Submitted:
19 February 2025
Posted:
20 February 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Method for Analysis of Metabolic Pathway
3. Why genetic code and amino acid synthetic pathway coevolve?
4. Conditions making it possible to study on the origin and evolutionary process of genetic code
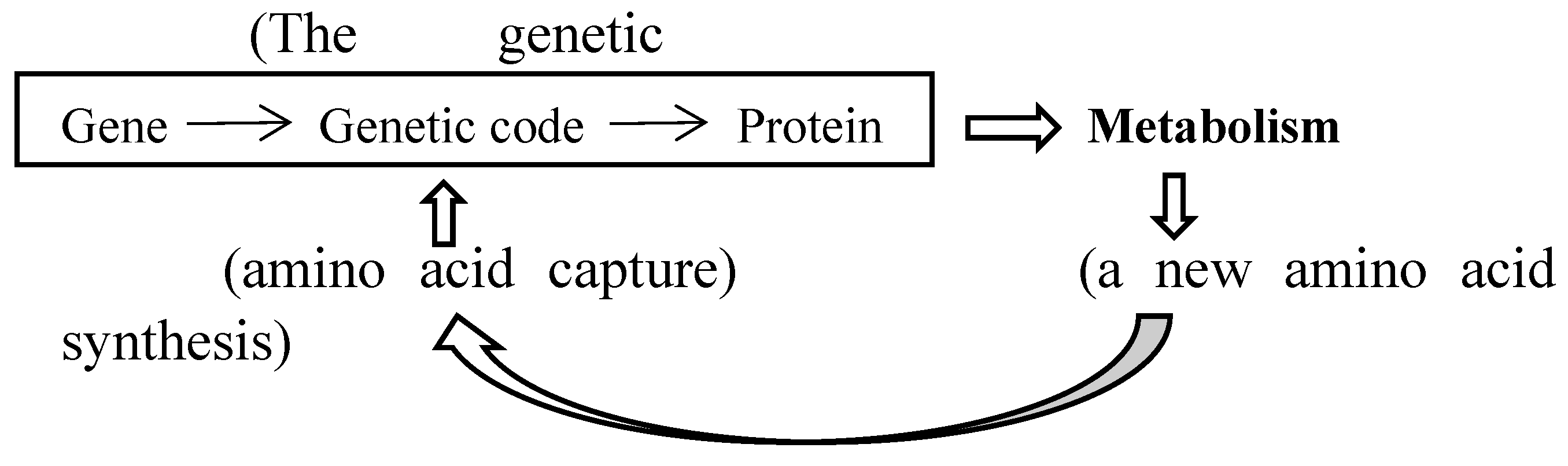
4.1. Formation of a new amino acid synthetic pathway leads to evolution of the genetic code
4.1.1. The order of amino acid capture into genetic code could be determined from analysis of modern metabolic map
4.1.2. Origin of the genetic code deduced from analyses of universal genetic code and modern metabolic pathways
5. The origin of the genetic code deduced from coevolution theory
6. Evolutionary pathway of the genetic code viewed from coevolution theory or amino acid metabolism
6.1. Analysis of Evolutionary Process of the Genetic Code
6.2. Characteristics and Differences of RNY code and SNS code
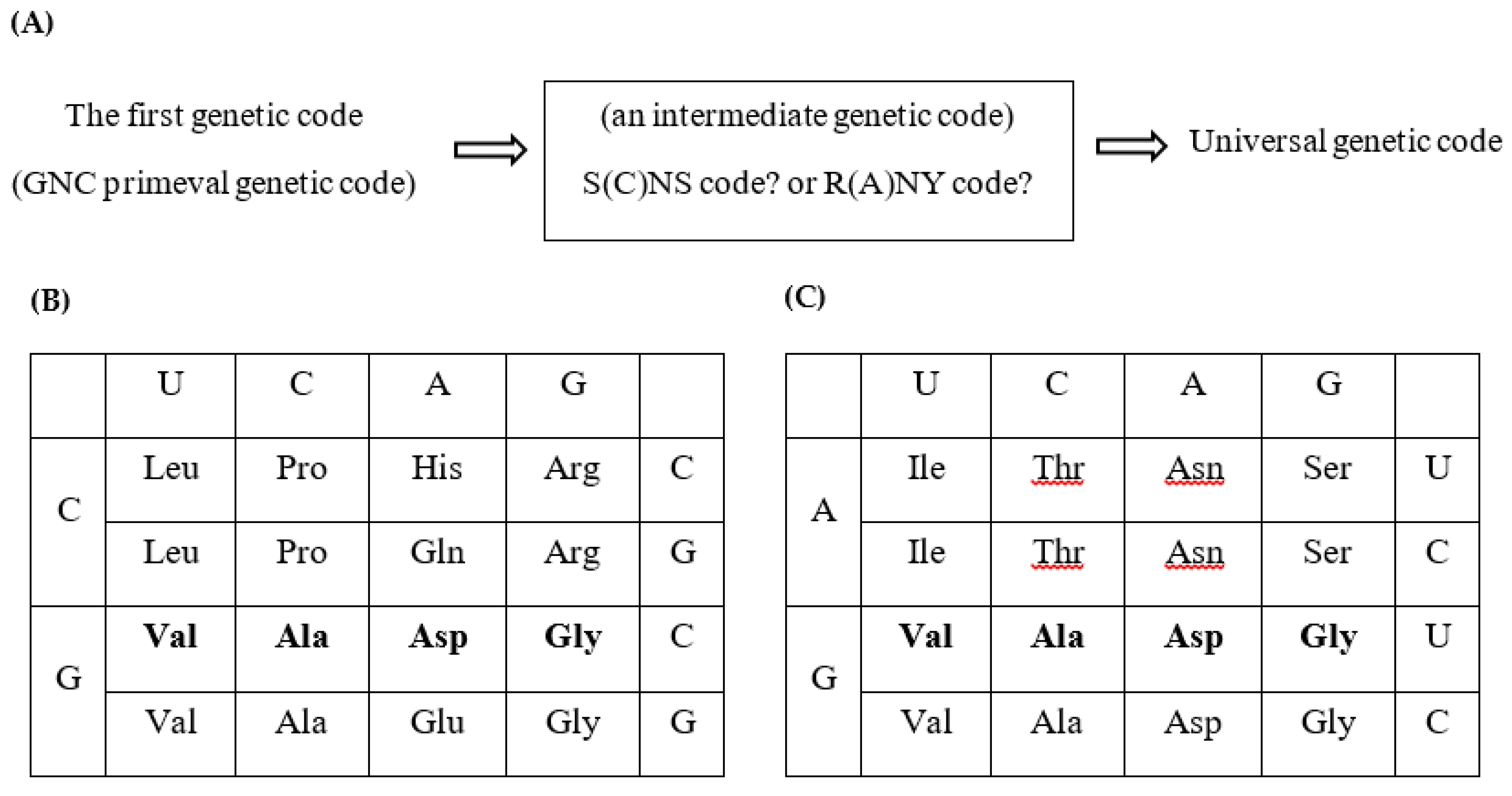
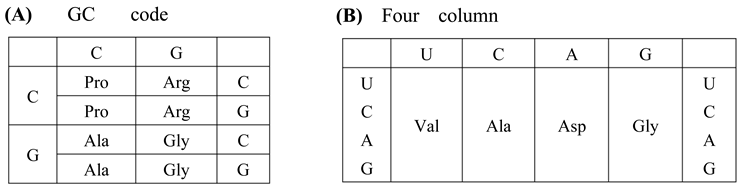
- Both SNS code and RNY code are consistent with the idea that the genetic code originated from the GNC primeval genetic code, because GNC code is contained in the two codes (Figure 4B,C).
- SNS code is symmetrical code between codon on sense strand and anticodon on antisense strand and is composed of ten amino acids and sixteen codons. Two acidic amino acids, Asp and Glu, and two basic amino acids, His and Arg, are contained in the SNS code. Two basic amino acids, His and Arg, and one highly hydrophobic amino acid, Leu, which are relatively complex amino acids, are contained in the SNS code. (Figure 4B).
- RNY code is also symmetrical code between codon and anticodon and is composed of eight relatively simple amino acids except Ile and sixteen codons. No basic amino acid and only one acidic amino acid, Asp, is contained in the RNY code (Figure 4C).
- 4. In the case of SNS code, it is considered that the order of codon capture was GNC--GNG--CNC--CNG codons. On the contrary, it is supposed that codons were captured in order of codons GNC--GNU--ANC--ANU to form RNY code.
6.3. Evidences showing that SNS code but not RNY code was used as an intermediate code
6.4. Decoding SNS Code and RNY Code by tRNA
7. Limits of the Coevolution Theory
7.1. The Reason Why ANN Code Was Introduced into SNN Code After SNN Code Was Formed
7.2. The reason why it could not be determined which one of two evolutionary pathways passed through based on only coevolution theory or amino acid metabolism
7.3. Examples appropriate to the coevolution theory
8. Discussion
References
- Wong, J.T. A co-evolution theory of the genetic code. Proc. Natl. Acad. Sci. USA 1975, 72, 1909–1912. [Google Scholar] [CrossRef] [PubMed]
- Wong, J.T.; Ng, S.K.; Mat, W.K.; Hu, T.; Xue, H. Coevolution Theory of the Genetic Code at Age Forty: Pathway to Translation and Synthetic Life. Life 2016, 6, 12. [Google Scholar] [CrossRef] [PubMed]
- Di Giulio, M. The coevolution theory of the origin of the genetic code. J Mol Evol. 1999, 48, 253–255. [Google Scholar] [CrossRef]
- Di Giulio, M. An extension of the coevolution theory of the origin of the genetic code. Biol Direct. 2008, 3, 37. [Google Scholar] [CrossRef]
- Di Giulio, M. Theories of the origin of the genetic code: Strong corroboration for the coevolution theory. Biosystems 2024, 239, 105217. [Google Scholar] [CrossRef] [PubMed]
- KEGG Pathway Dtabase. https://www.genome.jp/kegg/pathway.
- Frank, A.; Froese, T. The Standard Genetic Code can Evolve from a Two-Letter GC Code Without Information Loss or Costly Reassignments. Orig. Life Evol. Biosph. 2018, 48, 259–272. [Google Scholar] [CrossRef]
- Higgs, P.G. A four-column theory for the origin of the genetic code: Tracing the evolutionary pathways that gave rise to an optimized code. Biol Direct 2009, 24, 4–16. [Google Scholar] [CrossRef]
- Ikehara, K.; Omori, Y.; Arai, R.; Hirose, A. A novel theory on the origin of the genetic code: a GNC-SNS hypothesis. J. Mol. Evol. 2002, 54, 530–538. [Google Scholar] [CrossRef]
- Ikehara, K. Towards Revealing the Origin of life.—Presenting the GADV Hypothesis; Springer Nature, Gewerbestrasse: Cham, Switzerland, 2021. [Google Scholar]
- Ikehara, K. Protein ordered sequences are formed by random joining of amino acids in protein 0th-order structure, followed by evolutionary process. Orig. Life Evol. Biosph. 2014, 44, 279–281. [Google Scholar] [CrossRef]
- Ikehara, K. Why were [GADV]-amino acids and GNC codons selected and how was GNC primeval genetic code established? Genes 2023, 14, 375. [Google Scholar] [CrossRef]
- Taghavi, A.; van der Schoot, P.; Berryman, J.T. DNA partitions into triplets under tension in the presence of organic cations, with sequence evolutionary age predicting the stability of the triplet phase. Q. Rev. Biophys 2017, e15. [Google Scholar] [CrossRef] [PubMed]
- Ikehara, K. The origin of tRNA deduced from Pseudomonas aeruginosa 5’ anticodon-stem sequence: Anticodon stem-loop hypothesis. Orig. Life Evol. Biosph. 2019, 49, 61–75. [Google Scholar] [CrossRef]
- Lei, L.; Burton, Z.F. Evolution of Life on Earth: tRNA, Aminoacyl-tRNA Synthetases and the Genetic Code. Life 2020, 10, 21. [Google Scholar] [CrossRef] [PubMed]
- Watson, J.D.; Hopkins, N.H.; Roberts, J.W.; Steitz, J.A.; Weiner, A.M. Molecular Biology of the Gene. 4th ed. Vol. 1, The Benjamin/Cummings Publishing Company, Inc.: Menlo Park, CA, USA 1987.
- Zamudio, G.S.; José, M.V. On the Uniqueness of the Standard Genetic Code. Life 2017, 7, 7. [Google Scholar] [CrossRef]
- Ikehara, K.; Amada, F.; Yoshida, S.; Mikata, Y.; Tanaka, A. A possible origin of newly-born bacterial genes: Significance of GC-rich nonstop frame on antisense strand. Nucl. Acids Res. 1996, 24, 4249–4255. [Google Scholar] [CrossRef] [PubMed]
- Lei, L.; Burton, Z.F. Evolution of the genetic code. Transcription 2021, 12, 28–53. [Google Scholar] [CrossRef]
- Kim, Y.; Opron, K. Burton, Z.F. A tRNA- and Anticodon-Centric View of the Evolution of Aminoacyl-tRNA Synthetases, tRNAomes, and the Genetic Code. Life 2019, 9, 37. [Google Scholar] [CrossRef]
- Rogers, S.O. Evolution of the genetic code based on conservative changes of codons, amino acids, and aminoacyl tRNA synthetases, J. Theor. Biol. 2019, 466, 1–10. [Google Scholar] [CrossRef]
- Foltan, J.S. tRNA genes and the genetic code. J. Theor. Biol. 2008, 253, 469–482. [Google Scholar] [CrossRef]
- Jackman., J.E.; Alfonzo, J.D. Transfer RNA modifications: nature's combinatorial chemistry playground. Wiley Interdiscip 2013, 4, 35–48. [Google Scholar] [CrossRef]
- Yared, M.J.; Marcelot, A.; Barraud, P. Beyond the Anticodon: tRNA Core Modifications and Their Impact on Structure, Translation and Stress Adaptation. Genes 2024, 15, 374. [Google Scholar] [CrossRef] [PubMed]


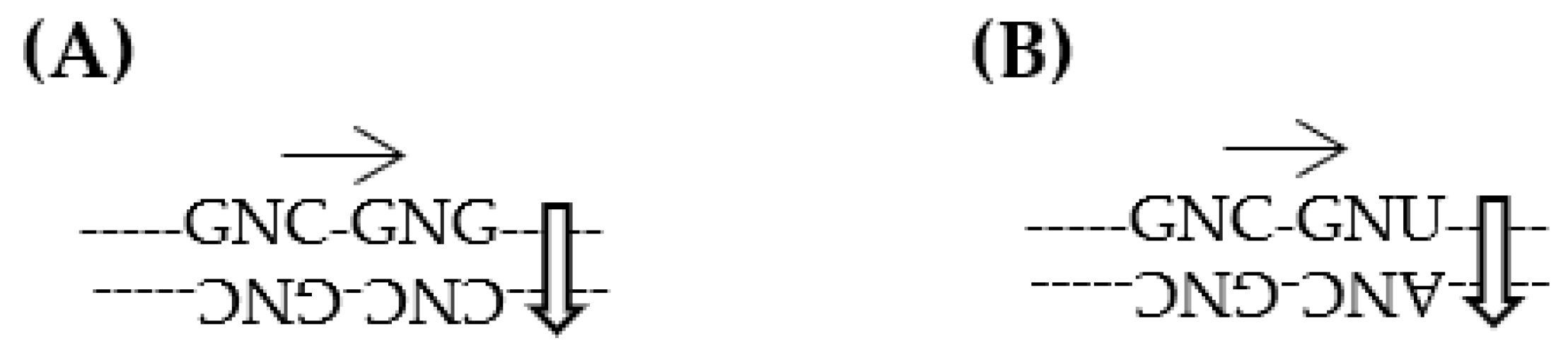
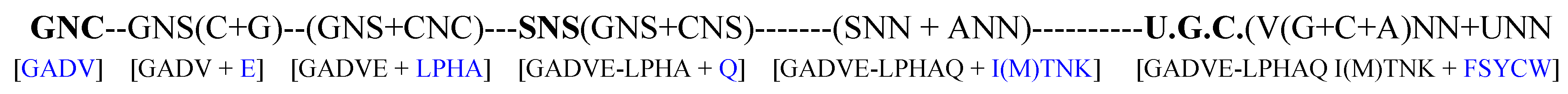
 |
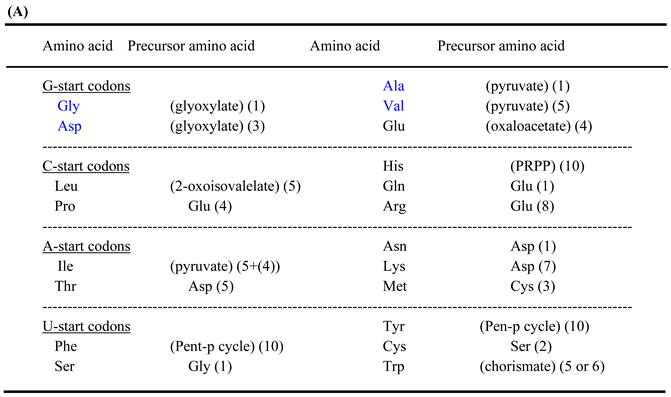 |
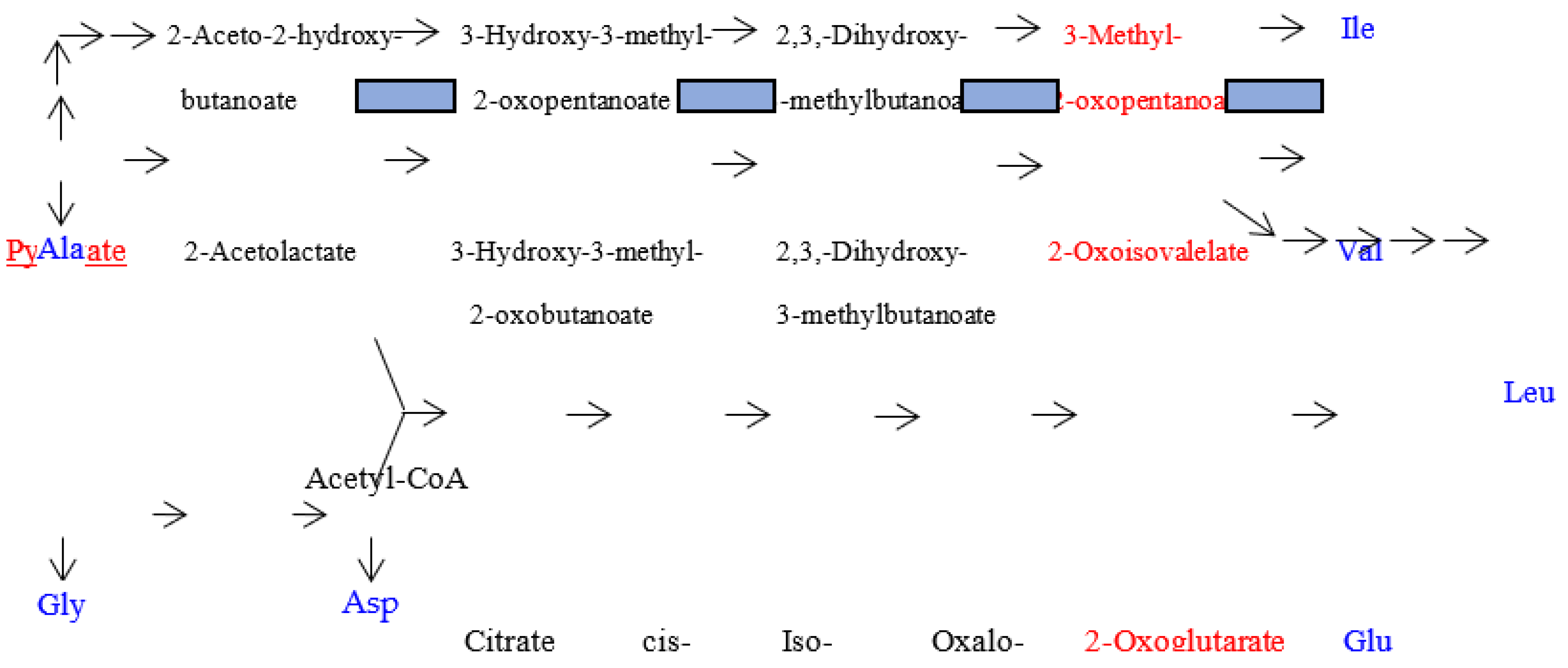
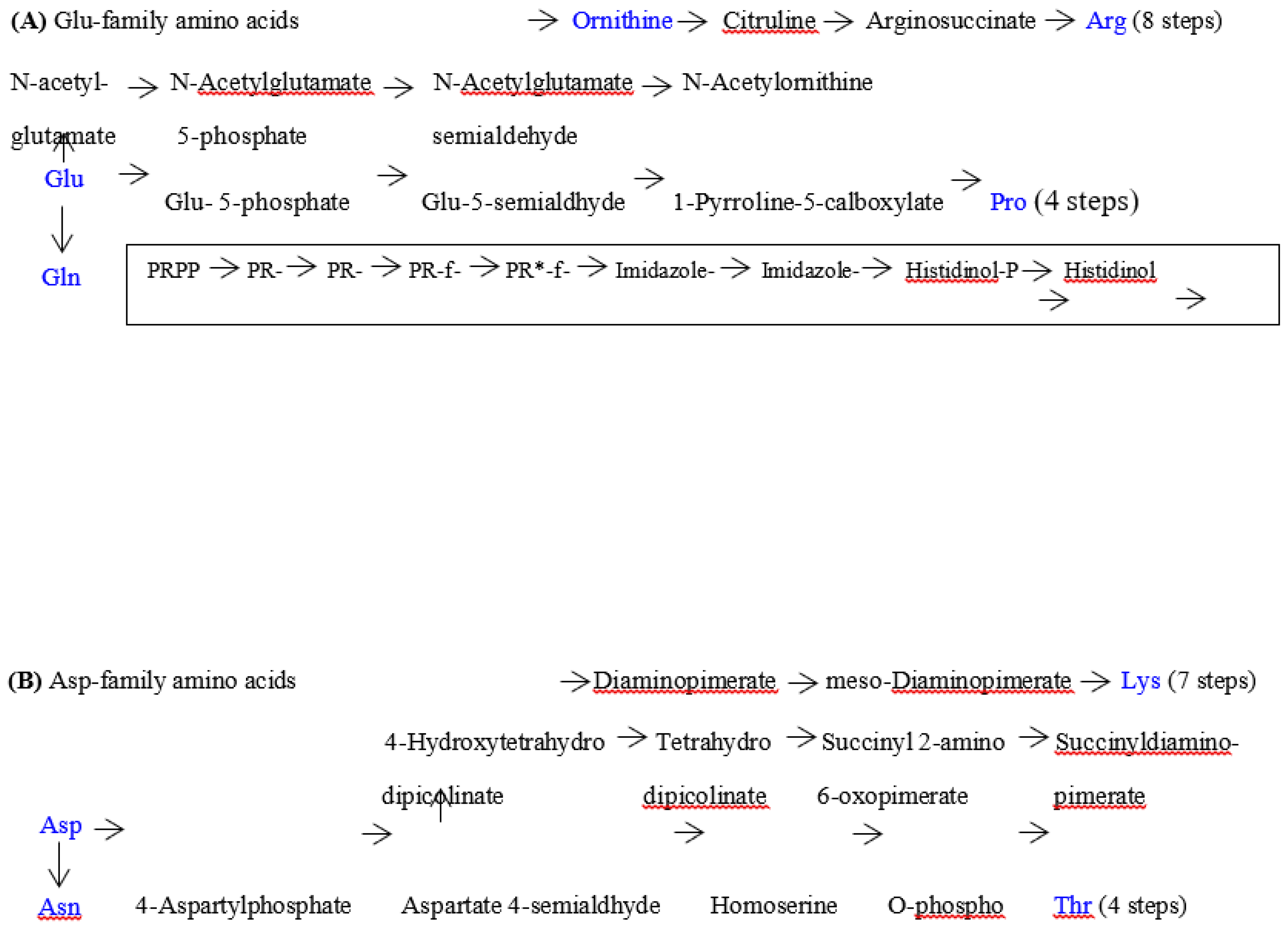
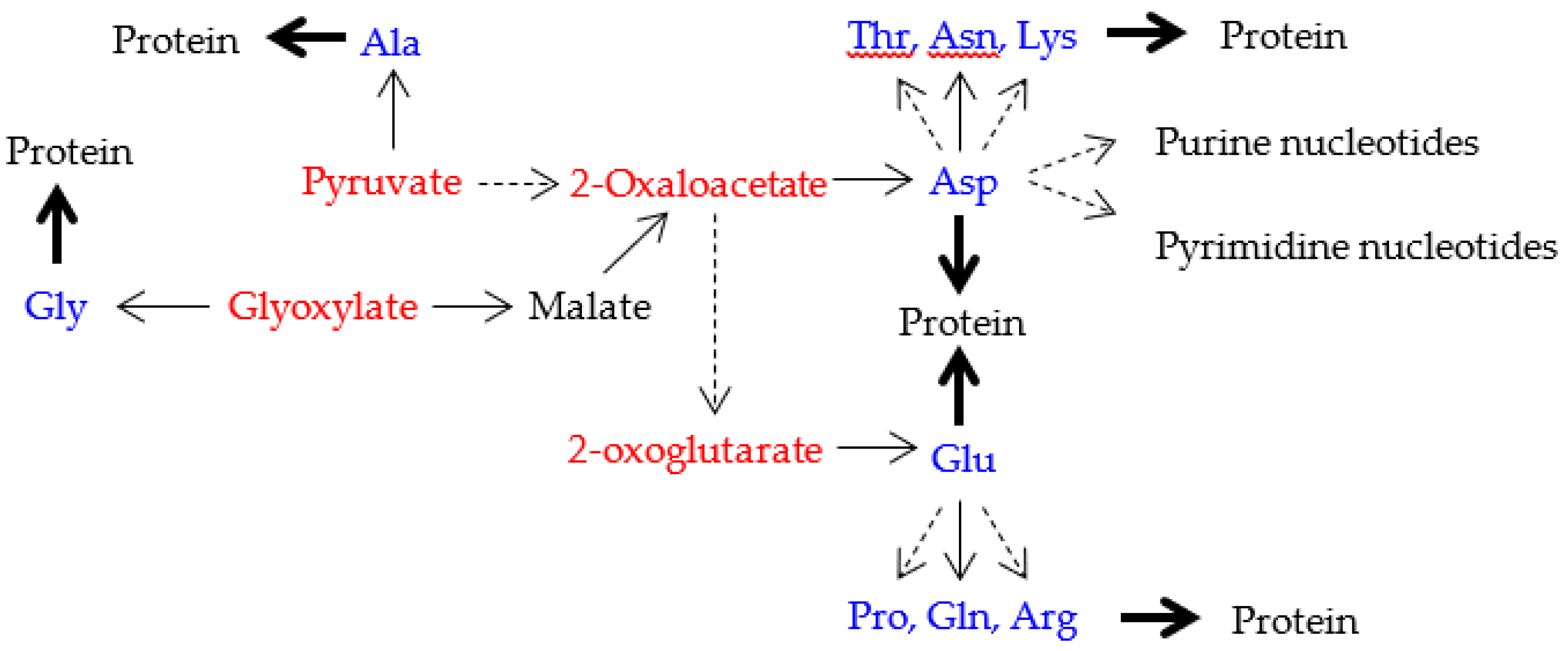
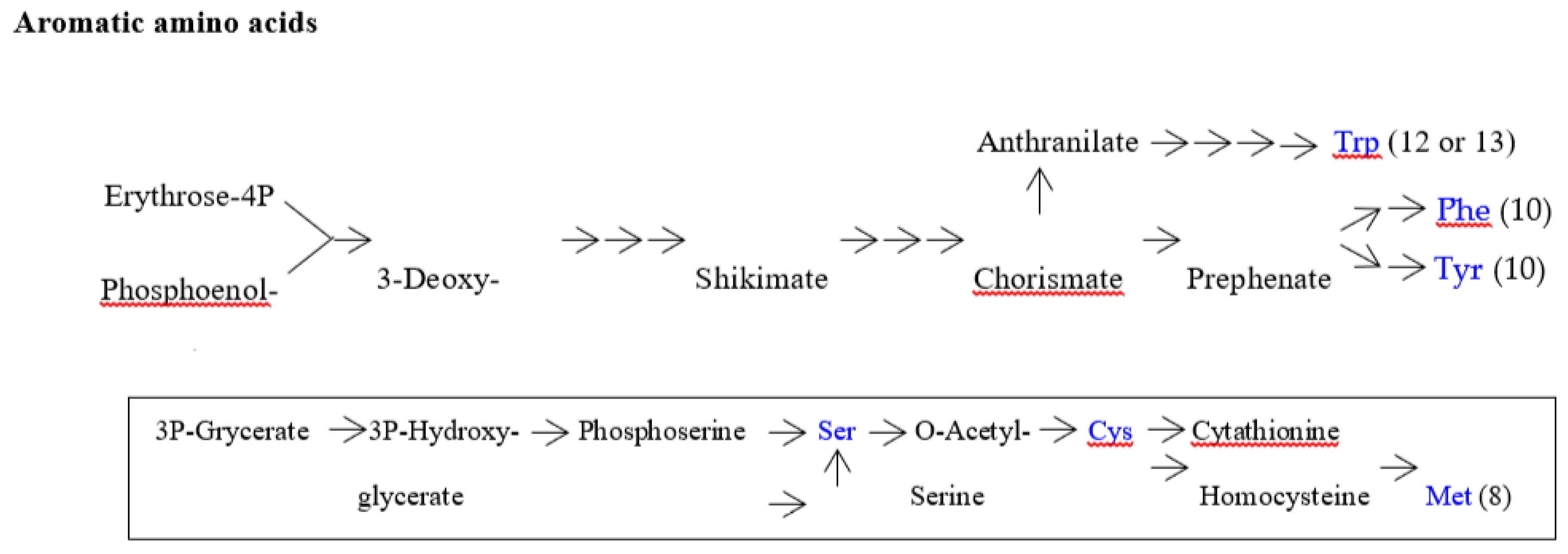
| Note: Some characteristics can bee seen in the above table. (1) amino acid metabolic pathways of all the five amino acids (Gly, Ala, Asp, Val and Glu) using a G-start codon do not use any precursor amino acid. (2) Reaction steps from pyruvate more than 8 are used in the synthetic pathways of two hydrophobic amino acids (Leu and Ile) with a long side chain. (3) Many reaction steps (more than 10) counted from the initial precursor molecules are also used in the synthetic pathways of three aromatic amino acids (Phe, Tyr and Trp). (4) Intermediate molecule (2-oxoisovalerate) for Val synthesis is used in Leu synthetic pathway and a series of enzymes on the Val synthetic pathway are used for Ile synthesis. (5) PRPP is used at the initial step of His synthetic pathway. (6) Glu and Asp are used as a precursor amino acid in synthetic pathways for three amino acids encoded by C-start codon and A-start codon, respectively. Pent-p cycle is a abbreviation of pentose-phosphate cycle. |
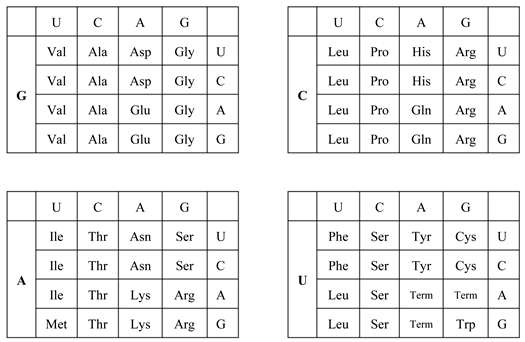
| (B) The universal genetic code table summarized for each base at the first codon position. (Upper left) G-start codon table. (Upper right) C-start codon table. (Lower left) A-start codon table. (Lower right) U-start codon table. Term in the U-start codon table indicates termination codon. |
 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




