Submitted:
24 December 2024
Posted:
25 December 2024
You are already at the latest version
Abstract
Lung cancer is a leading cause of cancer-related mortality worldwide, largely due to its heterogeneity and intrinsic drug resistance. Malignant pleural effusions (MPE) provide diverse tumor cell populations ideal for studying these complexities. Although chemotherapy and tar-geted therapies can be initially effective, subpopulations of cancer cells with phenotypic plasticity often survive treatment, eventually developing resistance. Here, we integrated single-cell isola-tion and three-dimensional (3D) spheroid culture to dissect subclonal heterogeneity and drug responses, aiming to inform precision medicine approaches. Using A549 lung cancer cells, we established a cisplatin-resistant line and isolated three resistant subclones (Holoclone, Meroclone, Paraclone) via single-cell sorting. In 3D spheroids, Docetaxel and Alimta displayed higher IC50 values than in 2D cultures, suggesting that 3D models better reflect clinical dosing. Additionally, MPE-derived Holoclone and Paraclone subclones exhibited distinct sensitivities to Giotrif and Capmatinib, revealing their heterogeneous drug responses. Molecular analyses confirmed ele-vated MDR-1, ABCG2, cancer stem cells (CSC) markers (OCT4, SOX2, CD44, CD133), and epi-thelial-mesenchymal transition (EMT) markers (E-cadherin downregulation, increased Vimentin, N-cadherin, Twist) in resistant subclones, correlating with enhanced migration and invasion. This integrated approach clarifies the interplay between heterogeneity, CSC/EMT phenotypes, and drug resistance, providing a valuable tool for predicting therapeutic responses and guiding per-sonalized, combination-based lung cancer treatments.
Keywords:
1. Introduction
2. Results
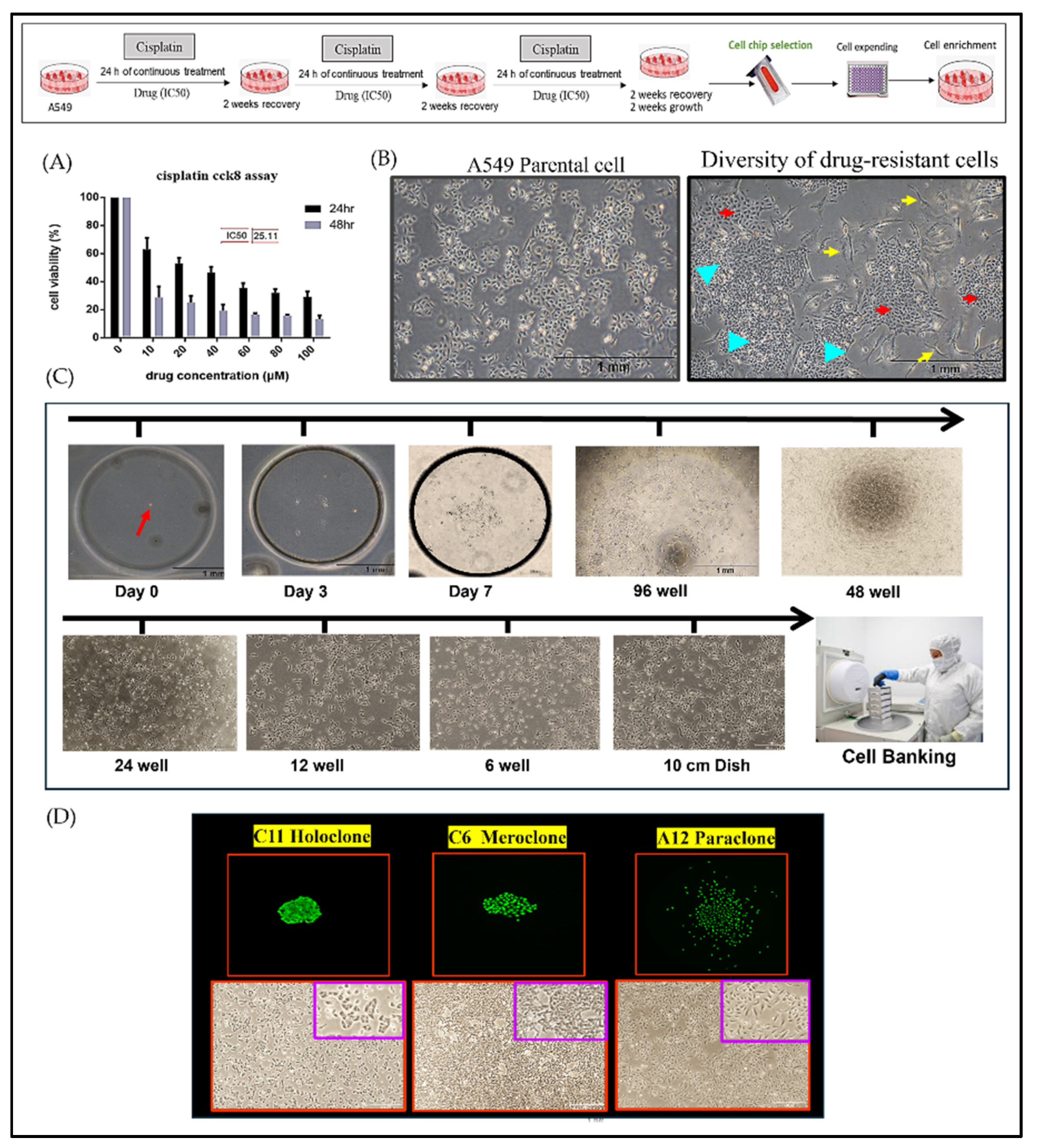
2.1. Integration of Single-Cell Culture Technology for the Establishment and Preservation of Phenotypically Heterogeneous Monoclonal Cell Lines
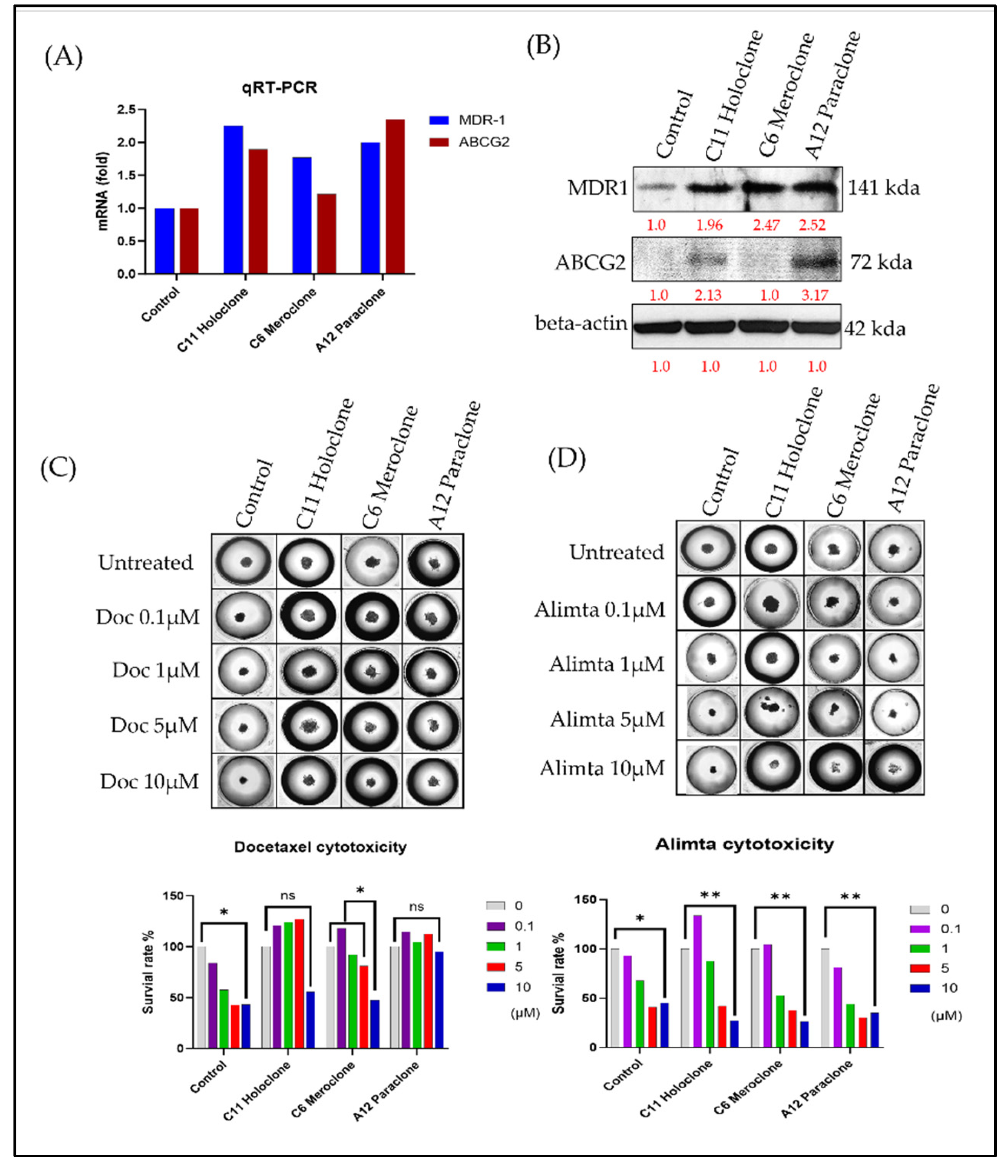
2.2. Phenotypic Heterogeneity of Different Drug-Resistant Subclonal Cells and Their Impact on Chemotherapy Drug Sensitivity.
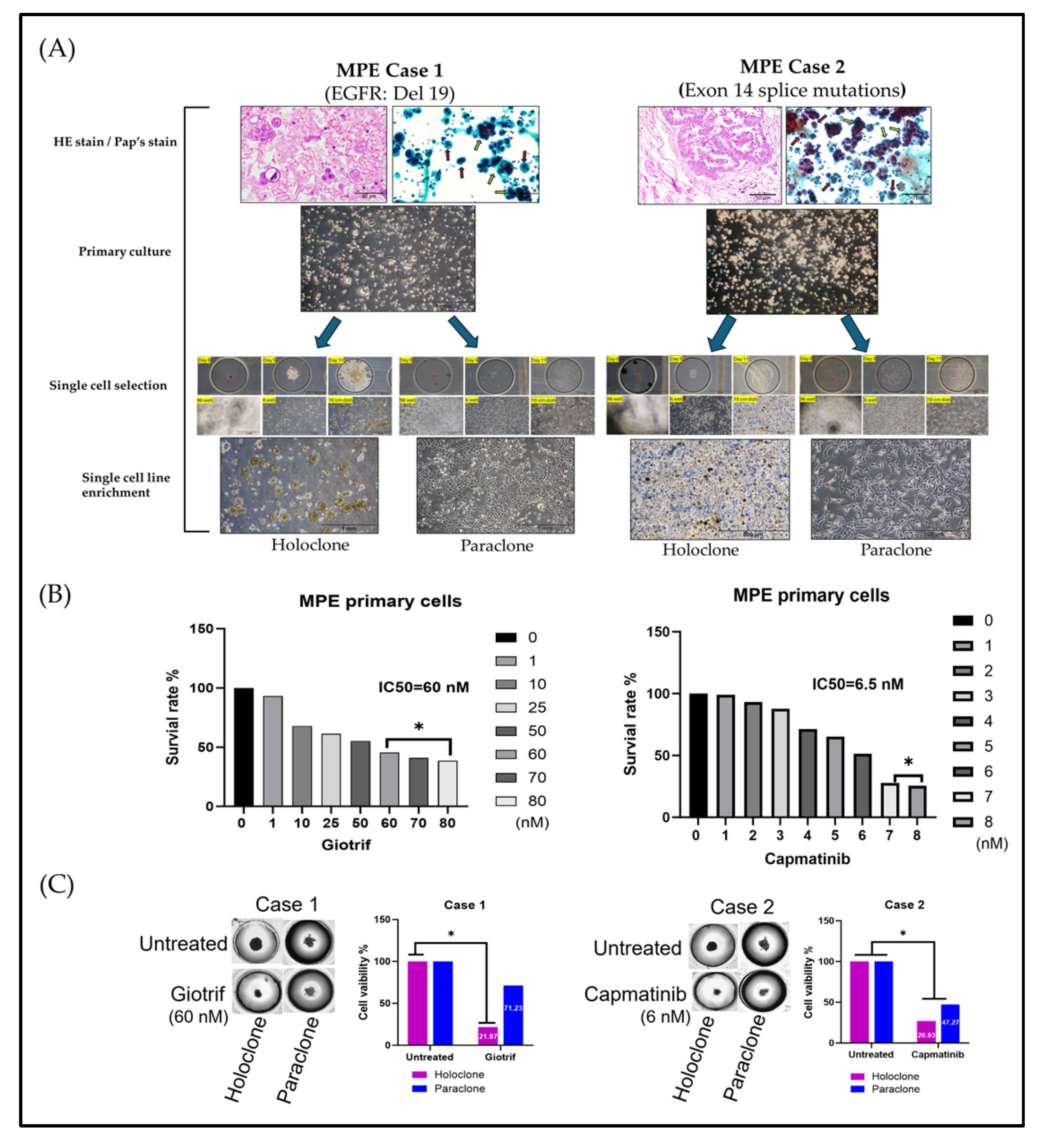
2.3. Evidence of Cellular Phenotypic Heterogeneity and Tumor Spheroid Drug Prediction in Primary MPE Cultures
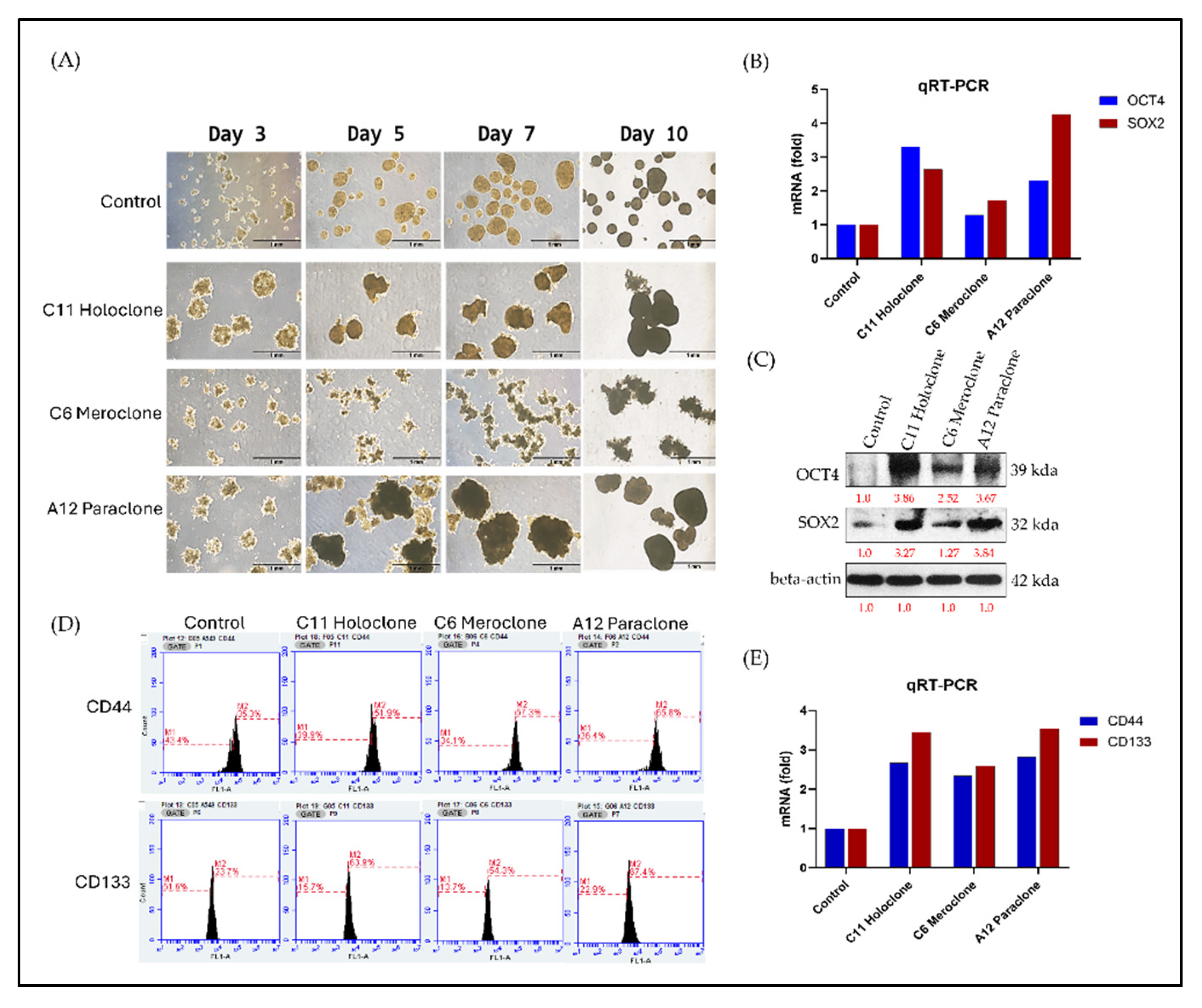
2.4. Multilevel Analysis of Stem Cell Characteristics and Biomarker Expression in Drug-Resistant Subclonal Cells
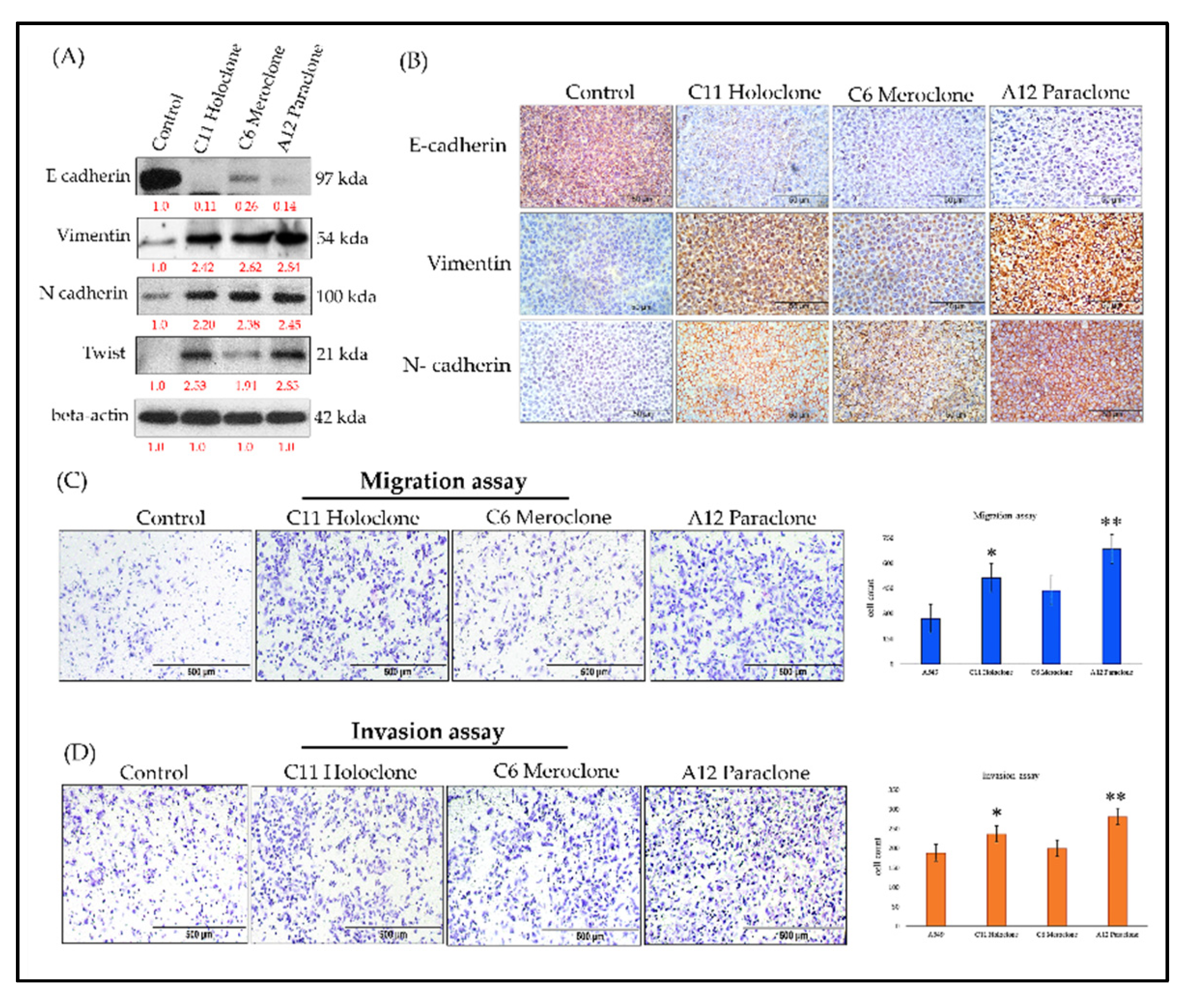
2.5. Molecular Characterization of EMT Transformation and Enhanced Invasive Capability in Drug-Resistant Subclonal Cells
3. Discussion
4. Materials and Methods
4.1. Cell Culture, Drug-Resistant Cancer Cell Lines and MPE Sample Prepare
4.2. Single-Cell Isolation and Culture Procedure
4.3. D Tumorsphere Culture
4.4. Growth Curves of 3D Tumorsphere Detected by CCK8 Assay
4.5. Quantitative Real-Time-PCR (qRT-PCR)
4.6. Western Blot Analysis
4.7. Migration and Invasion Assay
4.8. Flow Cytometry
4.9. Immunohistochemistry
4.10. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- F. Bray, J. Ferlay, I. Soerjomataram, R. L. Siegel, L. A. Torre, and A. Jemal, “Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries,” CA Cancer J Clin, vol. 68, no. 6, pp. 394-424, Nov 2018. [CrossRef]
- Y. Liu and Z. Zheng, “Understanding the global cancer statistics 2022: growing cancer burden,” Sci China Life Sci, vol. 67, no. 10, pp. 2274-2276, Oct 2024. [CrossRef]
- Q. Li et al., “Disparities in 36 cancers across 185 countries: secondary analysis of global cancer statistics,” Front Med, vol. 18, no. 5, pp. 911-920, Oct 2024. [CrossRef]
- Z. F. Lim and P. C. Ma, “Emerging insights of tumor heterogeneity and drug resistance mechanisms in lung cancer targeted therapy,” J Hematol Oncol, vol. 12, no. 1, p. 134, Dec 9 2019. [CrossRef]
- G. R. Bhat et al., “Cancer cell plasticity: from cellular, molecular, and genetic mechanisms to tumor heterogeneity and drug resistance,” Cancer Metastasis Rev, vol. 43, no. 1, pp. 197-228, Mar 2024. [CrossRef]
- Zhang, K. Zhang, K. Miao, H. Sun, and C. X. Deng, “Tumor heterogeneity reshapes the tumor microenvironment to influence drug resistance,” Int J Biol Sci, vol. 18, no. 7, pp. 3019-3033, 2022. [CrossRef]
- S. Li et al., “Overcoming multi-drug resistance in SCLC: a synergistic approach with venetoclax and hydroxychloroquine targeting the lncRNA LYPLAL1-DT/BCL2/BECN1 pathway,” Mol Cancer, vol. 23, no. 1, p. 243, Oct 31 2024. [CrossRef]
- Y. H. Huang et al., “Attenuation of PI3K-Akt-mTOR Pathway to Reduce Cancer Stemness on Chemoresistant Lung Cancer Cells by Shikonin and Synergy with BEZ235 Inhibitor,” Int J Mol Sci, vol. 25, no. 1, Jan 3 2024. [CrossRef]
- S. Brosseau et al., “YAP/TEAD involvement in resistance to paclitaxel chemotherapy in lung cancer,” Mol Cell Biochem, Mar 1 2024. [CrossRef]
- J. Canon et al., “The clinical KRAS(G12C) inhibitor AMG 510 drives anti-tumour immunity,” Nature, vol. 575, no. 7781, pp. 217-223, Nov 2019. [CrossRef]
- M. J. Nokin et al., “RAS-ON inhibition overcomes clinical resistance to KRAS G12C-OFF covalent blockade,” Nat Commun, vol. 15, no. 1, p. 7554, Aug 30 2024. [CrossRef]
- Mohanty et al., “Acquired resistance to KRAS G12C small-molecule inhibitors via genetic/nongenetic mechanisms in lung cancer,” Sci Adv, vol. 9, no. 41, p. eade3816, Oct 13 2023. [CrossRef]
- M. Kurimchak et al., “The drug efflux pump MDR1 promotes intrinsic and acquired resistance to PROTACs in cancer cells,” Sci Signal, vol. 15, no. 749, p. eabn2707, Aug 30 2022,. [CrossRef]
- H. Tian et al., “Genetic trajectory and clonal evolution of multiple primary lung cancer with lymph node metastasis,” Cancer Gene Ther, vol. 30, no. 3, pp. 507-520, Mar 2023,. [CrossRef]
- Russo et al., “Tracking Clonal Evolution of EGFR-Mutated Non-Small Cell Lung Cancer Through Liquid Biopsy: Management of C797S Acquired Mutation,” Clin Lung Cancer, vol. 24, no. 7, pp. 660-665, Nov 2023. [CrossRef]
- P. C. Moore, K. W. Henderson, and M. Classon, “The epigenome and the many facets of cancer drug tolerance,” Adv Cancer Res, vol. 158, pp. 1-39, 2023. [CrossRef]
- G. S. Franca et al., “Cellular adaptation to cancer therapy along a resistance continuum,” Nature, vol. 631, no. 8022, pp. 876-883, Jul 2024. [CrossRef]
- J. Chmielecki et al., “Candidate mechanisms of acquired resistance to first-line osimertinib in EGFR-mutated advanced non-small cell lung cancer,” Nat Commun, vol. 14, no. 1, p. 1070, Feb 27 2023. [CrossRef]
- W. Yao et al., “Epithelial-mesenchymal plasticity (EMP) in wound healing: Exploring EMT mechanisms, regulatory network, and therapeutic opportunities,” Heliyon, vol. 10, no. 14, p. e34269, Jul 30 2024. [CrossRef]
- M. Sisto and S. Lisi, “Epigenetic Regulation of EMP/EMT-Dependent Fibrosis,” Int J Mol Sci, vol. 25, no. 5, Feb 28 2024. [CrossRef]
- B. Mishra and S. S. Nishank, “Therapeutic targeting approach on epithelial-mesenchymal plasticity to combat cancer metastasis,” Med Oncol, vol. 40, no. 7, p. 190, May 29 2023,. [CrossRef]
- J. Haerinck, S. Goossens, and G. Berx, “The epithelial-mesenchymal plasticity landscape: principles of design and mechanisms of regulation,” Nat Rev Genet, vol. 24, no. 9, pp. 590-609, Sep 2023. [CrossRef]
- H. Acloque, J. Yang, and E. Theveneau, “Epithelial-to-mesenchymal plasticity from development to disease: An introduction to the special issue,” Genesis, vol. 62, no. 1, p. e23581, Feb 2024,. [CrossRef]
- G. Toyokawa et al., “Tumor plasticity and therapeutic resistance in oncogene-addicted non-small cell lung cancer: from preclinical observations to clinical implications,” Crit Rev Oncol Hematol, vol. 184, p. 103966, Apr 2023. [CrossRef]
- Z. D. Shi et al., “Tumor cell plasticity in targeted therapy-induced resistance: mechanisms and new strategies,” Signal Transduct Target Ther, vol. 8, no. 1, p. 113, Mar 11 2023. [CrossRef]
- [26], S. S. Ramisetty et al., “Leveraging Cancer Phenotypic Plasticity for Novel Treatment Strategies,” J Clin Med, vol. 13, no. 11, Jun 5 2024. [CrossRef]
- P. C. McDonald and S. Dedhar, “Persister cell plasticity in tumour drug resistance,” Semin Cell Dev Biol, vol. 156, pp. 1-10, Mar 15 2024. [CrossRef]
- J. He, Z. Qiu, J. Fan, X. Xie, Q. Sheng, and X. Sui, “Drug tolerant persister cell plasticity in cancer: A revolutionary strategy for more effective anticancer therapies,” Signal Transduct Target Ther, vol. 9, no. 1, p. 209, Aug 14 2024. [CrossRef]
- J. Zhang, X. Xu, H. Deng, L. Liu, Y. Xiang, and J. Feng, “Overcoming cancer drug-resistance calls for novel strategies targeting abnormal alternative splicing,” Pharmacol Ther, vol. 261, p. 108697, Sep 2024,. [CrossRef]
- X. Xu et al., “PARP1 promotes EGFR-TKI drug-resistance via PI3K/AKT pathway in non-small-cell lung cancer,” Cancer Chemother Pharmacol, vol. 94, no. 2, pp. 209-221, Aug 2024. [CrossRef]
- K. Zhang et al., “A Microfluidic Chip-Based Automated System for Whole-Course Monitoring the Drug Responses of Organoids,” Anal Chem, vol. 96, no. 24, pp. 10092-10101, Jun 18 2024. [CrossRef]
- H. Yang et al., “Integrating cfDNA liquid biopsy and organoid-based drug screening reveals PI3K signaling as a promising therapeutic target in colorectal cancer,” J Transl Med, vol. 22, no. 1, p. 132, Feb 3 2024. [CrossRef]
- G. Piro et al., “Pancreatic Cancer Patient-Derived Organoid Platforms: A Clinical Tool to Study Cell- and Non-Cell-Autonomous Mechanisms of Treatment Response,” Front Med (Lausanne), vol. 8, p. 793144, 2021. [CrossRef]
- X. Li et al., “Assay establishment and validation of a high-throughput organoid-based drug screening platform,” Stem Cell Res Ther, vol. 13, no. 1, p. 219, May 26 2022,. [CrossRef]
- S. H. Lee et al., “Prediction of TKI response in EGFR-mutant lung cancer patients-derived organoids using malignant pleural effusion,” NPJ Precis Oncol, vol. 8, no. 1, p. 111, May 21 2024. [CrossRef]
- Z. Zhou et al., “Isolation and identification of cancer stem cells from PC3 human prostate carcinoma cell line,” Int J Clin Exp Pathol, vol. 10, no. 8, pp. 8377-8382, 2017. [Online]. Available: https://www.ncbi.nlm.nih.gov/pubmed/31966689.
- E. Meacham and S. J. Morrison, “Tumour heterogeneity and cancer cell plasticity,” Nature, vol. 501, no. 7467, pp. 328-37, Sep 19 2013. -. [CrossRef]
- F. Luond, S. Tiede, and G. Christofori, “Breast cancer as an example of tumour heterogeneity and tumour cell plasticity during malignant progression,” Br J Cancer, vol. 125, no. 2, pp. 164-175, Jul 2021. [CrossRef]
- V. Almendro, A. Marusyk, and K. Polyak, “Cellular heterogeneity and molecular evolution in cancer,” Annu Rev Pathol, vol. 8, pp. 277-302, Jan 24 2013. [CrossRef]
- R. Thoma, M. Zimmermann, I. Agarkova, J. M. Kelm, and W. Krek, “3D cell culture systems modeling tumor growth determinants in cancer target discovery,” Adv Drug Deliv Rev, vol. 69-70, pp. 29-41, Apr 2014. [CrossRef]
- M. Kumar et al., “A High-Throughput Neurosphere-Based Colony Formation Assay to Test Drug and Radiation Sensitivity of Different Patient-Derived Glioblastoma Lines,” Cells, vol. 13, no. 23, Dec 3 2024. [CrossRef]
- J. A. Engelken, T. Butelmann, F. Tribukait-Riemenschneider, and V. P. Shastri, “Towards a 3D-Printed Millifluidic Device for Investigating Cellular Processes,” Micromachines (Basel), vol. 15, no. 11, Oct 31 2024. [CrossRef]
- S. Cordeiro et al., “Breaking the mold: 3D cell cultures reshaping the future of cancer research,” Front Cell Dev Biol, vol. 12, p. 1507388, 2024,. [CrossRef]
- S. M. Shaffer et al., “Rare cell variability and drug-induced reprogramming as a mode of cancer drug resistance,” Nature, vol. 546, no. 7658, pp. 431-435, Jun 15 2017. [CrossRef]
- W. A. Flavahan, E. Gaskell, and B. E. Bernstein, “Epigenetic plasticity and the hallmarks of cancer,” Science, vol. 357, no. 6348, Jul 21 2017. [CrossRef]
- L. Wang et al., “Malignant pleural effusion facilitates the establishment and maintenance of tumor organoid biobank with multiple patient-derived lung tumor cell sources,” Exp Hematol Oncol, vol. 13, no. 1, p. 115, Nov 15 2024. [CrossRef]
- Y. M. Chang et al., “Interleukin-8/CXCR1 Signaling Contributes to the Progression of Pulmonary Adenocarcinoma Resulting in Malignant Pleural Effusion,” Cells, vol. 13, no. 11, Jun 3 2024. [CrossRef]
- Q. Ye, J. Wang, B. Wang, M. Zhao, Z. Wu, and X. Liu, “Establishment of an in Vitro Three-Dimensional Vascularized Micro-Tumor Model and Screening of Chemotherapeutic Drugs,” Technol Cancer Res Treat, vol. 23, p. 15330338241286755, Jan-Dec 202. [CrossRef]
- H. Ryoo, H. Kimmel, E. Rondo, and G. H. Underhill, “Advances in high throughput cell culture technologies for therapeutic screening and biological discovery applications,” Bioeng Transl Med, vol. 9, no. 3, p. e10627, May 2024. [CrossRef]
- J. Petiti, S. Caria, L. Revel, M. Pegoraro, and C. Divieto, “Standardized Protocol for Resazurin-Based Viability Assays on A549 Cell Line for Improving Cytotoxicity Data Reliability,” Cells, vol. 13, no. 23, Nov 26 2024. [CrossRef]
- A. A. Olijnik et al., “Generating human bone marrow organoids for disease modeling and drug discovery,” Nat Protoc, vol. 19, no. 7, pp. 2117-2146, Jul 2024. [CrossRef]
- H. C. Hung et al., “Enhancement of Tumorigenicity, Spheroid Niche, and Drug Resistance of Pancreatic Cancer Cells in Three-Dimensional Culture System,” J Cancer, vol. 15, no. 8, pp. 2292-2305, 2024. [CrossRef]
- I. Gopallawa et al., “Applications of Organoids in Advancing Drug Discovery and Development,” J Pharm Sci, vol. 113, no. 9, pp. 2659-2667, Sep 2024. [CrossRef]
- M. Swayden, J. Iovanna, and P. Soubeyran, “Pancreatic cancer chemo-resistance is driven by tumor phenotype rather than tumor genotype,” Heliyon, vol. 4, no. 12, p. e01055, Dec 2018. [CrossRef]
- J. Plava, M. Cihova, M. Burikova, M. Matuskova, L. Kucerova, and S. Miklikova, “Recent advances in understanding tumor stroma-mediated chemoresistance in breast cancer,” Mol Cancer, vol. 18, no. 1, p. 67, Mar 30 2019. [CrossRef]
- Z. Dong et al., “Genetic and phenotypic profiling of single living circulating tumor cells from patients with microfluidics,” Proc Natl Acad Sci U S A, vol. 121, no. 19, p. e2315168121, May 7 2024. [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



