, 11(11), 777–788. [Google Scholar] [CrossRef]
- Uday, G.; Shardha, H. B.; Priyanka, K.; Jagirdhar, S. Advances in Plant Proteomics toward Improvement of Crop Productivity and Stress Resistance. Plant Proteomics: Implications in Growth, Quality Improvement, and Stress Resilience 2024, 11–29. [Google Scholar]
- Ulfig, A.; Jakob, U. Cellular oxidants and the proteostasis network: balance between activation and destruction. Trends in Biochemical Sciences 2024. [Google Scholar] [CrossRef]
- Ullah, F. U. M.; Rho, S.; Lee, M. Y. Predictive modeling for ubiquitin proteins through advanced machine learning technique. Heliyon 2024, 10(12). [Google Scholar]
- van der Wal, L.; Bezstarosti, K.; Demmers, J. A. A ubiquitinome analysis to study the functional roles of the proteasome associated deubiquitinating enzymes USP14 and UCH37. Journal of Proteomics 2022, 262, 104592. [Google Scholar] [CrossRef]
- Voutsadakis, I. A. Ubiquitin-and ubiquitin-like proteins-conjugating enzymes (E2s) in breast cancer. Molecular biology reports 2013, 40(2), 2019–2034. [Google Scholar] [CrossRef]
- Wang, C.; Zhang, Y.; Chen, W.; Wu, Y.; Xing, D. New-generation advanced PROTACs as potential therapeutic agents in cancer therapy. Molecular Cancer 2024, 23(1), 110. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Maldonado, M. A. The ubiquitin-proteasome system and its role in inflammatory and autoimmune diseases. Cell Mol Immunol 2006, 3(4), 255–261. [Google Scholar]
- Wang, J.; Xiang, Y.; Fan, M.; Fang, S.; Hua, Q. The Ubiquitin–Proteasome System in Tumor Metabolism. Cancers 2023, 15(8), 2385. [Google Scholar] [CrossRef]
- Wang, Z.; Inuzuka, H.; Zhong, J.; Wan, L.; Fukushima, H.; Sarkar, F. H.; Wei, W. Tumor suppressor functions of FBW7 in cancer development and progression. FEBS letters 2012, 586(10), 1409–1418. [Google Scholar] [CrossRef]
- Xu, T.; Wang, Q.; Wang, Q.; Sun, L. Mass spectrometry-intensive top-down proteomics: an update on technology advancements and biomedical applications. Analytical Methods 2024. [Google Scholar] [CrossRef]
- Yan, X.; Liu, X.; Zhao, C.; Chen, G. Q. Applications of synthetic biology in medical and pharmaceutical fields. Signal transduction and targeted therapy 2023, 8(1), 199. [Google Scholar] [CrossRef] [PubMed]
- Yang, Q.; Zhao, J.; Chen, D.; Wang, Y. E3 ubiquitin ligases: styles, structures and functions. Molecular biomedicine 2021, 2, 1–17. [Google Scholar] [CrossRef] [PubMed]
- Yu, F.; Yu, C.; Li, F.; Zuo, Y.; Wang, Y.; Yao, L.; Ye, L. Wnt/β-catenin signaling in cancers and targeted therapies. Signal Transduction and Targeted Therapy 2021, 6(1), 307. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.; Sidhu, S. S. Development of inhibitors in the ubiquitination cascade. FEBS letters 2014, 588(2), 356–367. [Google Scholar] [CrossRef]
- Zhou, Y.; Chen, Z.; Liu, S.; Liu, S.; Liao, Y.; Du, A.; Xia, Z. A Cullin 5-based complex serves as an essential modulator of ORF9b stability in SARS-CoV-2 replication. Signal Transduction and Targeted Therapy 2024, 9(1), 159. [Google Scholar] [CrossRef]
- Zou, Q.; Liu, M.; Liu, K.; Zhang, Y.; North, B. J.; Wang, B. E3 ubiquitin ligases in cancer stem cells: key regulators of cancer hallmarks and novel therapeutic opportunities. Cellular Oncology 2023, 46(3), 545–570. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



