Submitted:
10 December 2024
Posted:
11 December 2024
You are already at the latest version
Abstract
Daunomycin is a chemotherapeutic agent widely used for the treatment of leukemia, but its toxicity toward healthy dividing cells limits its clinical use and its production by fermentation. Here we describe the development of a specialized cultivation medium for daunomycin production, including a shift to oil rather than sugar as the primary carbon source. This achieved an almost threefold increase in daunomycin yields, reaching 5.5–6.0 g/L. Daunomycin produced in the oil-based medium was predominantly found in the solid sediment, whereas that produced in the sugar-based medium was mostly soluble. The oil-based medium thus induces an autonomous daunomycin-resistance mechanism involving biogenic nanoparticle formation. Characterization of the nanoparticles confirmed the incorporation of iron and daunomycin, indicating this approach has the potential to mitigate cytotoxicity while improving yields. The presence of proteins associated with iron homeostasis and oxidative stress responses revealed the ability of the production strain to adapt to high iron concentrations. Our findings provide insight into the mechanisms of biogenic nanoparticle formation and the optimization of cultivation processes. Further investigation will help to refine microbial production systems for daunomycin and also broaden the application of similar strategies for the synthesis of other therapeutically important compounds.
Keywords:
1. Introduction
2. Materials and methods
2.1 Chemicals
2.2. Bacterial strain and isolation
2.3. Triclosan-induced mutation
2.4 Olive pomace oil-resistant strain development
2.5. Protoplast formation
2.6. Working bank preparation
2.7. Inoculum preparation
2.8. Production medium
2.9. Effect of carbon and nitrogen sources on daunomycin production
2.10. Effect of iron on daunomycin production
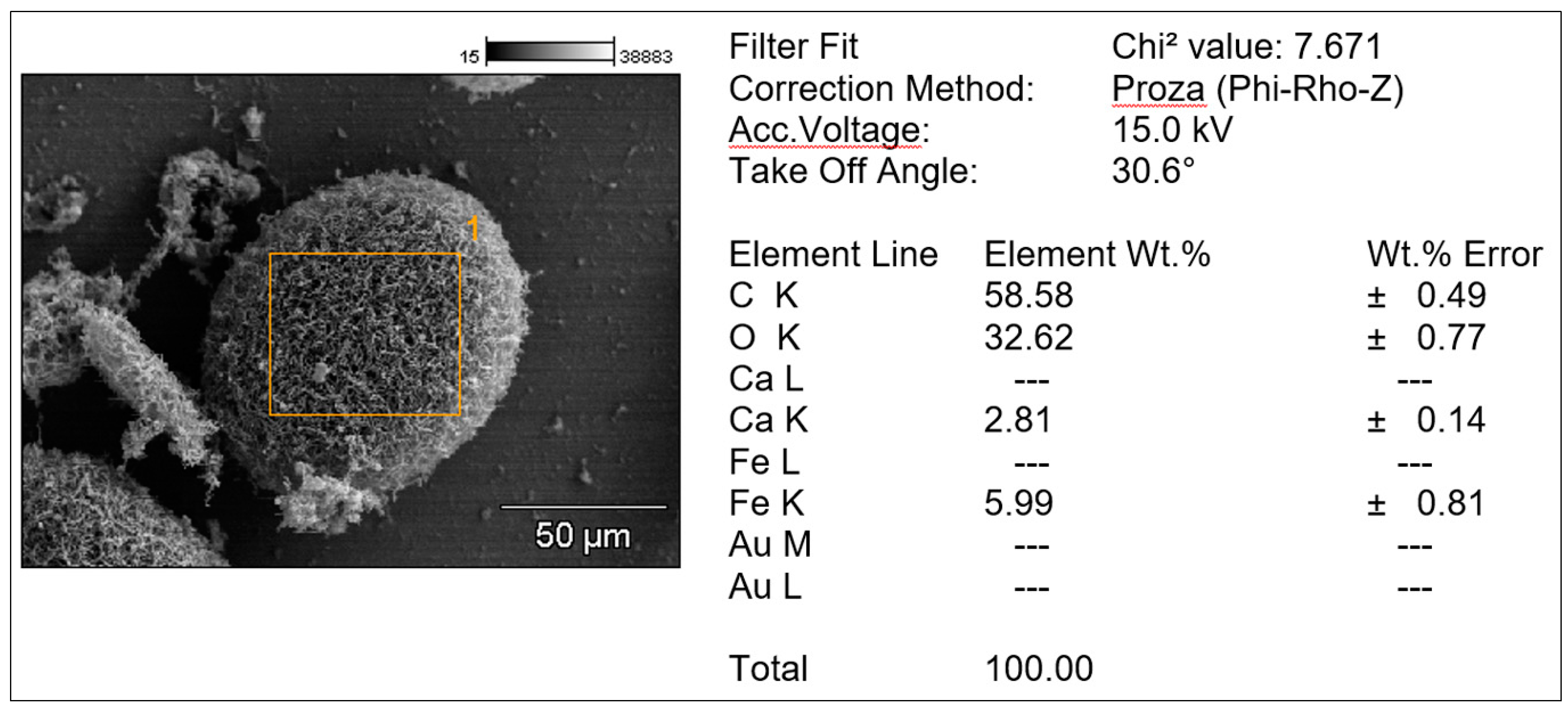
2.11. Direct detection of iron
2.12. X-ray diffraction of iron-containing samples
2.13. Monitoring of daunomycin production
2.14. Changes in media color and physical parameters
2.15. Sediment friability
2.16. Observation of morphological changes during fermentation
2.16.1. Optical microscopy
2.16.2. Scanning electron microscopy
2.16.3. Transmission electron microscopy
2.17. Polyacrylamide gel electrophoresis
2.18. Measurement of pH, dissolved oxygen, and redox potential during fermentation
2.19. Extraction and sample preparation
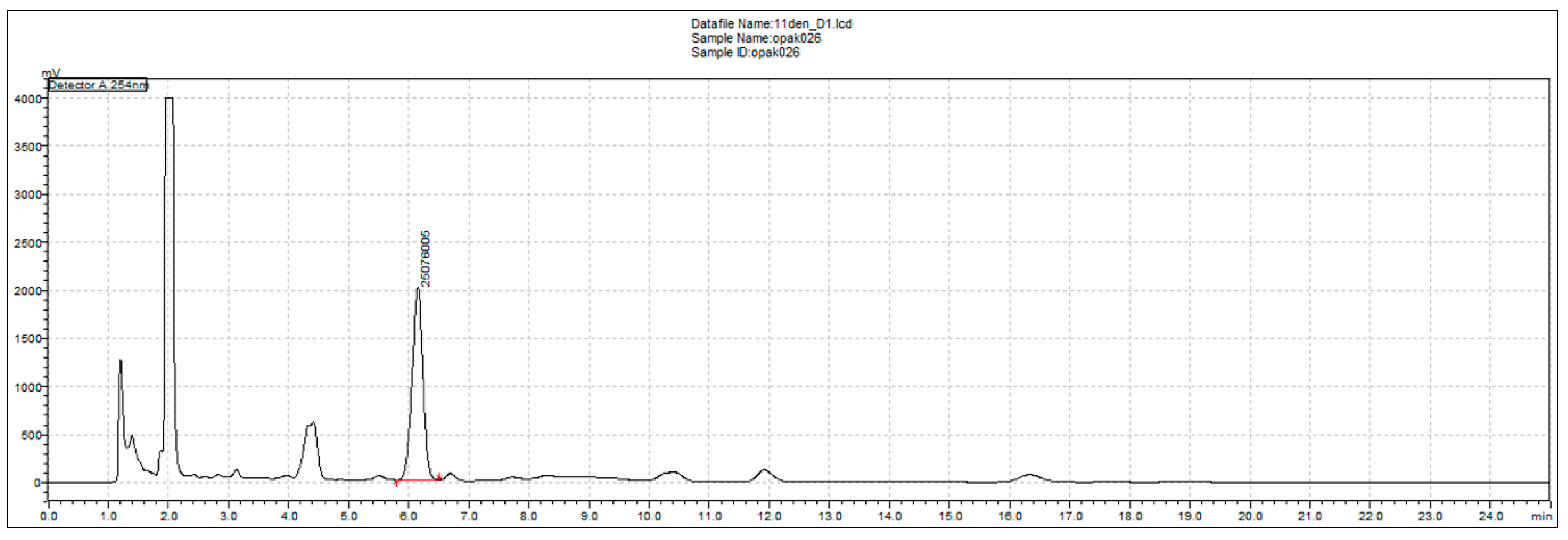
2.20. HPLC
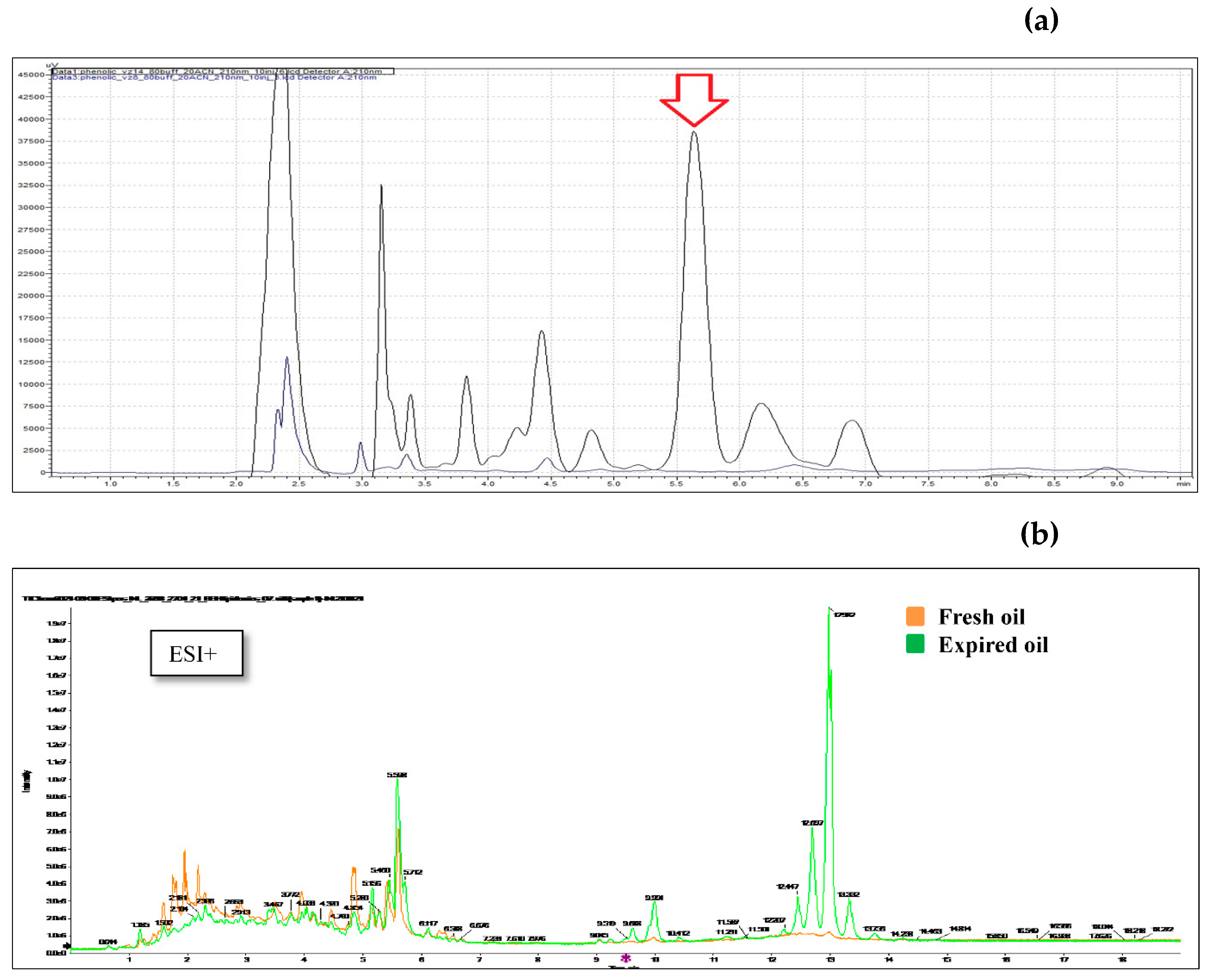
2.21. Olive oil extraction and HPLC
3. Results and discussion
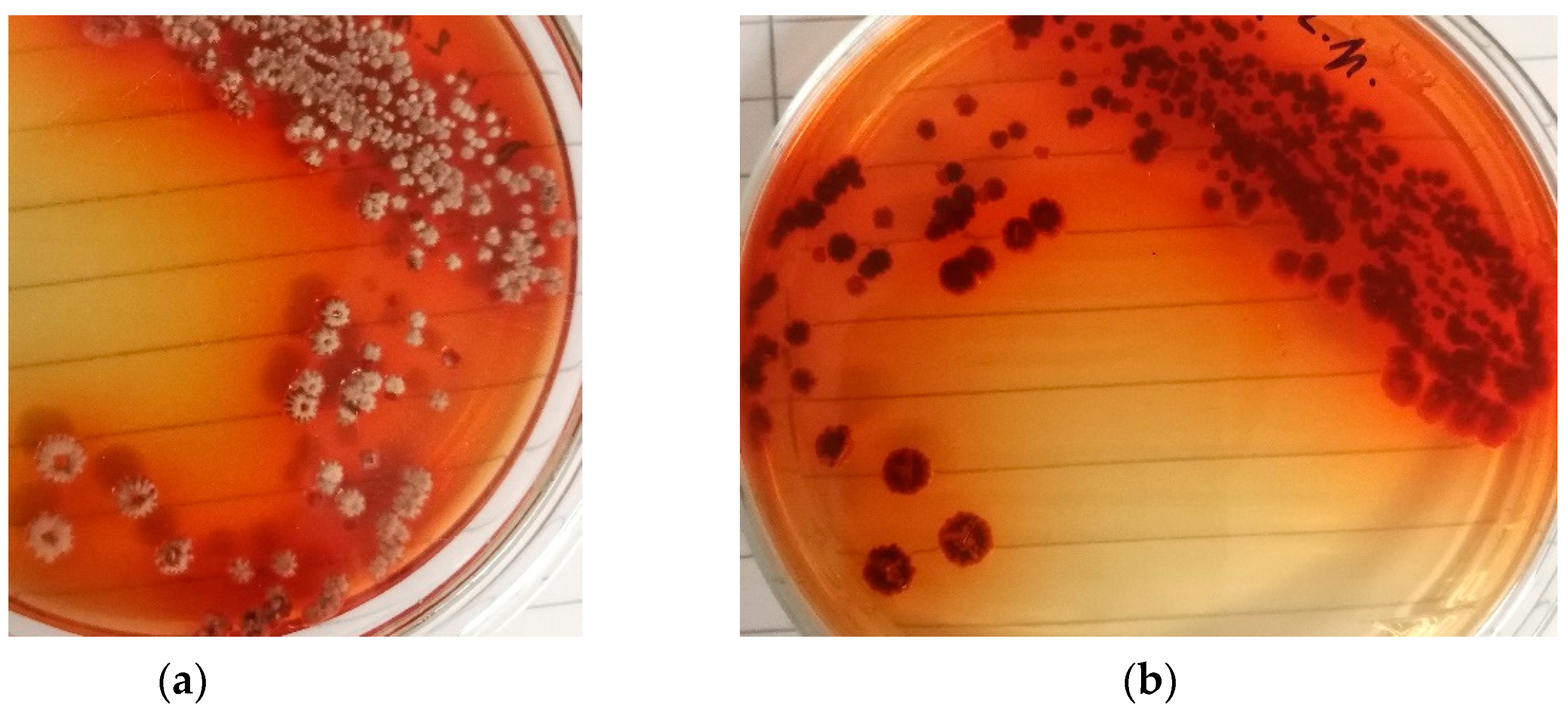
3.1. Production strain
3.2. Optimization of the production medium
3.2.1. Optimization of physical parameters
3.2.2 Replacement of glucose with oils
| Duration of cultivation [h] | N [samples] | Yield [mg/L] | pH | ||
|---|---|---|---|---|---|
| Average | SD | Average | SD | ||
| Olive oil refined | |||||
| 120 | 8 | 1201.3 | 141.7 | 6.03 | 0.03 |
| 168 | 8 | 1991.7 | 597.1 | 6.15 | 0.05 |
| 216 | 8 | 2220.9 | 492.9 | 8.64 | 0.13 |
| 264 | 8 | 2964.4 | 555.2 | 8.07 | 0.22 |
| Olive oil pomace | |||||
| 120 | 8 | 1710.9 | 86.7 | 6.93 | 0.02 |
| 168 | 8 | 2986.3 | 176.9 | 6.84 | 0.03 |
| 216 | 8 | 4603.7 | 228.9 | 6.87 | 0.03 |
| 264 | 8 | 5448.1 | 108.5 | 6.91 | 0.05 |
| Rapeseed oil | |||||
| 120 | 8 | 861.3 | 32.7 | 6.08 | 0.06 |
| 168 | 8 | 1128.5 | 49.3 | 6.14 | 0.05 |
| 216 | 8 | 1110.8 | 207.4 | 8.51 | 0.20 |
| 264 | 8 | 1089.4 | 88.1 | 7.97 | 0.58 |
| Hemp oil | |||||
| 120 | 8 | 61.1 | 17.7 | 6.74 | 0.56 |
| 168 | 8 | 125.7 | 11.4 | 7.45 | 0.75 |
| 216 | 8 | 168.2 | 42.5 | 8.11 | 0.11 |
| 264 | 8 | 152.6 | 41.8 | 8.32 | 0.08 |
3.2.3. Influence of soy source on daunomycin production
3.2.4 Influence of yeast source on daunomycin production
3.2.5. Optimal media composition for daunomycin production
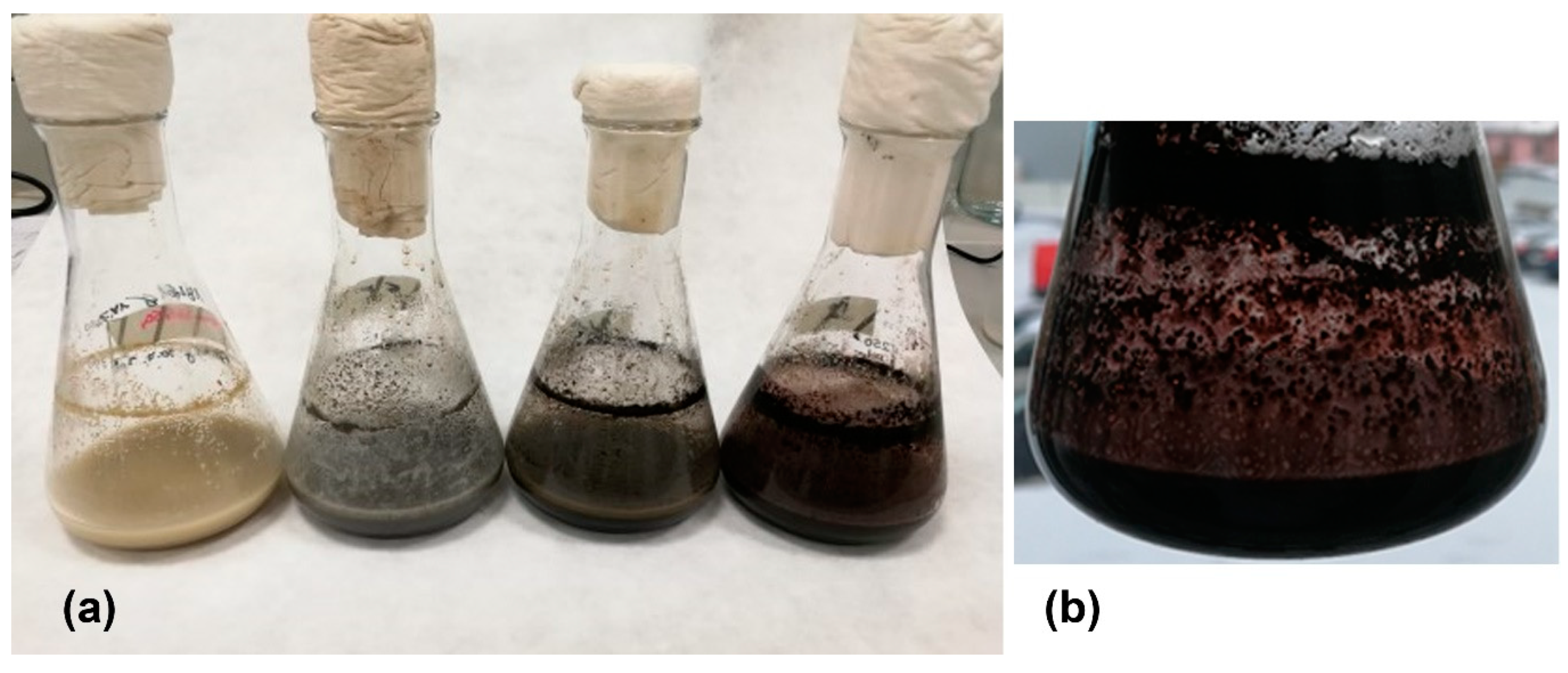
3.3. Changes in media color and physical parameters
3.4. The precipitation of iron-containing complexes
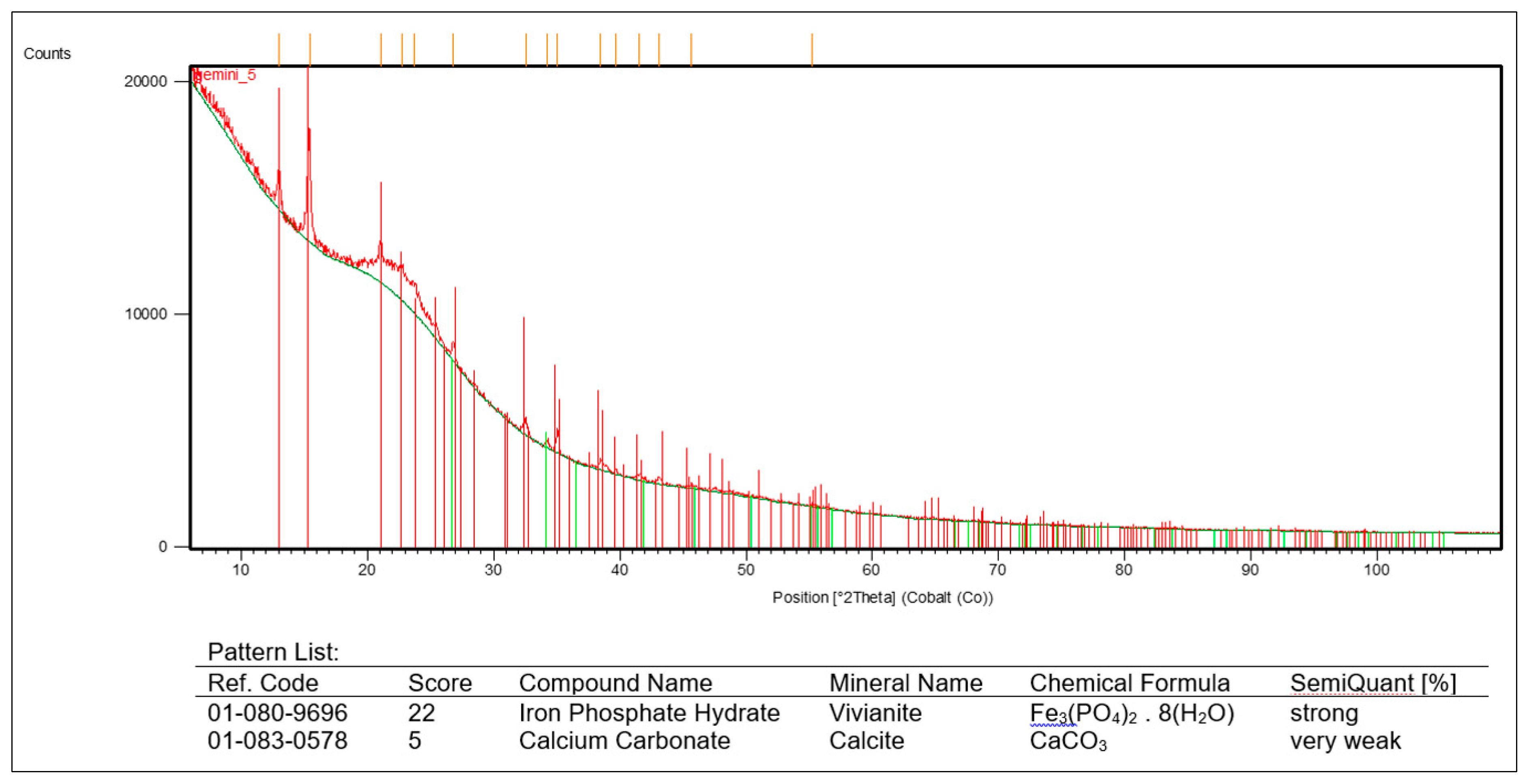
3.5. The presence of vivianite
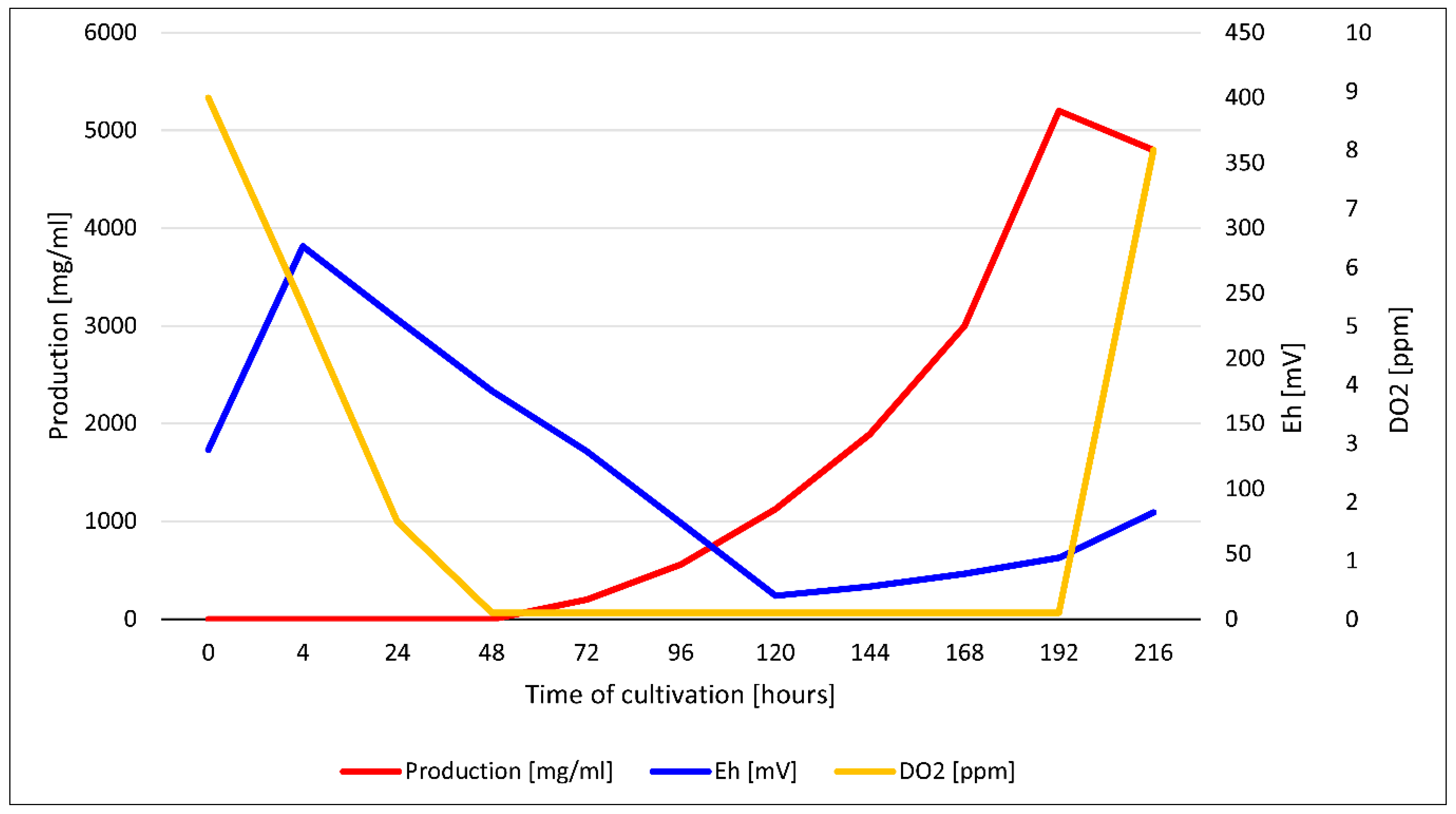
3.6. Measurement of pH, DO, and redox potential during fermentation
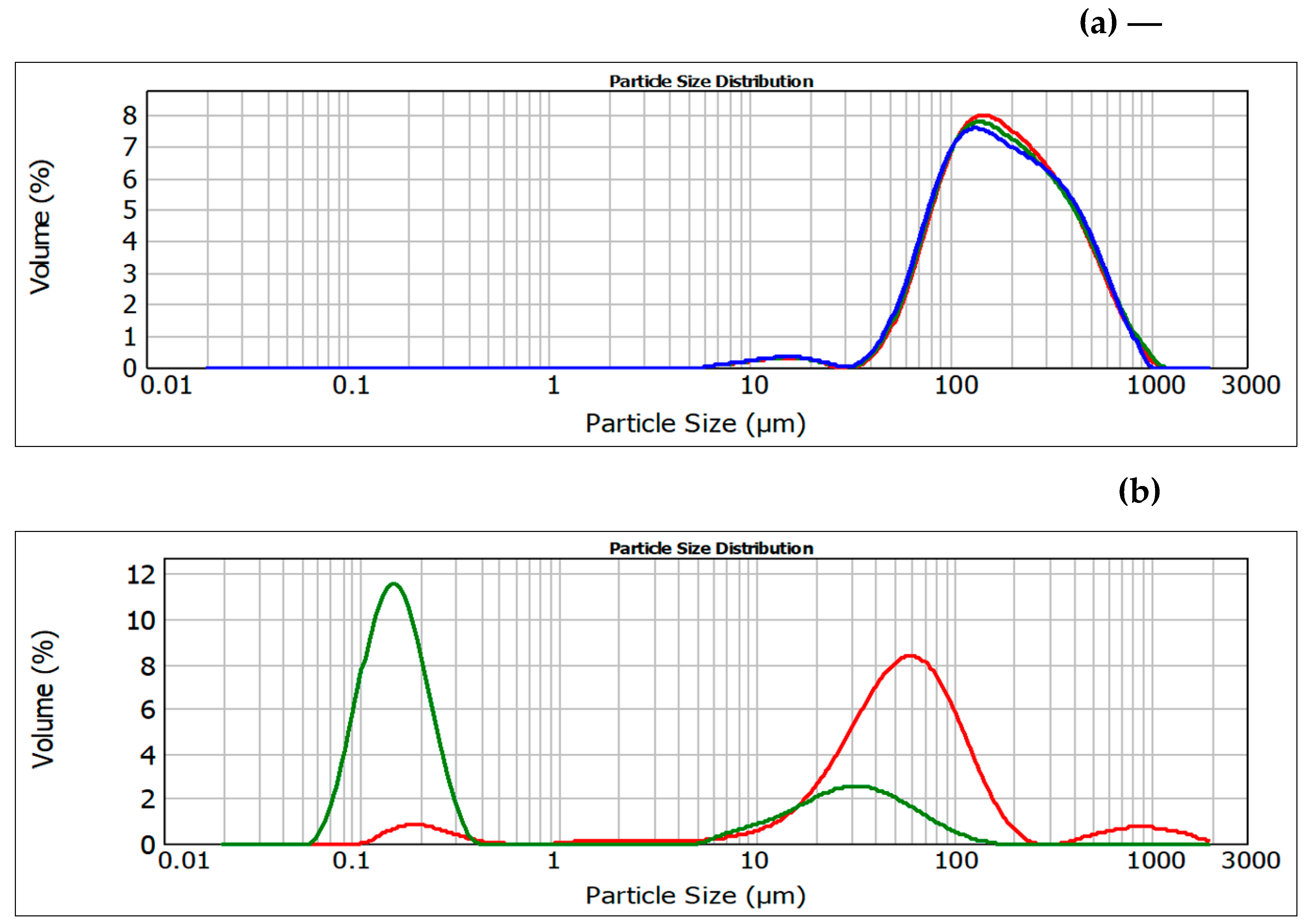
3.7. Sediment friability
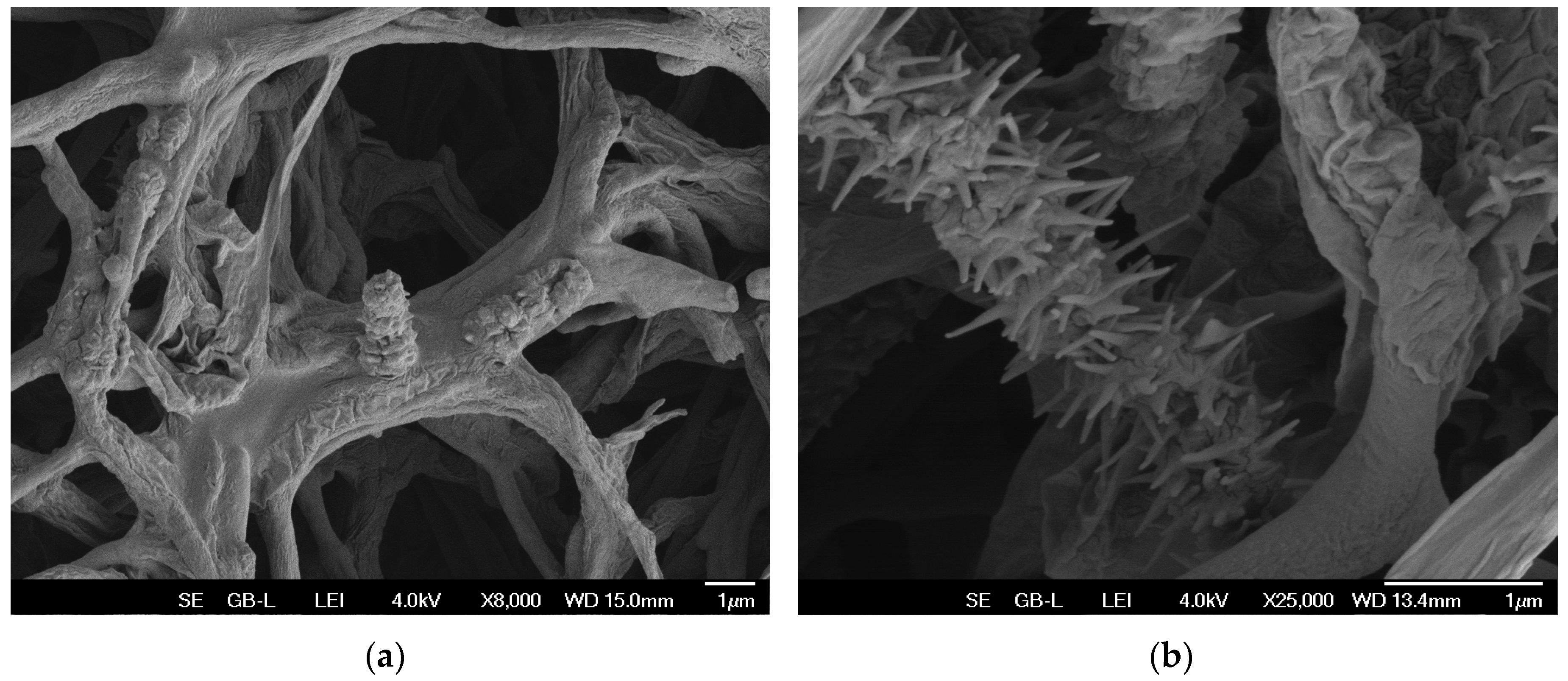
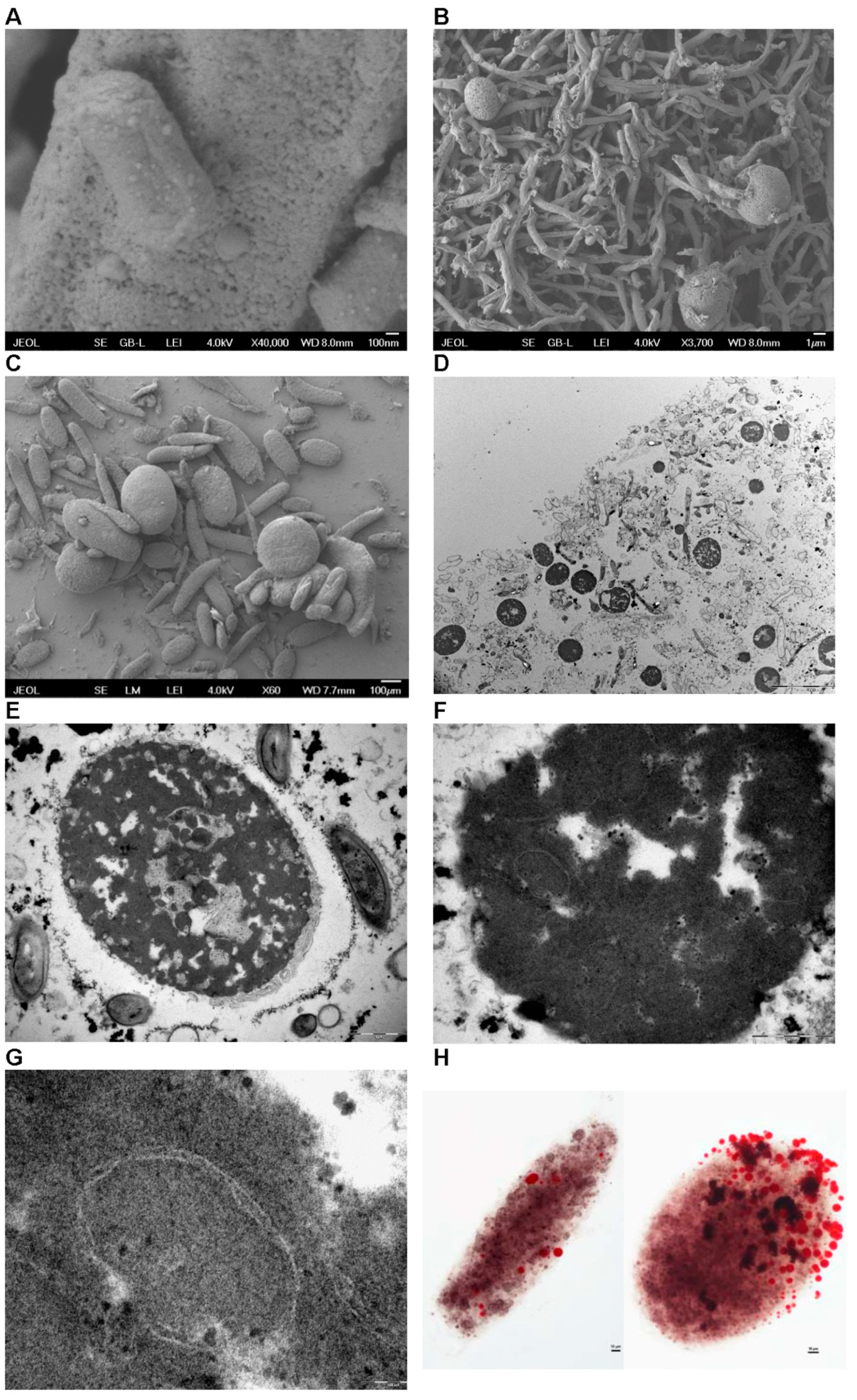
3.8. Observation of morphological changes during fermentation
3.9. Detection and identification of proteins
4. Conclusions
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Thomas, G.J. Synthesis of Anthracyclines Related to Daunomycin. In: Recent Progress in the Chemical Synthesis of Antibiotics; Lukacs, G.; Ohno, M., Eds.; Springer, Berlin, Heidelberg, 1990; pp. 467-496. [CrossRef]
- Thirumaran, R.; Prendergast, G.C.; Gilman, P.B. Chapter 7 - Cytotoxic Chemotherapy in Clinical Treatment of Cancer. In: Cancer Immunotherapy; Prendergast, G.C.; Jaffee, E.M.; Eds.; Academic Press, Burlington, USA, 2007. pp. 101-116. [CrossRef]
- Srinivasan, P.; Palani, S.N.; Prasad, R. Daunorubicin efflux in Streptomyces peucetius modulates biosynthesis by feedback regulation. FEMS Microbiol. Lett. 2010, 305, 18–27. [Google Scholar] [CrossRef] [PubMed]
- Vasanthakumar, A.; Kattusamy, K.; Prasad, R. Regulation of daunorubicin biosynthesis in Streptomyces peucetius – feed forward and feedback transcriptional control. J. Basic Microbiol. 2013, 53, 636–644. [Google Scholar] [CrossRef] [PubMed]
- Taatjes, D.J.; Gaudiano, G.; Resing, K.; Koch, T.H. Redox pathway leading to the alkylation of DNA by the anthracycline, antitumor drugs adriamycin and daunomycin. J. Med. Chem. 1997, 40, 1276–86. [Google Scholar] [CrossRef] [PubMed]
- Kaiserová, H.; Simunek, T.; Sterba, M.; den Hartog, G.J.; Schröterová, L.; Popelová, O.; Gersl, V.; Kvasnicková, E.; Bast, A. New iron chelators in anthracycline-induced cardiotoxicity. Cardiovasc. Toxicol. 2007, 7, 145–50. [Google Scholar] [CrossRef]
- Cai, J.; Deng, T.; Shi, J.; Chen, C.; Wang, Z.; Liu, Y. Daunorubicin resensitizes Gram-negative superbugs to the last-line antibiotics and prevents the transmission of antibiotic resistance. iScience 2023, 26, 106809. [Google Scholar] [CrossRef] [PubMed]
- Yuan, T.; Yin, C.; Zhu, C.; Zhu, B.; Hu, Y. Improvement of antibiotic productivity by knock-out of dauW in Streptomyces coeruleobidus. Microbiol. Res. 2011, 166, 539–547. [Google Scholar] [CrossRef]
- Malla, S.; Niraula, N.P.; Liou, K.; Sohng, J.K. Improvement in doxorubicin productivity by overexpression of regulatory genes in Streptomyces peucetius. Res. Microbiol. 2010, 161, 109–117. [Google Scholar] [CrossRef]
- Afonso, IS.; Cardoso, B.; Nobrega, G.; Minas, G.; Ribeiro, J.E.; Lima, R.A. Green synthesis of nanoparticles from olive oil waste for environmental and health applications: A review. J. Environ. Chem. Eng. 2024, 12, 114022. [Google Scholar] [CrossRef]
- Nunes, M.A.; Costa, A.S.G.; Bessada, S.; Santos, J.; Puga, H.; Alves, R.C.; Freitas, V.; Oliveira, M.B.P.P. Olive pomace as a valuable source of bioactive compounds: A study regarding its lipid- and water-soluble components. Sci. Total Environ. 2018, 644, 229–236. [Google Scholar] [CrossRef]
- Mashaghi, S.; Jadidi, T.; Koenderink, G.; Mashaghi, A. Lipid nanotechnology. Int J Mol Sci. 2013, 14, 4242–4282. [Google Scholar] [CrossRef] [PubMed]
- Lombardo, D.; Calandra, P.; Pasqua, L.; Magazù, S. Self-Assembly of Organic Nanomaterials and Biomaterials: The Bottom-Up Approach for Functional Nanostructures Formation and Advanced Applications. Materials 2020, 13, 1048. [Google Scholar] [CrossRef]
- Kargari Aghmiouni, D.; Khoee, S. Dual-Drug Delivery by Anisotropic and Uniform Hybrid Nanostructures: A Comparative Study of the Function and Substrate–Drug Interaction Properties. Pharmaceutics 2023, 15, 1214. [Google Scholar] [CrossRef] [PubMed]
- Perron, N.R.; Brumaghim, J.L. A review of the antioxidant mechanisms of polyphenol compounds related to iron binding. Cell Biochem. Biophys. 2009, 53, 75–100. [Google Scholar] [CrossRef] [PubMed]
- Hsueh, C.C.; Wu, C.C.; Chen, B.Y. Polyphenolic compounds as electron shuttles for sustainable energy utilization. Biotechnol. Biofuels 2019, 12, 1–26. [Google Scholar] [CrossRef] [PubMed]
- Amini, S.M.; Akbari, A. Metal nanoparticles synthesis through natural phenolic acids. IET Nanobiotech. 2019, 13, 771–777. [Google Scholar] [CrossRef] [PubMed]
- Anand, S.; Sowbhagya, R.; Ansari, M.A.; Alzohairy, M.A.; Alomary, M.N.; Almalik, A.I.; Ahmad, W.; Tripathi, T.; Elderdery, A.Y. Polyphenols and Their Nanoformulations: Protective Effects against Human Diseases. Life 2022, 12, 1639. [Google Scholar] [CrossRef] [PubMed]
- Studart, A.R.; Amstad, E.; Gauckler, L.J. Colloidal stabilization of nanoparticles in concentrated suspensions. 2017; 23, 1081–1090. [Google Scholar] [CrossRef]
- El-Moslamy, S.H.; Elnouby, M.S.; Rezk, A.H; El-Fakharany, E.M. Scaling-up strategies for controllable biosynthetic ZnO NPs using cell free-extract of endophytic Streptomyces albus: characterization, statistical optimization, and biomedical activities evaluation. Sci. Rep. 2023, 13, 3200. [Google Scholar] [CrossRef]
- Anani, O.A.; Inobeme, A.; Osarenotor, O.; Olisaka, F.N.; Aidonojie, P.A.; Olatunji, E.O.; Aishatu, H.I. Application of Microorganisms as Biofactories to Produce Biogenic Nanoparticles for Environmental Cleanup: Currents Advances and Challenges. Curr. Nanosci. 2023, 6, 770–782. [Google Scholar] [CrossRef]
- Kulkarni, D.; Sherkar, R.; Shirsathe, C.; Sonwane, R.; Varpe, N.; Shelke, S.; More, M.P.; Pardeshi, S.R.; Dhaneshwar, G.; Junnuthula, V.; Dyawanapelly, S. Biofabrication of nanoparticles: sources, synthesis, and biomedical applications. Front. Bioeng. Biotechnol. 2023, 11, 1159193. [Google Scholar] [CrossRef]
- McMurry, L.M.; Oethinger, M.; Levy, S.B. Triclosan targets lipid synthesis. Nature 1998, 394, 531–532. [Google Scholar] [CrossRef]
- Hopwood, D.A.; Wright, H.M. Factors affecting recombinant frequency in protoplast fusions of Streptomyces coelicolour. J. Gen. Microbiol. 1979, 11, 137–143. [Google Scholar] [CrossRef] [PubMed]
- Hopwood, D.A.; Wright, H.M. Protoplast Fusion in Streptomyces: Fusions Involving Ultraviolet-irradiated Protoplasts. Microbiology 1981, 126, 21–27. [Google Scholar] [CrossRef]
- Okanishi, M.; Suzuki, K.; Umezawa, H. Formation and reversion of streptomycete protoplasts: cultural conditions and morphological study. J. Gen. Microbiol. 1974, 80, 389–400. [Google Scholar] [CrossRef]
- Hopwood, D.A.; Wright, H.M. Bacterial protoplast fusion: recombination in fused protoplasts of Streptomyces coelicolour. Mol. Gen. Genet. 1978, 162, 307–317. [Google Scholar] [CrossRef]
- Hopwood, D.A.; Wright, H.M.; Bibb, M.J.; Cohen, S.N. Genetic recombination through protoplast fusion in Streptomyces. Nature 1977, 268, 171–174. [Google Scholar] [CrossRef] [PubMed]
- Laemmli, U.K. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 1970, 227, 680–685. [Google Scholar] [CrossRef]
- Hurkova, K.; Uttl, L.; Rubert, J.; Navratilova, K.; Kocourek, V.; Stranska-Zachariasova, M.; Paprstein, F.; Hajslova, J. Cranberries versus lingonberries: A challenging authentication of similar Vaccinium fruit. Food Chem. 2019, 284, 162–170. [Google Scholar] [CrossRef] [PubMed]
- Blumauerová, M.; Pokorný, V.; Stastná, J.; Hostálek, Z.; Vanĕk, Z. Development mutants of Streptomyces coeruleorubidus, a producer of anthracyclines: isolation and preliminary characterization. Folia Microbiol. (Praha) 1978, 23, 177–82. [Google Scholar] [CrossRef]
- Mateos, R.; Sarria, B.; Bravo, L. Nutritional and other health properties of olive pomace oil. Crit. Rev. Food Sci. Nutrition 2019, 60, 3506–3521. [Google Scholar] [CrossRef]
- Čepo, V.D.; Radić, K.; Jurmanović, S.; Jug, M.; Grdić Rajković, M.; Pedisić, S.; Moslavac, T.; Albahari, P. Valorization of Olive Pomace-Based Nutraceuticals as Antioxidants in Chemical, Food, and Biological Models. Molecules 2018, 23, 2070. [Google Scholar] [CrossRef]
- Zhang, M.N.; Huang, G.R.; Jiang, J.X. Iron binding capacity of dephytinised soy protein isolate hydrolysate as influenced by the degree of hydrolysis and enzyme type. J. Food Sci. Technol. 2014, 51, 994–999. [Google Scholar] [CrossRef]
- Cruz, C.; Santos, J.; Santos, F.; Silva, G.; Cruz, E.; Ramos, G.; Nascimento, J. Texturized soy protein as an alternative low-cost media for bacteria cultivation. Bacterial Empire 2020, 3, 74–76. [Google Scholar] [CrossRef]
- Malik, N.; Simarani, K.; Aziz, M.A. Soybean as an alternative nutrient medium for Bacillus subtilis growth. Malaysian Applied Biology 2022, 51, 67–74. [Google Scholar] [CrossRef]
- Lynch, S.R.; Dassenko, S.A.; Cook, J.D.; Juillerat, M.A.; Hurrell, R.F. Inhibitory Effect of a Soybean-Protein-Related Moiety on Iron Absorption in Humans13. Am. J. Clin. Nutr. 1994, 60, 567–572. [Google Scholar] [CrossRef] [PubMed]
- Xu, X.; Sutak, R.; Richardson, D.R. Iron chelation by clinically relevant anthracyclines: alteration in expression of iron-regulated genes and atypical changes in intracellular iron distribution and trafficking. Mol. Pharmacol. 2008, 73, 833–44. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Zeng, Q.; Liu, X.; Chen, P.; Guo, X.; Ma, L.Z.; Dong, H.; Huang, Y. Iron reduction by diverse actinobacteria under oxic and pH-neutral conditions and the formation of secondary minerals. Chem. Geol. 2019, 525, 390–399. [Google Scholar] [CrossRef]
- He, Z.; Chang, J.; Feng, Y.; Wang, S.; Yuan, Q.; Liang, D.; Liu, J.; Li, N. Carbon nanotubes accelerates the bio-induced vivianite formation. Sci. Total Environ. 2022, 20, 844–157060. [Google Scholar] [CrossRef]
- Miot, J.; Benzerara, K.; Morin, G.; Bernard, S.; Beyssac, O.; Larquet, E.; Kappler, A.; Guyot, F. Transformation of vivianite by anaerobic nitrate-reducing iron-oxidizing bacteria. Geobiology. 2009, 7, 373–84. [Google Scholar] [CrossRef]
- Dong, J.; Ning, J.; Tian, Y.; Li, H.; Chen, H.; Guan, W. The involvement of multiple ABC transporters in daunorubicin efflux in Streptomyces coeruleorubidus. Microb. Biotechnol. 2024, 17, e70023. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, J.N.T.; Harbison, A.M. Scanning Electron Microscopy Sample Preparation and Imaging. Methods Mol. Biol. 2017, 1606, 71–84. [Google Scholar] [CrossRef]
- Rivera, M. Bacterioferritin: Structure, Dynamics, and Protein-Protein Interactions at Play in Iron Storage and Mobilization. Acc. Chem. Res. 2017, 50, 331–340. [Google Scholar] [CrossRef] [PubMed]
- Mahaseth, T.; Kuzminov, A. Potentiation of hydrogen peroxide toxicity: From catalase inhibition to stable DNA-iron complexes. Mutat. Res. Rev. Mutat. Res. 2017; 773, 274–281. [Google Scholar] [CrossRef]
- Kim, Y.; Roe, J.H.; Park, J.H.; Cho, Y.J.; Lee, K.L. Regulation of iron homeostasis by peroxide-sensitive CatR, a Fur-family regulator in Streptomyces coelicolour. J. Microbiol. 2021, 59, 1083–1091. [Google Scholar] [CrossRef] [PubMed]
- Toone, E.J. Advances in Enzymology and Related Areas of Molecular Biology, Protein Evolution; Wiley-Interscience, 2006. [CrossRef]
- Sirivech, S.; Driskell, J.; Frieden, E. NADH-FMN oxidoreductase activity and iron content of organs from riboflavin and iron-deficient rats. J Nutr. 1977, 107, 739–745. [Google Scholar] [CrossRef] [PubMed]
- Martelin, E.; Lapatto, R.; Raivio, K.O. Regulation of xanthine oxidoreductase by intracellular iron. Am. J. Physiol. Cell Physiol. 2002, 283, C1722–8. [Google Scholar] [CrossRef]
- Schnepf, M.I. A study of iron binding by soy protein. ETD collection for University of Nebraska-Lincoln. AAI8423827, 1984.
- Yang, R.; Zhu, L.; Meng, D.; Wang, Q.; Zhou, K.; Wang, Z.; Zhou, Z. Proteins from leguminous plants: from structure, property to the function in encapsulation/binding and delivery of bioactive compounds. Crit. Rev. Food Sci. Nutrition 2021, 62, 5203–5223. [Google Scholar] [CrossRef]
- Cascales, E.; Bernadac, A.; Gavioli, M.; Lazzaroni, J.C.; Lloubes, R. Pal lipoprotein of Escherichia coli plays a major role in outer membrane integrity. J. Bacteriol. 2002, 184, 754–759. [Google Scholar] [CrossRef] [PubMed]
- Ohe, T.; Watanabe, Y. Purification and properties of xanthine dehydrogenase from Streptomyces cyanogenus. J. Biochem. 1979, 86, 45–53. [Google Scholar] [PubMed]
- Leimkühler, S.; Iobbi-Nivol, Ch. Bacterial molybdoenzymes: old enzymes for new purposes. FEMS Microbiol. Rev. 2016, 40, 1–18. [Google Scholar] [CrossRef]
- Kantola, J.; Kunnari, T.; Hautala, A.; Hakala, J.; Ylihonko, K.; Mäntsälä, P. Elucidation of anthracyclinone biosynthesis by stepwise cloning of genes for anthracyclines from three different Streptomyces spp. Microbiology (Reading) 2000, 146, 155–163. [Google Scholar] [CrossRef]
- Youn, H.D.; Kim, E.J.; Roe, J.H.; Hah, Y.C.; Kang, S.O. A novel nickel-containing superoxide dismutase from Streptomyces spp. Biochem. J. 1996, 318, 889–896. [Google Scholar] [CrossRef]
- LeMaréchal, P.; Decottignies, P.; Marchand, CH.; Degrouard, J.; Jaillard, D.; Dulermo, T.; Froissard, M.; Smirnov, A.; Chapuis, V.; Virolle, M. Comparative Proteomic Analysis of Streptomyces lividans Wild-Type and ppk Mutant Strains Reveals the Importance of Storage Lipids for Antibiotic Biosynthesis. Appl. Environ. Microbiol. 2013, 79, 5907–5917 https://. [Google Scholar] [CrossRef] [PubMed]
- Chevrette, M.G.; Carlson, C.M.; Ortega, H.E.; Thomas, C.; Ananiev, G.E.; Barns, K.J.; Book, A.J.; Cagnazzo, J.; Carlos, C.; Flanigan, W.; Grubbs, K.J.; Horn, H.A.; Hoffmann, F.M.; Klassen, J.L.; Knack, J.J.; Lewin, G.R.; McDonald, B.R.; Muller, L.; Melo, W.G.P.; Pinto-Tomás, A.A.; Schmitz, A.; Wendt-Pienkowski, E.; Wildman, S.; Zhao, M.; Zhang, F.; Bugni, T.S.; Andes, D.R.; Pupo, M.T.; Currie, C.R. The antimicrobial potential of Streptomyces from insect microbiomes. Nat. Commun. 2019, 10, 516. [Google Scholar] [CrossRef] [PubMed]
- Van Moll, L.; De Smet, J.; Cos, P.; Van Campenhout, L. Microbial symbionts of insects as a source of new antimicrobials: a review. Crit. Rev. Microbiol. 2021, 47, 562–579. [Google Scholar] [CrossRef]
- De Smet, J.; Cos, P.; Van Campenhout, L. Microbial symbionts of insects as a source of new antimicrobials: a review. Crit. Rev. Microbiol. 2021, 47, 562–579. [Google Scholar] [CrossRef]
- Diarra U, Osborne-Naikatini T and Subramani R (2024) Actinomycetes associated with hymenopteran insects: a promising source of bioactive natural products. Front. Microbiol. 2024, 15, 1303010. [CrossRef]
- Dettner, K. Potential Pharmaceuticals from Insects and Their Co-Occurring Microorganisms. In: Insect Biotechnology. Biologically-Inspired Systems.; Vilcinskas, A. (eds); Springer, Dordrecht, 2011; Volume 2, pp. 95-119. [CrossRef]
- Bode, H.B. Insect-Associated Microorganisms as a Source for Novel Secondary Metabolites with Therapeutic Potential. In: Insect Biotechnology. Biologically-Inspired Systems.; Vilcinskas, A. (eds); Springer, Dordrecht, 2011; Volume 2, pp. 77-93. [CrossRef]



| Duration of cultivation [h] | N [samples] | Yield [mg/L] | pH | ||
|---|---|---|---|---|---|
| Average | SD | Average | SD | ||
| Baker’s yeast – 1 g/L | |||||
| 120 | 8 | 1116.5 | 656.9 | 7.66 | 0.53 |
| 168 | 8 | 1747.8 | 571.6 | 7.83 | 0.45 |
| 216 | 8 | 2138.6 | 512.5 | 8.47 | 0.27 |
| 264 | 8 | 2047.9 | 440.7 | 7.61 | 0.27 |
| Baker’s yeast – 5 g/L | |||||
| 120 | 12 | 967.8 | 858.5 | 7.96 | 0.64 |
| 168 | 12 | 1877.0 | 862.5 | 7.84 | 0.58 |
| 216 | 12 | 2162.7 | 299.3 | 8.49 | 0.25 |
| 264 | 12 | 2108.6 | 285.8 | 7.81 | 0.32 |
| Baker’s yeast – 10 g/L | |||||
| 120 | 8 | 1710.9 | 86.7 | 6.93 | 0.02 |
| 168 | 8 | 2986.3 | 176.9 | 6.84 | 0.03 |
| 216 | 8 | 4603.7 | 228.9 | 6.87 | 0.03 |
| 264 | 8 | 5448.1 | 108.5 | 6.91 | 0.05 |
| Duration of cultivation [h] | N [samples] | Yield [mg/L] | pH | ||
|---|---|---|---|---|---|
| Average | SD | Average | SD | ||
| Baker’s yeast | |||||
| 120 | 8 | 1710.9 | 86.7 | 6.93 | 0.02 |
| 168 | 8 | 2986.3 | 176.9 | 6.84 | 0.03 |
| 216 | 8 | 4603.7 | 228.9 | 6.87 | 0.03 |
| 264 | 8 | 5448.1 | 108.5 | 6.91 | 0.05 |
| Brewer’s yeast | |||||
| 120 | 7 | 600.6 | 154.7 | 6.87 | 0.04 |
| 168 | 7 | 1141.2 | 293.2 | 6.78 | 0.03 |
| 216 | 7 | 3353.8 | 279.8 | 6.70 | 0.02 |
| 264 | 7 | 4240.1 | 325.1 | 6.75 | 0.03 |
| Duration of cultivation [h] | N [samples] | Yield [mg/L] | pH | ||
|---|---|---|---|---|---|
| Average | SD | Average | SD | ||
| 120 | 11 | 1389.0 | 350.7 | 6.40 | 0.17 |
| 168 | 11 | 3070.2 | 724.6 | 6.62 | 0.13 |
| 216 | 11 | 4728.2 | 726.1 | 6.72 | 0.10 |
| 264 | 11 | 5466.4 | 681.4 | 7.24 | 0.41 |
| Group One: | Group Two: |
|
|
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




