Submitted:
26 November 2024
Posted:
28 November 2024
You are already at the latest version
Abstract
In 2022, the World Health Organization reported that tuberculosis (TB) was the second highest cause of death globally from a single infectious agent, following COVID-19. Development of new antitubercular agents with a novel mechanism of action for use in the complex TB therapy is considered one of the key approaches for fighting TB. In this study we examined the gene expression profile of M. smegmatis when exposed to a promising antituberculosis agent quinoxaline 1,4-dioxide (QdNO) 7-chloro-2-(ethoxycarbonyl)-3-methyl-6-(piperazin-1-yl)quinoxaline-1,4-dioxide-1 (LCTA-3368). We looked at how the bacterial response changed with different doses (1/4 × MIC, 1/2 × MIC, and 1 x MIC) and durations (30 min and 90 min) of treatment with the drug. Our analysis revealed significant upregulation of the expression of genes involved in DNA repair and replication processes, and changes in the expression of 95 genes encoding proteins with oxidoreductase activity. This confirmed the assumption about the mechanism of antibacterial action of QdNOs associated with the formation of free radicals leading to DNA damage.
Keywords:
1. Introduction
2. Materials and Methods
2.1. Microbial Cultures and Growth Conditions
2.2. Total RNA Extraction
2.3. Library Preparation and RNA Sequencing
2.4. Bioinformatics Analysis
3. Results
3.1. Whole-Transcriptome Analysis
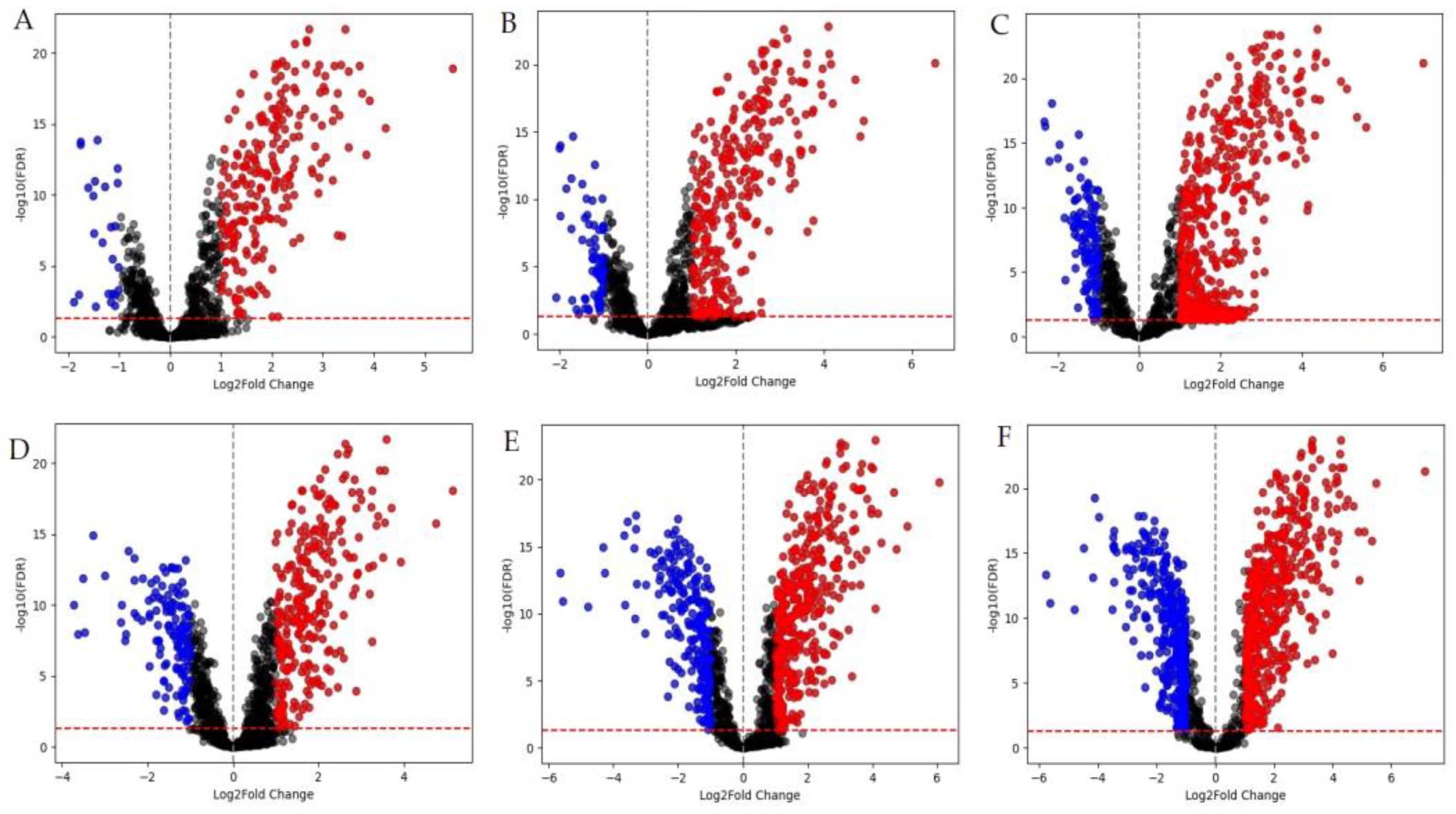
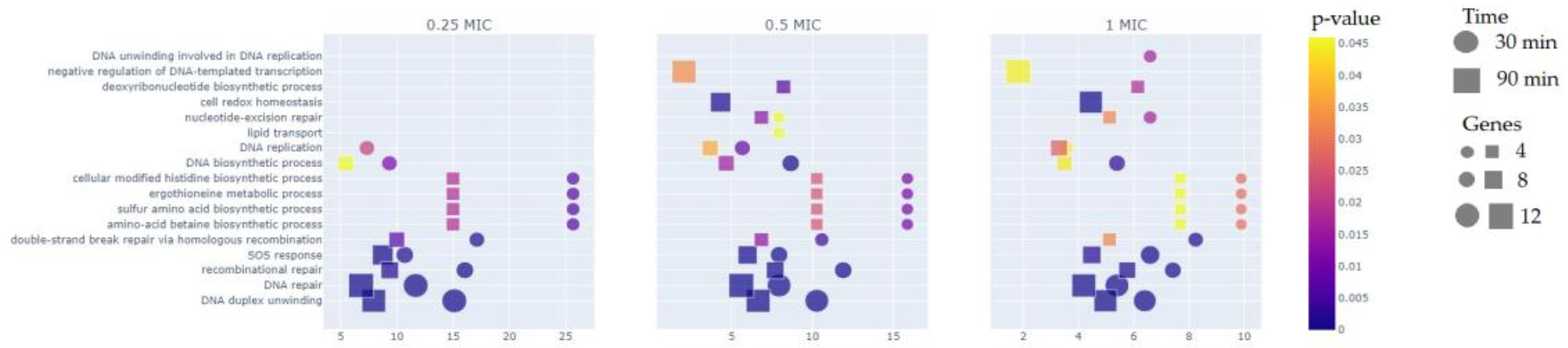
3.2. Functional Groups of Differentially Expressed Genes
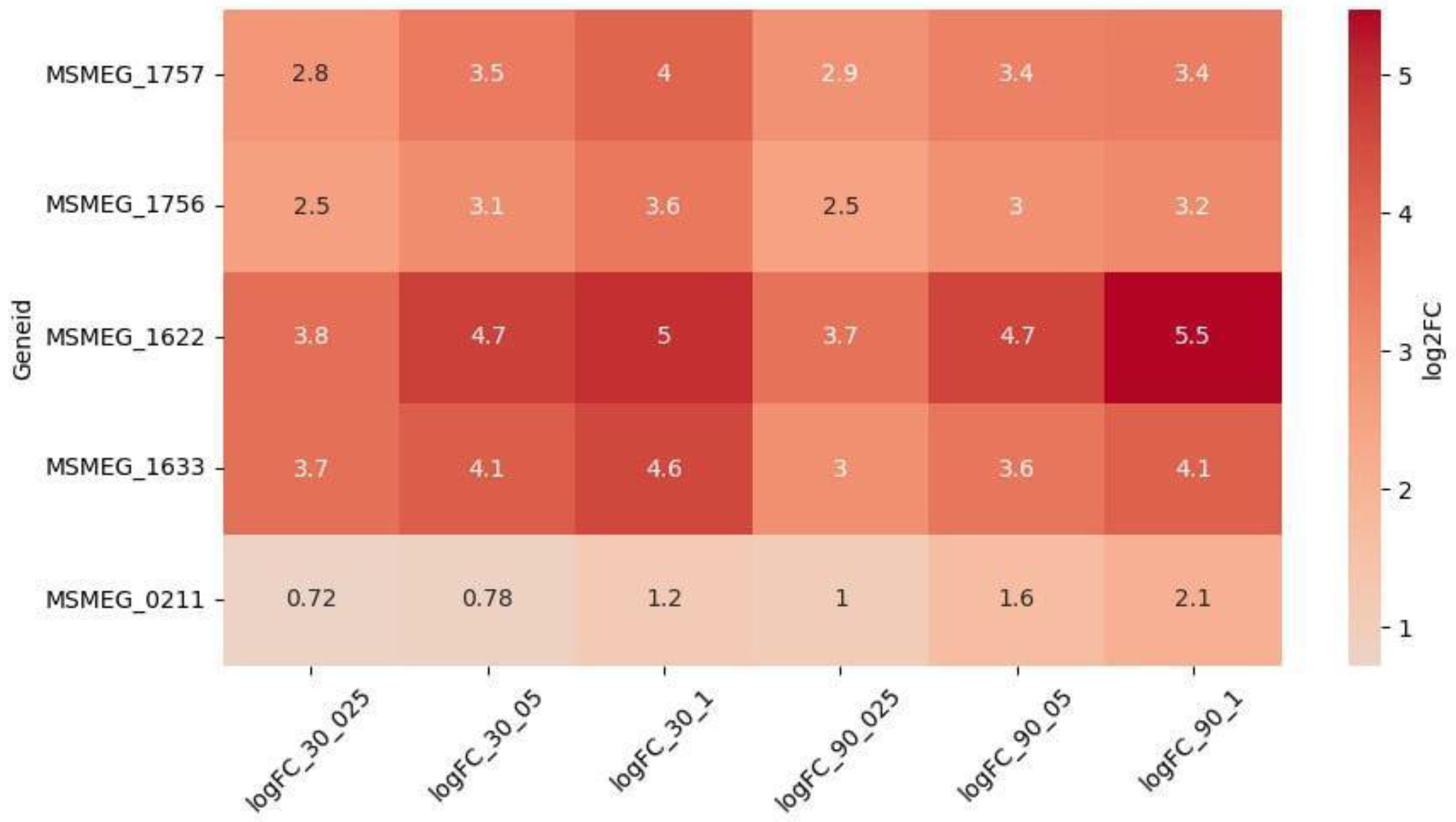
3.2.1. Reparation Genes
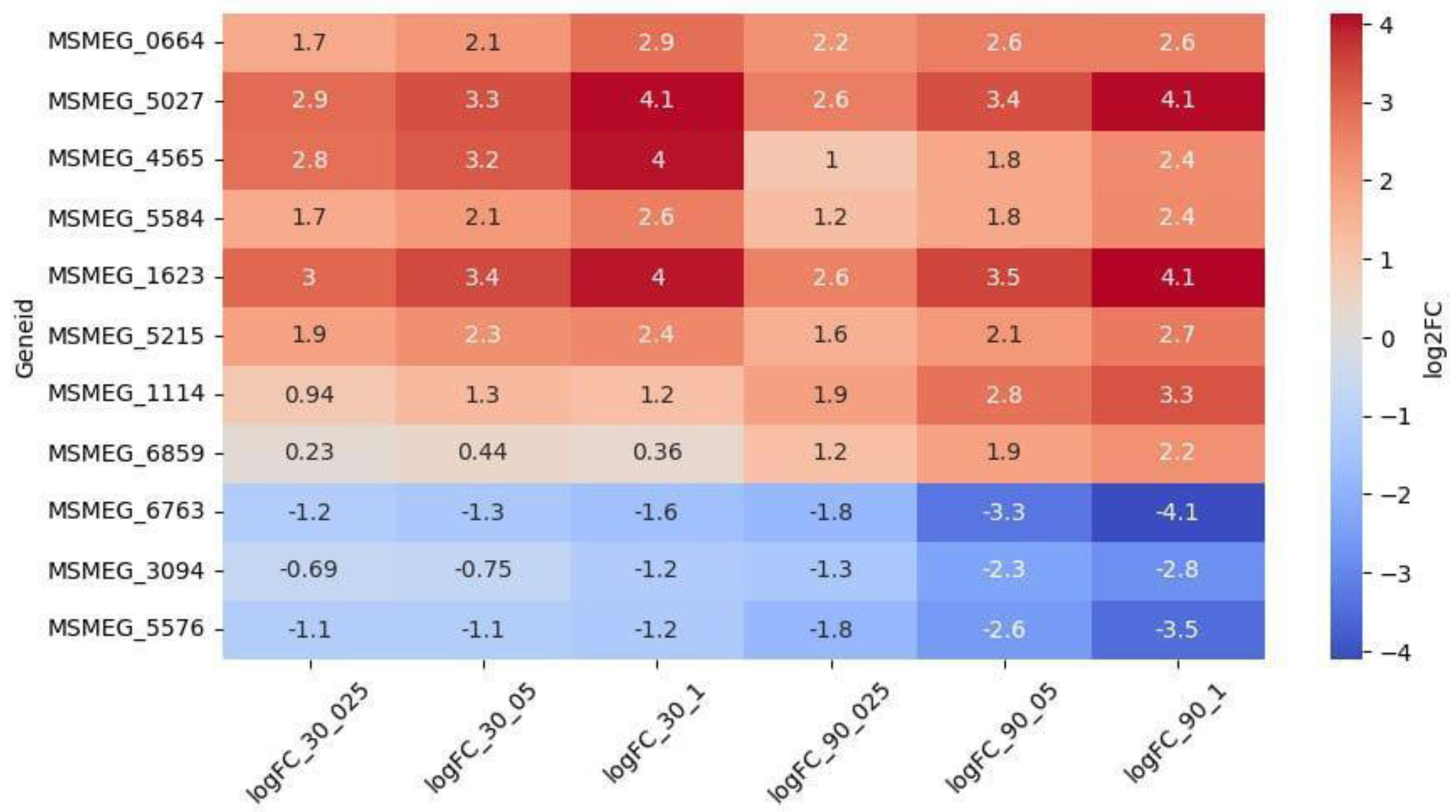
3.2.2. Oxidoreductases
Other Genes
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Global Tuberculosis Report 2023 Available online: https://www.who.int/teams/global-tuberculosis-programme/tb-reports/global-tuberculosis-report-2023.
- Cheng, G.; Sa, W.; Cao, C.; Guo, L.; Hao, H.; Liu, Z.; Wang, X.; Yuan, Z. Quinoxaline 1,4-Di-N-Oxides: Biological Activities and Mechanisms of Actions. Front Pharmacol 2016, 7, 64. [Google Scholar] [CrossRef] [PubMed]
- Zanetti, S.; Sechi, L.A.; Molicotti, P.; Cannas, S.; Bua, A.; Deriu, A.; Carta, A.; Paglietti, G. In Vitro Activity of New Quinoxalin 1,4-Dioxide Derivatives against Strains of Mycobacterium Tuberculosis and Other Mycobacteria. Int J Antimicrob Agents 2005, 25, 179–181. [Google Scholar] [CrossRef] [PubMed]
- Ortega, M.A.; Montoya, M.E.; Jaso, A.; Zarranz, B.; Tirapu, I.; Aldana, I.; Monge, A. Antimycobacterial Activity of New Quinoxaline-2-Carbonitrile and Quinoxaline-2-Carbonitrile 1,4-Di-N-Oxide Derivatives. Pharmazie 2001, 56, 205–207. [Google Scholar] [CrossRef] [PubMed]
- Villar, R.; Vicente, E.; Solano, B.; Pérez-Silanes, S.; Aldana, I.; Maddry, J.A.; Lenaerts, A.J.; Franzblau, S.G.; Cho, S.-H.; Monge, A.; et al. In Vitro and in Vivo Antimycobacterial Activities of Ketone and Amide Derivatives of Quinoxaline 1,4-Di-N-Oxide. J Antimicrob Chemother 2008, 62, 547–554. [Google Scholar] [CrossRef]
- Crawford, P.W.; Scamehorn, R.G.; Hollstein, U.; Ryan, M.D.; Kovacic, P. Cyclic Voltammetry of Phenazines and Quinoxalines Including Mono- and Di-N-Oxides. Relation to Structure and Antimicrobial Activity. Chem Biol Interact 1986, 60, 67–84. [Google Scholar] [CrossRef]
- Chowdhury, G.; Kotandeniya, D.; Daniels, J.S.; Barnes, C.L.; Gates, K.S. Enzyme-Activated, Hypoxia-Selective DNA Damage by 3-Amino-2-Quinoxalinecarbonitrile 1,4-Di-N-Oxide. Chem Res Toxicol 2004, 17, 1399–1405. [Google Scholar] [CrossRef]
- Frolova, S.G.; Vatlin, A.A.; Maslov, D.A.; Yusuf, B.; Buravchenko, G.I.; Bekker, O.B.; Klimina, K.M.; Smirnova, S.V.; Shnakhova, L.M.; Malyants, I.K.; et al. Novel Derivatives of Quinoxaline-2-Carboxylic Acid 1,4-Dioxides as Antimycobacterial Agents: Mechanistic Studies and Therapeutic Potential. Pharmaceuticals (Basel) 2023, 16, 1565. [Google Scholar] [CrossRef]
- Cooper, C.B. Development of Mycobacterium Tuberculosis Whole Cell Screening Hits as Potential Antituberculosis Agents. J Med Chem 2013, 56, 7755–7760. [Google Scholar] [CrossRef]
- King, G.M. Uptake of Carbon Monoxide and Hydrogen at Environmentally Relevant Concentrations by Mycobacteria†. Appl Environ Microbiol 2003, 69, 7266–7272. [Google Scholar] [CrossRef]
- Junnotula, V.; Sarkar, U.; Sinha, S.; Gates, K.S. Initiation of DNA Strand Cleavage by 1,2,4-Benzotriazine 1,4-Dioxide Antitumor Agents: Mechanistic Insight from Studies of 3-Methyl-1,2,4-Benzotriazine 1,4-Dioxide. J Am Chem Soc 2009, 131, 1015–1024. [Google Scholar] [CrossRef]
- Briffotaux J, Liu S, Gicquel B. Genome-Wide Transcriptional Responses of Mycobacterium to Antibiotics. Front Microbiol. [CrossRef]
- Vatlin, A.A.; Shitikov, E.A.; Shahbaaz, M.; Bespiatykh, D.A.; Klimina, K.M.; Christoffels, A.; Danilenko, V.N.; Maslov, D.A. Transcriptomic Profile of Mycobacterium Smegmatis in Response to an Imidazo[1,2-b][1,2,4,5]Tetrazine Reveals Its Possible Impact on Iron Metabolism. Front Microbiol 2021, 12, 724042. [Google Scholar] [CrossRef] [PubMed]
- Logsdon, M.M.; Aldridge, B.B. Stable Regulation of Cell Cycle Events in Mycobacteria: Insights From Inherently Heterogeneous Bacterial Populations. Front Microbiol 2018, 9, 514. [Google Scholar] [CrossRef] [PubMed]
- Babraham Bioinformatics - FastQC A Quality Control Tool for High Throughput Sequence Data Available online: https://www.bioinformatics.babraham.ac.uk/projects/fastqc/.
- Ewels, P.; Magnusson, M.; Lundin, S.; Käller, M. MultiQC: Summarize Analysis Results for Multiple Tools and Samples in a Single Report. Bioinformatics 2016, 32, 3047–3048. [Google Scholar] [CrossRef] [PubMed]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A Flexible Trimmer for Illumina Sequence Data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef]
- Kim, D.; Paggi, J.M.; Park, C.; Bennett, C.; Salzberg, S.L. Graph-Based Genome Alignment and Genotyping with HISAT2 and HISAT-Genotype. Nat Biotechnol 2019, 37, 907–915. [Google Scholar] [CrossRef]
- Okonechnikov, K.; Conesa, A.; García-Alcalde, F. Qualimap 2: Advanced Multi-Sample Quality Control for High-Throughput Sequencing Data. Bioinformatics 2016, 32, 292–294. [Google Scholar] [CrossRef]
- Liao, Y.; Smyth, G.K.; Shi, W. featureCounts: An Efficient General Purpose Program for Assigning Sequence Reads to Genomic Features. Bioinformatics 2014, 30, 923–930. [Google Scholar] [CrossRef]
- Robinson, M.D.; McCarthy, D.J.; Smyth, G.K. edgeR: A Bioconductor Package for Differential Expression Analysis of Digital Gene Expression Data. Bioinformatics 2010, 26, 139–140. [Google Scholar] [CrossRef]
- R: The R Project for Statistical Computing Available online: https://www.r-project.org/index.html.
- Python 3.13 Documentation — DevDocs Available online: https://devdocs.io/python/.
- Matplotlib Documentation — Matplotlib 3.9.2 Documentation Available online: https://matplotlib.org/stable/index.html.
- NumPy Documentation Available online: https://numpy.org/doc/.
- GOpiscator 0.1.5 Available online: https://pypi.org/project/gopiscator/.
- Xu, F; Cheng, G; Hao, H; Wang, Y; Wang, X; Chen, D; Peng, D; Liu, Z; Yuan, Z; Dai, M. Mechanisms of Antibacterial Action of Quinoxaline 1,4-di-N-oxides against Clostridium perfringens and Brachyspira hyodysenteriae. Front Microbiol. 2016, 7 - 1948. [CrossRef]
- Suter, W.; Rosselet, A.; Knüsel, F. Mode of Action of Quindoxin and Substituted Quinoxaline-Di-N-Oxides on Escherichia Coli. Antimicrob Agents Chemother 1978, 13, 770–783. [Google Scholar] [CrossRef]
- DuPlessis, E.R.; Pellett, J.; Stankovich, M.T.; Thorpe, C. Oxidase Activity of the Acyl-CoA Dehydrogenases. Biochemistry 1998, 37, 10469–10477. [Google Scholar] [CrossRef]
- Padavattan, S.; Jos, S.; Gogoi, H.; Bagautdinov, B. Crystal Structure of Enoyl-CoA Hydratase from Thermus Thermophilus HB8. Acta Crystallogr F Struct Biol Commun 2021, 77, 148–155. [Google Scholar] [CrossRef] [PubMed]
- Savolainen, K.; Bhaumik, P.; Schmitz, W.; Kotti, T.J.; Conzelmann, E.; Wierenga, R.K.; Hiltunen, J.K. Alpha-Methylacyl-CoA Racemase from Mycobacterium Tuberculosis. Mutational and Structural Characterization of the Active Site and the Fold. J Biol Chem 2005, 280, 12611–12620. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Zhang, H.; Huang, L.; Pan, Y.; Li, J.; Chen, D. Deoxidation rates play a critical role in DNA damage mediated by important synthetic drugs, quinoxaline 1,4-dioxides. Chemical Research in Toxicology 2015, 28(3), 470–481. [Google Scholar] [CrossRef] [PubMed]
- Haoxian, An; Yonggang, Li; Yanshen, Li; Shanmin, Gong; Ya’ning, Zhu; Xinru, Li; Shuang, Zhou; Yongning, Wu. Advances in Metabolism and Metabolic Toxicology of Quinoxaline 1,4-Di-N-oxides. Chemical Research in Toxicology 2024, 37 (4), 528-539. [CrossRef]
- Gupta, R.; Barkan, D.; Redelman-Sidi, G.; Shuman, S.; Glickman, M.S. Mycobacteria Exploit Three Genetically Distinct DNA Double-Strand Break Repair Pathways. Mol Microbiol 2011, 79, 316–330. [Google Scholar] [CrossRef] [PubMed]
- Sharma, A.; Nair, D.T. Cloning, Expression, Purification, Crystallization and Preliminary Crystallographic Analysis of MsDpo4: A Y-Family DNA Polymerase from Mycobacterium Smegmatis. Acta Crystallogr Sect F Struct Biol Cryst Commun 2011, 67, 812–816. [Google Scholar] [CrossRef]
- Dos Vultos, T.; Mestre, O.; Tonjum, T.; Gicquel, B. DNA Repair in Mycobacterium Tuberculosis Revisited. FEMS Microbiol Rev 2009, 33, 471–487. [Google Scholar] [CrossRef]
- Riccardo, Miggiano; Castrese, Morrone; Franca, Rossi; Menico, Rizzi. Targeting Genome Integrity in Mycobacterium Tuberculosis: From Nucleotide Synthesis to DNA Replication and Repair. Molecules 2020, 25(5), 1205. [CrossRef]
- Chen, X.; Bradley, N.P.; Lu, W.; Wahl, K.L.; Zhang, M.; Yuan, H.; Hou, X.-F.; Eichman, B.F.; Tang, G.-L. Base Excision Repair System Targeting DNA Adducts of Trioxacarcin/LL-D49194 Antibiotics for Self-Resistance. Nucleic Acids Res 2022, 50, 2417–2430. [Google Scholar] [CrossRef]
- Khanam, T.; Rai, N.; Ramachandran, R. Mycobacterium Tuberculosis Class II Apurinic/Apyrimidinic-Endonuclease/3’-5’ Exonuclease III Exhibits DNA Regulated Modes of Interaction with the Sliding DNA β-Clamp. Mol Microbiol 2015, 98, 46–68. [Google Scholar] [CrossRef]
- Daniel, Nätt; Annika, Thorsell. Stress-induced transposon reactivation: a mediator or an estimator of allostatic load? Environ Epigenet. [CrossRef]
- Frolova, S.G. , Klimina K.M., Kumar R., Vatlin A.A., Salunke D.B., Kendrekar P., Danilenko V.N., Maslov D.A. Identification of Mutations Conferring Tryptanthrin Resistance to Mycobacterium Smegmatis. Antibiotics, 2021, 10(1), 6.
- Ganley, B.; Chowdhury, G.; Bhansali, J.; Daniels, J. S.; Gates, K. S. Redox-activated, hypoxia-selective DNA cleavage by quinoxaline 1,4-di-N-oxide. Bioorg. Med. Chem 2001, 9, 2395–2401, 2395. [Google Scholar]
- Ashwani, Kumar; Aisha, Farhana; Loni, Guidry; Vikram, Saini; Mary, Hondalus; Adrie, JC. Steyn Redox homeostasis in mycobacteria: the key to tuberculosis control? Expert Rev Mol Med. 2011, 13. [Google Scholar] [CrossRef]
- Ganley, B.; Chowdhury, G.; Bhansali, J.; Daniels, J. S.; Gates, K. S. . Redox-activated, hypoxia-selective DNA cleavage by quinoxaline 1,4-di-N-oxide. Bioorg. Med. Chem 2001, 9, 2395–2401. [Google Scholar] [CrossRef]
- Gurumurthy, M.; Rao, M.; Mukherjee, T. ; Rao, et al. A novel F420-dependent anti-oxidant mechanism protects Mycobacterium tuberculosis against oxidative stress and bactericidal agents. Mol. Microbiol 2013, 87, 744–755. [CrossRef]
- Greening, C.; Jirapanjawat, T.; Afroze, S.; Ney, B.; Scott, C.; Pandey, G.; Lee, BM.; Russell, RJ.; Jackson, CJ.; Oakeshott, JG.; Taylor, MC.; Warden, AC. ; Mycobacterial F420H2-Dependent Reductases Promiscuously Reduce Diverse Compounds through a Common Mechanism. Front. Microbiol. 2017, 8, 1000. [Google Scholar] [CrossRef]
- Vatlin, A.A.; Frolova, S.G.; Bekker, O.B.; Danilenko, V.N. Studying the mechanism of action of new derivatives of quinoxalin-1,4-dioxide on the model organism Mycobacterium smegmatis. RUDN Journal of Ecology and Life Safety 2024, 32(1), 41–50. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



