Submitted:
07 February 2025
Posted:
11 February 2025
You are already at the latest version
Abstract
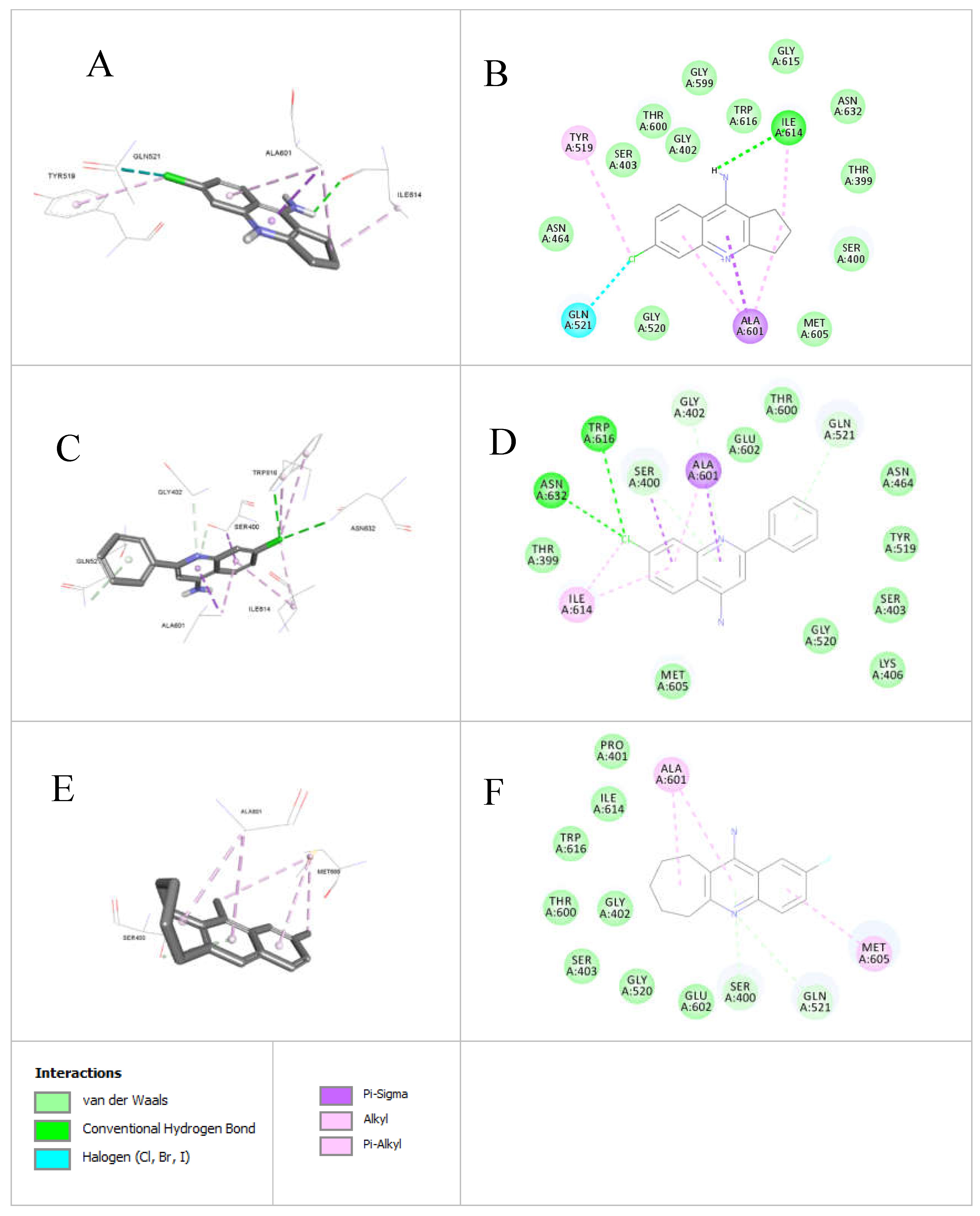
4-Aminoquinoline derivatives were synthesized on a small scale using a novel microwave-assisted method, and scaled up in sealed tubes. The synthesis adhered to green chemistry principles, employing a solvent-free approach for both the reaction and purification. The purification was achieved through simple washing, no need for column chromatography. All reactions were conducted at temperatures between 90–150 ºC within 90–120 minutes, achieving yields of up to 95%. The products were characterized using FT-IR, 1H- and 13C-NMR spectroscopy, and HR-MS spectrometry. Antibacterial and antifungal activity testing revealed that four compounds exhibited moderate antibacterial activity. Compound 6-chlorocyclopentaquinolinamine demonstrated a strong MIC of 0.125 mM against MRSA, while compound 2-fluorocycloheptaquinolinamine showed a MIC of 0.25 mM against S. pyogenes. A Structure-Activity Relationship (SAR) docking study was conducted within the Penicillin Binding Protein (PBP) binding site. Docking analysis of anti-MRSA com-pounds 7-chlorophenylquinolinamine, 6-chlorocyclopentaquinolinamine, and 2-fluorocycloheptaquinolinamine in the MRSA PBP2a binding pocket (PDB: 4DK1) revealed that 6-chlorocyclopentaquinolinamine and 7-chlorophenylquinolinamine interacted via hydrophobic (ALA601, ILE614), hydrogen bonding (GLN521), and halogen interactions (TYR519, THR399). Compound 6-chlorocyclopentaquinolinamine exhibited superior MRSA inhibition (20 mm inhibition zone vs. 12.5 mm for 7-chlorophenylquinolinamine), attributed to additional π-alkyl interactions and favorable docking parameters, including higher Ligscore2 (4.03), PLP1 (59.15), and Dock Score (34.31). In contrast, compound 2-fluorocycloheptaquinolinamine exhibited weaker activity due to its bulky structure, limited interactions, and less favorable docking scores.
Keywords:
1. Introduction
2. Materials and Methods
2.1. Chemicals and Materials
2.2. Synthesis
General Procedure (GP)
2.3. Antibacterial Antifungal Determination Method
2.3.1. Microorganisims
2.3.2. Well Diffusion Method
2.3.3. Minimum Inhibitory Concentration (MIC)
2.3.4. Minimum Bactericidal Concentration (MBC)
2.4. Molecular Docking Methodology
2.4.1. Software Packages
- CS ChemDraw® Ultra. Cambridge Soft Corp. www.cambridgesoft.com. as a 2D chemical drawing tool.
- BIOVIA Discovery Studio® 4.1 (DS 4.1) implemented with Standalone Application, https://www.3dsbiovia.com for ligand docking protocols.
2.4.2. Ligand Preparation and Molecular DockingProtein Preparation
Ligand Preparation
Binding Site Definition
- PBP2a Binding Site:
- 3NKH Integrase Enzyme Binding Site:
2.5. Docking Procedure
2.5.1. Scoring and Evaluation
2.5.2. Ligand Pose Analysis
3. Results and Discussion
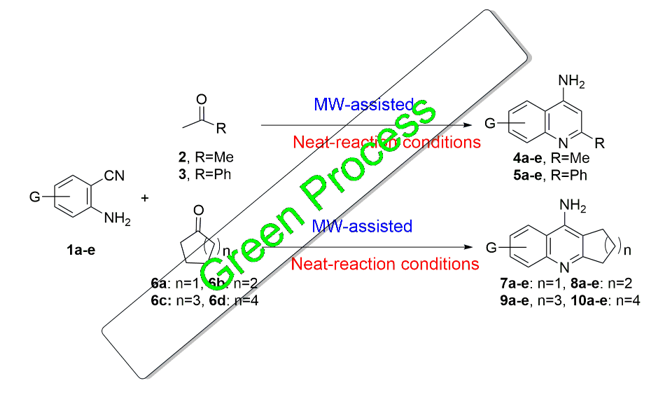
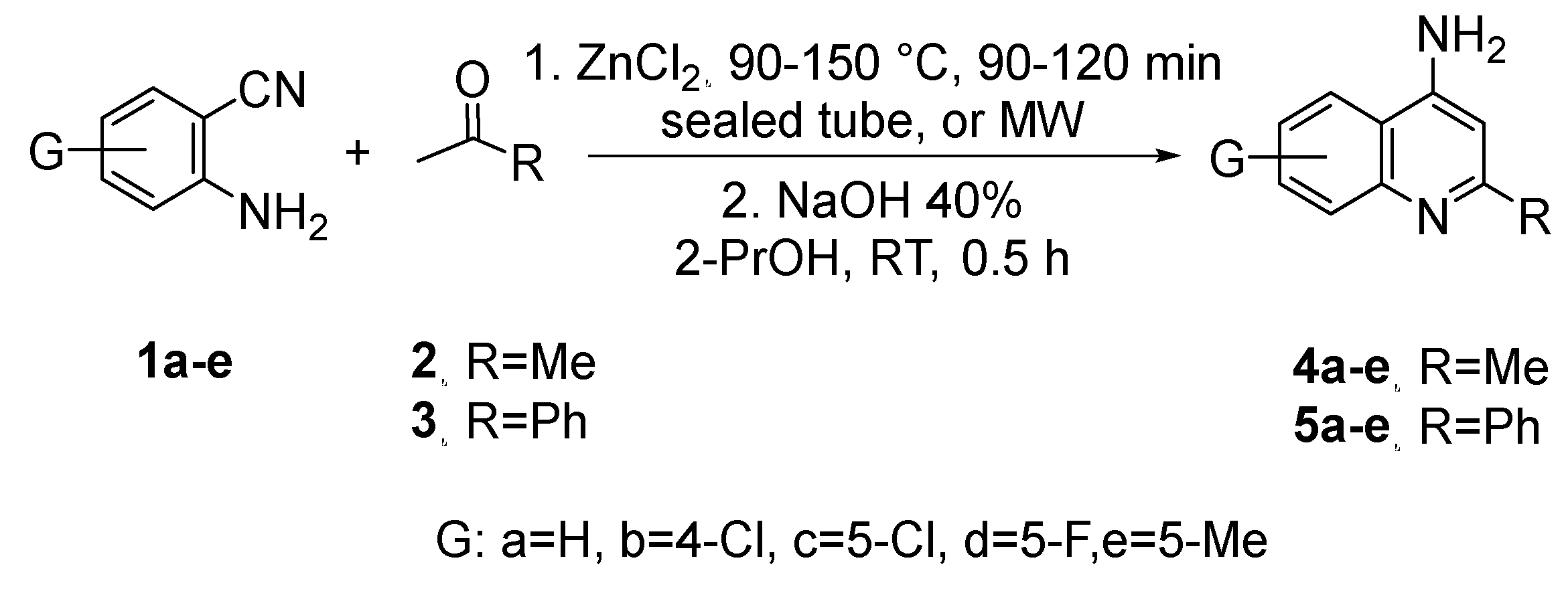
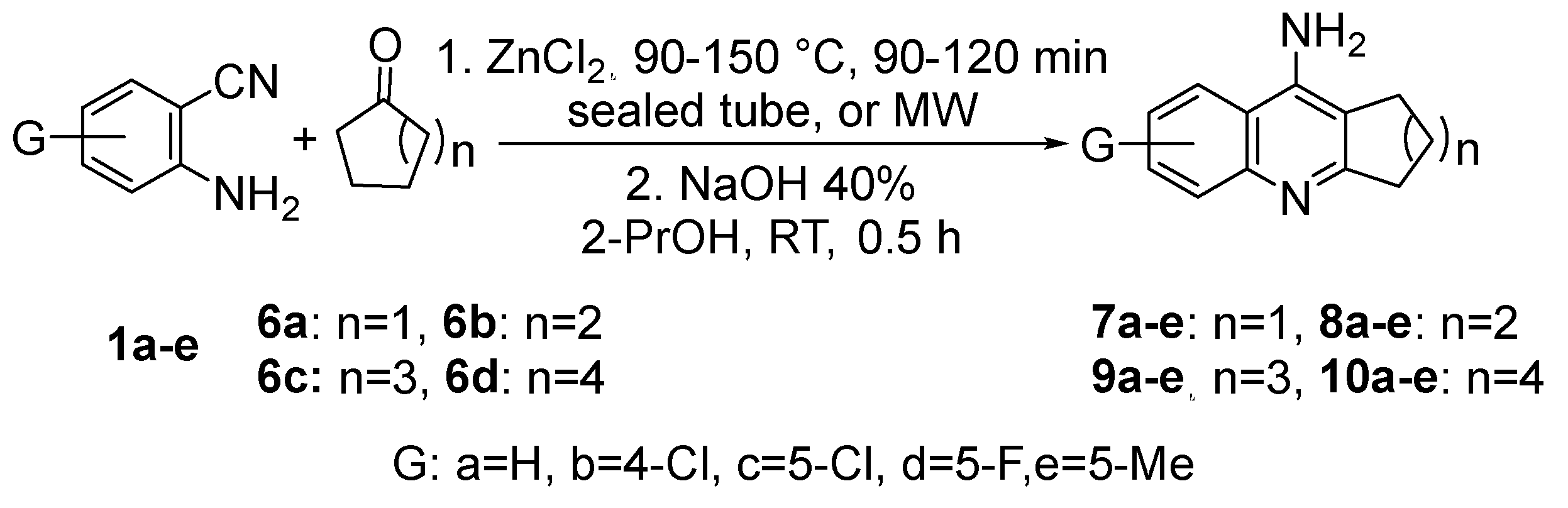
3.1. Synthesis and Characterization
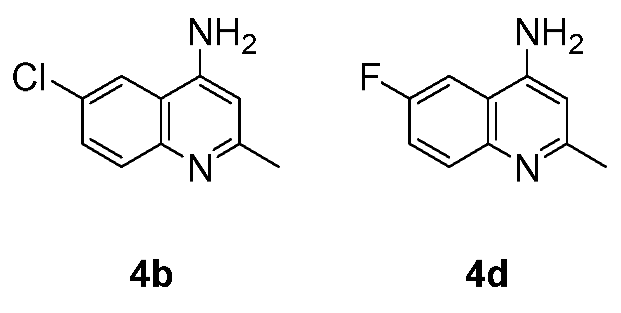
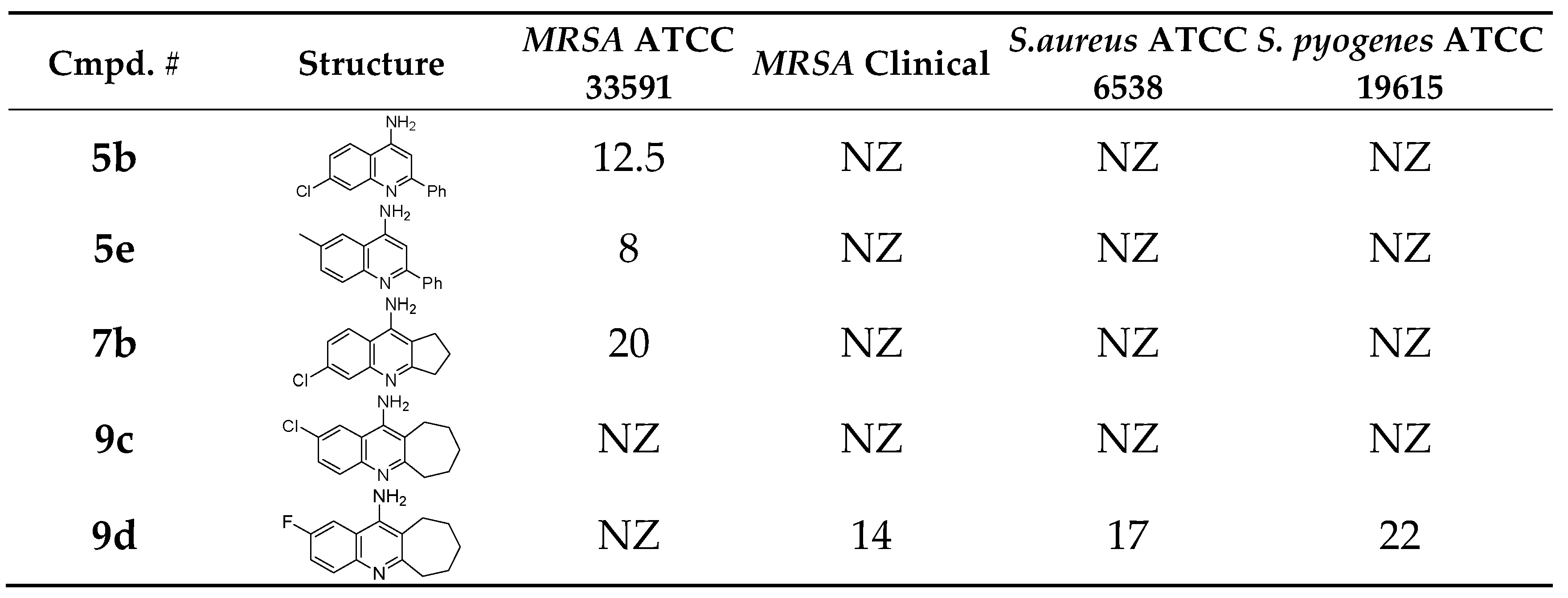
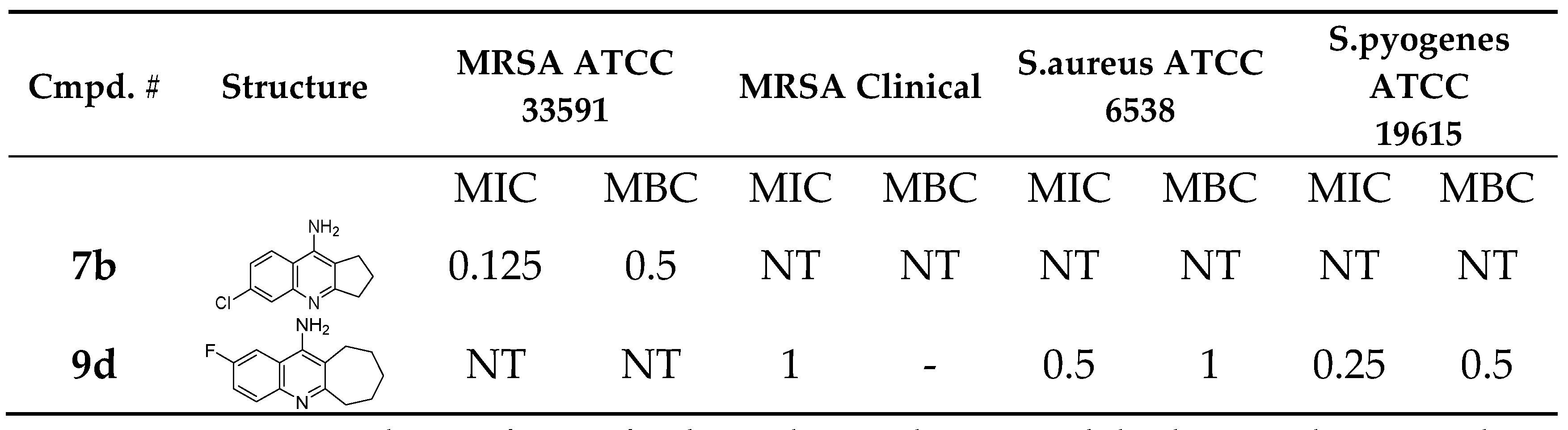
3.2. Antibacterial and Antifungal Activity
3.3. Results of the Molecular Docking Experiment
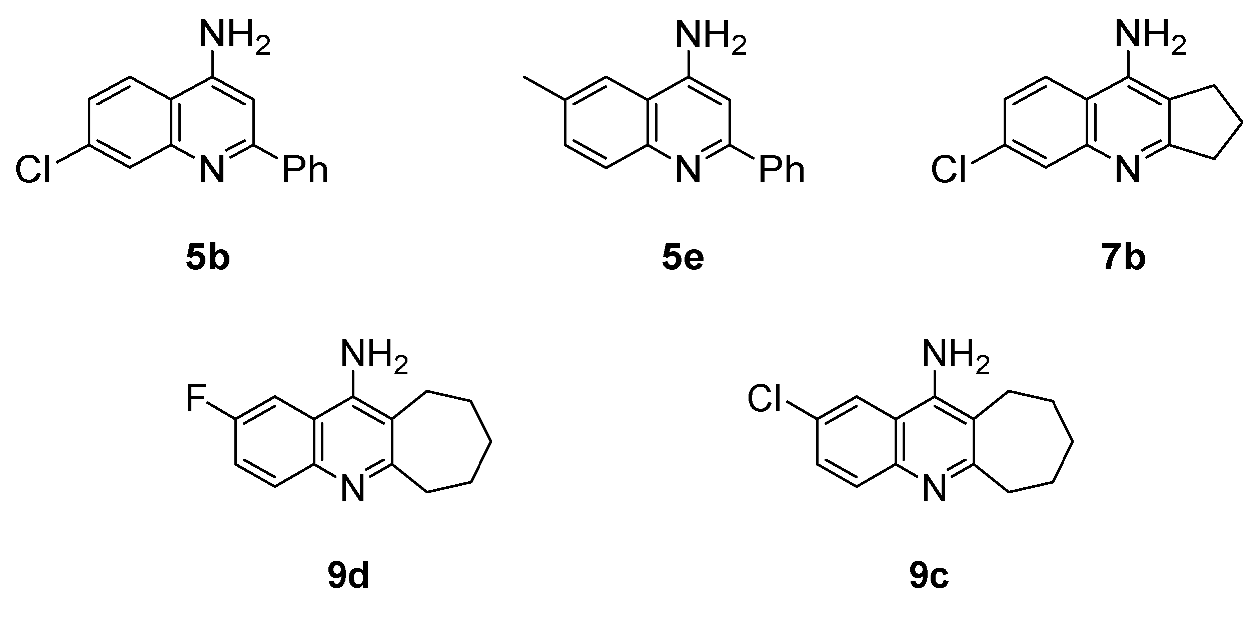
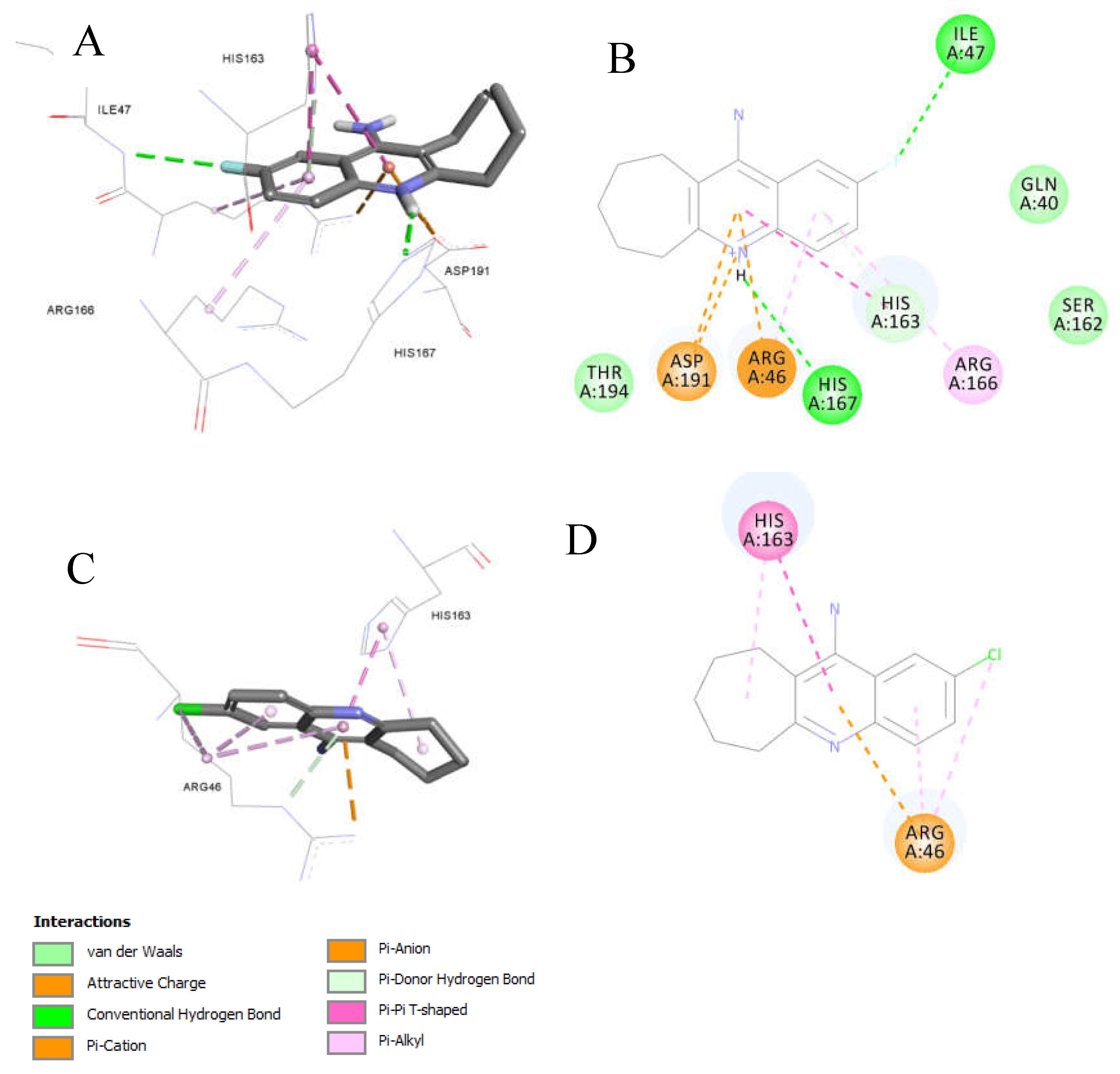
3.3.1. Docking of Anti-MRSA Active Compounds into the Binding Pocket of MRSA PBP2a Discussion
3.3.2. Docking of the Anti-MRSA Active Compounds Into the Binding Pocket MRSA Integrase
3.3.3. Ligand Pose Analysis Inside the Binding Pocket of MRSA-Penicillin Binding Protein 2a (PBP2a)
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Sheldon, R.A. E Factors, Green Chemistry and Catalysis: An Odyssey. Chemical Communications 2008, 3352–3365. [Google Scholar] [CrossRef] [PubMed]
- Polshettiwar, V.; Varma, R.S. Microwave-Assisted Organic Synthesis and Transformations Using Benign Reaction Media. Acc Chem Res 2008, 41, 629–639. [Google Scholar] [CrossRef]
- Van Praet, S.; Preegel, G.; Rammal, F.; Sels, B.F. One-Pot Consecutive Reductive Amination Synthesis of Pharmaceuticals: From Biobased Glycolaldehyde to Hydroxychloroquine. ACS Sustain Chem Eng 2022, 10, 6503–6508. [Google Scholar] [CrossRef]
- Nainwal, L.M.; Tasneem, S.; Akhtar, W.; Verma, G.; Khan, M.F.; Parvez, S.; Shaquiquzzaman, M.; Akhter, M.; Alam, M.M. Green Recipes to Quinoline: A Review. Eur J Med Chem 2019, 164, 121–170. [Google Scholar] [CrossRef]
- Ray, R.L. Alkaloids—The World’s Pain Killers. J Chem Educ 1960, 37, 451. [Google Scholar] [CrossRef]
- Matada, B.S.; Pattanashettar, R.; Yernale, N.G. A Comprehensive Review on the Biological Interest of Quinoline and Its Derivatives. Bioorg Med Chem 2021, 32, 115973. [Google Scholar] [CrossRef]
- Kaur, K.; Jain, M.; Reddy, R.P.; Jain, R. Quinolines and Structurally Related Heterocycles as Antimalarials. Eur J Med Chem 2010, 45, 3245–3264. [Google Scholar] [CrossRef]
- Browning, D.J. Pharmacology of Chloroquine and Hydroxychloroquine. In Hydroxychloroquine and Chloroquine Retinopathy; Browning, D.J., Ed.; Springer New York: New York, NY, 2014; ISBN 978-1-4939-0597-3. [Google Scholar]
- PANDEY, A. V; BISHT, H.; BABBARWAL, V.K.; SRIVASTAVA, J.; PANDEY, K.C.; CHAUHAN, V.S. Mechanism of Malarial Haem Detoxification Inhibition by Chloroquine. Biochemical Journal 2001, 355, 333–338. [Google Scholar] [CrossRef] [PubMed]
- Kimura, T.; Takabatake, Y.; Takahashi, A.; Isaka, Y. Chloroquine in Cancer Therapy: A Double-Edged Sword of Autophagy. Cancer Res 2013, 73, 3–7. [Google Scholar] [CrossRef]
- Gautret, P.; Lagier, J.-C.; Parola, P.; Hoang, V.T.; Meddeb, L.; Mailhe, M.; Doudier, B.; Courjon, J.; Giordanengo, V.; Vieira, V.E.; et al. Hydroxychloroquine and Azithromycin as a Treatment of COVID-19: Results of an Open-Label Non-Randomized Clinical Trial. Int J Antimicrob Agents 2020, 56, 105949. [Google Scholar] [CrossRef] [PubMed]
- Tzekov, R. Ocular Toxicity Due to Chloroquine and Hydroxychloroquine:Electrophysiological and Visual Function Correlates. Documenta Ophthalmologica 2005, 110, 111–120. [Google Scholar] [CrossRef] [PubMed]
- Verbaanderd, C.; Maes, H.; Schaaf, M.B.; Sukhatme, V.P.; Pantziarka, P.; Sukhatme, V.; Agostinis, P.; Bouche, G. Repurposing Drugs in Oncology (ReDO) - Chloroquine and Hydroxychloroquine as Anti-Cancer Agents. Ecancermedicalscience 2017, 11. [Google Scholar] [CrossRef] [PubMed]
- Zhou, W.; Wang, H.; Yang, Y.; Chen, Z.-S.; Zou, C.; Zhang, J. Chloroquine against Malaria, Cancers and Viral Diseases. Drug Discov Today 2020, 25, 2012–2022. [Google Scholar] [CrossRef] [PubMed]
- Lei, Z.-N.; Wu, Z.-X.; Dong, S.; Yang, D.-H.; Zhang, L.; Ke, Z.; Zou, C.; Chen, Z.-S. Chloroquine and Hydroxychloroquine in the Treatment of Malaria and Repurposing in Treating COVID-19. Pharmacol Ther 2020, 216, 107672. [Google Scholar] [CrossRef]
- Nardi, N.; Baumgarten, L.G.; Dreyer, J.P.; Santana, E.R.; Winiarski, J.P.; Vieira, I.C. Nanocomposite Based on Green Synthesis of Gold Nanoparticles Decorated with Functionalized Multi-Walled Carbon Nanotubes for the Electrochemical Determination of Hydroxychloroquine. J Pharm Biomed Anal 2023, 236, 115681. [Google Scholar] [CrossRef]
- Abumansour, H.; Abusara, O.H.; Khalil, W.; Abul-Futouh, H.; Ibrahim, A.I.M.; Harb, M.K.; Abulebdah, D.H.; Ismail, W.H. Biological Evaluation of Levofloxacin and Its Thionated Derivatives: Antioxidant Activity, Aldehyde Dehydrogenase Enzyme Inhibition, and Cytotoxicity on A549 Cell Line. Naunyn Schmiedebergs Arch Pharmacol 2024. [Google Scholar] [CrossRef]
- Zhang, N.; Xie, H.; Lu, W.; Li, F.; Li, J.; Guo, Z. Chloroquine Sensitizes Hepatocellular Carcinoma Cells to Chemotherapy via Blocking Autophagy and Promoting Mitochondrial Dysfunction; 2017; Vol. 10;
- Latarissa, I.R.; Barliana, M.I.; Meiliana, A.; Lestari, K. Potential of Quinine Sulfate for Covid-19 Treatment and Its Safety Profile: Review. Clin Pharmacol 2021, 13, 225–234. [Google Scholar] [CrossRef]
- Zhao, Z.; Song, J.; Shang, Y.; Shan, M.; Tang, L.; Pang, T.; Ye, Q.; Cheng, G.; Jiang, G. Design, Synthesis, Biological Evaluation and Docking Study of A New Series of 9-Aminoacridine Derivatives as Anti-Inflammatory Agents. ChemistrySelect 2024, 9, e202302387. [Google Scholar] [CrossRef]
- Bauer, A.W.; Kirby, W.M.M.; Sherris, J.C.; Turck, M. Antibiotic Susceptibility Testing by a Standardized Single Disk Method. Am J Clin Pathol 1966, 45, 493–496. [Google Scholar] [CrossRef]
- Gedam, V.; Punyapwar, S.; Raut, P.; Chahande, A. Rapid In Vitro Proliferation of Tinospora Cordifolia Callus Biomass from Stem and Leaf with Phytochemical and Antimicrobial Assessment: 7th IconSWM—ISWMAW 2017, Volume 2. In; 2019; pp. 421–431 ISBN 978-981-13-2783-4.
- Ibrahim, A.I.M.; Abul-Futouh, H.; Bourghli, L.M.S.; Abu-Sini, M.; Sunoqrot, S.; Ikhmais, B.; Jha, V.; Sarayrah, Q.; Abulebdah, D.H.; Ismail, W.H. Design and Synthesis of Thionated Levofloxacin: Insights into a New Generation of Quinolones with Potential Therapeutic and Analytical Applications. Curr Issues Mol Biol 2022, 44, 4626–4638. [Google Scholar] [CrossRef]
- Wu, G.; Robertson, D.H.; Brooks III, C.L.; Vieth, M. Detailed Analysis of Grid-Based Molecular Docking: A Case Study of CDOCKER—A CHARMm-Based MD Docking Algorithm. J Comput Chem 2003, 24, 1549–1562. [Google Scholar] [CrossRef]
- Krammer, A.; Kirchhoff, P.D.; Jiang, X.; Venkatachalam, C.M.; Waldman, M. LigScore: A Novel Scoring Function for Predicting Binding Affinities. J Mol Graph Model 2005, 23, 395–407. [Google Scholar] [CrossRef] [PubMed]
- Gehlhaarl, D.K.; Verkhivkerl, G.M.; Rejtol, P.A.; Sherman, C.J.; Fogel, D.B.; Fogel, L.J.; Freer’, S.T. Molecular Recognition of the Inhibitor AC-1343 by HIV-l Protease: Conformationally Flexible Docking by Evolutionary Programming. Chem Biol 1995, 2, 317–324. [Google Scholar] [CrossRef] [PubMed]
- Jain, A.N. Scoring Functions for Protein-Ligand Docking. Curr Protein Pept Sci 2006, 7, 407–420. [Google Scholar] [CrossRef]
- Muegge, I.; Martin, Y.C. A General and Fast Scoring Function for Protein-Ligand Interactions: A Simplified Potential Approach. J Med Chem 1999, 42, 791–804. [Google Scholar] [CrossRef] [PubMed]
- Bæk, K.T.; Gründling, A.; Mogensen, R.G.; Thøgersen, L.; Petersen, A.; Paulander, W.; Frees, D. β-Lactam Resistance in Methicillin-Resistant Staphylococcus Aureus USA300 Is Increased by Inactivation of the ClpXP Protease. Antimicrob Agents Chemother 2014, 58, 4593–4603. [Google Scholar] [CrossRef] [PubMed]
- Lovering, A.L.; Gretes, M.C.; Safadi, S.S.; Danel, F.; De Castro, L.; Page, M.G.P.; Strynadka, N.C.J. Structural Insights into the Anti-Methicillin-Resistant Staphylococcus Aureus (MRSA) Activity of Ceftobiprole. Journal of Biological Chemistry 2012, 287, 32096–32102. [Google Scholar] [CrossRef]
- Purrello, S.M.; Daum, R.S.; Edwards, G.F.S.; Lina, G.; Lindsay, J.; Peters, G.; Stefani, S. Meticillin-Resistant Staphylococcus Aureus (MRSA) Update: New Insights into Bacterial Adaptation and Therapeutic Targets. J Glob Antimicrob Resist 2014, 2, 61–69. [Google Scholar] [CrossRef]
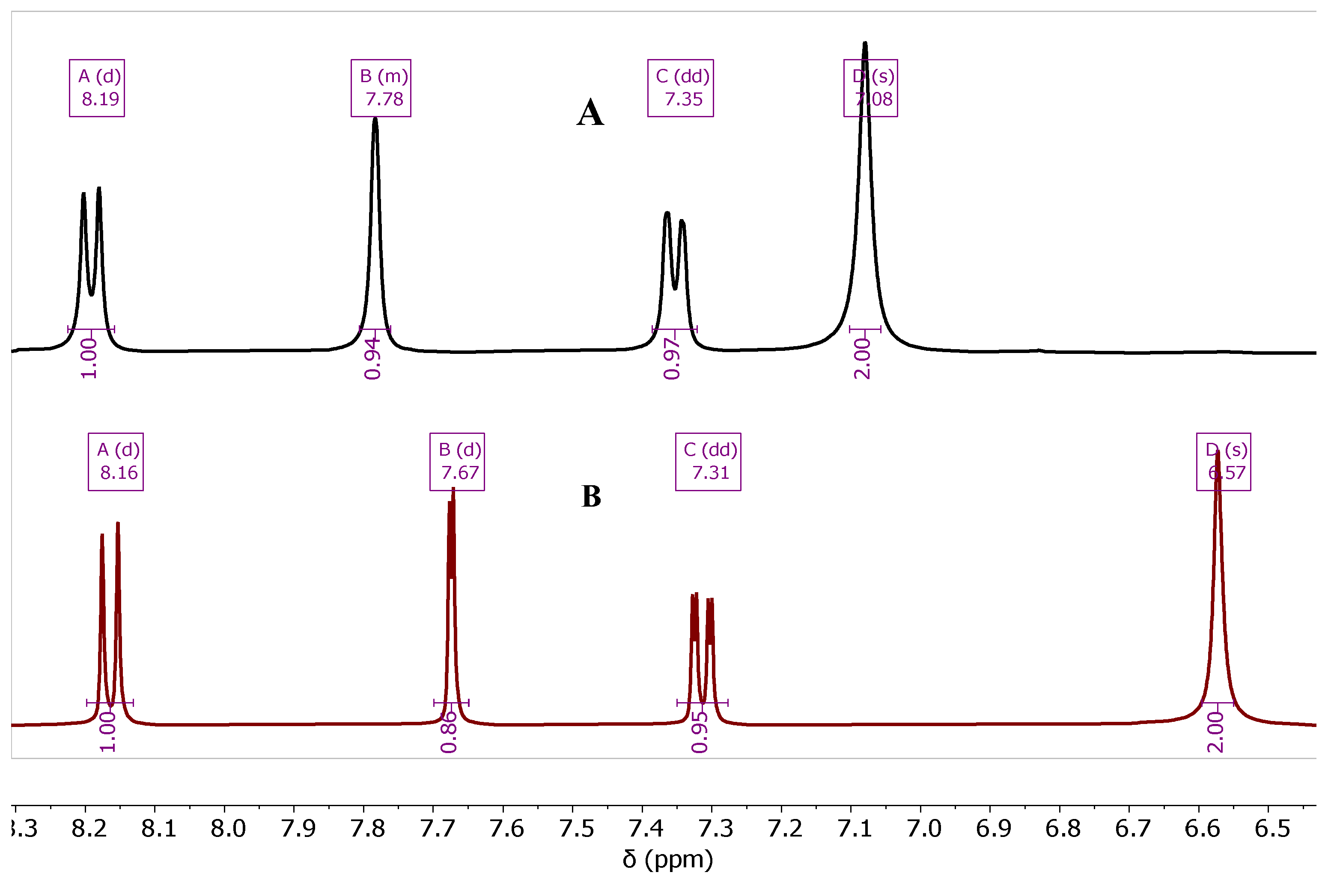
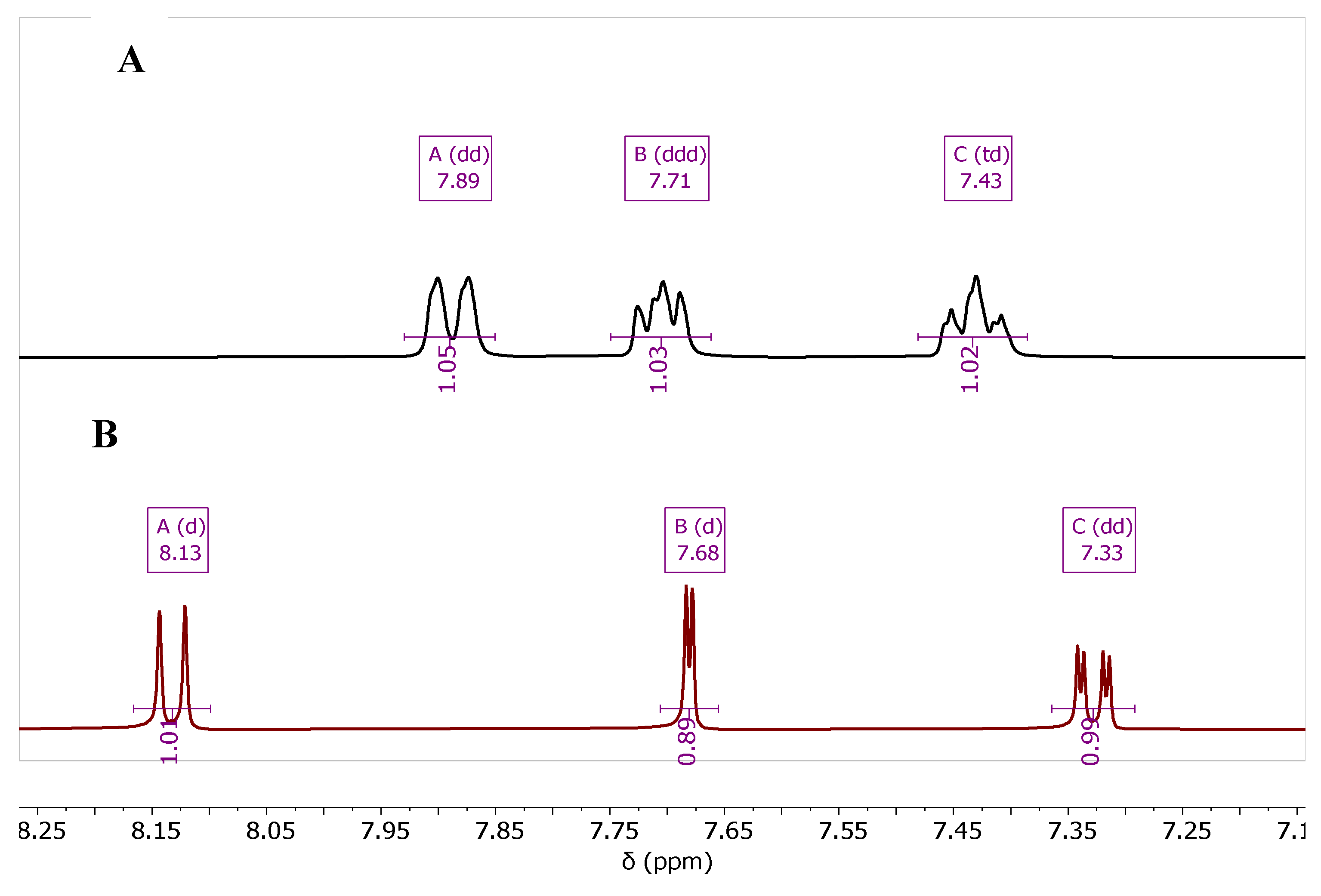
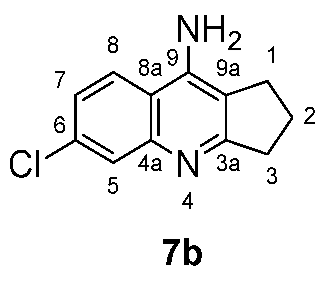
| Cmpd. # | NH2 | H-5 | H-7 | H-8 |
| 7b Zn-bound | 7.08 | 7.78 | 7.35 | 8.18 |
| 7b Zn-free | 6.57 | 7.67 | 7.31 | 8.16 |
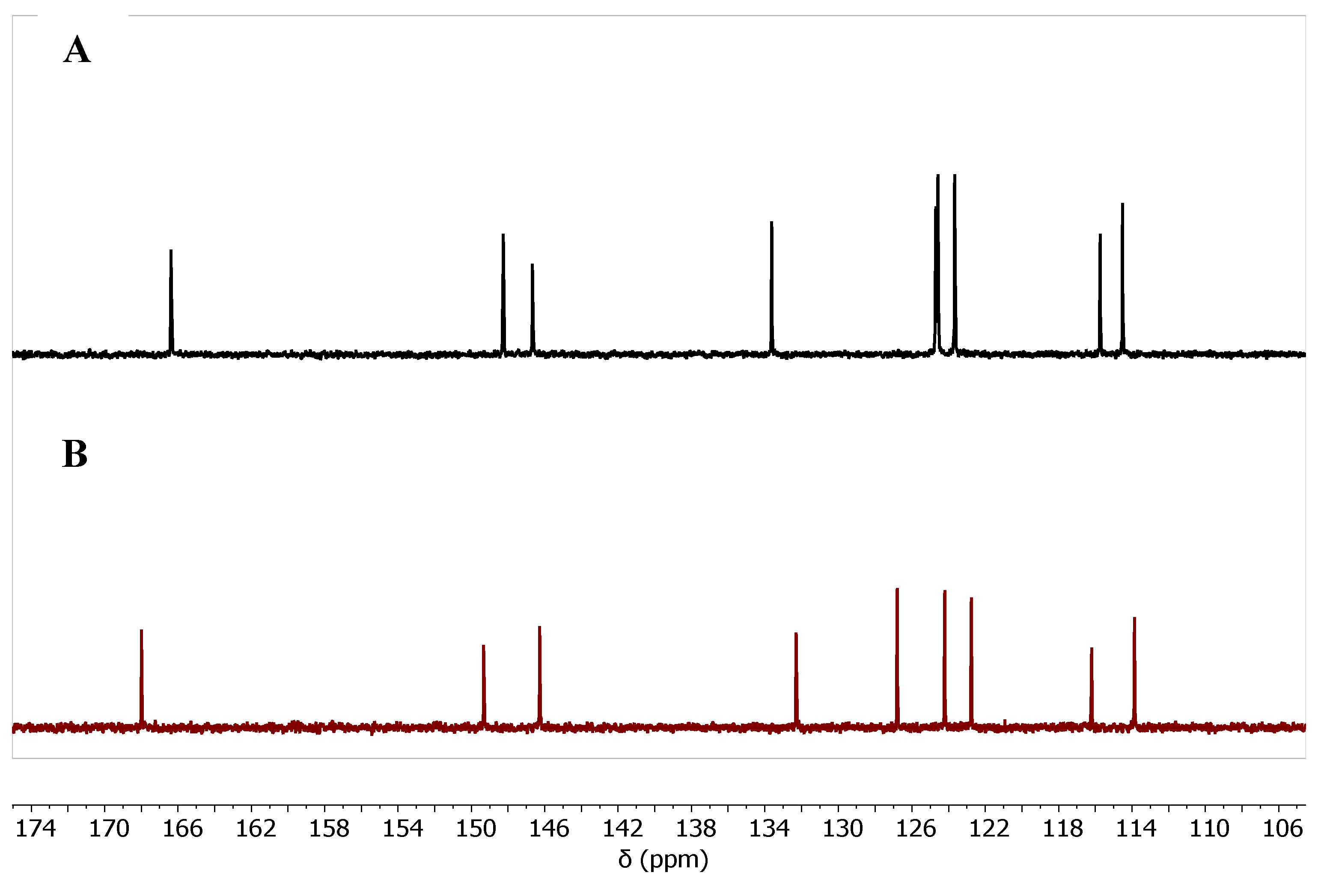
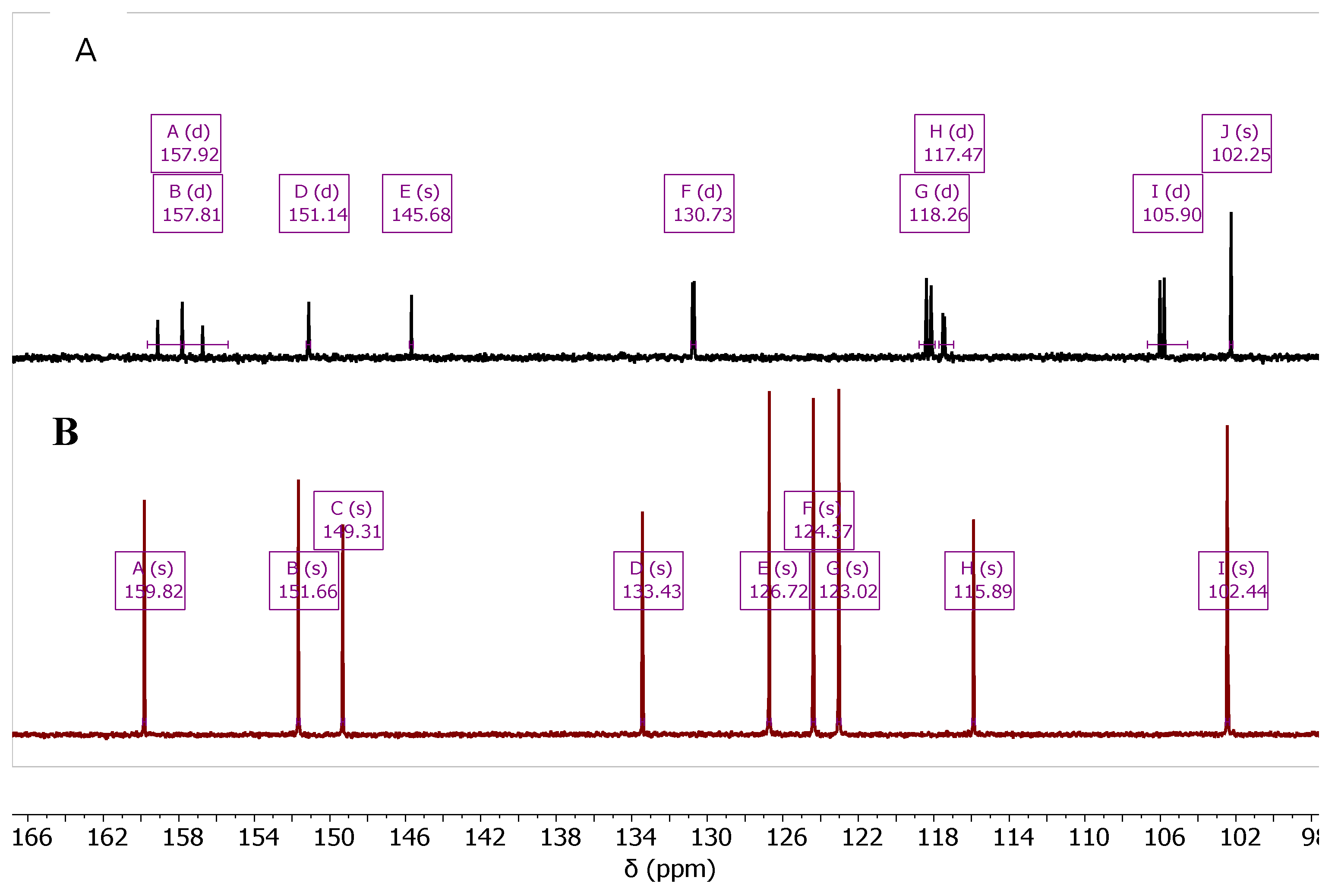
| Cmpd. # | C3a | C4a | C5 | C6 | C7 | C8 | C8a | C9 | C9a |
| 7b Zn-bound | 166.37 | 146.68 | 124.69 | 133.64 | 115.73 | 123.65 | 114.51 | 148.26 | 114.51 |
| 7b Zn-free | 168.00 | 146.28 | 126.79 | 132.29 | 122.76 | 124.21 | 116.20 | 149.33 | 113.86 |
 |
 |
| Cmpd. # | Ligscore1 dreiding | Ligscore2 dreiding | PLP1 | PLP2 | Jain | PMF | Dock Score | Ligand Internal energy |
| 7b | 1.5 | 4.03 | 59.15 | 60.01 | -0.3 | 9.93 | 34.317 | -1.916 |
| 9d | 0.69 | 2.05 | 51.16 | 65.23 | 2.15 | -16.25 | 8.764 | -1.148 |
| Cmpd. # | Ligscore1 dreiding | Ligscore2 dreiding | PLP1 | PLP2 | Jain | PMF | DockScore | Ligand Internal energy |
| 5e | 0.78 | 2.4 | 16.85 | 20.76 | -1.18 | 56.81 | 13.094 | -0.294 |
| 9d | 2.42 | 3.09 | 24.22 | 42.31 | 2.44 | 78.12 | 20.617 | -1.148 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




