Submitted:
06 January 2025
Posted:
07 January 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results
2.1. Clinical Profile of the Patients
2.2. Overview of Analysis Results for Each Library
2.3. Variants Detected in Tumor Suppressor Genes
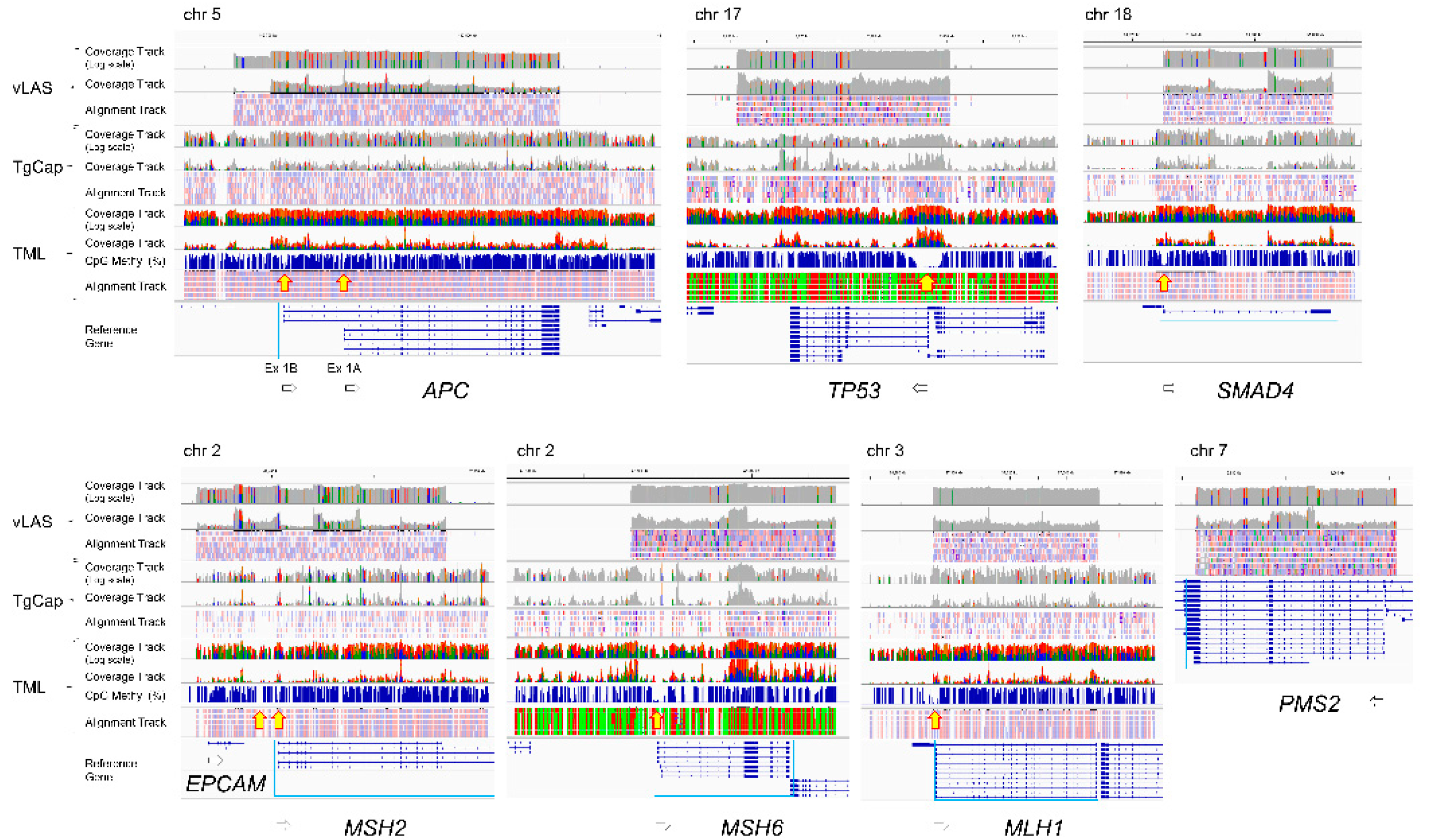
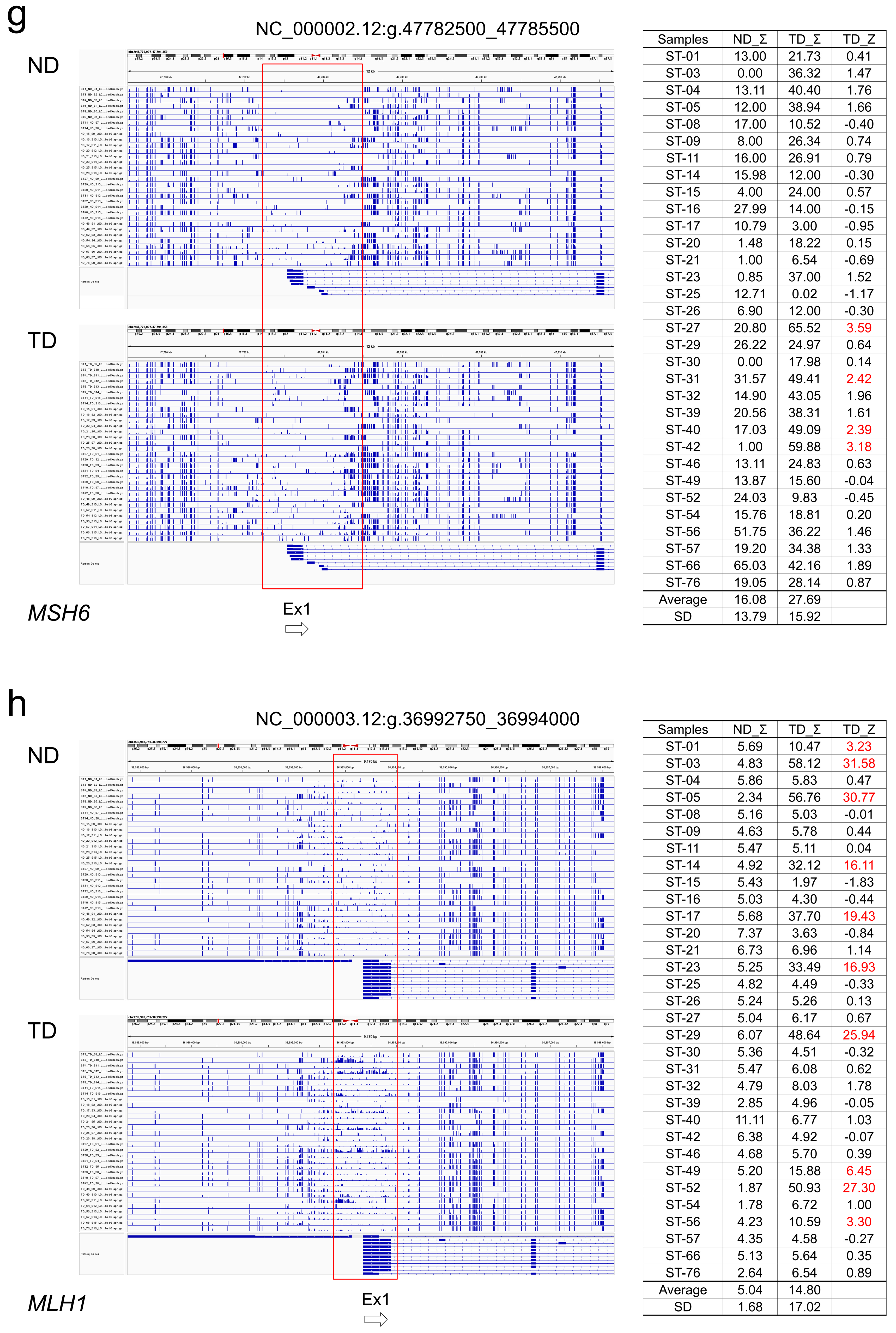
2.4. CpG Methylation Alterations in Tumor and Normal Colon Tissue
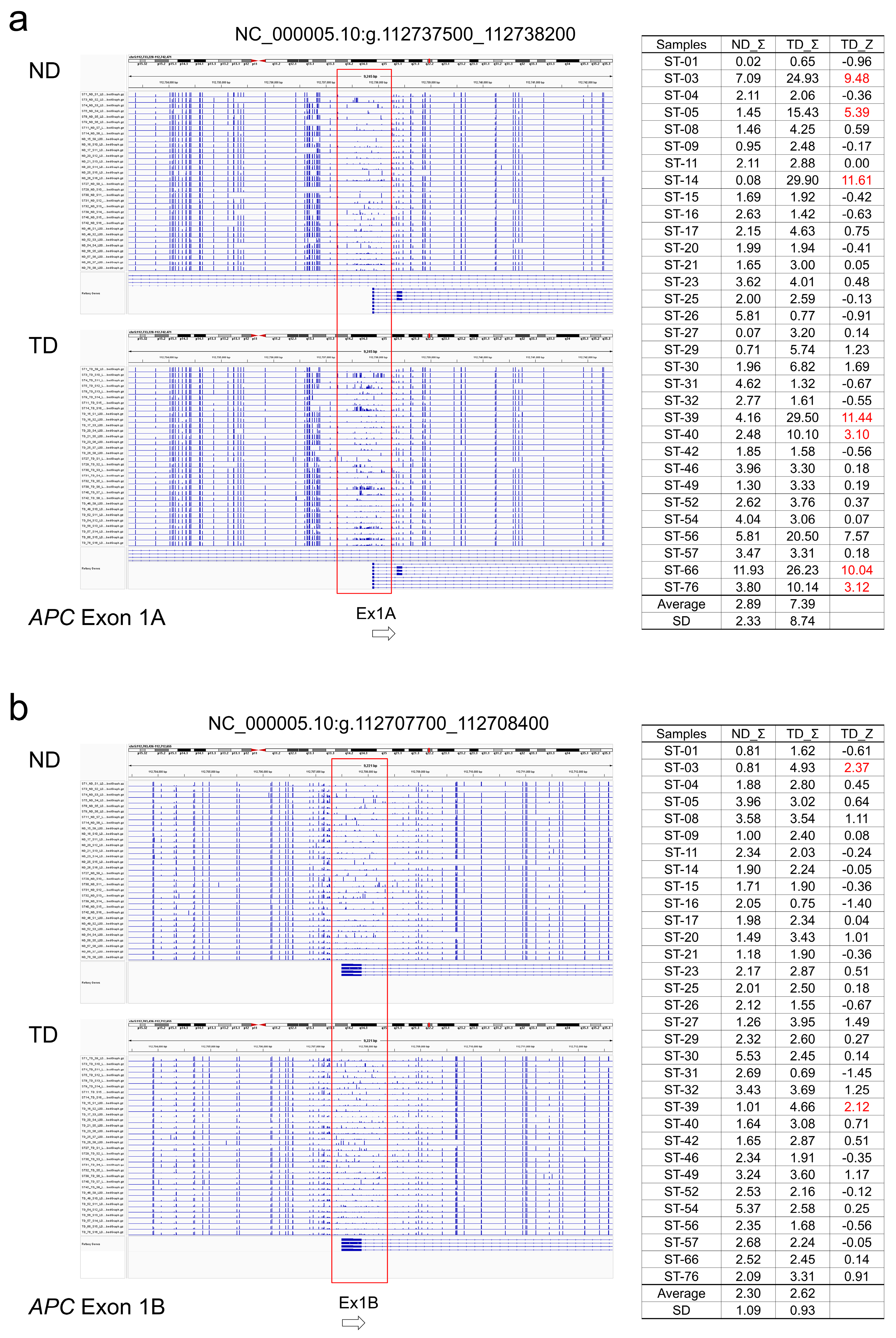
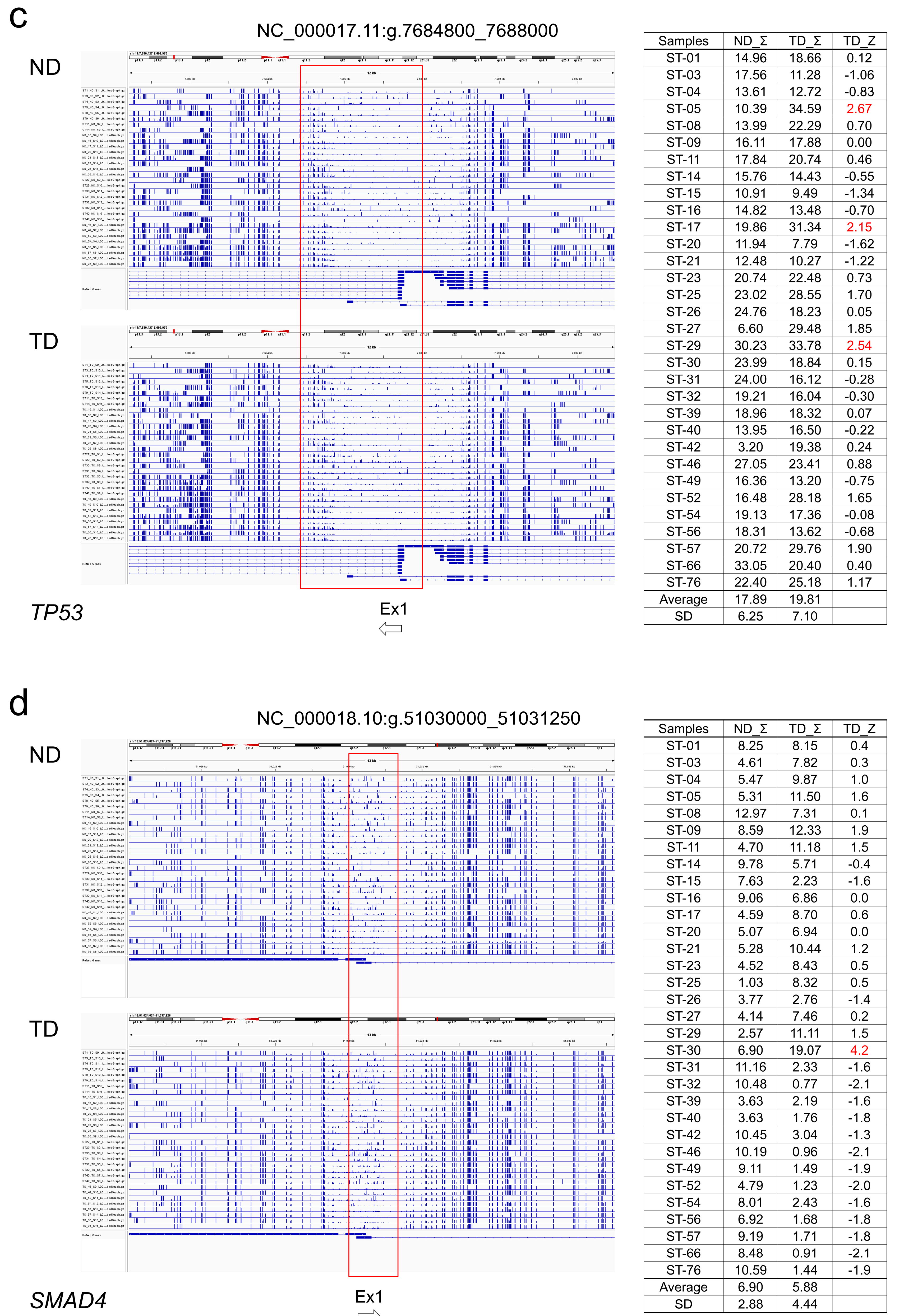
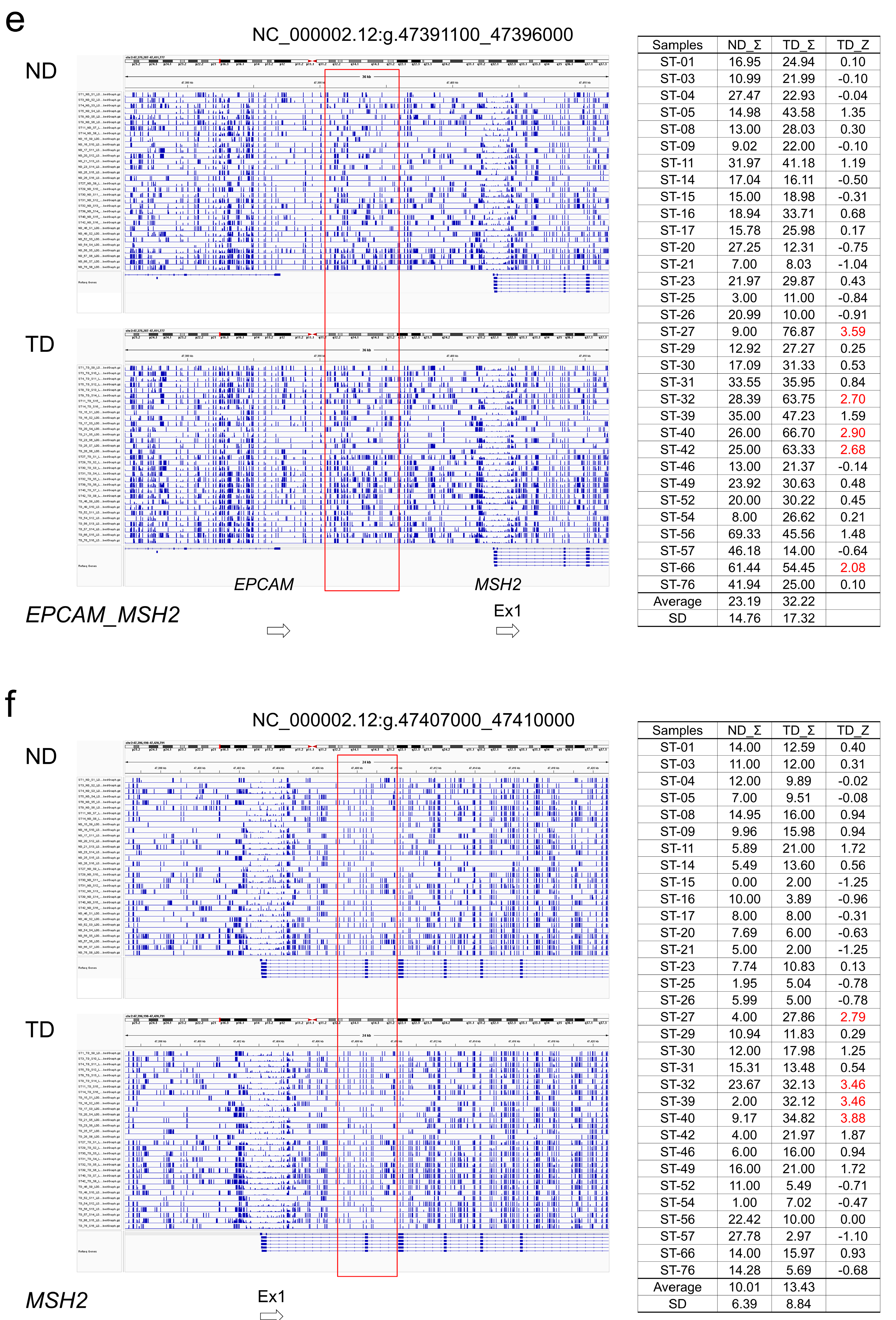
2.5. Association of Somatic DNA Variants with CpG Methylation in Tumor Suppressor Genes
3. Discussion
3.1. DNA Variants in APC, TP53, SMAD4 and MMR Genes
3.2. DNA Methylation in Tumor Suppression Genes
3.3. Association of DNA Variants with CpG Methylation in Tumor Suppressor Genes
3.4. Limitation of This Study
4. Materials and Methods
4.1. Patient and Sample Collection
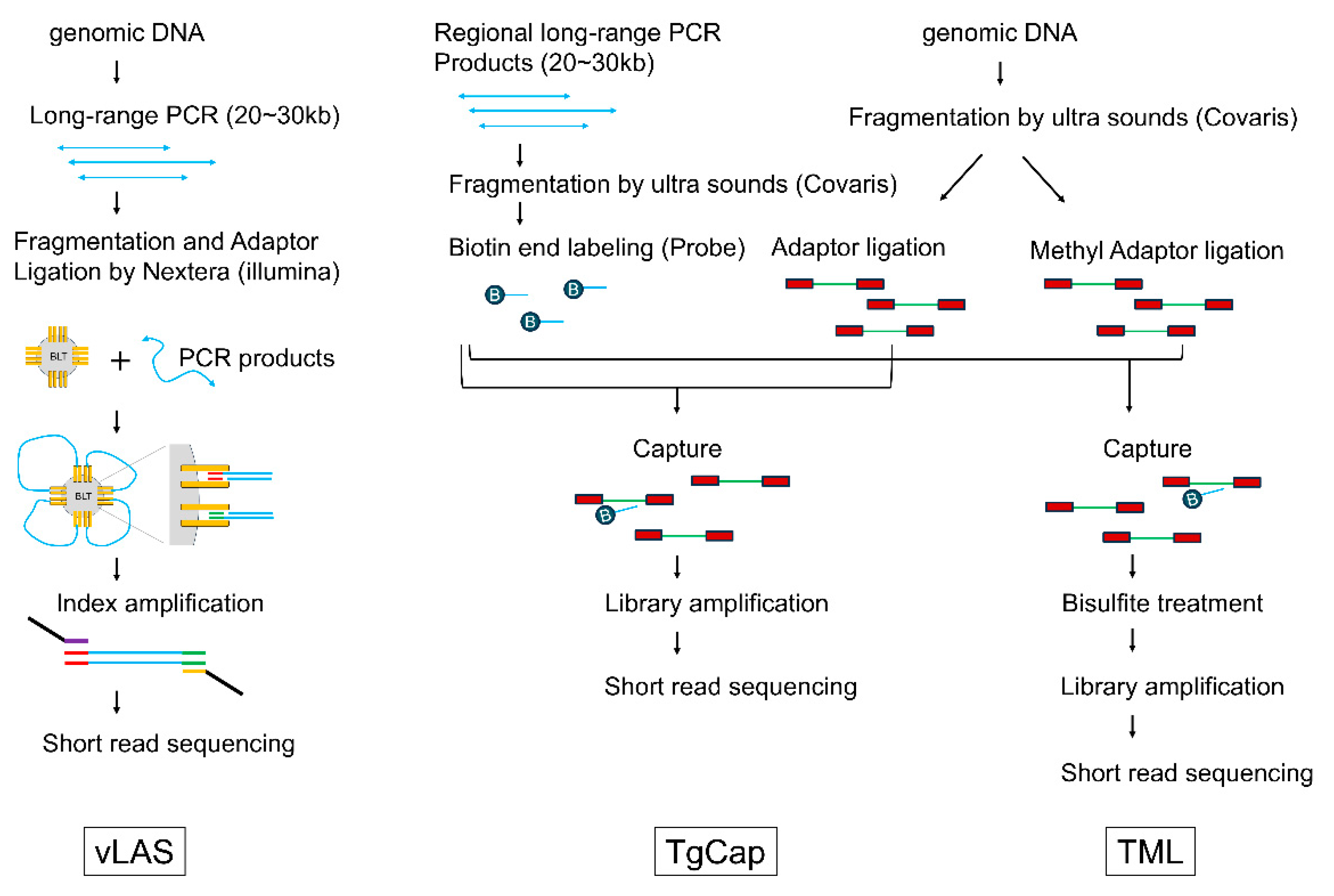
4.2. Library Preparations and Sequencing
4.3. DATA Analysis Pipeline
4.4. Annotation of Variants
4.5. Evaluation of CpG Methylation
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| LOH | loss of heterozygosity |
| vLAS | very long amplicon sequencing |
| NGS | next-generation sequencing |
| TgCap | targeted genome capture |
| TML | targeted methyl landscape |
| MSI-H | microsatellite instability high |
| MSS | Microsatellite stable |
| VAF | variant allele frequency |
| MMR | mismatch repair |
References
- Carnevali, I.W.; Cini, G.; Libera, L.; Sahnane, N.; Facchi, S.; Viel, A.; Sessa, F.; Tibiletti, M.G. , Promoter Methylation Could Be the Second Hit in Lynch Syndrome Carcinogenesis. Genes (Basel) 2023, 14. [Google Scholar] [CrossRef] [PubMed]
- Di Ruscio, A.; Welner, R.S.; Tenen, D.G.; Amabile, G. , The second hit of DNA methylation. Mol Cell Oncol 2016, 3, e1093690. [Google Scholar] [CrossRef] [PubMed]
- Yamamoto, E.; Suzuki, H.; Takamaru, H.; Yamamoto, H.; Toyota, M.; Shinomura, Y. , Role of DNA methylation in the development of diffuse-type gastric cancer. Digestion 2011, 83, 241–249. [Google Scholar] [CrossRef] [PubMed]
- Segditsas, S.; Sieber, O.M.; Rowan, A.; Setien, F.; Neale, K.; Phillips, R.K.; Ward, R.; Esteller, M.; Tomlinson, I.P. , Promoter hypermethylation leads to decreased APC mRNA expression in familial polyposis and sporadic colorectal tumours, but does not substitute for truncating mutations. Exp Mol Pathol 2008, 85, 201–206. [Google Scholar] [CrossRef]
- Ding, Z.; Jiang, T.; Piao, Y.; Han, T.; Han, Y.; Xie, X. , Meta-analysis of the association between APC promoter methylation and colorectal cancer. Onco Targets Ther 2015, 8, 211–222. [Google Scholar]
- Poosari, A.; Nutravong, T.; Namwat, W.; Wasenang, W.; Sa-Ngiamwibool, P.; Ungareewittaya, P. , The relationship between P16. Scientific reports 2022, 12, 10337. [Google Scholar]
- Herrero, A.B.; Rojas, E.A.; Misiewicz-Krzeminska, I.; Krzeminski, P.; Gutiérrez, N.C. , Molecular Mechanisms of p53 Deregulation in Cancer: An Overview in Multiple Myeloma. Int J Mol Sci 2016, 17. [Google Scholar] [CrossRef]
- Li, J.; Ma, X.; Chakravarti, D.; Shalapour, S.; DePinho, R.A. , Genetic and biological hallmarks of colorectal cancer. Genes Dev 2021, 35, (11–12), 787. [Google Scholar] [CrossRef]
- Fodde, R. , The APC gene in colorectal cancer. Eur J Cancer 2002, 38, 867–871. [Google Scholar] [CrossRef]
- Zhu, G.; Pei, L.; Xia, H.; Tang, Q.; Bi, F. , Role of oncogenic KRAS in the prognosis, diagnosis and treatment of colorectal cancer. Mol Cancer 2021, 20, 143. [Google Scholar] [CrossRef]
- Müller, M.F.; Ibrahim, A.E.; Arends, M.J. , Molecular pathological classification of colorectal cancer. Virchows Arch 2016, 469, 125–134. [Google Scholar] [CrossRef] [PubMed]
- Molnár, B.; Galamb, O.; Péterfia, B.; Wichmann, B.; Csabai, I.; Bodor, A.; Kalmár, A.; Szigeti, K.A.; Barták, B.K.; Nagy, Z.B.; Valcz, G.; Patai, Á.; Igaz, P.; Tulassay, Z. , Gene promoter and exon DNA methylation changes in colon cancer development - mRNA expression and tumor mutation alterations. BMC Cancer 2018, 18, 695. [Google Scholar] [CrossRef] [PubMed]
- Tomicic, M.T.; Dawood, M.; Efferth, T. , Epigenetic Alterations Upstream and Downstream of p53 Signaling in Colorectal Carcinoma. Cancers (Basel) 2021, 13. [Google Scholar] [CrossRef] [PubMed]
- Dobre, M.; Salvi, A.; Pelisenco, I.A.; Vasilescu, F.; De Petro, G.; Herlea, V.; Milanesi, E. , Crosstalk Between DNA Methylation and Gene Mutations in Colorectal Cancer. Front Oncol 2021, 11, 697409. [Google Scholar] [CrossRef] [PubMed]
- Monti, P.; Perfumo, C.; Bisio, A.; Ciribilli, Y.; Menichini, P.; Russo, D.; Umbach, D.M.; Resnick, M.A.; Inga, A.; Fronza, G. , Dominant-negative features of mutant TP53 in germline carriers have limited impact on cancer outcomes. Mol Cancer Res 2011, 9, 271–279. [Google Scholar] [CrossRef]
- Alankarage, D.; Enriquez, A.; Steiner, R.D.; Raggio, C.; Higgins, M.; Milnes, D.; Humphreys, D.T.; Duncan, E.L.; Sparrow, D.B.; Giampietro, P.F.; Chapman, G.; Dunwoodie, S.L. , Myhre syndrome is caused by dominant-negative dysregulation of SMAD4 and other co-factors. Differentiation 2022, 128, 1–12. [Google Scholar] [CrossRef]
- Wei, Y.; Sun, Z.; Wang, Y.; Xie, Z.; Xu, S.; Xu, Y.; Zhou, X.; Bi, J.; Zhu, Z. , Methylation in the TP53 promoter is associated with ischemic stroke. Mol Med Rep 2019, 20, 1404–1410. [Google Scholar] [CrossRef]
- Yuan, L.; Wang, L.; Du, X.; Qin, L.; Yang, M.; Zhou, K.; Wu, M.; Yang, Y.; Zheng, Z.; Xiang, Y.; Qu, X.; Liu, H.; Qin, X.; Liu, C. , The DNA methylation of FOXO3 and TP53 as a blood biomarker of late-onset asthma. J Transl Med 2020, 18, 467. [Google Scholar] [CrossRef]
- Ura, H.; Togi, S.; Niida, Y. , Targeted Double-Stranded cDNA Sequencing-Based Phase Analysis to Identify Compound Heterozygous Mutations and Differential Allelic Expression. Biology (Basel) 2021, 10. [Google Scholar] [CrossRef]
- Ura, H.; Togi, S.; Niida, Y. , Target-capture full-length double-strand cDNA sequencing for alternative splicing analysis. RNA Biol 2021, 1–8. [Google Scholar] [CrossRef]
- Togi, S.; Ura, H.; Niida, Y. , Optimization and Validation of Multimodular, Long-Range PCR-Based Next-Generation Sequencing Assays for Comprehensive Detection of Mutation in Tuberous Sclerosis Complex. J Mol Diagn 2021, 23, 424–446. [Google Scholar] [CrossRef] [PubMed]
- Niida, Y.; Togi, S.; Ura, H. , Streamlining Genetic Diagnosis With Long-Range Polymerase Chain Reaction (PCR)-Based Next-Generation Sequencing for Type I and Type II Collagenopathies. Cureus 2023, 15, e50482. [Google Scholar] [CrossRef] [PubMed]
- Nomura, F.; Shimizu, A.; Togi, S.; Ura, H.; Niida, Y. , SNP Array Screening and Long Range PCR-Based Targeted Next Generation Sequencing for Autosomal Recessive Disease with Consanguinity: Insight from a Case of Xeroderma Pigmentosum Group C. Genes (Basel) 2023, 14. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Durbin, R. , Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics 2010, 26, 589–595. [Google Scholar] [CrossRef]
- Cibulskis, K.; Lawrence, M.S.; Carter, S.L.; Sivachenko, A.; Jaffe, D.; Sougnez, C.; Gabriel, S.; Meyerson, M.; Lander, E.S.; Getz, G. , Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nat Biotechnol 2013, 31, 213–219. [Google Scholar] [CrossRef]
- Wilm, A.; Aw, P.P.; Bertrand, D.; Yeo, G.H.; Ong, S.H.; Wong, C.H.; Khor, C.C.; Petric, R.; Hibberd, M.L.; Nagarajan, N. , LoFreq: a sequence-quality aware, ultra-sensitive variant caller for uncovering cell-population heterogeneity from high-throughput sequencing datasets. Nucleic Acids Res 2012, 40, 11189–11201. [Google Scholar] [CrossRef]
- Van der Auwera, G.A.; Carneiro, M.O.; Hartl, C.; Poplin, R.; Del Angel, G.; Levy-Moonshine, A.; Jordan, T.; Shakir, K.; Roazen, D.; Thibault, J.; Banks, E.; Garimella, K.V.; Altshuler, D.; Gabriel, S.; DePristo, M.A. , From FastQ data to high confidence variant calls: the Genome Analysis Toolkit best practices pipeline. Curr Protoc Bioinformatics 2013, 43, 11.10.1–111033. [Google Scholar] [CrossRef]
- Cingolani, P.; Platts, A.; Wang le, L.; Coon, M.; Nguyen, T.; Wang, L.; Land, S.J.; Lu, X.; Ruden, D.M. , A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3. Fly 2012, 6, 80–92. [Google Scholar] [CrossRef]
- Sherry, S.T.; Ward, M.H.; Kholodov, M.; Baker, J.; Phan, L.; Smigielski, E.M.; Sirotkin, K. , dbSNP: the NCBI database of genetic variation. Nucleic Acids Res 2001, 29, 308–311. [Google Scholar] [CrossRef]
- Landrum, M.J.; Lee, J.M.; Riley, G.R.; Jang, W.; Rubinstein, W.S.; Church, D.M.; Maglott, D.R. , ClinVar: public archive of relationships among sequence variation and human phenotype. Nucleic Acids Res 2014, 42, (Database issue), D980–5. [Google Scholar] [CrossRef]
- Ye, K.; Schulz, M.H.; Long, Q.; Apweiler, R.; Ning, Z. , Pindel: a pattern growth approach to detect break points of large deletions and medium sized insertions from paired-end short reads. Bioinformatics 2009, 25, 2865–2871. [Google Scholar] [CrossRef] [PubMed]
- Krueger, F.; Andrews, S.R. , Bismark: a flexible aligner and methylation caller for Bisulfite-Seq applications. Bioinformatics 2011, 27, 1571–1572. [Google Scholar] [CrossRef] [PubMed]
- de Sainte Agathe, J.M.; Filser, M.; Isidor, B.; Besnard, T.; Gueguen, P.; Perrin, A.; Van Goethem, C.; Verebi, C.; Masingue, M.; Rendu, J.; Cossée, M.; Bergougnoux, A.; Frobert, L.; Buratti, J.; Lejeune, É.; Le Guern, É.; Pasquier, F.; Clot, F.; Kalatzis, V.; Roux, A.F.; Cogné, B.; Baux, D. , SpliceAI-visual: a free online tool to improve SpliceAI splicing variant interpretation. Human genomics 2023, 17, 7. [Google Scholar] [CrossRef]
- Thorvaldsdottir, H.; Robinson, J.T.; Mesirov, J.P. , Integrative Genomics Viewer (IGV): high-performance genomics data visualization and exploration. Briefings in bioinformatics 2013, 14, 178–192. [Google Scholar] [CrossRef] [PubMed]
- Horak, P.; Griffith, M.; Danos, A.M.; Pitel, B.A.; Madhavan, S.; Liu, X.; Chow, C.; Williams, H.; Carmody, L.; Barrow-Laing, L.; Rieke, D.; Kreutzfeldt, S.; Stenzinger, A.; Tamborero, D.; Benary, M.; Rajagopal, P.S.; Ida, C.M.; Lesmana, H.; Satgunaseelan, L.; Merker, J.D.; Tolstorukov, M.Y.; Campregher, P.V.; Warner, J.L.; Rao, S.; Natesan, M.; Shen, H.; Venstrom, J.; Roy, S.; Tao, K.; Kanagal-Shamanna, R.; Xu, X.; Ritter, D.I.; Pagel, K.; Krysiak, K.; Dubuc, A.; Akkari, Y.M.; Li, X.S.; Lee, J.; King, I.; Raca, G.; Wagner, A.H.; Li, M.M.; Plon, S.E.; Kulkarni, S.; Griffith, O.L.; Chakravarty, D.; Sonkin, D. , Standards for the classification of pathogenicity of somatic variants in cancer (oncogenicity): Joint recommendations of Clinical Genome Resource (ClinGen), Cancer Genomics Consortium (CGC), and Variant Interpretation for Cancer Consortium (VICC). Genet Med 2022, 24, 986–998. [Google Scholar] [CrossRef]
| Patient | Age | Sex | Locus | KRAS | BRAF | MSI |
|---|---|---|---|---|---|---|
| ST-01 | 62 | F | T | - | - | MSS |
| ST-03 | 79 | M | R | G12D | - | MSS |
| ST-04 | 55 | F | S | G13D | - | MSS |
| ST-05 | 90 | F | T | - | - | MSS |
| ST-08 | 86 | F | C | - | - | MSS |
| ST-09 | 73 | M | A | G12D | - | MSS |
| ST-11 | 67 | M | R | - | - | MSS |
| ST-14 | 67 | F | S | G12D | - | MSS |
| ST-15 | 80 | M | C | G13D | - | MSS |
| ST-16 | 70 | M | D | - | - | MSS |
| ST-17 | 84 | M | A | - | V600E | MSI-H |
| ST-20 | 81 | F | C | G13D | - | MSS |
| ST-21 | 64 | F | S | - | - | MSS |
| ST-23 | 85 | F | A | - | V600E | MSI-H |
| ST-25 | 64 | M | R | - | - | MSS |
| ST-26 | 74 | F | A | - | - | MSS |
| ST-27 | 77 | M | D | G12D | - | MSS |
| ST-29 | 74 | M | AR | - | - | MSI-H |
| ST-30 | 61 | F | A | G13C | - | MSS |
| ST-31 | 84 | M | A | G12V | - | MSS |
| ST-32 | 55 | F | A | G12D | - | MSS |
| ST-39 | 73 | F | T | - | - | MSS |
| ST-40 | 85 | M | R | - | V600E | MSS |
| ST-42 | 83 | F | C | G13C | - | MSS |
| ST-46 | 68 | M | D | G12D | - | MSS |
| ST-49 | 82 | F | A | - | V600E | MSI-H |
| ST-52 | 88 | M | S | - | - | MSS |
| ST-54 | 75 | M | S+T(LST) | - | - | MSS |
| ST-56 | 70 | M | S+ML | - | - | MSS |
| ST-57 | 48 | F | R | - | - | MSI-H |
| ST-66 | 64 | F | R+LM | - | - | MSS |
| ST-76 | 59 | M | S | - | - | MSS |
| Tumor | APC | TP53 | SMAD4 | ||||
| DNA | Variants | MZS | MZS | Variants | MZS | Variants | MZS |
| Sample | (VAF) | Ex1A | Ex1B | (VAF) | (VAF) | ||
| ST-01 | R252* (0.33) N1818fs (0.40) |
- | - | R209Q (0.13) LO, DN | - | - | - |
| ST-03 | - | VH | H | Y181N (0.25) V | - | - | - |
| ST-04 | E763* (0.34) | - | - | R234H (0.51) LO, DN | - | - | - |
| ST-05 | R71C (0.10) V S457* (0.09) T1556fs (0.04) |
H | - | - | H | - | - |
| ST-08 | S943* (0.59) P1497fs (0.22)C T1556fs (0.20)C |
- | - | R209Q (0.67) LO, DN | - | N64fs (0.66) | - |
| ST-09 | T1368fs (0.24) S1415fs (0.42) |
- | - | - | - | - | - |
| ST-11 | Y935fs (0.44) | - | - | S176C (0.09) V | - | R361C (0.12) LO | - |
| ST-14 | V452fs (0.24) E1237* (0.34) |
EH | - | E246G (0.31) V | - | - | - |
| ST-15 | E1397* (0.52) | - | - | Y387fs (0.17) Q97E (0.23) V |
- | - | - |
| ST-16 | Q1367* (0.95) | - | - | E255fs (0.91) | - | - | - |
| ST-17 | L1449N (0.40) V | - | - | - | H | - | - |
| ST-20 | Ser1539* (0.92) | - | - | - | - | - | - |
| ST-21 | - | - | - | c.-32delA (0.44) | - | - | - |
| ST-23 | - | - | - | V118D (0.82) V | - | C363R (0.77) V | - |
| ST-25 | G635fs (0.15) | - | - | Ile123fs (0.06) | - | - | - |
| ST-26 | S245* (0.62) S2146L (0.15) V |
- | - | G206S (0.88) LO | - | - | - |
| ST-27 | 1357* (0.33) | - | - | E255G (0.33) V | - | - | - |
| ST-29 | S1163_Y1166del (0.13) E1265fs (0.23) S2295_R2301del (0.10) R2347fs (0.11) S2512fs (0.11) |
- | - | L383fs(0.54) R243W (0.14) LO, DN |
H | - | - |
| ST-30 | S874* (0.35) R1435* (0.31) |
- | - | R234H (0.49) T, DN R228W (0.12) T |
- | - | H |
| ST-31 | R876* (0.63) R1450* (0.17) |
- | - | R209Q (0.86) LO, DN | - | - | - |
| ST-32 | R1314fs (0.91) | - | - | - | - | - | - |
| ST-39 | - | EH | H | V118D (0.73) V | - | C363R (0.81) V | - |
| ST-40 | - | H | - | V134L (0.43) LO | - | A118V (0.35) V | - |
| ST-42 | E901* (0.30) T1556fs (0.18) |
- | - | Q126* (0.22) | - | - | - |
| ST-46 | D1394fs (0.43) | - | - | c.258G>A T86= (0.27) Sp | - | - | - |
| ST-49 | - | - | - | - | - | - | - |
| ST-52 | V1405fs (0.63) | - | - | - | - | - | - |
| ST-54 | - | - | - | E17* (0.31) | - | - | - |
| ST-56 | S1421fs (0.59) S2607F (0.29) V |
VH | - | G227E (0.50) V | - | L495H (0.33) V | - |
| ST-57 | - | - | - | - | - | - | - |
| ST-66 | S1315fs (0.67) | EH | - | C102Y (0.45) V | - | - | - |
| ST-76 | c.835-8A>G (0.60) Sp Q2742* (0.25) |
H | - | Y387fs (0.645) R209Q (0.50) LO, DN |
- | C401Y (0.25) V A456V (0.40) V |
- |
| Tumor | MSH2 | MSH6 | MLH1 | PMS2 | ||||
| DNA | Variants | MZS | MZS | Variants | MZS | Variants | MZS | Variants |
| Sample | (VAF) | 5’UTR | In1_2 | (VAF) | (VAF) | (VAF) | ||
| ST-01 | - | - | - | - | - | - | H | - |
| ST-03 | - | - | - | - | - | - | EH | M676V (0.42) V |
| ST-04 | - | - | - | - | - | - | - | - |
| ST-05 | - | - | - | - | - | - | EH | - |
| ST-08 | - | - | - | - | - | - | - | - |
| ST-09 | Q893* (0.18) | - | - | - | - | - | - | - |
| ST-11 | - | - | - | - | - | - | - | - |
| ST-14 | - | - | - | - | - | - | EH | - |
| ST-15 | - | - | - | - | - | - | - | - |
| ST-16 | - | - | - | - | - | - | - | - |
| ST-17 MH | - | - | - | - | - | N444fs (0.34) | EH | - |
| ST-20 | - | - | - | - | - | 1731+2T>C (0.25) | - | - |
| ST-21 | - | - | - | - | - | - | - | - |
| ST-23 MH | - | - | - | - | - | - | EH | - |
| ST-25 | - | - | - | - | - | Q197H (0.10) V | - | - |
| ST-26 | - | - | - | - | - | F656C (0.09) V | - | - |
| ST-27 | - | H | H | - | H | - | - | - |
| ST-29 MH | Q4* (0.25) | - | - | S625C (0.27) VI1170fs (0.25) | - | - | EH | - |
| ST-30 | - | - | - | - | - | - | - | - |
| ST-31 | - | - | - | - | H | - | - | - |
| ST-32 | - | H | H | - | H | - | - | - |
| ST-39 | - | - | H | - | - | - | - | - |
| ST-40 | - | H | H | - | H | - | - | - |
| ST-42 | - | H | - | - | H | - | - | - |
| ST-46 | - | - | - | - | - | - | - | - |
| ST-49 MH | - | - | - | - | - | - | VH | - |
| ST-52 | - | - | - | - | - | - | EH | - |
| ST-54 | - | - | - | - | - | - | - | - |
| ST-56 | - | - | - | - | - | M587I (0.25) V | H | - |
| ST-57 MH | - | - | - | - | - | c.306+1G>A (0.60) | - | - |
| ST-66 | - | H | - | - | - | - | - | - |
| ST-76 | N919D (0.50) V | - | - | E807* (0.40)K1315R (0.50) V | - | - | - | - |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




