Submitted:
15 January 2025
Posted:
16 January 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results
2.1. Selection of DNA Aptamers Against Human ICAM-1
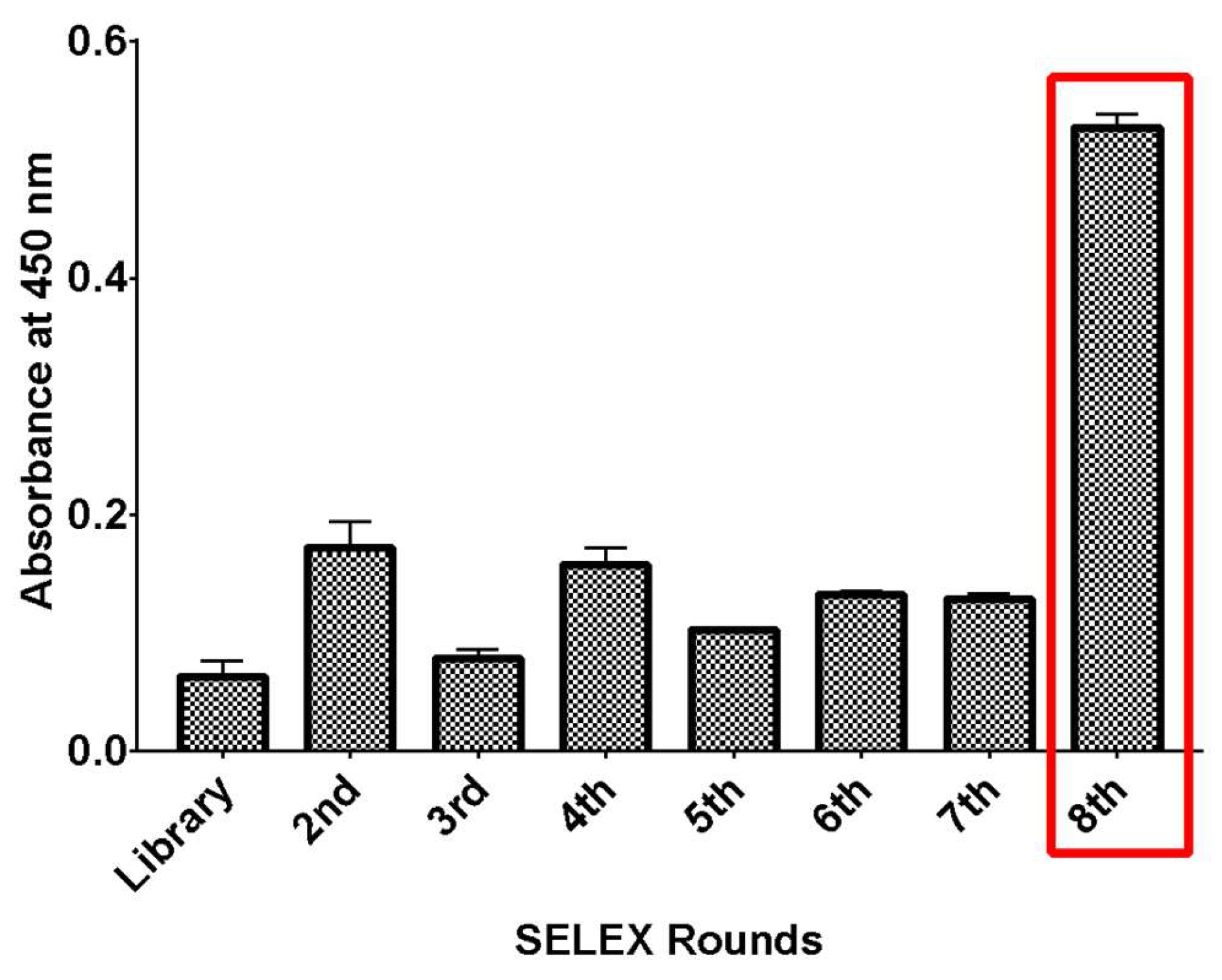
2.2. SELEX Enrichment Analysis
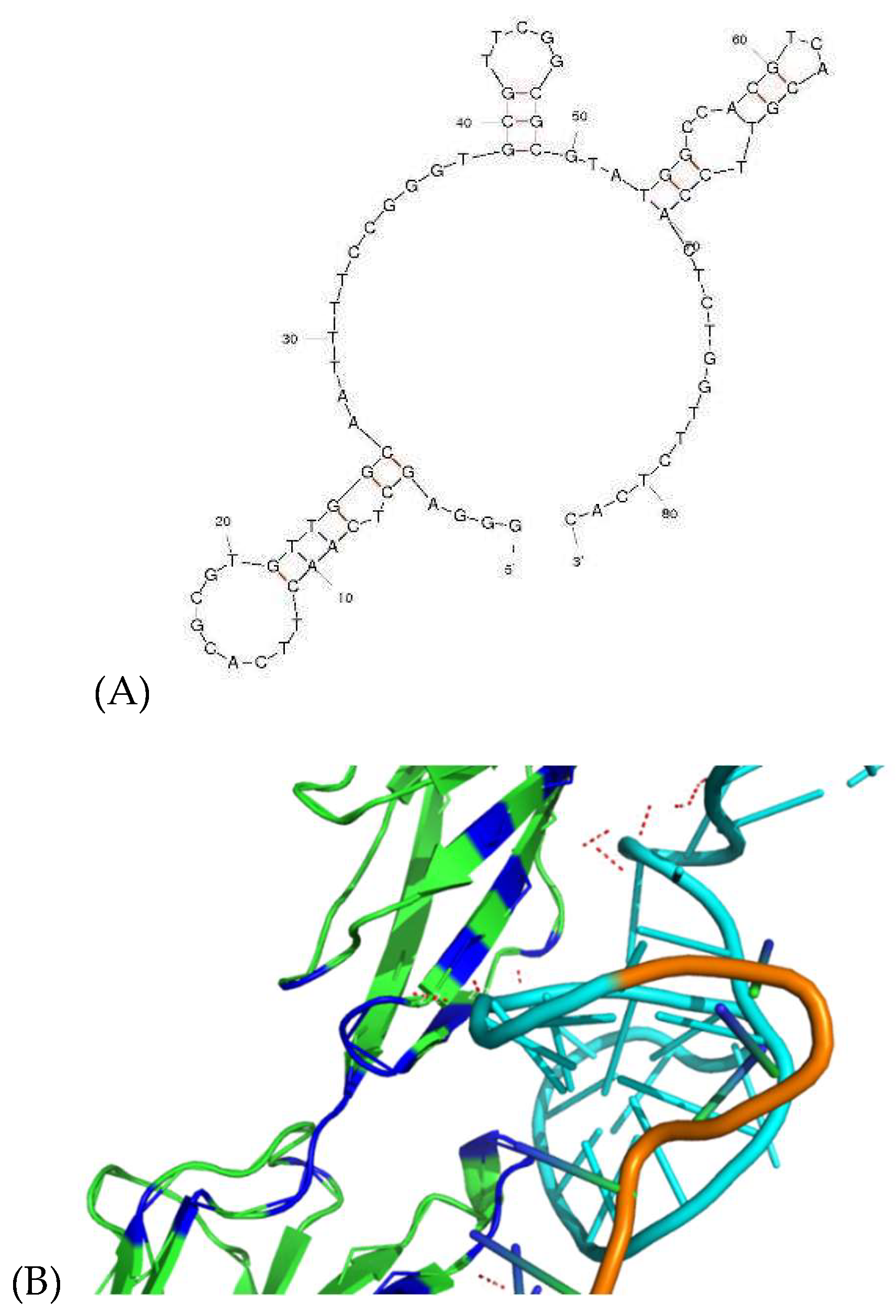
2.3. Sequence and Structure Analysis of Aptamer
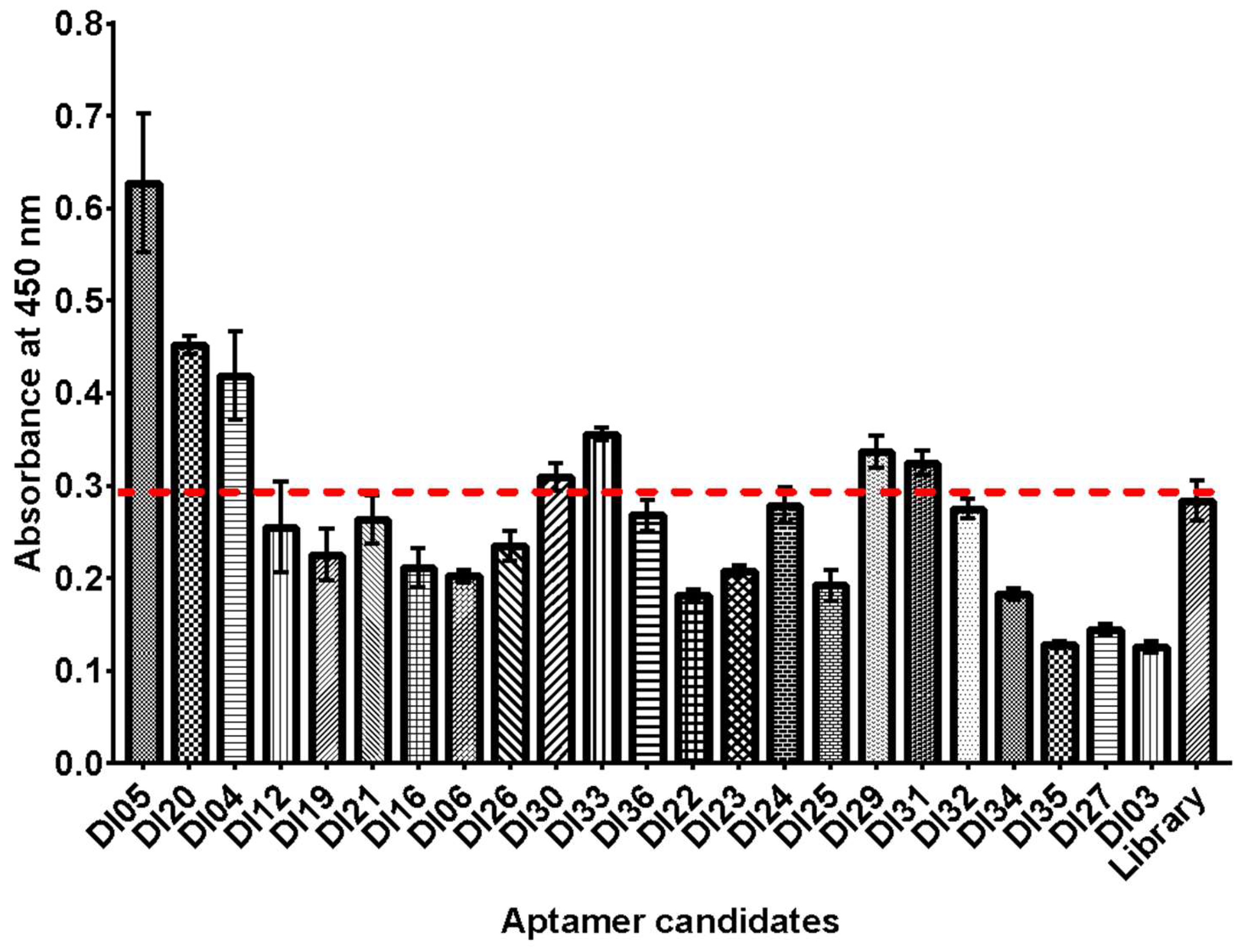
2.4. Binding Analysis of Isolated Aptamer
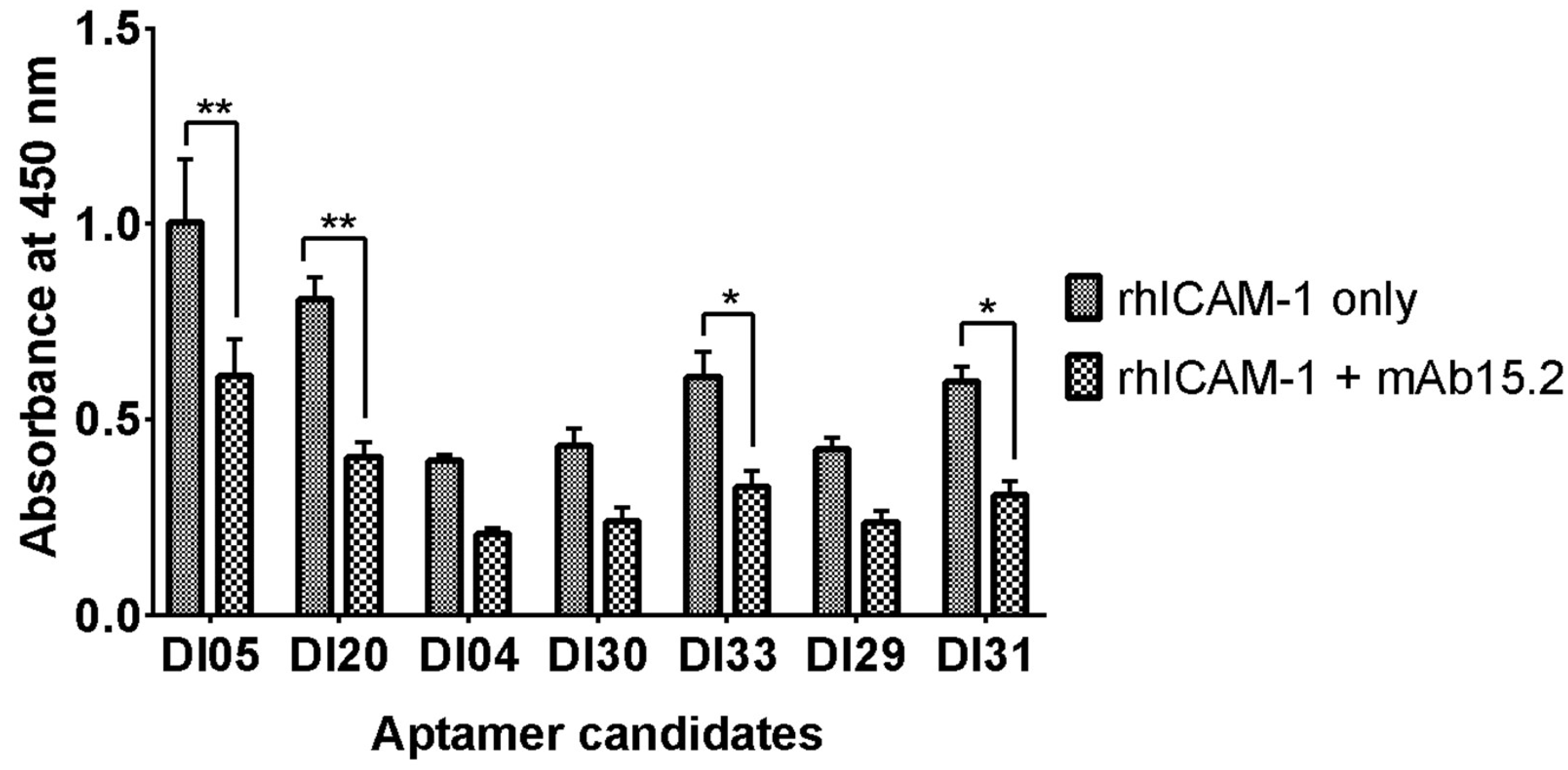
2.5. Binding Orientation of Aptamer to rhICAM-1.
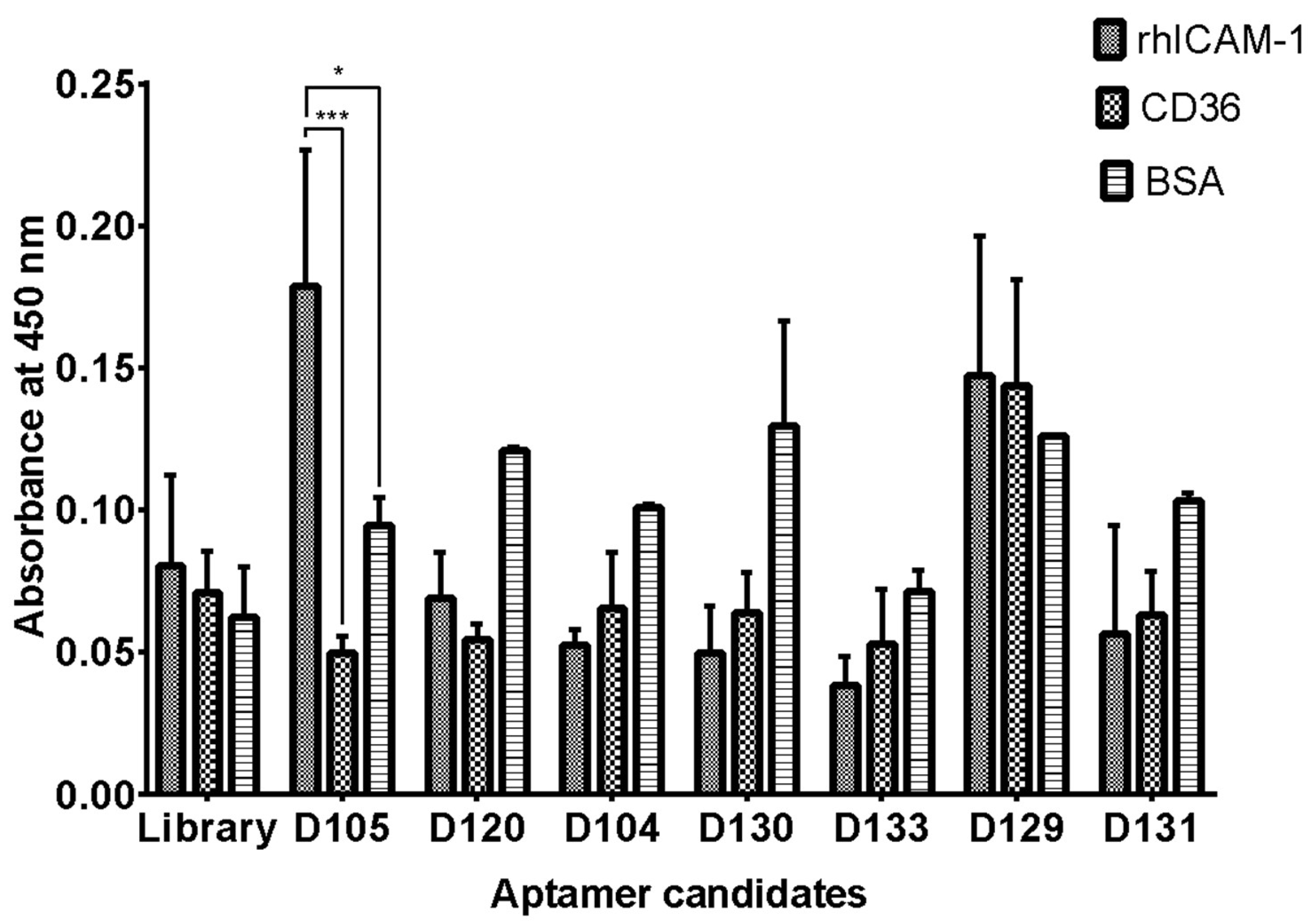
2.6. Specificity of Isolated DNA Aptamer
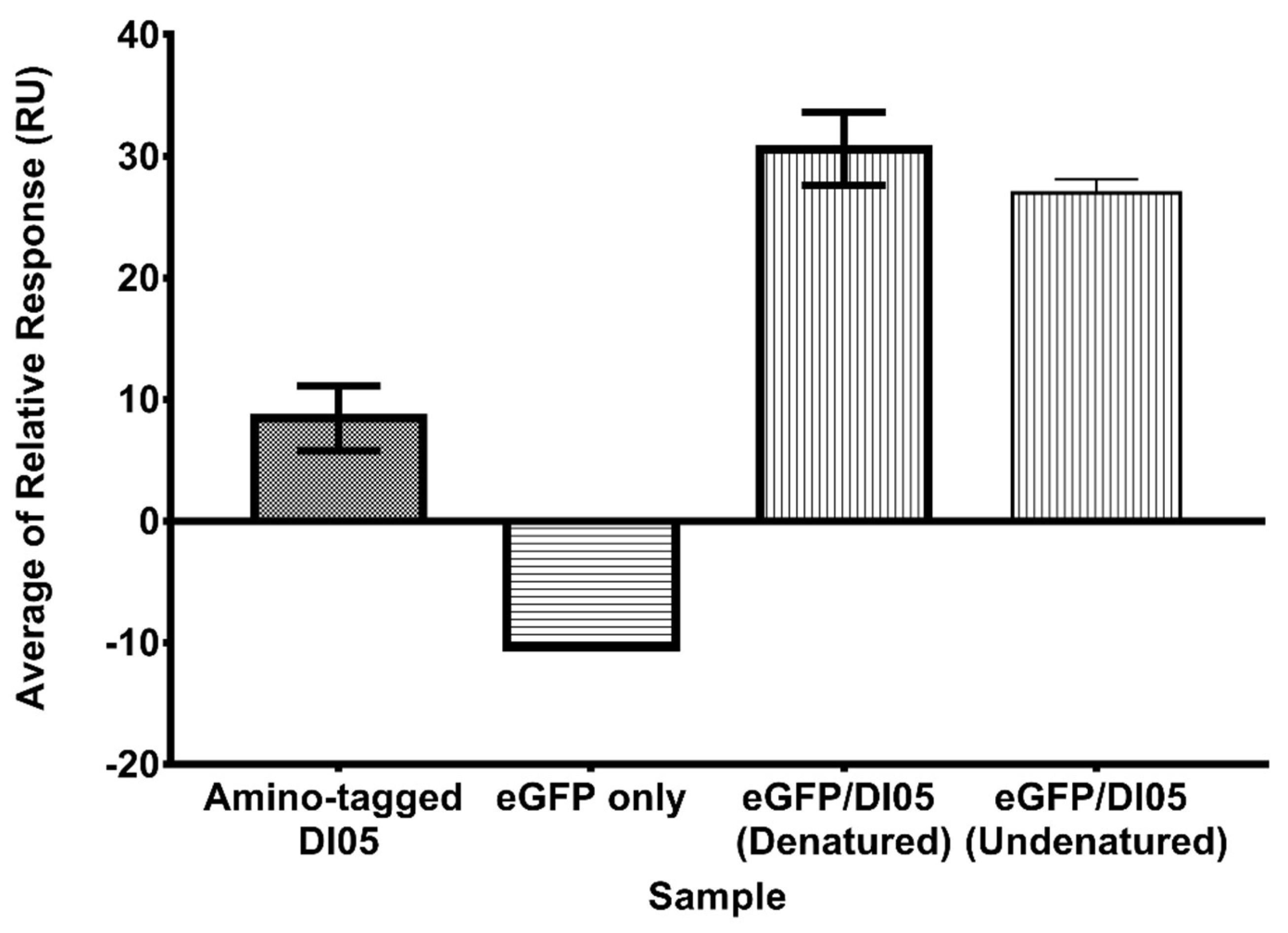
2.7. The eGFP-Conjugated Aptamer Increased the Binding Affinity
2.8. Molecular Docking of DI05 to ICAM-1 Protein Using HADDOCK
| ICAM-1 residue | DI05 residue | Type of interaction |
|---|---|---|
| Arg49 | Gua48 (2.0, 2.3, 2.5) Ade52 (2.3) | Hydrogen |
| Gua50, Cyt49 | Hydrophobic | |
| Thr20 | Tyh51 (2.7)Ade52 (2.6) | Hydrogen & Hydrophobic |
| Cyt59 | Hydrophobic | |
| Ser16, Gly14 | Tyh67 (2.7, 2.2) | Hydrogen |
| Ser7 | Thy61 = 1.8 | |
| Arg166 | Thy80 = 2.4 | Hydrogen & Hydrophobic |
| Pro45 | Gua41, Gua48 | Hydrophobic |
| Leu43 | Cyt47 | |
| Leu44 | Gua48 | |
| Arg49 | Gua49, Gua50 | |
| Ser5, Pro6 | Thy51 | |
| Val51 | Ade52 | |
| Leu18 | Gua52, Ade52 | |
| Pro167 | Cyt59 | |
| Gly169 | Ade58, Cyt59 | |
| Glu171 | Cyt57, Cyt79, Tyh78 | |
| Leu11, Val17 | Thy67 | |
| Gly15 | ||
| Cyt68 | ||
| Leu170, Ile10 | Thy66 | |
| Leu172 | Thy78 | |
| Cyt79 = 2.8 | Hydrogen (PyMOL) & Hydrophobic |
3. Discussion
4. Materials and Methods
4.1. Preparation of Selection Matrix
4.2. SELEX Experiment
4.3. Amplification of Bound Aptamer
4.4. Cloning and Sequencing
4.5. Predicting Secondary Structure
4.6. Enzyme-Linked Apta-Sorbent Assay (ELASA)
4.7. Specificity of Isolated DNA Aptamers
4.8. Generation of Protein-Aptamer Conjugate
4.9. Surface Plasmon Resonance (SPR)
4.10. Molecular Docking
4.11. Statistical Analysis
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Zhou J, Rossi J: Aptamers as targeted therapeutics: current potential and challenges. Nat Rev Drug Discov 2017, 16, 181–202. [CrossRef] [PubMed]
- Fu Z, Xiang J: Aptamers, the Nucleic Acid Antibodies, in Cancer Therapy. Int J Mol Sci 2020, 21.
- Heo K, Min S-W, Sung HJ, Kim HG, Kim HJ, Kim YH, Choi BK, Han S, Chung S, Lee ES et al: An aptamer-antibody complex (oligobody) as a novel delivery platform for targeted cancer therapies. Journal of Controlled Release 2016, 229, 1–9. [CrossRef]
- Ni S, Zhuo Z, Pan Y, Yu Y, Li F, Liu J, Wang L, Wu X, Li D, Wan Y et al: Recent Progress in Aptamer Discoveries and Modifications for Therapeutic Applications. ACS Applied Materials & Interfaces 2021, 13, 9500–9519.
- Kang S, Hah SS: Improved ligand binding by antibody-aptamer pincers. Bioconjug Chem 2014, 25, 1421–1427. [CrossRef] [PubMed]
- Skovsgaard MB, Mortensen MR, Palmfeldt J, Gothelf KV: Aptamer-Directed Conjugation of DNA to Therapeutic Antibodies. Bioconjug Chem 2019, 30, 2127–2135. [CrossRef] [PubMed]
- Yeo JE, Wickramaratne S, Khatwani S, Wang Y-C, Vervacke J, Distefano MD, Tretyakova NY: Synthesis of site-specific DNA-protein conjugates and their effects on DNA replication. ACS Chem Biol 2014, 9, 1860–1868. [CrossRef] [PubMed]
- Giepmans BNG, Adams SR, Ellisman MH, Tsien RY: The Fluorescent Toolbox for Assessing Protein Location and Function. 2006, 312, 217–224. Science, 2006; 312, 217–224.
- Silamut K, Phu NH, Whitty C, Turner GD, Louwrier K, Mai NT, Simpson JA, Hien TT, White NJ: A quantitative analysis of the microvascular sequestration of malaria parasites in the human brain. Am J Pathol 1999, 155, 395–410. [CrossRef] [PubMed]
- Willimann K, Matile H, Weiss NA, Imhof BA: In vivo sequestration of Plasmodium falciparum-infected human erythrocytes: a severe combined immunodeficiency mouse model for cerebral malaria. J Exp Med 1995, 182, 643–653. [CrossRef] [PubMed]
- Mustaffa KMF, Storm J, Whittaker M, Szestak T, Craig AG: In vitro inhibition and reversal of Plasmodium falciparum cytoadherence to endothelium by monoclonal antibodies to ICAM-1 and CD36. Malar J 2017, 16, 279. [CrossRef]
- Chames P, Van Regenmortel M, Weiss E, Baty D: Therapeutic antibodies: successes, limitations and hopes for the future. Br J Pharmacol 2009, 157, 220–233. [CrossRef]
- Arruebo M, Valladares M, González-Fernández Á: Antibody-Conjugated Nanoparticles for Biomedical Applications. Journal of Nanomaterials 2009, 2009, 439389. [CrossRef]
- Kikin O, D'Antonio L, Bagga PS: QGRS Mapper: a web-based server for predicting G-quadruplexes in nucleotide sequences. Nucleic Acids Research 2006, 34 (Suppl. 2), W676–W682. [CrossRef] [PubMed]
- Liu J, You M, Pu Y, Liu H, Ye M, Tan W: Recent developments in protein and cell-targeted aptamer selection and applications. Curr Med Chem 2011, 18, 4117–4125. [CrossRef]
- Conference Proceedings – 4th International Conference on Molecular Diagnostics and Biomarker Discovery: Antibody Technology. BMC Proceedings 2019, 13, 10. [CrossRef]
- Zumrut HE, Ara MN, Maio GE, Van NA, Batool S, Mallikaratchy PR: Ligand-guided selection of aptamers against T-cell Receptor-cluster of differentiation 3 (TCR-CD3) expressed on Jurkat. E6 cells. Anal Biochem 2016, 512, 1–7. [Google Scholar] [CrossRef]
- Wilner SE, Wengerter B, Maier K, de Lourdes Borba Magalhães M, Del Amo DS, Pai S, Opazo F, Rizzoli SO, Yan A, Levy M: An RNA alternative to human transferrin: a new tool for targeting human cells. Mol Ther Nucleic Acids 2012, 1, e21. [CrossRef] [PubMed]
- Gowda DC, Wu X: Parasite Recognition and Signaling Mechanisms in Innate Immune Responses to Malaria. Front Immunol 2018, 9, 3006. [CrossRef]
- van Buul JD, van Rijssel J, van Alphen FPJ, Hoogenboezem M, Tol S, Hoeben KA, van Marle J, Mul EPJ, Hordijk PL: Inside-out regulation of ICAM-1 dynamics in TNF-alpha-activated endothelium. PLoS One 2010, 5, e11336. [CrossRef]
- Volanti C, Gloire G, Vanderplasschen A, Jacobs N, Habraken Y, Piette J: Downregulation of ICAM-1 and VCAM-1 expression in endothelial cells treated by photodynamic therapy. Oncogene 2004, 23, 8649–8658. [CrossRef] [PubMed]
- Şener BB, Yiğit D, Bayraç AT, Bayraç C: Inhibition of cell migration and invasion by ICAM-1 binding DNA aptamers. Anal Biochem 2021, 628, 114262. [CrossRef]
- Dursun AD, Dogan S, Kavruk M, Busra Tasbasi B, Sudagidan M, Deniz Yilmaz M, Yilmaz B, Ozalp VC, Tuna BG: Surface plasmon resonance aptasensor for soluble ICAM-1 protein in blood samples. Analyst 2022, 147, 1663–1668. [CrossRef]
- Jo N, Mailhos C, Ju M, Cheung E, Bradley J, Nishijima K, Robinson GS, Adamis AP, Shima DT: Inhibition of platelet-derived growth factor B signaling enhances the efficacy of anti-vascular endothelial growth factor therapy in multiple models of ocular neovascularization. Am J Pathol 2006, 168, 2036–2053. [CrossRef]
- Lennartz F, Adams Y, Bengtsson A, Olsen RW, Turner L, Ndam NT, Ecklu-Mensah G, Moussiliou A, Ofori MF, Gamain B et al: Structure-Guided Identification of a Family of Dual Receptor-Binding PfEMP1 that Is Associated with Cerebral Malaria. Cell Host & Microbe 2017, 21, 403–414.
- Nik Abdul Aziz Nik Kamarudin, Judy Nur Aisha Sat, Nur Fatihah Mohd Zaidi, Mustaffa KMF: Evolution of specific RNA aptamers via SELEX targeted recombinant human CD36 protein: A candidate therapeutic target in severe malaria Asian Pacific Journal of Tropical Biomedicine 2019.
- Mukherjee M, Manonmani HK, Bhatt P: Aptamer as capture agent in enzyme-linked apta-sorbent assay (ELASA) for ultrasensitive detection of Aflatoxin B1. Toxicon 2018, 156, 28–33. [CrossRef] [PubMed]
- van Zundert GCP, Rodrigues JPGLM, Trellet M, Schmitz C, Kastritis PL, Karaca E, Melquiond ASJ, van Dijk M, de Vries SJ, Bonvin AMJJ: The HADDOCK2. 2 Web Server: User-Friendly Integrative Modeling of Biomolecular Complexes. Journal of Molecular Biology 2016, 428, 720–725. [Google Scholar] [CrossRef] [PubMed]
- Sabri MZ, Abdul Hamid AA, Sayed Hitam SM, Abdul Rahim MZ: In Silico Screening of Aptamers Configuration against Hepatitis B Surface Antigen. Adv Bioinformatics 2019, 2019, 6912914. [CrossRef]
- Jeddi I, Saiz L: Three-dimensional modeling of single stranded DNA hairpins for aptamer-based biosensors. Sci Rep 2017, 7, 1178. [CrossRef] [PubMed]
| SELEX Cycle |
Amount of ssDNA (pmol) | Volume of Selection Matrix (µl) | Incubation time (min) | Washing Time | Number of PCR cycles |
| 1 | 500 | 80 | 30 | 2 | 7 |
| 2 | 50 | 80 | 30 | 2 | 15 |
| 3 | 50 | 80 | 30 | 2 | 17 |
| 4 | 50 | 40 | 30 | 4 | 19 |
| 5 | 50 | 40 | 30 | 4 | 17 |
| 6 | 50 | 40 | 15 | 4 | 17 |
| 7 | 50 | 20 | 15 | 8 | 23 |
| 8 | 50 | 20 | 15 | 8 | 23 |
| Sequence ID | Sequence | Frequency | Group | Length QGRS | G-Score |
|---|---|---|---|---|---|
| DI22 | TCATAGCATTGAGCTATGCTGCGTTAAGCATGAAAAATAT | 1/25 | A/T-rich family | - | - |
| DI05 / DI15 | TTGGCAATTTTCCGGGTGCGTTCGGCGCGTATGGCCACGT | 2/25 | G/C-rich family | 32 | 29 |
| DI12 | GTTGGGTATTATGGGGTGGGTGGTTGGGGGTTGGTTTGGT | 1/25 | G-rich family | 25 | 58 |
| DI24 | ATGGGTAAAAACTGGGGGTGGGTGGTGGGGGTTGGTTGGG | 1/25 | G-rich family | 38 | 61 |
| DI31 | CCAGAGTAGCTCAGGGGTTGGGGTTAGGGGCTTGGTCCTT | 1/25 | G-rich family | 20 | 31 |
| DI03 | AAGGTTCATTGGGGGTGGGCTGGTTGGGGGTTGGTCGGGT | 1/25 | G-rich family | 29 | 58 |
| DI04 | CCGCTTTCCAACAGCAACAGAATCCCACGCAAGAACCATC | 1/25 | Other | - | - |
| DI19 | TGGCTTAGGGCCAGACATAATTCAACACTTGGCATATATA | 1/25 | Other | - | - |
| DI21 | CGTCCCCCTCGCATTGGGCATCCACCATAACCTTCTTCTC | 1/25 | Other | - | - |
| DI16 | CCTACCCTTAGACACTAACCCACAGTCCGTAGTCAGACTC | 1/25 | Other | - | - |
| DI26 | AGTCCTACGCCCCCAACGGTATAGAGTGAAACACAAGAA | 1/25 | Other | - | - |
| DI30 | AGCTTTCACAGAATCTGACCCCGCCGTAGATTGTCACAAAGTC | 1/25 | Other | - | - |
| DI36 | TTTACCCTATAATGTCCCAATTACCACGATCCTTAC | 1/25 | Other | - | - |
| DI29 | GCATTCTACTACATAAAACTATTGGATCTAATCTGACCTC | 1/25 | Other | - | - |
| DI32 | TTGCACTCTCCTAACTACAACTTGCCCTTGGATTCTGTGT | 1/25 | Other | - | - |
| DI34 | AACTATTGGCTTCCATTTGAACATGGAAAAAACTGGGCTG | 1/25 | Other | - | - |
| DI06 | AACCCCATGCGTCACCTACTCAGTTGTTGTCACCTCTCTG | 1/25 | Other | - | - |
| DI27 | AATCGCTGACGGCTTGGGGGCTTTGTCTAAAGGGGGAT | 1/25 | T/G-rich family | 23 | 21 |
| DI23 | TTTCATGCCCGAAGTAGGTTGGGTAGGCTTGGTCTTCGTG | 1/25 | T/G-rich family | 16 | 30 |
| DI25 | AGCAACGGCTTAGCGGGGTTTCATTATTTTCTTGCTTCTA | 1/25 | T-rich family | - | - |
| DI35 | AATTGTTCTTAACGTTGCAGCGGACTAATTTTGTGTCTGA | 1/25 | T-rich family | - | - |
| DI33 | GGTCAATTTTGGCGTTTTAACCTTACCGCGACCTTCCTTC | 1/25 | T-rich family | - | - |
| DI20 / DI28 | TGTCAGGCTGGACGACTAGATCAGTTTCGTTGTGCTATCT | 2/25 | T-rich family | - | - |
| KD (M) | Rmax (RU) | offset (RU) | Chi² (RU²) | |
| 108 ± 22.69 nM | 22.86 | -5.604 | 0.376 | eGFP-conjugated DI05 |
| 252 ± 46.26 nM | 61.19 | 3.417 | 1 | Unconjugated DI05 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




