Submitted:
04 March 2025
Posted:
04 March 2025
You are already at the latest version
Abstract
Low-density DNA microarrays are pivotal in molecular diagnostics due to their cost-effectiveness, high sensitivity, and ease of use. However, reliable spot localization remains a challenge, primarily due to variations in spot positions resulting from printing and alignment inaccuracies, as well as the presence of image artifacts. Traditional intensity-based methods are often inadequate, particularly when target spots exhibit weak fluorescence signals. In this study, we propose a rapid spot localization method that combines spot template matching with point pattern matching, enhanced through vectorized programming and the use of square (box) templates. By replacing circular templates with box templates, separable filtering and moving average techniques can be applied, significantly reducing computational overhead without compromising detection performance. The proposed method not only overcomes the intensive computation associated with normalized cross-correlation but also improves overall efficiency, making it highly suitable for high-throughput and resource-constrained diagnostic applications. Validation was performed using images obtained from HPV genotyping of actual patient samples on a commercial DNA microarray, confirming the method’s practical applicability in a clinical setting.
Keywords:
1. Introduction
2. Materials and Methods
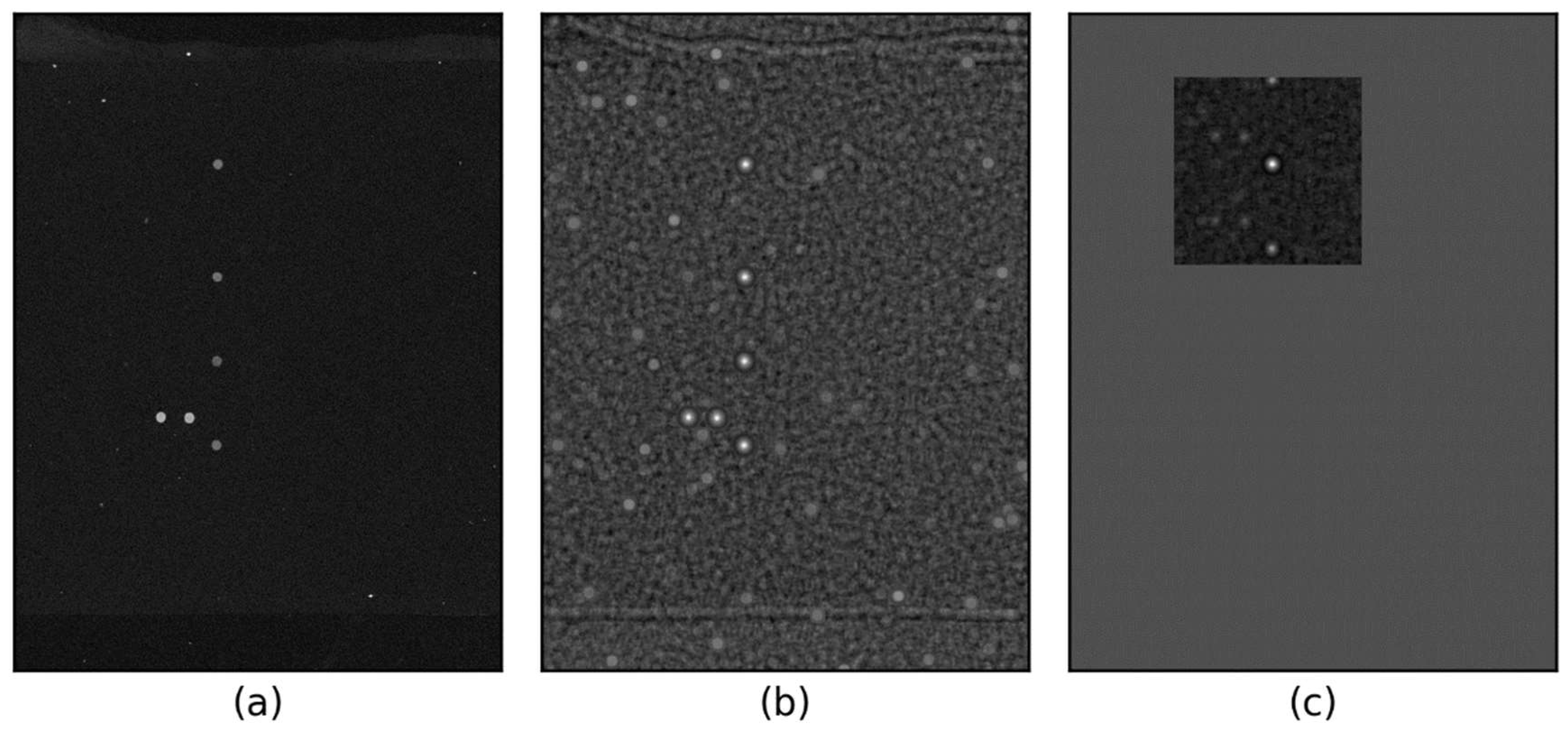
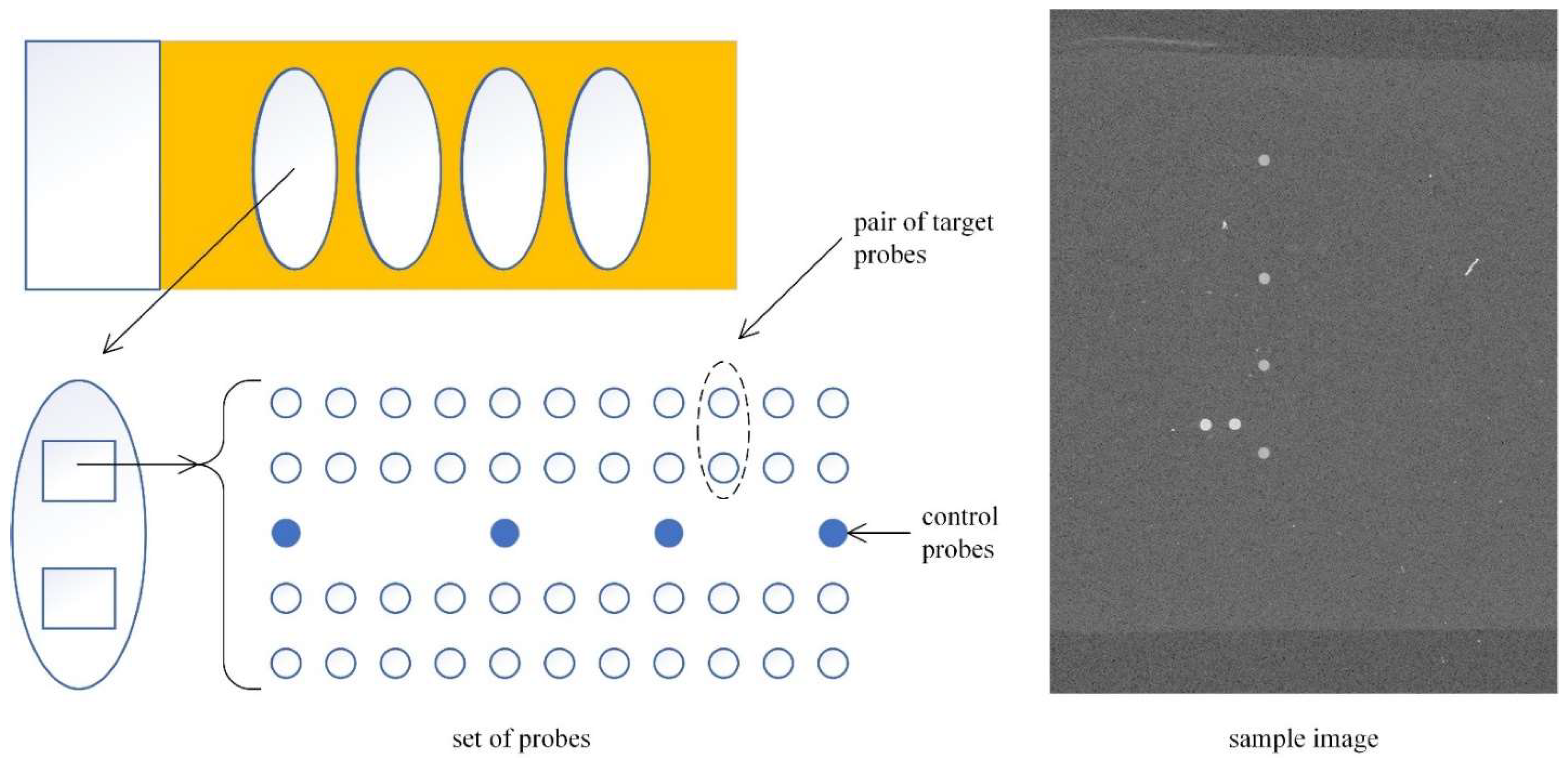
2.1. Experimental Images
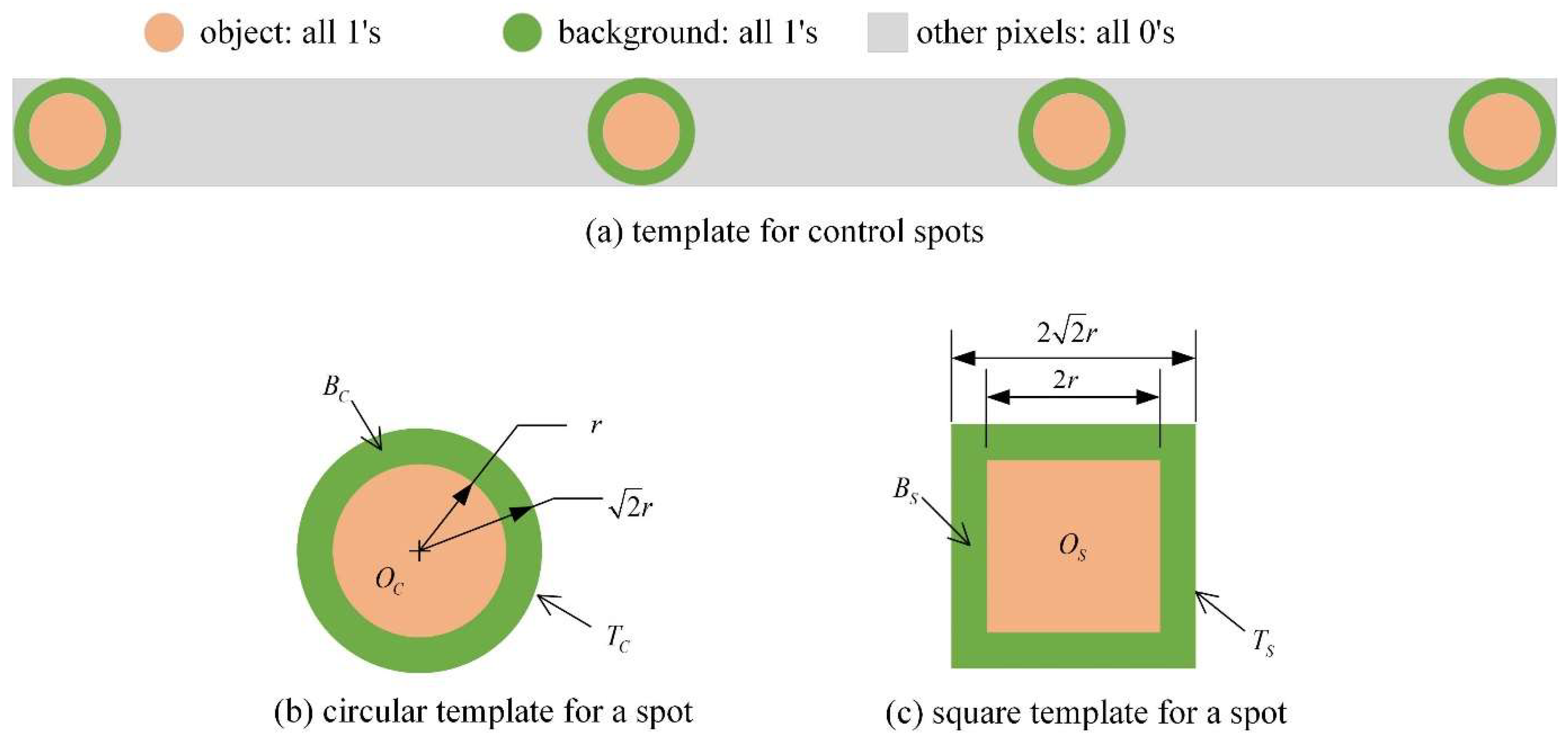
2.2. Template Matching
2.3. Spot Template Matching Response Calculation
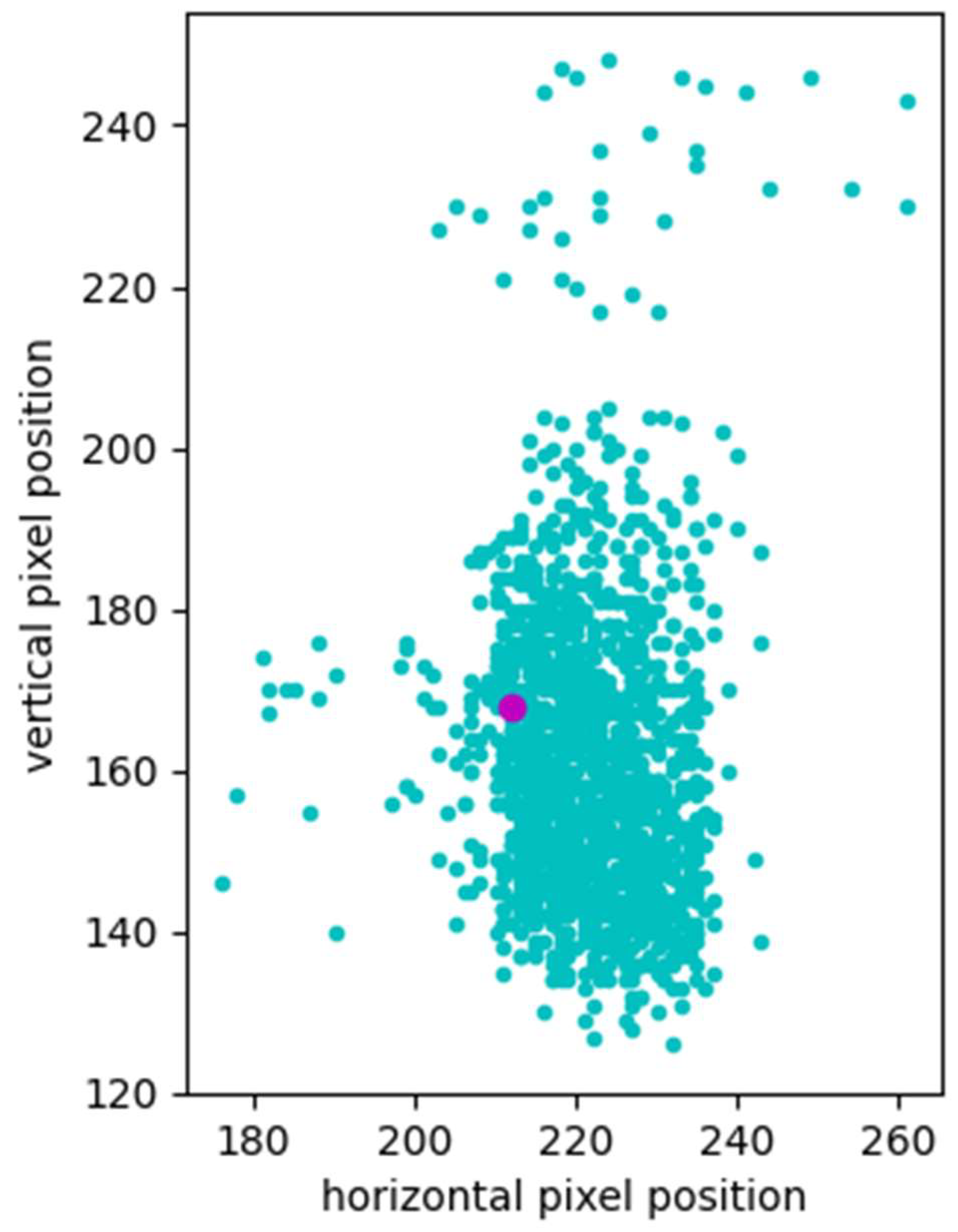
2.4. Verification of Control Spot Localization Performance
3. Results
3.1. Computation Time Analysis
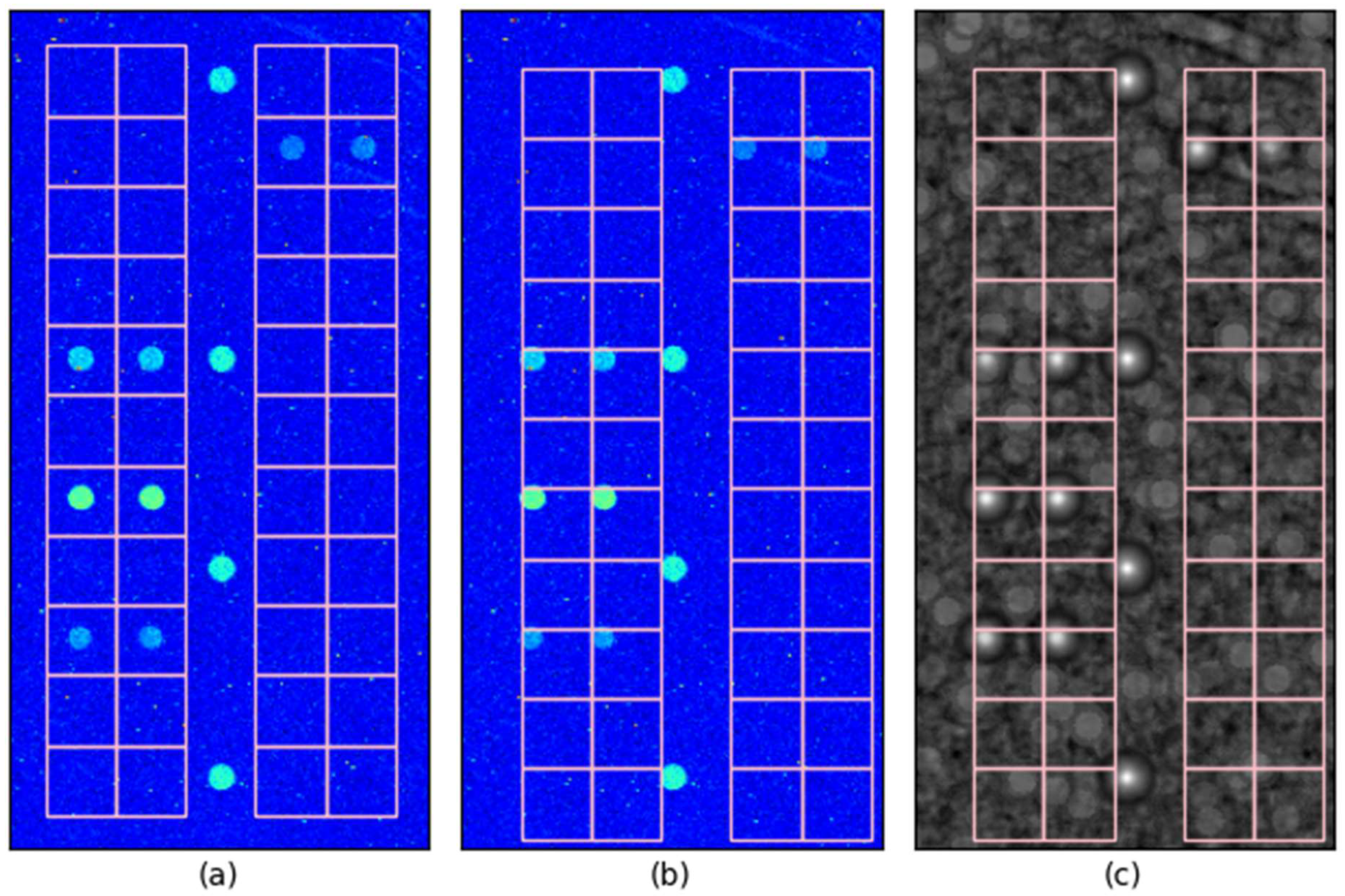
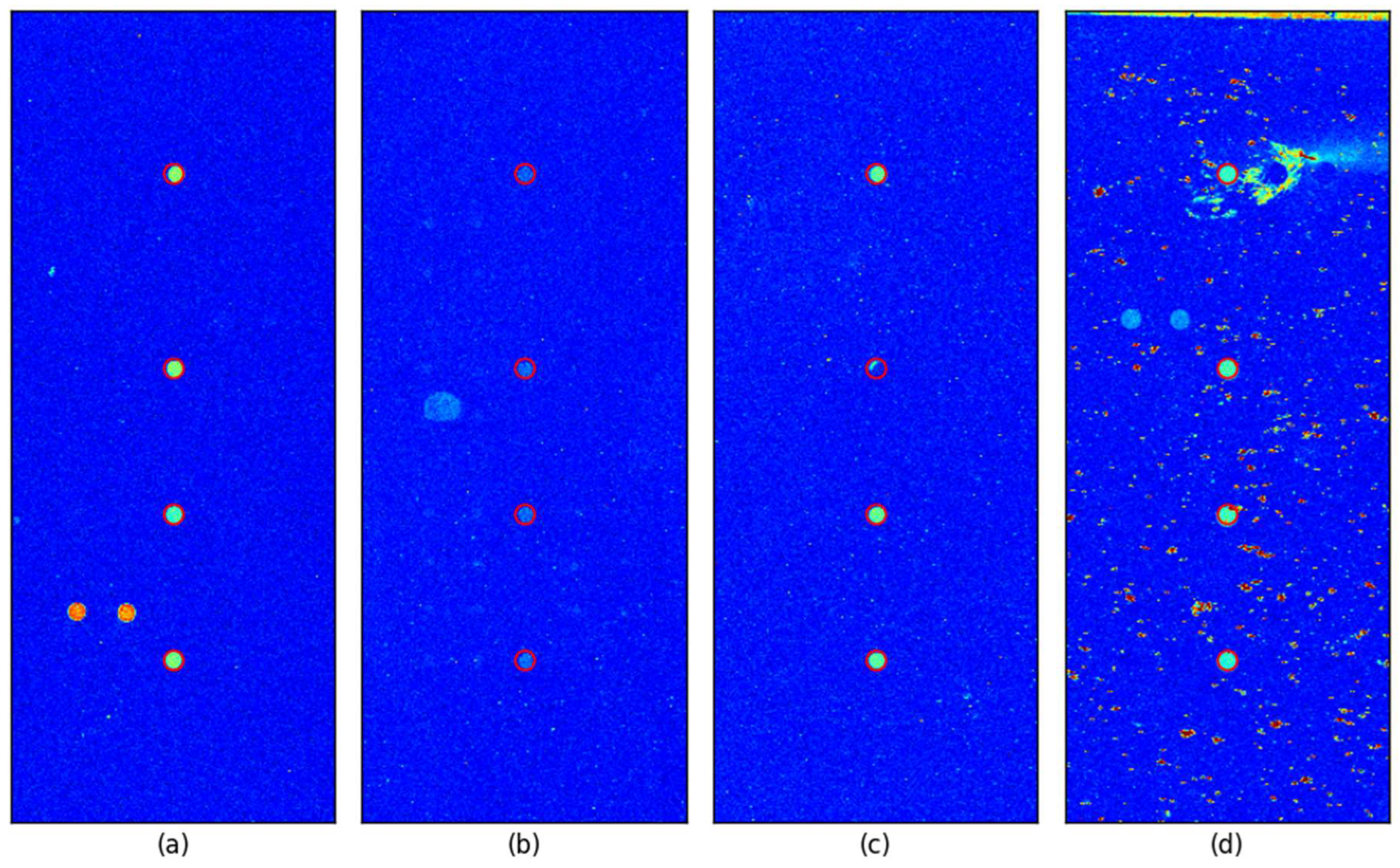
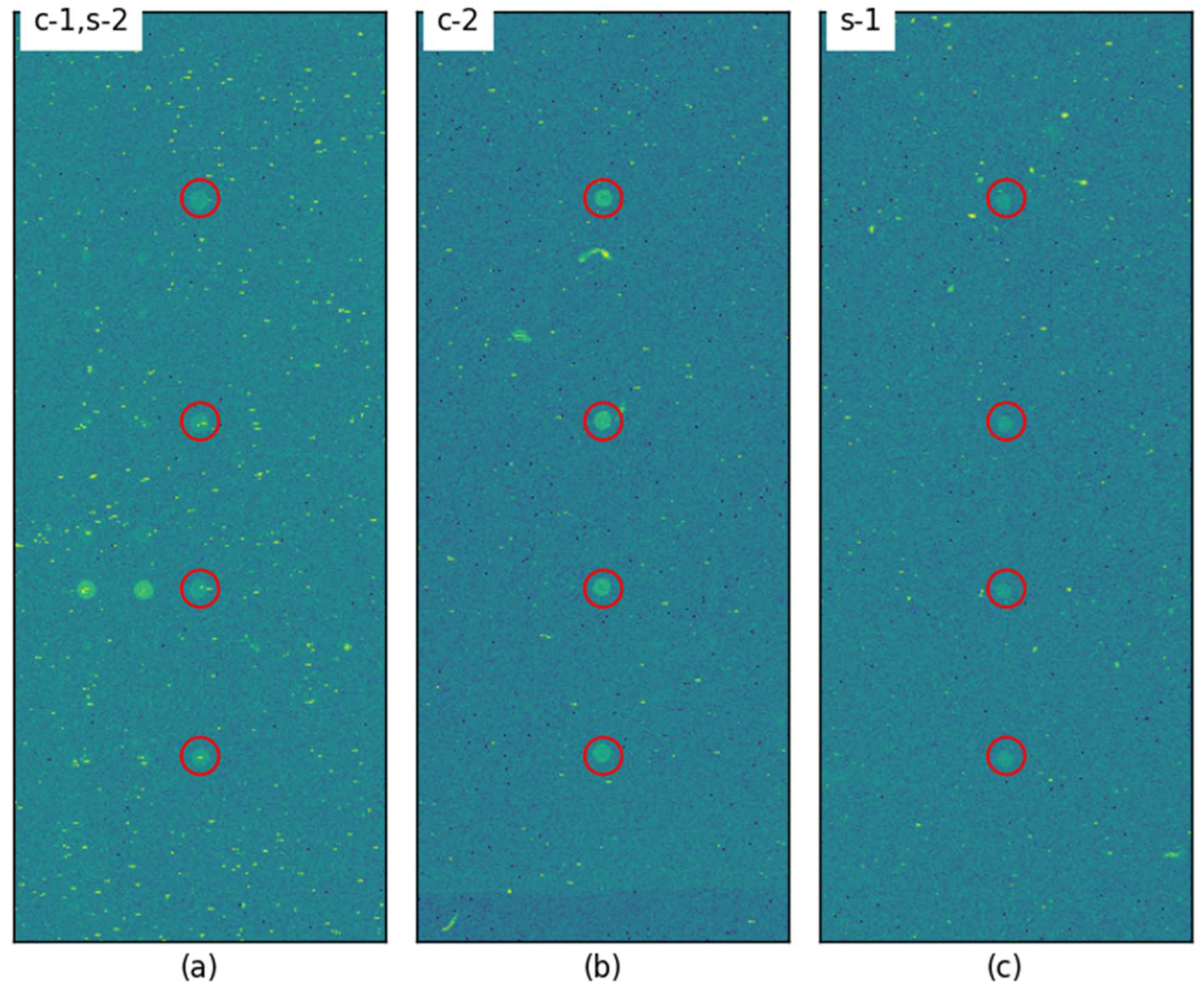
3.2. Visual Inspection of Spot Locating
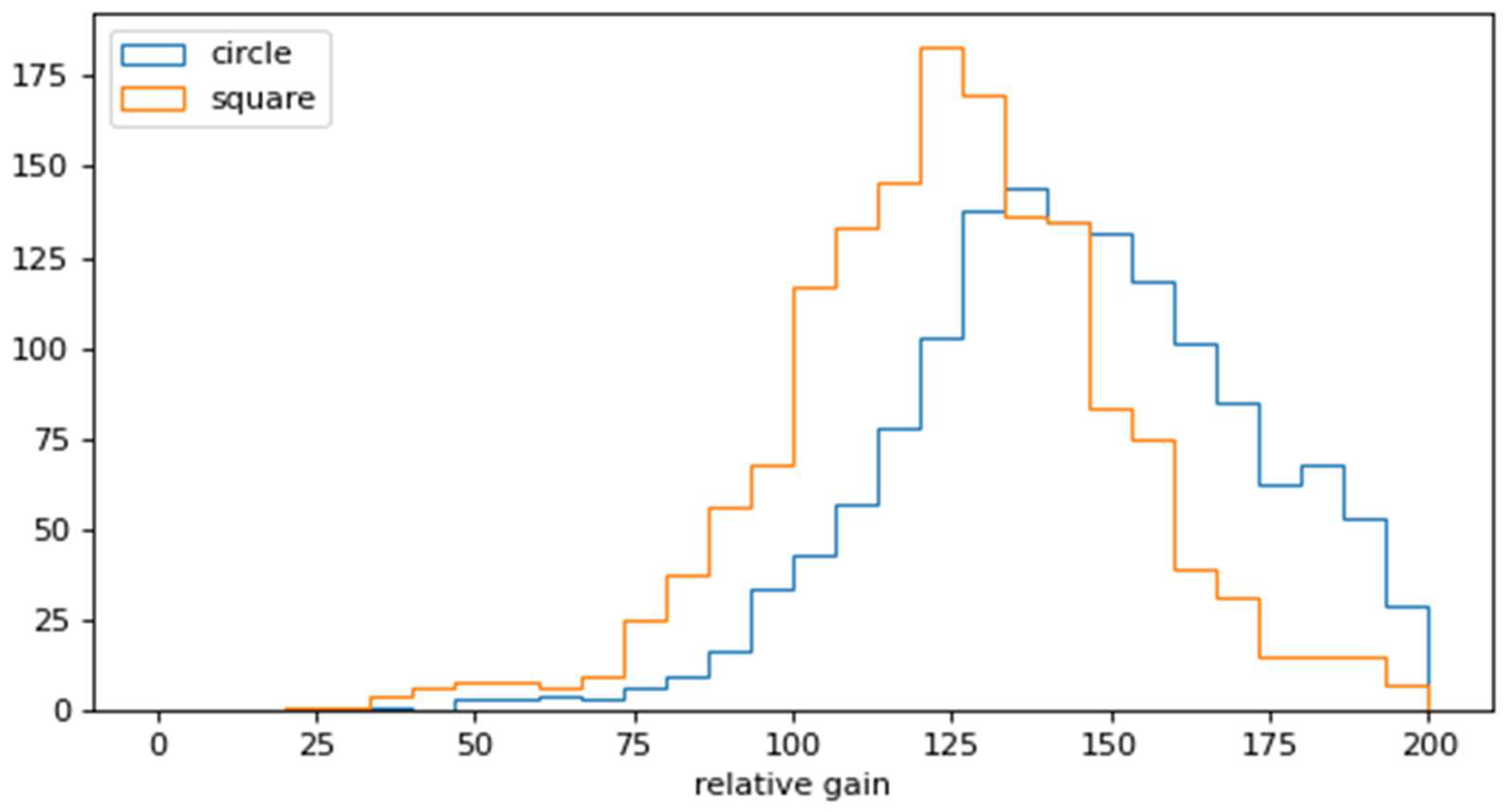
3.3. Optimal Template Size Selection
4. Discussion
5. Conclusions
Author Contributions
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Ikonnikova A, Morozova A, Antonova O, Ochneva A, Fedoseeva E, Abramova O, Emelyanova M, Filippova M, Morozova I, Zorkina Y, Syunyakov T, Andryushchenko A, Andreuyk D, Kostyuk G, Gryadunov D. Evaluation of the Polygenic Risk Score for Alzheimer’s Disease in Russian Patients with Dementia Using a Low-Density Hydrogel Oligonucleotide Microarray. International Journal of Molecular Sciences. 2023 Jan;24(19):14765.
- Das A, Santhosh S, Giridhar M, Behr J, Michel T, Schaudy E, Ibáñez-Redín G, Lietard J, Somoza MM. Dipodal Silanes Greatly Stabilize Glass Surface Functionalization for DNA Microarray Synthesis and High-Throughput Biological Assays. Anal Chem. 2023 Oct 17;95(41):15384–93.
- Shaskolskiy B, Kandinov I, Kravtsov D, Vinokurova A, Gorshkova S, Filippova M, Kubanov A, Solomka V, Deryabin D, Dementieva E, Gryadunov D. Hydrogel Droplet Microarray for Genotyping Antimicrobial Resistance Determinants in Neisseria gonorrhoeae Isolates. Polymers. 2021 Jan;13(22):3889.
- Waldmüller S, Freund P, Mauch S, Toder R, Vosberg HP. Low-density DNA microarrays are versatile tools to screen for known mutations in hypertrophic cardiomyopathy. Human Mutation. 2002;19(5):560–9.
- de Longueville F, Surry D, Meneses-Lorente G, Bertholet V, Talbot V, Evrard S, Chandelier N, Pike A, Worboys P, Rasson JP, Le Bourdellès B, Remacle J. Gene expression profiling of drug metabolism and toxicology markers using a low-density DNA microarray. Biochemical Pharmacology. 2002 Jul 1;64(1):137–49.
- Ma X, Li Y, Liang Y, Liu Y, Yu L, Li C, Liu Q, Chen L. Development of a DNA microarray assay for rapid detection of fifteen bacterial pathogens in pneumonia. BMC Microbiol. 2020 Dec;20(1):177.
- Abanda B, Paguem A, Achukwi MD, Renz A, Eisenbarth A. Development of a Low-Density DNA Microarray for Detecting Tick-Borne Bacterial and Piroplasmid Pathogens in African Cattle. Trop Med Infect Dis. 2019 Apr 12;4(2):64.
- Galbiati S, Damin F, Di Carlo G, Ferrari M, Cremonesi L, Chiari M. Development of new substrates for high-sensitive genotyping of minority mutated alleles. ELECTROPHORESIS. 2008;29(23):4714–22.
- Damin F, Galbiati S, Soriani N, Burgio V, Ronzoni M, Ferrari M, Chiari M. Analysis of KRAS, NRAS and BRAF mutational profile by combination of in-tube hybridization and universal tag-microarray in tumor tissue and plasma of colorectal cancer patients. PLOS ONE. 2018 18;13(12):e0207876.
- Davies SW, Seale DA. DNA Microarray Stochastic Model. IEEE Trans.on Nanobioscience. 2005 Sep;4(3):248–54.
- Ryu M, Kim JD, Min BG, Kim J, Kim YY. Probe classification of on-off type DNA microarray images with a nonlinear matching measure. J Biomed Opt. 2006;11(1):014027.
- Ryu M, Dae Kim J, Goo Min B, Pang MG, Kim J. Nonlinear matching measure for the analysis of on-off type DNA microarray images. J Biomed Opt. 2004;9(3):432.
- Sako H, Fujio M, Furukawa N. The constellation matching and its application. In: Proceedings 2001 International Conference on Image Processing (Cat No01CH37205) [Internet]. 2001 [cited 2025 Feb 11]. p. 790–3 vol.1. Available from: https://ieeexplore.ieee.org/abstract/document/959164.
- Neal FB, Russ JC. Measuring Shape. CRC Press; 2012. 443 p.
- Russ, JC. The Image Processing Handbook. 5th ed. Boca Raton: CRC Press; 2006. 832 p.
- Qin L, Rueda L, Ali A, Ngom A. Spot Detection and Image Segmentation in DNA??Microarray Data: Applied Bioinformatics. 2005;4(1):1–11.
- Jones, NL. Fast annual daylighting simulation and high dynamic range image processing using NumPy. Science and Technology for the Built Environment. 2024 Apr 20;30(4):327–40.
- Watkinson N, Tai P, Nicolau A, Veidenbaum A. NumbaSummarizer: A Python Library for Simplified Vectorization Reports. In: 2020 IEEE International Parallel and Distributed Processing Symposium Workshops (IPDPSW) [Internet]. New Orleans, LA, USA: IEEE; 2020 [cited 2025 Feb 18]. p. 1–7. Available from: https://ieeexplore.ieee.org/document/9150361/.
- Maeda Y, Fukushima N, Matsuo H. Taxonomy of Vectorization Patterns of Programming for FIR Image Filters Using Kernel Subsampling and New One. Applied Sciences. 2018 Jul 26;8(8):1235.
- Szeliski R. Computer Vision: Algorithms and Applications, 2nd Edition.



| operation | PC (Windows 11) | Raspberry Pi 4 (Debian 12) | ||
| time (μsec) | ratio | time (msec) | ratio | |
| for-loop square | 4180 | 82 | 3540 | 6000 |
| square | 51 | 1 | 0.59 | 1 |
| square root | 61 | 1 | 1.58 | 3 |
| image add | 69 | 1 | 0.58 | 1 |
| square convolution | 428 | 8 | 3.03 | 5 |
| circle convolution | 3830 | 75 | 36.5 | 62 |
| square locating | 3100 | 61 | 26.8 | 45 |
| Radius(pixels) | 5 | 6 | 7 | 8 | 9 |
| Minimum (%) | 0.4 | 38.2 | 28.9 | 26.3 | 8.3 |
| Max. pos. error (pixels) | 5.83 | 0 | 4.00 | 5.00 | 5.00 |
| Radius(pixels) | 4 | 5 | 6 | 7 | 8 |
| Minimum (%) | 0.0 | 8.7 | 24.4 | 6.3 | 4.6 |
| Max. pos. error (pixels) | 5.83 | 4.47 | 3.61 | 5.00 | 6.08 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




