Submitted:
11 March 2025
Posted:
13 March 2025
You are already at the latest version
Abstract
To date, Leucobacter species have been identified from diverse sources with various ecological and functional roles. However, the genomic features and pathogenic potential of antibiotic-resistant Leucobacter strains remains understudied. Here, we isolated the Leucobacter sp. HNU-1 from tropical Hainan Province, China, and found it can induce diapause in Caenorhabditis elegans following ingestion, while exhibiting no significant effects on the nematode's lifespan, survival rate, locomotion, and intestinal epithelial cells. This bacterium demonstrates resistance to multiple antibiotics, including kanamycin, streptomycin, sulfonamides, and vancomycin. On LB medium, Leucobacter sp. HNU-1 forms yellow, opaque colonies with a smooth, moist surface, regular edges, a convex center, and no surrounding halo, with diameters ranging from 2 to 3 mm. Furthermore, we performed whole-genome sequencing using third-generation high-throughput sequencing technology. De novo assembly revealed a genome size of 3,375,033 bp, with a GC content of 70.37%. A total of 3,270 functional genes, accounting for 88.98% of the genome, were annotated, along with six potential CRISPR sequences and other genetic elements. Genomic and bioinformatic analyses further identified antibiotics-related genes. This research provides a theoretical foundation for investigating antibiotic-resistant environmental bacteria in tropical environments and offers new insights into potential therapeutic strategies for microbial infections and host-microbe interactions.
Keywords:
1. Introduction
2. Results
2.1. The Developmental Toxicity of Leucobacter sp. HNU-1 to Caenorhabditis elegans
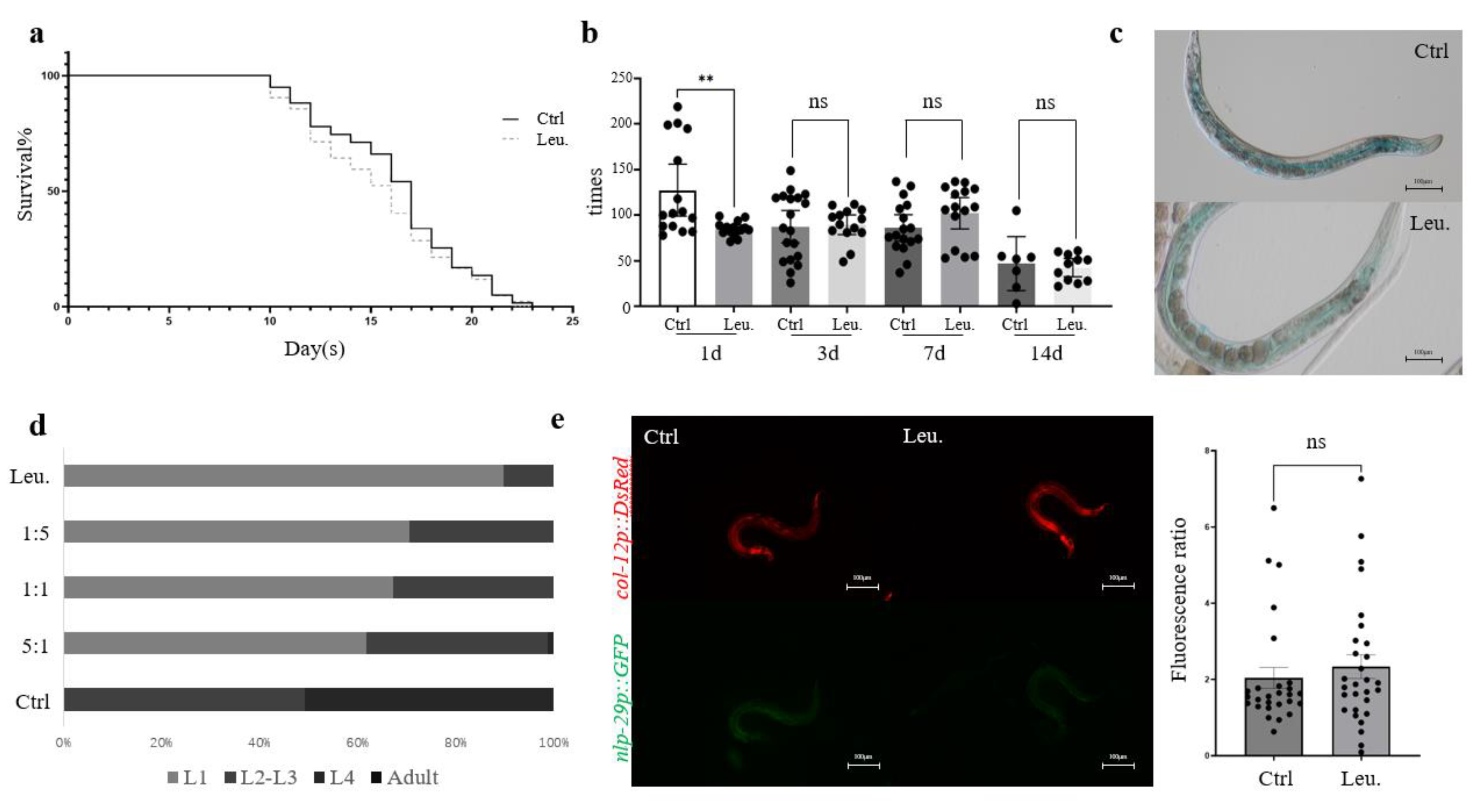
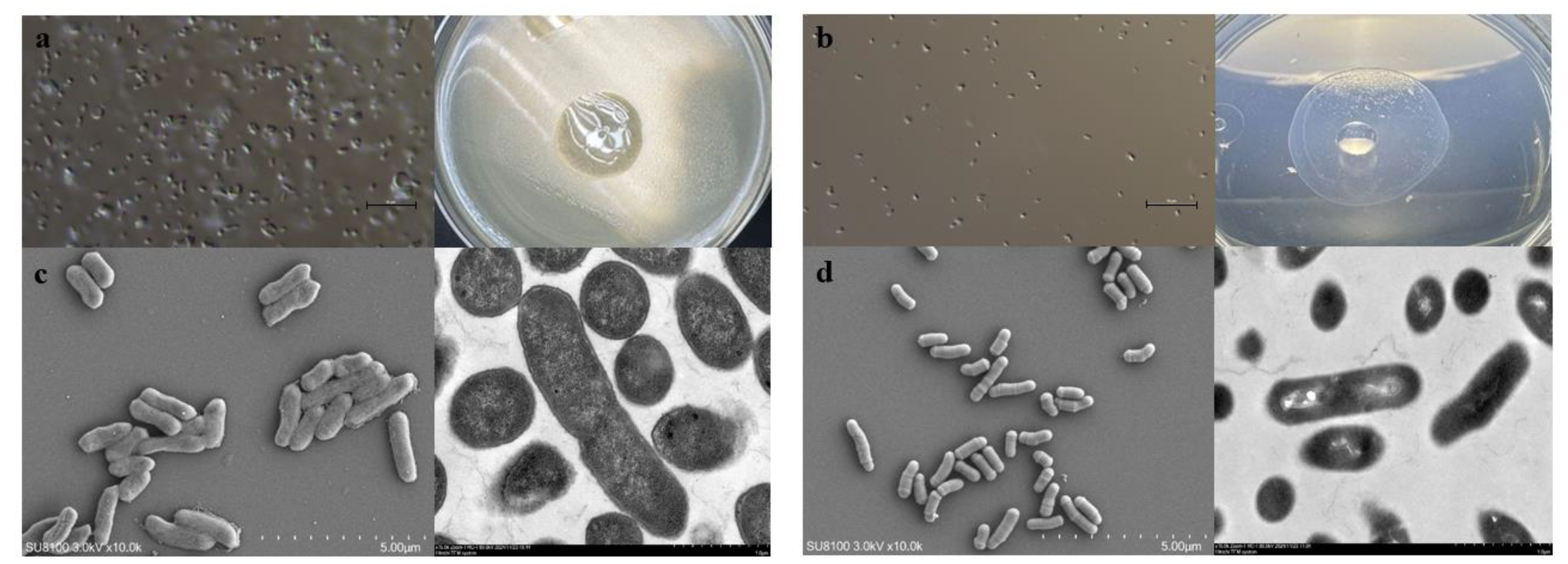
2.2. Morphological Identification of Leucobacter
2.3. Biochemical Tests of HNU-1
| Biochemical Test | Result |
| Semi-solid agar motility | + |
| Ornithine decarboxylase | - |
| Lysine decarboxylase | + |
| Citrate utilization | - |
| H₂S production | - |
| Urease | - |
| Indole test | + |
| Methyl red (MR) test | + |
| Voges-Proskauer (VP) test | - |
| Phenylalanine deamination | - |
| Mannitol utilization | + |
| Inositol utilization | - |
| Sorbitol utilization | + |
| Maltose utilization | + |
| Ribitol utilization | - |
| Raffinose utilization | - |
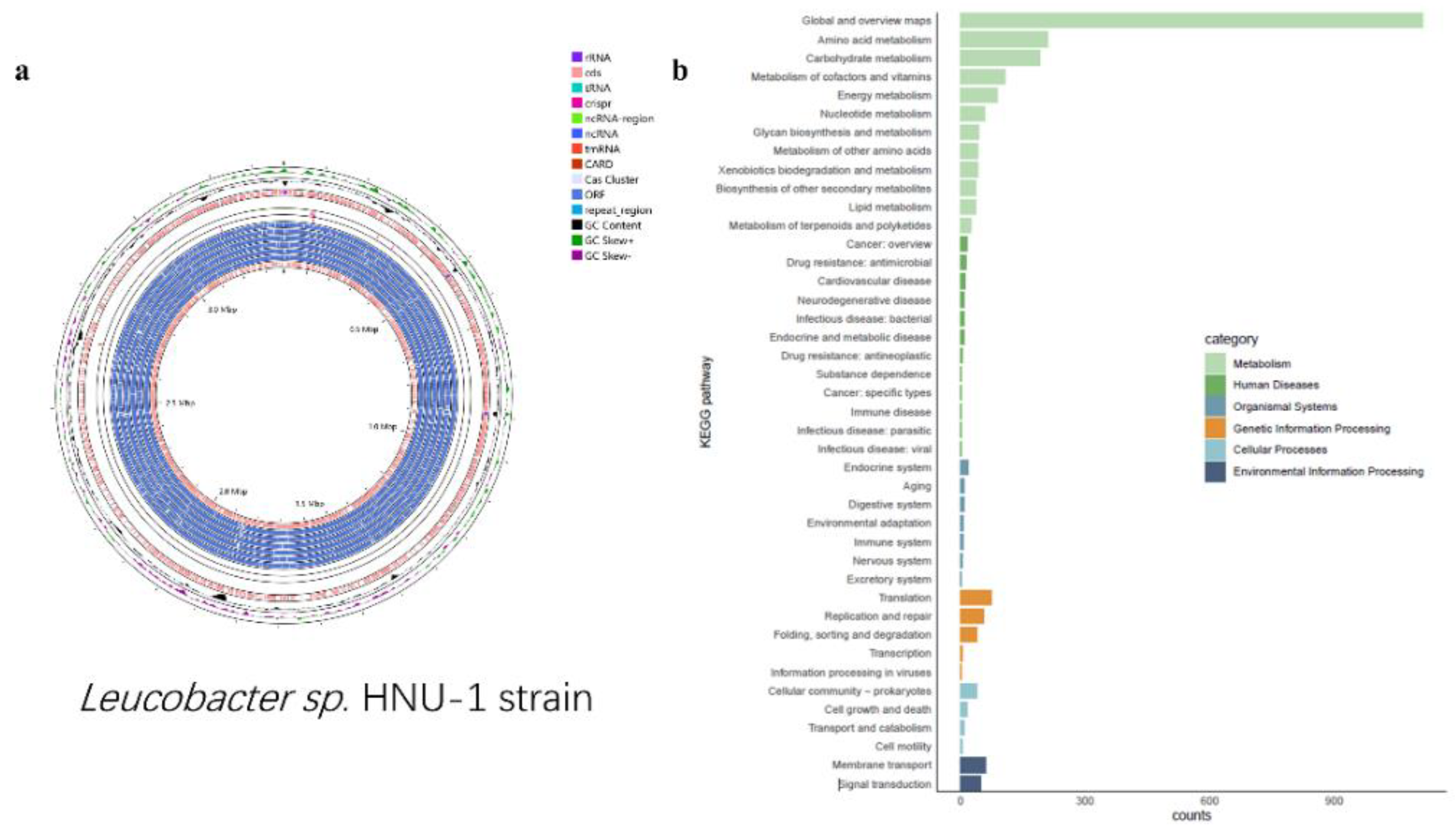
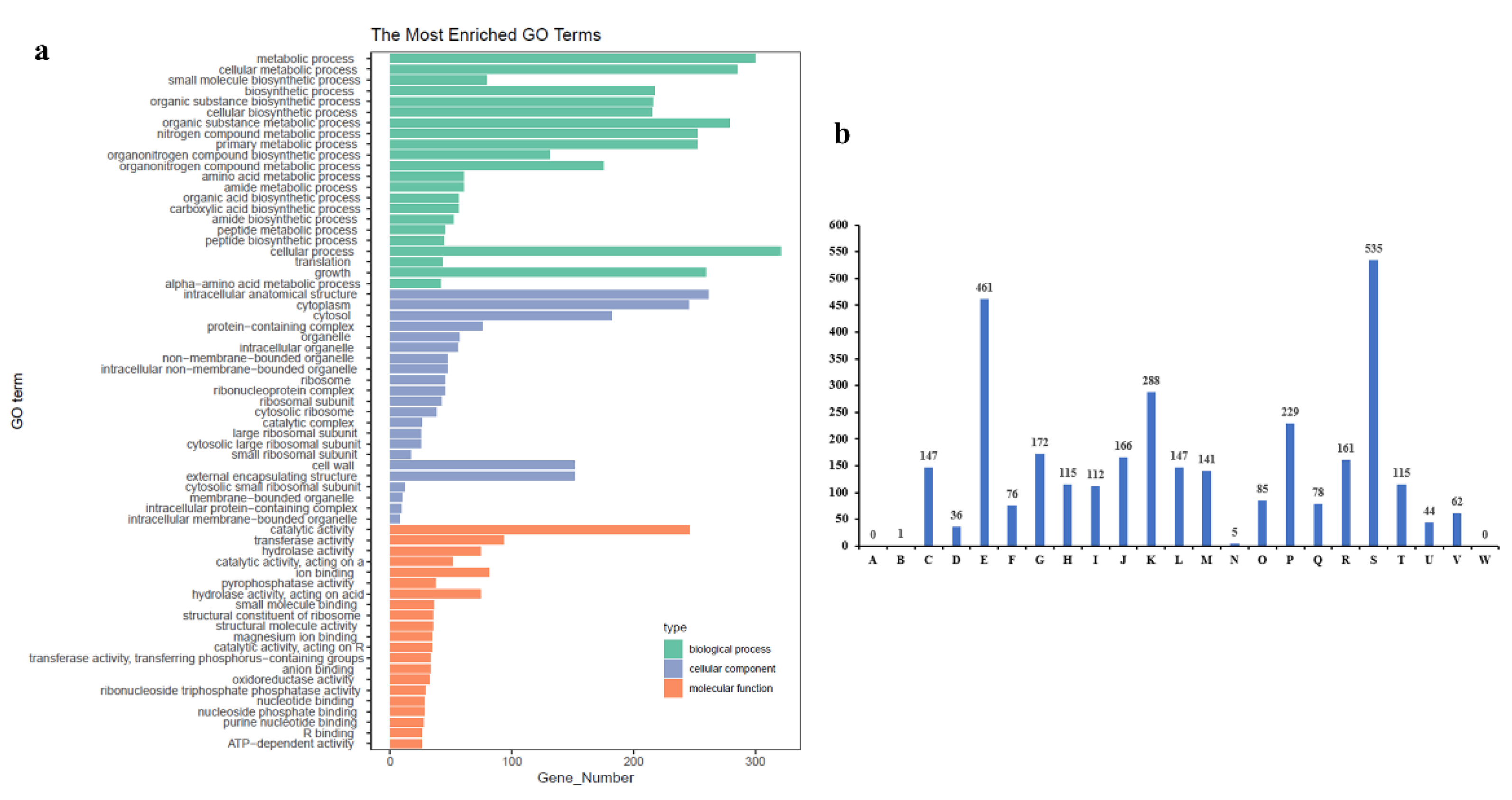
2.4. 16S Sequencing, Genome Assembly, and KEGG, GO, and COG Pathway Annotation of HNU-1
2.5. Carbohydrate-Active Enzymes Pathway Annotation of HNU-1
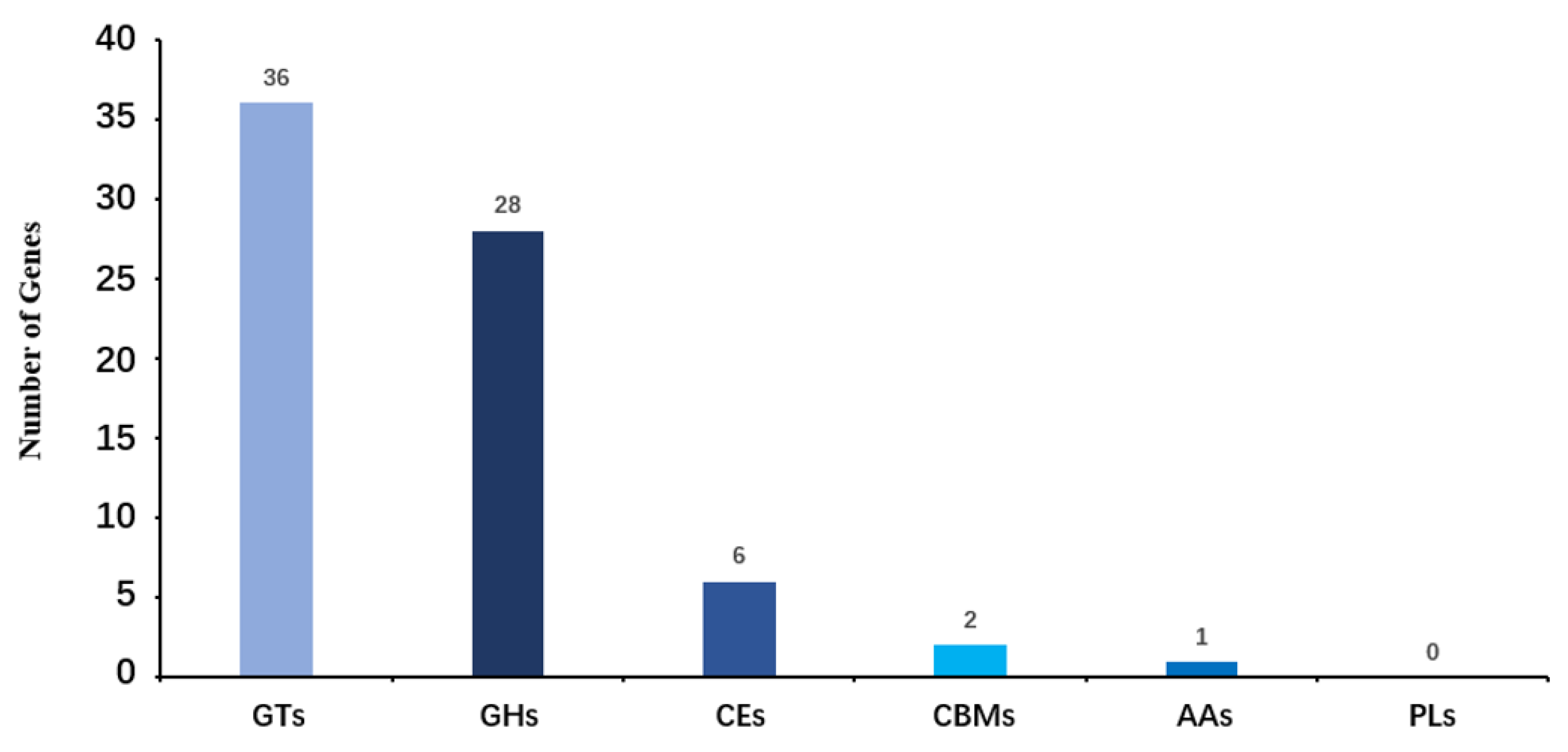
2.6. CARD Antibiotic Resistance Prediction and K-B Method Antimicrobial Susceptibility Testing of HNU-1
2.7. Construction of the Phylogenetic Tree of HNU-1
2.8. Comparative Genomics of HNU-1 and Six Leucobacter Species, and ANI Analysis of All Leucobacter Genomes Available on NCBI
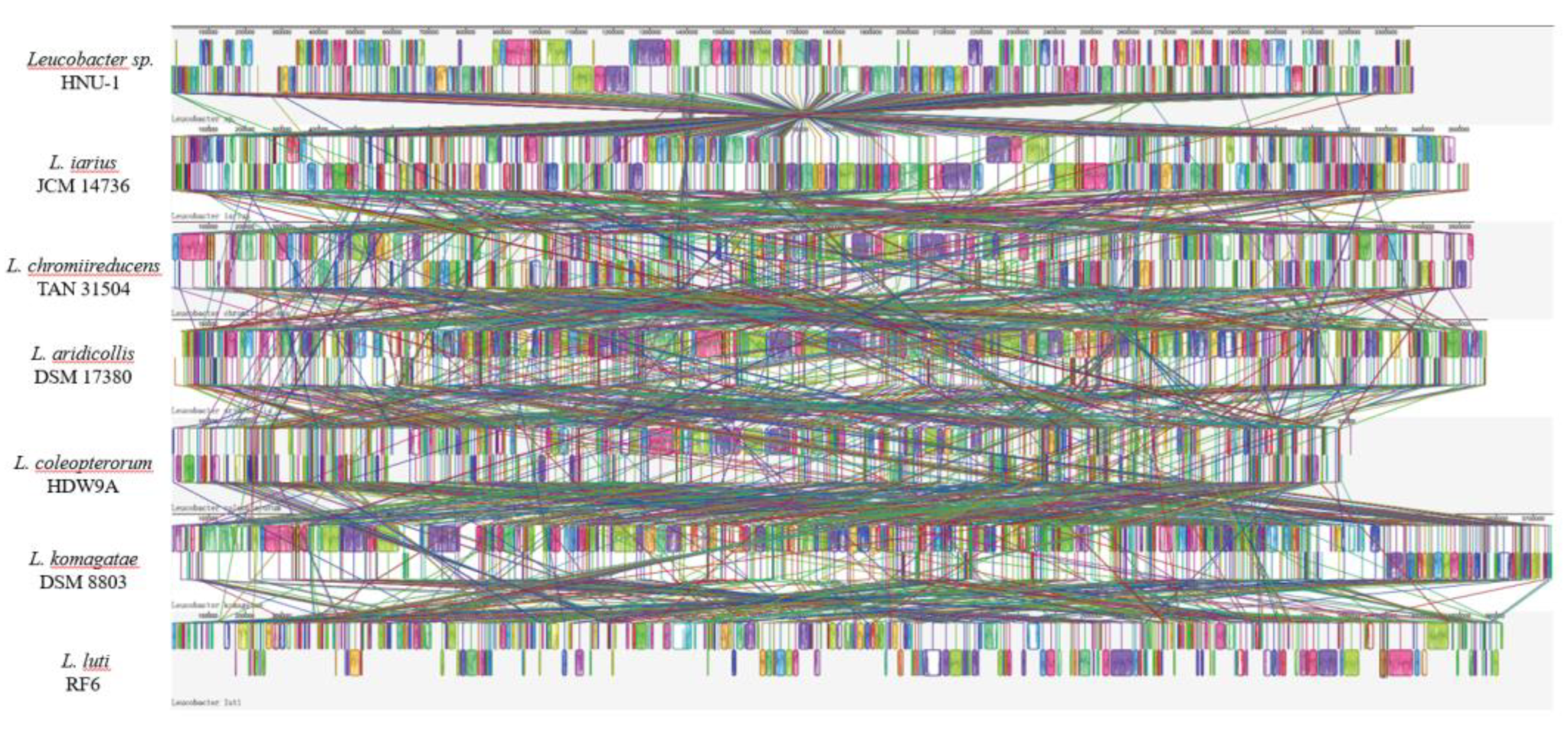
2.9. Bacterial Synteny Analysis of HNU-1 and Six Leucobacter Species

3. Discussion
4. Materials and Methods
4.1. Isolation of the HNU-1 and Extraction of Genomic DNA
4.2. Characterization and Molecular Identification of Leucobacter sp. HNU-1
4.3. C. elegans Strains and Maintenance
4.4. Preparation of Leucobacter sp. (HNU-1) and OP50 Mixed Suspension
4.5. Lifespan Observation of C. elegans
4.6. Analysis of the Fluorescence Intensity in Worms
4.7. Complete Genome Sequencing and Annotation
4.9. Biochemical Identification of the Isolated Strain Using 16 Biochemical Tests
4.10. Prediction of Antibiotic Synthesis Gene Clusters and Analysis of Associated Genes
4.11. Antibiotic Susceptibility Testing Using the K-B Disk Diffusion
4.12. Complete Nucleotide Sequence and Strain Accession Numbers
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- TAKEUCHI, M.; WEISS, N.; SCHUMANN, P.; YOKOTA, A.; TAKEUCHI, M.; WEISS, N.; SCHUMANN, P.; YOKOTA, A., Leucobacter komagatae gen. nov., sp. nov., a New Aerobic Gram-Positive, Nonsporulating Rod with 2,4-Diaminobutyric Acid in the Cell Wall. International Journal of Systematic and Evolutionary Microbiology 1996/10/01, 46, (4).
- Bates, K. A.; King, K. C. , <em>Leucobacter</em>. Trends in Microbiology 2021, 29, 1046–1047. [Google Scholar] [PubMed]
- Tan, M.-W.; Rahme, L. G.; Sternberg, J. A.; Tompkins, R. G.; Ausubel, F. M. , Pseudomonas aeruginosa killing of Caenorhabditis elegans used to identify P. aeruginosa virulence factors. Proceedings of the National Academy of Sciences 1999, 96, 2408–2413. [Google Scholar]
- Gao, H.; Qi, G.; Yin, R.; Zhang, H.; Li, C.; Zhao, X. , Bacillus cereus strain S2 shows high nematicidal activity against Meloidogyne incognita by producing sphingosine. Scientific Reports 2016, 6, 28756. [Google Scholar]
- Dinić, M.; Jakovljević, S.; Đokić, J.; Popović, N.; Radojević, D.; Strahinić, I.; Golić, N. , Probiotic-mediated p38 MAPK immune signaling prolongs the survival of Caenorhabditis elegans exposed to pathogenic bacteria. Scientific Reports 2021, 11, 21258. [Google Scholar]
- Lee, S. S.; Kennedy, S.; Tolonen, A. C.; Ruvkun, G. DAF-16 Target Genes That Control C. elegans Life-Span and Metabolism. Science 2003, 300, 644-647.
- Lin, Y.-C.; Uemori, K.; de Briel, D. A.; Arunpairojana, V.; Yokota, A. Zimmermannella helvola gen. nov., sp. nov., Zimmermannella alba sp. nov., Zimmermannella bifida sp. nov., Zimmermannella faecalis sp. nov. and Leucobacter albus sp. nov., novel members of the family Microbacteriaceae. International Journal of Systematic and Evolutionary Microbiology 2004, 54, 1669-1676.
- Morais, P. V.; Francisco, R.; Branco, R.; Chung, A. P.; da Costa, M. S. Leucobacter chromiireducens sp. nov, and Leucobacter aridicollis sp. nov., Two New Species Isolated From a Chromium Contaminated Environment. Systematic and Applied Microbiology 2004, 27, 646-652.
- Morais, P. V.; Paulo, C.; Francisco, R.; Branco, R.; Paula Chung, A.; da Costa, M. S. Leucobacter luti sp. nov., and Leucobacter alluvii sp. nov., two new species of the genus Leucobacter isolated under chromium stress. Systematic and Applied Microbiology 2006, 29, 414-421.
- Shin, N.-R.; Kim, M.-S.; Jung, M.-J.; Roh, S. W.; Nam, Y.-D.; Park, E.-J.; Bae, J.-W. Leucobacter celer sp. nov., isolated from Korean fermented seafood. International Journal of Systematic and Evolutionary Microbiology 2011, 61, 2353-2357.
- Yun, J.-H.; Roh, S. W.; Kim, M.-S.; Jung, M.-J.; Park, E.-J.; Shin, K.-S.; Nam, Y.-D.; Bae, J.-W. Leucobacter salsicius sp. nov., from a salt-fermented food. International Journal of Systematic and Evolutionary Microbiology 2011, 61, 502-506.
- Ue, H., Leucobacter exalbidus sp. nov., an actinobacterium isolated from a mixed culture from compost. The Journal of General and Applied Microbiology 2011, 57, 27-33.
- Behrendt, U.; Ulrich, A.; Schumann, P.Leucobacter tardus sp. nov., isolated from the phyllosphere of Solanum tuberosum L. International Journal of Systematic and Evolutionary Microbiology 2008, 58, 2574-2578.
- Her, J.; Lee, S.-S. Leucobacterhumi sp. nov., Isolated from Forest Soil. Current Microbiology 2015, 71, 235-242.
- Somvanshi, V. S.; Lang, E.; Schumann, P.; Pukall, R.; Kroppenstedt, R. M.; Ganguly, S.; Stackebrandt, E. Leucobacter iarius sp. nov., in the family Microbacteriaceae. International Journal of Systematic and Evolutionary Microbiology 2007, 57, 682-686.
- Muir, R. E.; Tan, M.-W. Leucobacter chromiireducens subsp. solipictus subsp. nov., a pigmented bacterium isolated from the nematode Caenorhabditis elegans, and emended description of L. chromiireducens. International Journal of Systematic and Evolutionary Microbiology 2007, 57, 2770-2776.
- Percudani, R. A Microbial Metagenome (Leucobacter sp.) in Caenorhabditis Whole Genome Sequences. Bioinformatics and Biology Insights 2013, 7, BBI.S11064.
- Hekimi, S.; Lakowski, B.; Barnes, T. M.; Ewbank, J. J. , Molecular genetics of life span in <em>C. elegans</em>: How much does it teach us? Trends in Genetics 1998, 14, 14–20. [Google Scholar]
- Park, H.-E. H.; Jung, Y.; Lee, S.-J. V. , Survival assays using Caenorhabditis elegans. Molecules and Cells 2017, 40, 90–99. [Google Scholar] [PubMed]
- Franco-Romero, A.; Morbidoni, V.; Milan, G.; Sartori, R.; Wulff, J.; Romanello, V.; Armani, A.; Salviati, L.; Conte, M.; Salvioli, S.; Franceschi, C.; Buonomo, V.; Swoboda, C. O.; Grumati, P.; Pannone, L.; Martinelli, S.; Jefferies, H. B. J.; Dikic, I.; van der Laan, J.; Cabreiro, F.; Millay, D. P.; Tooze, S. A.; Trevisson, E.; Sandri, M. C16ORF70/MYTHO promotes healthy aging in C.elegans and prevents cellular senescence in mammals. The Journal of Clinical Investigation 2024, 134, (15).
- Melentijevic, I.; Toth, M. L.; Arnold, M. L.; Guasp, R. J.; Harinath, G.; Nguyen, K. C.; Taub, D.; Parker, J. A.; Neri, C.; Gabel, C. V.; Hall, D. H.; Driscoll, M., C. elegans neurons jettison protein aggregates and mitochondria under neurotoxic stress. Nature 2017, 542, 367–371. [Google Scholar]
- O’Donnell, M. P.; Fox, B. W.; Chao, P.-H.; Schroeder, F. C.; Sengupta, P. , A neurotransmitter produced by gut bacteria modulates host sensory behaviour. Nature 2020, 583, 415–420. [Google Scholar]
- Ortiz, A.; Vega, N. M.; Ratzke, C.; Gore, J. , Interspecies bacterial competition regulates community assembly in the C. elegans intestine. The ISME Journal 2021, 15, 2131–2145. [Google Scholar]
- Liu, H.; Chen, P.; Yang, X.; Hao, F.; Tian, G.; Shan, Z.; Qi, B. Probiotics-sensing mechanism in neurons that initiates gut mitochondrial surveillance for pathogen defense. Cell Reports 2024, 43, (12).
- Wang, C.; Yang, M.; Liu, D.; Zheng, C. Metabolic rescue of α-synuclein-induced neurodegeneration through propionate supplementation and intestine-neuron signaling in C. elegans. Cell Reports 2024, 43, (3).
- Pujol, N.; Cypowyj, S.; Ziegler, K.; Millet, A.; Astrain, A.; Goncharov, A.; Jin, Y.; Chisholm, A. D.; Ewbank, J. J. , Distinct Innate Immune Responses to Infection and Wounding in the C. elegans Epidermis. Current Biology 2008, 18, 481–489. [Google Scholar]
- Ahmed, S. K.; Hussein, S.; Qurbani, K.; Ibrahim, R. H.; Fareeq, A.; Mahmood, K. A.; Mohamed, M. G. Antimicrobial resistance: Impacts, challenges, and future prospects. Journal of Medicine, Surgery, and Public Health 2024, 2, 100081.
- Kanehisa, M.; Goto, S. , KEGG: Kyoto Encyclopedia of Genes and Genomes. Nucleic Acids Research 2000, 28, 27–30. [Google Scholar] [CrossRef]
- Ashburner, M.; Ball, C. A.; Blake, J. A.; Botstein, D.; Butler, H.; Cherry, J. M.; Davis, A. P.; Dolinski, K.; Dwight, S. S.; Eppig, J. T.; Harris, M. A.; Hill, D. P.; Issel-Tarver, L.; Kasarskis, A.; Lewis, S.; Matese, J. C.; Richardson, J. E.; Ringwald, M.; Rubin, G. M.; Sherlock, G. Gene Ontology: tool for the unification of biology. Nature Genetics 2000, 25, 25–29. [Google Scholar]
- Galperin, M. Y.; Wolf, Y. I.; Makarova, K. S.; Vera Alvarez, R.; Landsman, D.; Koonin, E. V. , COG database update: focus on microbial diversity, model organisms, and widespread pathogens. Nucleic Acids Research 2021, (D1), D274–D281. [Google Scholar]
- Cantarel, B. L.; Coutinho, P. M.; Rancurel, C.; Bernard, T.; Lombard, V.; Henrissat, B. The Carbohydrate-Active EnZymes database (CAZy): an expert resource for Glycogenomics. Nucleic Acids Research 2009, 37, (suppl_1), D233-D238.
- Jia, B.; Raphenya, A. R.; Alcock, B.; Waglechner, N.; Guo, P.; Tsang, K. K.; Lago, B. A.; Dave, B. M.; Pereira, S.; Sharma, A. N.; Doshi, S.; Courtot, M.; Lo, R.; Williams, L. E.; Frye, J. G.; Elsayegh, T.; Sardar, D.; Westman, E. L.; Pawlowski, A. C.; Johnson, T. A.; Brinkman, F. S. L.; Wright, G. D.; McArthur, A. G. , CARD 2017: expansion and model-centric curation of the comprehensive antibiotic resistance database. Nucleic Acids Research 2017, (D1), D566–D573. [Google Scholar]
- Saitou, N.; Nei, M. , The neighbor-joining method: a new method for reconstructing phylogenetic trees. Molecular Biology and Evolution 1987, 4, 406–425. [Google Scholar] [PubMed]
- Nei, M.; Kumar, S. , Molecular Evolution and Phylogenetics. In Oxford University Press: 2000.
- Rodriguez-R, L. M.; Konstantinidis, K. T. , Estimating coverage in metagenomic data sets and why it matters. The ISME Journal 2014, 8, 2349–2351. [Google Scholar] [CrossRef]
- Goris, J.; Konstantinidis, K. T.; Klappenbach, J. A.; Coenye, T.; Vandamme, P.; Tiedje, J. M. , DNA–DNA hybridization values and their relationship to whole-genome sequence similarities. International Journal of Systematic and Evolutionary Microbiology 2007, 57, 81–91. [Google Scholar]
- Darling, A. C. E.; Mau, B.; Blattner, F. R.; Perna, N. T. , Mauve: Multiple Alignment of Conserved Genomic Sequence With Rearrangements. Genome Research 2004, 14, 1394–1403. [Google Scholar]
- Lee, D.; Zdraljevic, S.; Cook, D. E.; Frézal, L.; Hsu, J.-C.; Sterken, M. G.; Riksen, J. A. G.; Wang, J.; Kammenga, J. E.; Braendle, C.; Félix, M.-A.; Schroeder, F. C.; Andersen, E. C. Selection and gene flow shape niche-associated variation in pheromone response. Nature Ecology & Evolution 2019, 3, 1455-1463.
- Kang, W. K.; Florman, J. T.; Araya, A.; Fox, B. W.; Thackeray, A.; Schroeder, F. C.; Walhout, A. J. M.; Alkema, M. J. , Vitamin B12 produced by gut bacteria modulates cholinergic signalling. Nature Cell Biology 2024, 26, 72–85. [Google Scholar]
- Mallick, S.; Mishra, N.; Barik, B. K.; Negi, V. D. , <em>Salmonella</em> Typhimurium <em>fepB</em> negatively regulates <em>C. elegans</em> behavioral plasticity. Journal of Infection 2022, 84, 518–530. [Google Scholar]
- Amrit, F. R. G.; Ratnappan, R.; Keith, S. A.; Ghazi, A. , The C. elegans lifespan assay toolkit. Methods 2014, 68, 465–475. [Google Scholar] [PubMed]
- Naval-Sanchez, M.; Deshpande, N.; Tran, M.; Zhang, J.; Alhomrani, M.; Alsanie, W.; Nguyen, Q.; Nefzger, C. M. Benchmarking of ATAC Sequencing Data From BGI’s Low-Cost DNBSEQ-G400 Instrument for Identification of Open and Occupied Chromatin Regions. Frontiers in Molecular Biosciences 2022, 9.
- Deamer, D.; Akeson, M.; Branton, D. , Three decades of nanopore sequencing. Nature Biotechnology 2016, 34, 518–524. [Google Scholar] [PubMed]
- Kolmogorov, M.; Yuan, J.; Lin, Y.; Pevzner, P. A. , Assembly of long, error-prone reads using repeat graphs. Nature Biotechnology 2019, 37, 540–546. [Google Scholar]
- Cosma, B.-M.; Shirali Hossein Zade, R.; Jordan, E. N.; van Lent, P.; Peng, C.; Pillay, S.; Abeel, T. , Evaluating long-read de novo assembly tools for eukaryotic genomes: insights and considerations. GigaScience 2023, 12, giad100. [Google Scholar]
- Walker, B. J.; Abeel, T.; Shea, T.; Priest, M.; Abouelliel, A.; Sakthikumar, S.; Cuomo, C. A.; Zeng, Q.; Wortman, J.; Young, S. K.; Earl, A. M. , Pilon: An Integrated Tool for Comprehensive Microbial Variant Detection and Genome Assembly Improvement. PLOS ONE 2014, 9, e112963. [Google Scholar]
- Delcher, A. L.; Bratke, K. A.; Powers, E. C.; Salzberg, S. L. , Identifying bacterial genes and endosymbiont DNA with Glimmer. Bioinformatics 2007, 23, 673–679. [Google Scholar]
- Lowe, T. M.; Eddy, S. R. , tRNAscan-SE: A Program for Improved Detection of Transfer RNA Genes in Genomic Sequence. Nucleic Acids Research 1997, 25, 955–964. [Google Scholar]
- Lagesen, K.; Hallin, P.; Rødland, E. A.; Stærfeldt, H.-H.; Rognes, T.; Ussery, D. W. , RNAmmer: consistent and rapid annotation of ribosomal RNA genes. Nucleic Acids Research 2007, 35, 3100–3108. [Google Scholar]
- Chen, C.; Wu, Y.; Xia, R. , A painless way to customize Circos plot: From data preparation to visualization using TBtools. iMeta 2022, 1, e35. [Google Scholar]
- Richter, M.; Rosselló-Móra, R. , Shifting the genomic gold standard for the prokaryotic species definition. Proceedings of the National Academy of Sciences 2009, 106, 19126–19131. [Google Scholar]
- Wayne, L. G.; Brenner, D. J.; Colwell, R. R.; Grimont, P. A. D.; Kandler, O.; Krichevsky, M. I.; Moore, L. H.; Moore, W. E. C.; Murray, R. G. E.; Stackebrandt, E.; Starr, M. P.; Truper, H. G. , Report of the Ad Hoc Committee on Reconciliation of Approaches to Bacterial Systematics. International Journal of Systematic and Evolutionary Microbiology 1987, 37, 463–464. [Google Scholar]
- Kook, J.-K.; Park, S.-N.; Lim, Y. K.; Cho, E.; Jo, E.; Roh, H.; Shin, Y.; Paek, J.; Kim, H.-S.; Kim, H.; Shin, J. H.; Chang, Y.-H. , Genome-Based Reclassification of Fusobacterium nucleatum Subspecies at the Species Level. Current Microbiology 2017, 74, 1137–1147. [Google Scholar] [PubMed]
- McArthur Andrew, G.; Waglechner, N.; Nizam, F.; Yan, A.; Azad Marisa, A.; Baylay Alison, J.; Bhullar, K.; Canova Marc, J.; De Pascale, G.; Ejim, L.; Kalan, L.; King Andrew, M.; Koteva, K.; Morar, M.; Mulvey Michael, R.; O'Brien Jonathan, S.; Pawlowski Andrew, C.; Piddock Laura, J. V.; Spanogiannopoulos, P.; Sutherland Arlene, D.; Tang, I.; Taylor Patricia, L.; Thaker, M.; Wang, W.; Yan, M.; Yu, T.; Wright Gerard, D. , The Comprehensive Antibiotic Resistance Database. Antimicrobial Agents and Chemotherapy 2013, 57, 3348–3357. [Google Scholar] [CrossRef]

| Antibiotic | Start | Stop | AMR Gene Family | Best Identities |
Orientation |
| vancomycin | 418,532 | 419,158 | vanY;glycolpeptide resistance gene cluster | 31.71 | + |
| vancomycin | 659,487 | 660,260 | vanY;glycopeptide resistance gene cluster | 38.93 | + |
| sulfadiazine | 2,675,593 | 2,676,432 | sulfonamide resistant sul | 100 | - |
| spectinomycin | 2,676,937 | 2,677,716 | ANT(3'') | 100 | - |
| defensin | 1,174,301 | 1,174,543 | defensin resistant mprF | 100 | - |
| Category | Concentration/disc | Interpretation Standard (mm) | Diameter (mm) | Result | ||
| Resistant (R) | Intermediate (I) | Sensitive (S) | ||||
| Penicillin | 10U | ≤10 | 11-16 | ≥17 | 26.74 | S |
| Vancomycin | 30µg | ≤14 | 15-16 | ≥17 | 15.9 | I |
| Oxacillin | 1µg | ≤10 | 11-12 | ≥13 | 3 | R |
| Levofloxacin | 5µg | ≤13 | 14-17 | ≥18 | 18.26 | S |
| Clindamycin | 2µg | ≤14 | 15-19 | ≥20 | 12.1 | R |
| Erythromycin | 15µg | ≤13 | 14-22 | ≥23 | 15.22 | I |
| Polymyxin B | 300IU | ≤8 | 9-11 | ≥12 | 6.42 | R |
| Gentamicin | 10µg | ≤12 | 13-14 | ≥15 | 18.72 | S |
| Lincomycin | 2µg | ≤14 | 15-20 | ≥21 | 3 | R |
| Minocycline | 30µg | ≤15 | 16-18 | ≥19 | 32.16 | S |
| Tetracycline | 30µg | ≤11 | 12-14 | ≥15 | 12.26 | I |
| Chloramphenicol | 30µg | ≤12 | 13-17 | ≥18 | 29.56 | S |
| Imipenem | 10µg | ≤19 | 20-22 | ≥23 | 17.06 | R |
| Doxycycline | 30µg | ≤12 | 13-15 | ≥16 | 17.8 | S |
| Azithromycin | 15µg | ≤17 | 18-19 | ≥20 | 25.48 | S |
| Ceftriaxone | 30µg | ≤19 | 20-22 | ≥23 | 18.82 | R |
| Ceftazidime | 30µg | ≤14 | 15-17 | ≥18 | 19.72 | S |
| Cefoperazone | 75µg | ≤22 | 23-25 | ≥26 | 8 | R |
| Ciprofloxacin | 5µg | ≤15 | 16-20 | ≥21 | 12 | R |
| Norfloxacin | 10µg | ≤12 | 13-16 | ≥17 | 6.44 | R |
| Florfenicol | 30µg | ≤12 | 13-17 | ≥18 | 31.56 | S |
| Piperacillin | 100µg | ≤17 | 18-20 | ≥21 | 29.32 | S |
| Streptomycin | 10µg | ≤6 | 7-9 | ≥10 | 3 | R |
| Compound- Sulfonamides |
25µg | ≤10 | 11-15 | ≥16 | 3 | R |
| Ampicillin | 10µg | ≤13 | 14-16 | ≥17 | 24.14 | S |
| Kanamycin | 30µg | ≤13 | 14-17 | ≥18 | 3 | R |
| Amikacin | 30µg | ≤14 | 15-16 | ≥17 | 17.98 | S |
| Cefuroxim | 30µg | ≤14 | 15-17 | ≥18 | 18.48 | S |
| Cephalexin | 30µg | ≤14 | 15-17 | ≥18 | 22.64 | S |
| Cefamezin | 30µg | ≤14 | 15-17 | ≥18 | 24.42 | S |
| Item | Leucobacter sp. HNU-1 | Leucobacter iarius JCM 14736 | Leucobacter chromiireducens TAN 31504 | Leucobacter aridicollis DSM 17380 | Leucobacter coleopterorum HDW9A | Leucobacter komagatae DSM 8803 | Leucobacter luti RF6 |
| GeneBank assembly | PRJNA1137138 | GCA_039530105.1 | GCA_016758195.1 | GCA_013409595.1 | GCA_011382985.1 | GCA_006716085.1 | GCA_004217175.1 |
| Length of chromosome /bp | 3,375,033 | 3,524,626 | 3,537,946 | 3,573,416 | 3,215,551 | 3,752,337 | 3,618,231 |
| GC content/% | 70.37 | 70.57 | 68.93 | 67.32 | 60.32 | 66.63 | 69.46 |
| Number of CDSs | 3,268 | 3,188 | 3,096 | 3,257 | 2,980 | 3,375 | 3,089 |
| Number of rRNAs | 9 | 3 | 3 | 9 | 6 | 9 | 5 |
| Number of tRNAs | 52 | 53 | 54 | 52 | 45 | 50 | 46 |
| Number of CRISPRS | 6 | 13 | 1 | 2 | 2 | 2 | 3 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).




