Submitted:
19 November 2024
Posted:
20 November 2024
You are already at the latest version
Abstract
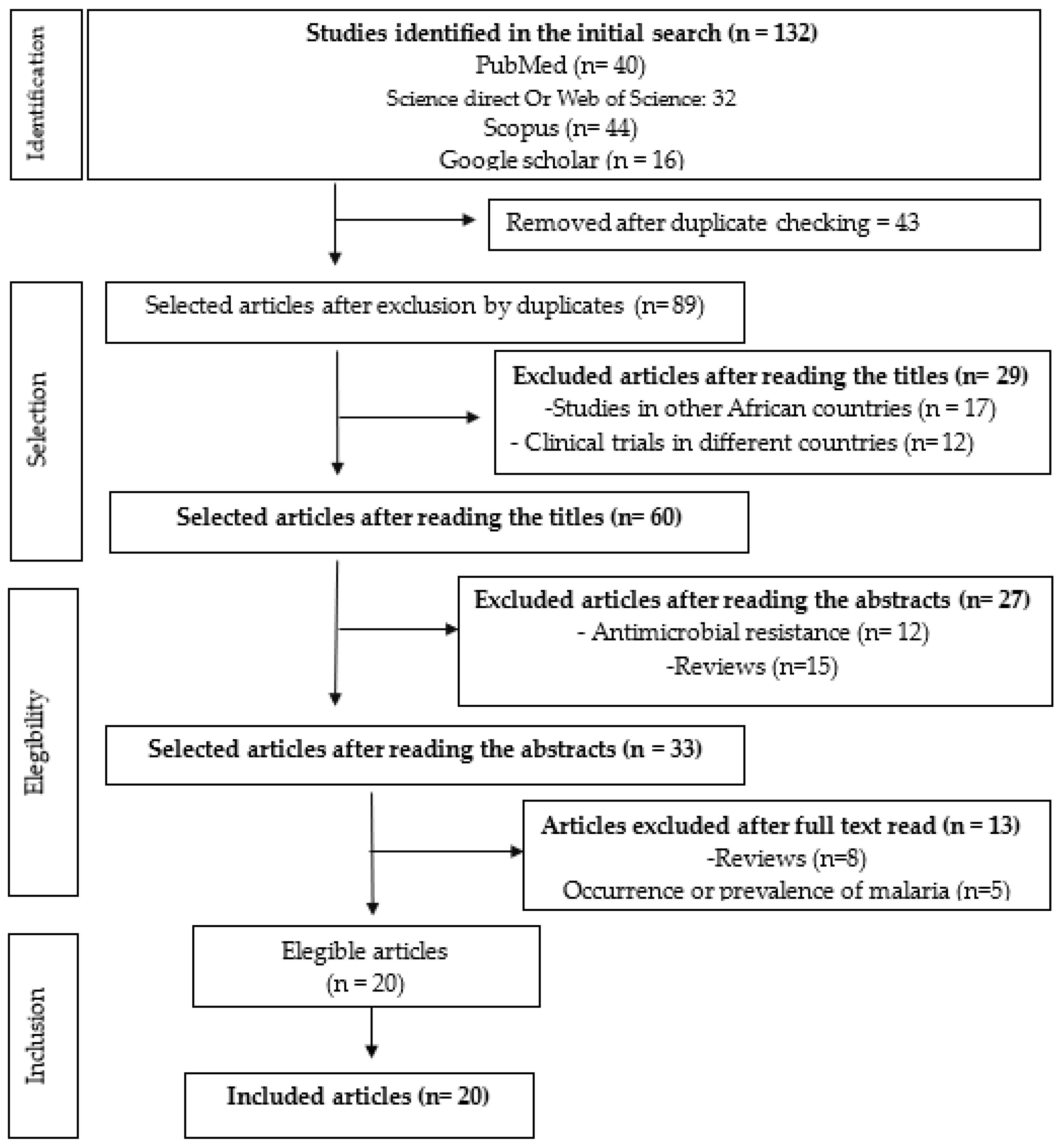
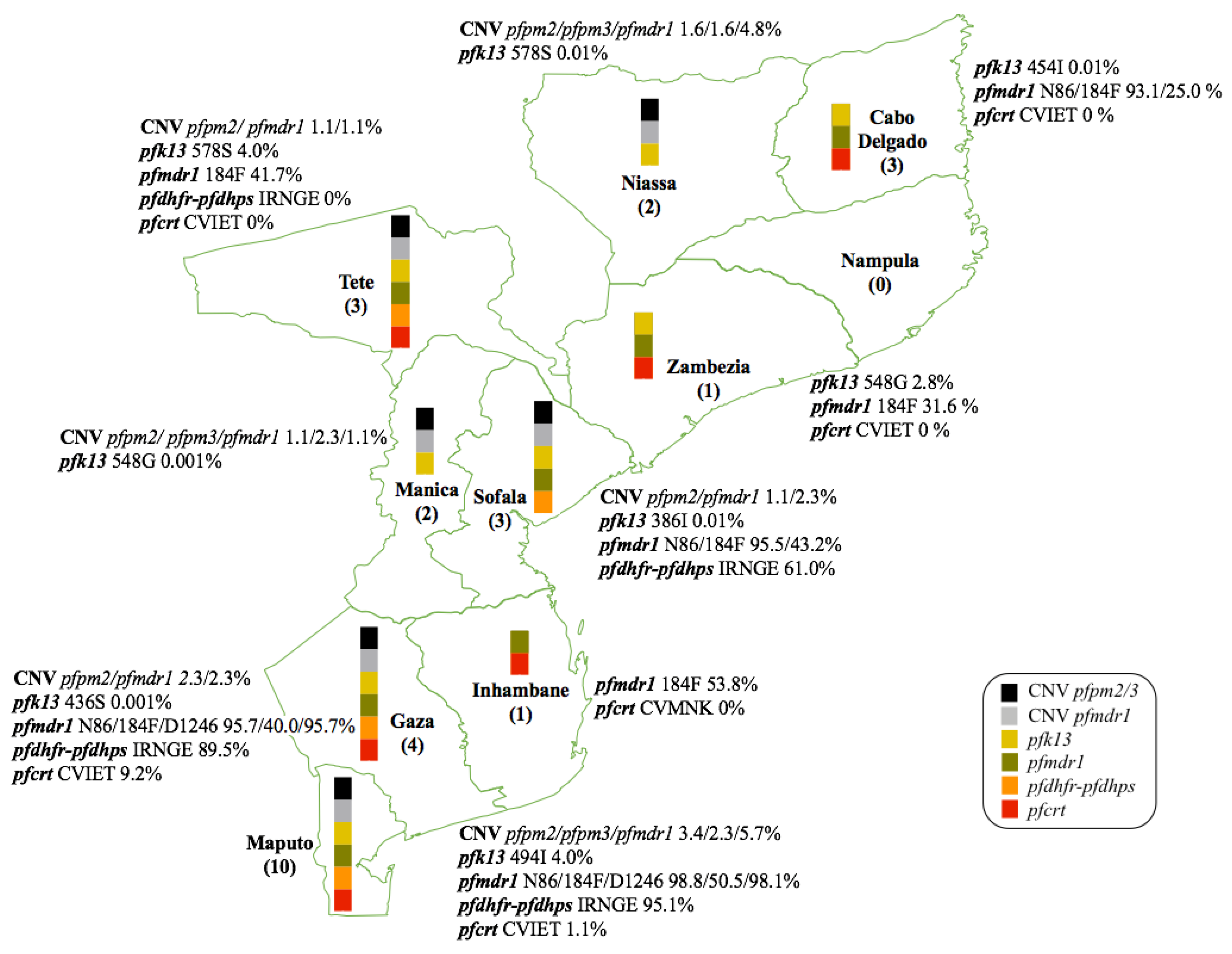
Malaria continues to be a significant public health burden in many tropical and subtropical regions. Mozambique ranks among the top countries affected by malaria, where it is a leading cause of morbidity and mortality, accounting for 29% of all hospital deaths in the general population and 42% of deaths amongst children under five. This review presents a comparative analysis of data on five critical genes associated with antimalarial drug resistance: pfmdr1, pfcrt, pfk13, pfdhfr and pfdhps, along with copy number variation (CNV) in genes pfmdr1 and pfpm2/3. These are genes associated with parasite response to currently used for antimalarials to treat uncomplicated P. falciparum malaria in Mozambique. The review synthesizes data collected from published studies conducted in Mozambique after the introduction of ACTs (2006) up to June 2024, highlighting the presence or absence of mutations in these genes across Mozambique. We aimed at mapping the prevalence and distribution of these molecular markers, across the country, in order to contribute to the development of targeted interventions to sustain the efficacy of malaria treatments in Mozambique. Four databases were used to access the articles: PubMed, Science direct, Scopus and Google scholar. The search strategy identified 132 studies addressing malaria and antimalarial resistance. Of these, 112 were excluded for various reasons, leaving 20 studies to be included in this review. Children and pregnant women represent the majority of target groups in studies on all types of antimalarials. Most studies (87.5%) were conducted in the provinces of Maputo and Gaza. The primary alleles re-ported were pfcrt CVMNK, and in the most recent data, its wild-type form was found in the majority of patients. Low prevalence of mutations in the pfk13 gene, were identified reflecting the effectiveness of ACTs. In pfk13 only mutation A578S was reported, in Niassa and Tete. Regarding CNVs were observed in studies carried out in the south of Mozambique, for pfmdr1 with a frequency of 1.1-5.1% and pfpm2 with a frequency of 1.1-3.4%. This review indicates that molecular markers linked to malaria resistance show considerable variation across provinces in Mozambique, with most up-to-date data accessible for Maputo and Gaza. In contrast, provinces such as Zambezia and Inhambane have limited data on several genes, while Nampula lacks data on all drug resistance markers. Our review reveals that gene mutations associated with antimalarial resistance, vary considerably by province in Mozambique, with more up-to-date data available for Maputo and Gaza. Other provinces, including Zambezia, Inhambane, and Cabo Delgado, have limited data on several genes, while Nampula entirely lacks data on all drug resistance molecular markers.
Keywords:
1. Introduction
2. Methods
2.1. Selection of Relevant Literature
2.2. Eligibility Criteria of Studies Include in the Review
2.3. Screening and Data Extraction
3. Results and Discussion
3.1. Basic Characteristics of Included Studies
3.2. Antimalarial Resistance Associated Polymorphisms
3.2.5. Copy Number Variations in pfmdr1 and pfpm2/3
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
Abbreviations
References
- Ippolito, M.M.; Moser, K.A.; Kabuya, J.-B.B.; Cunningham, C.; Juliano, J.J. Antimalarial Drug Resistance and Implications for the WHO Global Technical Strategy. Curr Epidemiol Rep 2021, 8, 46–62. [CrossRef]
- Oladipo, H.J.; Tajudeen, Y.A.; Oladunjoye, I.O.; Yusuff, S.I.; Yusuf, R.O.; Oluwaseyi, E.M.; AbdulBasit, M.O.; Adebisi, Y.A.; El-Sherbini, M.S. Increasing Challenges of Malaria Control in Sub-Saharan Africa: Priorities for Public Health Research and Policymakers. Annals of Medicine & Surgery 2022, 81. [CrossRef]
- Li, J.; Docile, H.J.; Fisher, D.; Pronyuk, K.; Zhao, L. Current Status of Malaria Control and Elimination in Africa: Epidemiology, Diagnosis, Treatment, Progress and Challenges. J Epidemiol Glob Health 2024. [CrossRef]
- WHO World Malaria Report 2023.Http://Www.Who.Int/Malaria/Publications/World-Malaria-Report-2023; 2023;
- Hemingway, J.; Shretta, R.; Wells, T.N.C.; Bell, D.; Djimdé, A.A.; Achee, N.; Qi, G. Tools and Strategies for Malaria Control and Elimination: What Do We Need to Achieve a Grand Convergence in Malaria? PLoS Biol 2016, 14, e1002380. [CrossRef]
- Plowe, C. V. Malaria Chemoprevention and Drug Resistance: A Review of the Literature and Policy Implications. Malar J 2022, 21, 104. [CrossRef]
- Rasmussen, C.; Alonso, P.; Ringwald, P. Current and Emerging Strategies to Combat Antimalarial Resistance. Expert Rev Anti Infect Ther 2022, 20, 353–372. [CrossRef]
- Thu, A.M.; Phyo, A.P.; Landier, J.; Parker, D.M.; Nosten, F.H. Combating Multidrug-resistant Plasmodium Falciparum Malaria. FEBS J 2017, 284, 2569–2578. [CrossRef]
- Wongsrichanalai, C.; Pickard, A.L.; Wernsdorfer, W.H.; Meshnick, S.R. Epidemiology of Drug-Resistant Malaria. Lancet Infect Dis 2002, 2, 209–218. [CrossRef]
- Raman, J.; Mauff, K.; Muianga, P.; Mussa, A.; Maharaj, R.; Barnes, K.I. Five Years of Antimalarial Resistance Marker Surveillance in Gaza Province, Mozambique, Following Artemisinin-Based Combination Therapy Roll Out. PLoS One 2011, 6, e25992. [CrossRef]
- Ecker, A.; Lehane, A.M.; Clain, J.; Fidock, D.A. PfCRT and Its Role in Antimalarial Drug Resistance. Trends Parasitol 2012, 28, 504–514. [CrossRef]
- Mohammed, A.; Ndaro, A.; Kalinga, A.; Manjurano, A.; Mosha, J.F.; Mosha, D.F.; van Zwetselaar, M.; Koenderink, J.B.; Mosha, F.W.; Alifrangis, M.; et al. Trends in Chloroquine Resistance Marker, Pfcrt-K76T Mutation Ten Years after Chloroquine Withdrawal in Tanzania. Malar J 2013, 12, 415. [CrossRef]
- Foguim, F.T.; Bogreau, H.; Gendrot, M.; Mosnier, J.; Fonta, I.; Benoit, N.; Amalvict, R.; Madamet, M.; Wein, S.; Pradines, B.; et al. Prevalence of Mutations in the Plasmodium Falciparum Chloroquine Resistance Transporter, PfCRT, and Association with Ex Vivo Susceptibility to Common Anti-Malarial Drugs against African Plasmodium Falciparum Isolates. Malar J 2020, 19, 201. [CrossRef]
- Bygbjerg, I.C.; Alifrangis, M.; Tomás, E.V.E.; Charlwood, D.; Thomsen, T.T.; Madsen, L.B.; Hansson, H.H. Rapid Selection of Plasmodium Falciparum Chloroquine Resistance Transporter Gene and Multidrug Resistance Gene-1 Haplotypes Associated with Past Chloroquine and Present Artemether-Lumefantrine Use in Inhambane District, Southern Mozambique. Am J Trop Med Hyg 2013, 88, 536–541. [CrossRef]
- Mehlotra, R.K.; Fujioka, H.; Roepe, P.D.; Janneh, O.; Ursos, L.M.B.; Jacobs-Lorena, V.; McNamara, D.T.; Bockarie, M.J.; Kazura, J.W.; Kyle, D.E.; et al. Evolution of a Unique Plasmodium Falciparum Chloroquine-Resistance Phenotype in Association with Pfcrt Polymorphism in Papua New Guinea and South America. Proceedings of the National Academy of Sciences 2001, 98, 12689–12694. [CrossRef]
- Wootton, J.C.; Feng, X.; Ferdig, M.T.; Cooper, R.A.; Mu, J.; Baruch, D.I.; Magill, A.J.; Su, X. Genetic Diversity and Chloroquine Selective Sweeps in Plasmodium Falciparum. Nature 2002, 418, 320–323. [CrossRef]
- Roper, C.; Maharaj, R.; Raman, J.; Kleinschmidt, I.; Little, F.; Barnes, K.I.; Cassam, Y. Five Years of Large-Scale Dhfr and Dhps Mutation Surveillance Following the Phased Implementation of Artesunate Plus Sulfadoxine-Pyrimethamine in Maputo Province, Southern Mozambique. Am J Trop Med Hyg 2010, 82, 788–794. [CrossRef]
- PMI Presitent Malaria Iniciative.
- Esu, E.; Tacoli, C.; Gai, P.; Berens-Riha, N.; Pritsch, M.; Loescher, T.; Meremikwu, M. Prevalence of the Pfdhfr and Pfdhps Mutations among Asymptomatic Pregnant Women in Southeast Nigeria. Parasitol Res 2018, 117, 801–807. [CrossRef]
- Naidoo, I.; Roper, C. Mapping ‘Partially Resistant’, ‘Fully Resistant’, and ‘Super Resistant’ Malaria. Trends Parasitol 2013, 29, 505–515. [CrossRef]
- Nhama, A.; Nhamússua, L.; Macete, E.; Bassat, Q.; Salvador, C.; Enosse, S.; Candrinho, B.; Carvalho, E.; Nhacolo, A.; Chidimatembue, A.; et al. In Vivo Efficacy and Safety of Artemether–Lumefantrine and Amodiaquine–Artesunate for Uncomplicated Plasmodium Falciparum Malaria in Mozambique, 2018. Malar J 2021, 20, 390. [CrossRef]
- Nsanzabana, C. Resistance to Artemisinin Combination Therapies (ACTs): Do Not Forget the Partner Drug! Trop Med Infect Dis 2019, 4, 26. [CrossRef]
- Li, Y. Artemisinin and Derivatives Pharmacodynamics, Toxicology, Pharmacokinetics, Mechanism of Action, Resistance, and Immune Regulation. In Artemisinin-Based and Other Antimalarials; Li, G., Li, Y., Li, Z., Zeng, M., Eds.; Academic Press: Cambridge, MA, USA, 2018; Chapter 5. In Artemisinin-Based and Other Antimalarials; Elsevier, 2018; pp. 197–351.
- Dhorda, M.; Amaratunga, C.; Dondorp, A.M. Artemisinin and Multidrug-Resistant Plasmodium Falciparum – a Threat for Malaria Control and Elimination. Curr Opin Infect Dis 2021, 34, 432–439. [CrossRef]
- Wicht, K.J.; Mok, S.; Fidock, D.A. Molecular Mechanisms of Drug Resistance in Plasmodium Falciparum Malaria. Annu Rev Microbiol 2020, 74, 431–454. [CrossRef]
- Phyo, A.P.; Ashley, E.A.; Anderson, T.J.C.; Bozdech, Z.; Carrara, V.I.; Sriprawat, K.; Nair, S.; White, M.M.; Dziekan, J.; Ling, C.; et al. Declining Efficacy of Artemisinin Combination Therapy Against P. Falciparum Malaria on the Thai–Myanmar Border (2003–2013): The Role of Parasite Genetic Factors. Clinical Infectious Diseases 2016, 63, 784–791. [CrossRef]
- Price, R.N.; Uhlemann, A.-C.; Brockman, A.; McGready, R.; Ashley, E.; Phaipun, L.; Patel, R.; Laing, K.; Looareesuwan, S.; White, N.J.; et al. Mefloquine Resistance in Plasmodium Falciparum and Increased Pfmdr1 Gene Copy Number. The Lancet 2004, 364, 438–447. [CrossRef]
- Eyase, F.L.; Akala, H.M.; Ingasia, L.; Cheruiyot, A.; Omondi, A.; Okudo, C.; Juma, D.; Yeda, R.; Andagalu, B.; Wanja, E.; et al. The Role of Pfmdr1 and Pfcrt in Changing Chloroquine, Amodiaquine, Mefloquine and Lumefantrine Susceptibility in Western-Kenya P. Falciparum Samples during 2008–2011. PLoS One 2013, 8, e64299. [CrossRef]
- Borges, S.; Cravo, P.; Creasey, A.; Fawcett, R.; Modrzynska, K.; Rodrigues, L.; Martinelli, A.; Hunt, P. Genomewide Scan Reveals Amplification of Mdr1 as a Common Denominator of Resistance to Mefloquine, Lumefantrine, and Artemisinin in Plasmodium Chabaudi Malaria Parasites. Antimicrob Agents Chemother 2011, 55, 4858–4865. [CrossRef]
- Preechapornkul, P.; Imwong, M.; Chotivanich, K.; Pongtavornpinyo, W.; Dondorp, A.M.; Day, N.P.J.; White, N.J.; Pukrittayakamee, S. Plasmodium Falciparum Pfmdr1 Amplification, Mefloquine Resistance, and Parasite Fitness. Antimicrob Agents Chemother 2009, 53, 1509–1515. [CrossRef]
- Sidhu, A.B.S.; Valderramos, S.G.; Fidock, D.A. Pfmdr1 Mutations Contribute to Quinine Resistance and Enhance Mefloquine and Artemisinin Sensitivity in Plasmodium Falciparum. Mol Microbiol 2005, 57, 913–926. [CrossRef]
- Póvoa, M.M.; Adagu, I.S.; Oliveira, S.G.; Machado, R.L.D.; Miles, M.A.; Warhurst, D.C. Pfmdr1Asn1042AspandAsp1246TyrPolymorphisms, Thought to Be Associated with Chloroquine Resistance, Are Present in Chloroquine-Resistant and -Sensitive Brazilian Field Isolates OfPlasmodium Falciparum. Exp Parasitol 1998, 88, 64–68. [CrossRef]
- Foote, S.J.; Kyle, D.E.; Martin, R.K.; Oduola, A.M.J.; Forsyth, K.; Kemp, D.J.; Cowman, A.F. Several Alleles of the Multidrug-Resistance Gene Are Closely Linked to Chloroquine Resistance in Plasmodium Falciparum. Nature 1990, 345, 255–258. [CrossRef]
- Lekana-Douki, J.B.; Boutamba, S.D.D.; Zatra, R.; Edou, S.E.Z.; Ekomy, H.; Bisvigou, U.; Toure-Ndouo, F.S. Increased Prevalence of the Plasmodium Falciparum Pfmdr1 86N Genotype among Field Isolates from Franceville, Gabon after Replacement of Chloroquine by Artemether–Lumefantrine and Artesunate–Mefloquine. Infection, Genetics and Evolution 2011, 11, 512–517. [CrossRef]
- Lopes, D.; Rungsihirunrat, K.; Nogueira, F.; Seugorn, A.; Gil, J.P.; do Rosário, V.E.; Cravo, P. Molecular Characterisation of Drug-Resistant Plasmodium Falciparum from Thailand. Malar J 2002, 1, 12. [CrossRef]
- Wurtz, N.; Fall, B.; Pascual, A.; Fall, M.; Baret, E.; Camara, C.; Nakoulima, A.; Diatta, B.; Fall, K.B.; Mbaye, P.S.; et al. Role of Pfmdr1 in In Vitro Plasmodium Falciparum Susceptibility to Chloroquine, Quinine, Monodesethylamodiaquine, Mefloquine, Lumefantrine, and Dihydroartemisinin. Antimicrob Agents Chemother 2014, 58, 7032–7040. [CrossRef]
- Dokomajilar, C.; Nsobya, S.L.; Greenhouse, B.; Rosenthal, P.J.; Dorsey, G. Selection of Plasmodium Falciparum Pfmdr1 Alleles Following Therapy with Artemether-Lumefantrine in an Area of Uganda Where Malaria Is Highly Endemic. Antimicrob Agents Chemother 2006, 50, 1893–1895. [CrossRef]
- Spring, M.D.; Lin, J.T.; Manning, J.E.; Vanachayangkul, P.; Somethy, S.; Bun, R.; Se, Y.; Chann, S.; Ittiverakul, M.; Sia-ngam, P.; et al. Dihydroartemisinin-Piperaquine Failure Associated with a Triple Mutant Including Kelch13 C580Y in Cambodia: An Observational Cohort Study. Lancet Infect Dis 2015, 15, 683–691. [CrossRef]
- Phuc, B.Q.; Rasmussen, C.; Duong, T.T.; Dong, L.T.; Loi, M.A.; Ménard, D.; Tarning, J.; Bustos, D.; Ringwald, P.; Galappaththy, G.L.; et al. Treatment Failure of Dihydroartemisinin/ Piperaquine for Plasmodium Falciparum Malaria, Vietnam. Emerg Infect Dis 2017, 23, 715–717. [CrossRef]
- Kahunu, G.M.; Thomsen, S.W.; Thomsen, L.W.; Mavoko, H.M.; Mulopo, P.M.; Hocke, E.F.; Nkoli, P.M.; Baraka, V.; Minja, D.T.R.; Mousa, A.; et al. Identification of the PfK13 Mutations R561H and P441L in the Democratic Republic of Congo. International Journal of Infectious Diseases 2024, 139, 41–49. [CrossRef]
- Schmedes, S.E.; Patel, D.; Dhal, S.; Kelley, J.; Svigel, S.S.; Dimbu, P.R.; Adeothy, A.-L.; Kahunu, G.M.; Nkoli, P.M.; Beavogui, A.H.; et al. Plasmodium Falciparum Kelch 13 Mutations, 9 Countries in Africa, 2014–2018. Emerg Infect Dis 2021, 27, 1902–1908. [CrossRef]
- Stokes, B.H.; Dhingra, S.K.; Rubiano, K.; Mok, S.; Straimer, J.; Gnädig, N.F.; Deni, I.; Schindler, K.A.; Bath, J.R.; Ward, K.E.; et al. Plasmodium Falciparum K13 Mutations in Africa and Asia Impact Artemisinin Resistance and Parasite Fitness. Elife 2021, 10. [CrossRef]
- Shamseer, L.; Moher, D.; Clarke, M.; Ghersi, D.; Liberati, A.; Petticrew, M.; Shekelle, P.; Stewart, L.A. Preferred Reporting Items for Systematic Review and Meta-Analysis Protocols (PRISMA-P) 2015: Elaboration and Explanation. BMJ 2015, 349, g7647–g7647. [CrossRef]
- Page, M.J.; McKenzie, J.E.; Bossuyt, P.M.; Boutron, I.; Hoffmann, T.C.; Mulrow, C.D.; Shamseer, L.; Tetzlaff, J.M.; Akl, E.A.; Brennan, S.E.; et al. The PRISMA 2020 Statement: An Updated Guideline for Reporting Systematic Reviews. Syst Rev 2021, 10, 89. [CrossRef]
- Allen, E.N.; Little, F.; Camba, T.; Cassam, Y.; Raman, J.; Boulle, A.; Barnes, K.I. Efficacy of Sulphadoxine-Pyrimethamine with or without Artesunate for the Treatment of Uncomplicated Plasmodium Falciparum Malaria in Southern Mozambique: A Randomized Controlled Trial. Malar J 2009, 8, 1–10. [CrossRef]
- Fernandes, N.; Figueiredo, P.; Do Rosário, V.E.; Cravo, P. Analysis of Sulphadoxine/Pyrimethamine Resistance-Conferring Mutations of Plasmodium Falciparum from Mozambique Reveals the Absence of the Dihydrofolate Reductase 164L Mutant. Malar J 2007, 6, 1–4. [CrossRef]
- da Silva, C.; Matias, D.; Dias, B.; Cancio, B.; Silva, M.; Viegas, R.; Chivale, N.; Luis, S.; Salvador, C.; Duarte, D.; et al. Anti-Malarial Resistance in Mozambique: Absence of Plasmodium Falciparum Kelch 13 (K13) Propeller Domain Polymorphisms Associated with Resistance to Artemisinins. Malar J 2023, 22. [CrossRef]
- Escobar, C.; Pateira, S.; Lobo, E.; Lobo, L.; Teodosio, R.; Dias, F.; Fernandes, N.; Arez, A.P.; Varandas, L.; Nogueira, F. Polymorphisms in Plasmodium Falciparum K13-Propeller in Angola and Mozambique after the Introduction of the ACTs. PLoS One 2015, 10, e0119215. [CrossRef]
- Kenangalem, E.; Poespoprodjo, J.R.; Douglas, N.M.; Burdam, F.H.; Gdeumana, K.; Chalfein, F.; Prayoga; Thio, F.; Devine, A.; Marfurt, J.; et al. Malaria Morbidity and Mortality Following Introduction of a Universal Policy of Artemisinin-Based Treatment for Malaria in Papua, Indonesia: A Longitudinal Surveillance Study. PLoS Med 2019, 16, e1002815. [CrossRef]
- Galatas, B.; Nhamussua, L.; Candrinho, B.; Mabote, L.; Cisteró, P.; Gupta, H.; Rabinovich, R.; Menéndez, C.; MacEte, E.; Saute, F.; et al. In-Vivo Efficacy of Chloroquine to Clear Asymptomatic Infections in Mozambican Adults: A Randomized, Placebo-Controlled Trial with Implications for Elimination Strategies. Sci Rep 2017, 7. [CrossRef]
- Raman, J.; Sharp, B.; Kleinschmidt, I.; Roper, C.; Streat, E.; Kelly, V.; Barnes, K.I. Differential Effect of Regional Drug Pressure on Dihydrofolate Reductase and Dihydropteroate Synthetase Mutations in Southern Mozambique. Am J Trop Med Hyg 2008, 78, 256–261. [CrossRef]
- Ocan, M.; Akena, D.; Nsobya, S.; Kamya, M.R.; Senono, R.; Kinengyere, A.A.; Obuku, E.A. Persistence of Chloroquine Resistance Alleles in Malaria Endemic Countries: A Systematic Review of Burden and Risk Factors. Malar J 2019, 18. [CrossRef]
- Mayor, A.; Serra-Casas, E.; Sanz, S.; Aponte, J.J.; Macete, E.; Mandomando, I.; Puyol, L.; Berzosa, P.; Dobaño, C.; Aide, P.; et al. Molecular Markers of Resistance to Sulfadoxine-Pyrimethamine during Intermittent Preventive Treatment for Malaria in Mozambican Infants. J Infect Dis 2008, 197, 1737–1742. [CrossRef]
- Chidimatembue, A.; Svigel, S.S.; Mayor, A.; Aíde, P.; Nhama, A.; Nhamussua, L.; Nhacolo, A.; Bassat, Q.; Salvador, C.; Enosse, S.; et al. Molecular Surveillance for Polymorphisms Associated with Artemisinin-Based Combination Therapy Resistance in Plasmodium Falciparum Isolates Collected in Mozambique, 2018. Malar J 2021, 20. [CrossRef]
- Gupta, H.; Galatas, B.; Chidimatembue, A.; Huijben, S.; Cisteró, P.; Matambisso, G.; Nhamussua, L.; Simone, W.; Bassat, Q.; Ménard, D.; et al. Effect of Mass Dihydroartemisinin–Piperaquine Administration in Southern Mozambique on the Carriage of Molecular Markers of Antimalarial Resistance. PLoS One 2020, 15, e0240174. [CrossRef]
- Gupta, H.; Macete, E.; Bulo, H.; Salvador, C.; Warsame, M.; Carvalho, E.; Ménard, D.; Ringwald, P.; Bassat, Q.; Enosse, S.; et al. Drug-Resistant Polymorphisms and Copy Numbers in Plasmodium Falciparum, Mozambique, 2015. Emerg Infect Dis 2018, 24, 40–48. [CrossRef]
- Mang’era, C.M.; Mbai, F.N.; Omedo, I.A.; Mireji, P.O.; Omar, S.A. Changes in Genotypes of Plasmodium Falciparum Human Malaria Parasite Following Withdrawal of Chloroquine in Tiwi, Kenya. Acta Trop 2012, 123, 202–207. [CrossRef]
- Mwanza, S.; Joshi, S.; Nambozi, M.; Chileshe, J.; Malunga, P.; Kabuya, J.-B.B.; Hachizovu, S.; Manyando, C.; Mulenga, M.; Laufer, M. The Return of Chloroquine-Susceptible Plasmodium Falciparum Malaria in Zambia. Malar J 2016, 15, 584. [CrossRef]
- Laufer, M.K.; Takala-Harrison, S.; Dzinjalamala, F.K.; Stine, O.C.; Taylor, T.E.; Plowe, C.V. Return of Chloroquine-Susceptible Falciparum Malaria in Malawi Was a Reexpansion of Diverse Susceptible Parasites. J Infect Dis 2010, 202, 801–808. [CrossRef]
- Bwire, G.M.; Ngasala, B.; Mikomangwa, W.P.; Kilonzi, M.; Kamuhabwa, A.A.R. Detection of Mutations Associated with Artemisinin Resistance at K13-Propeller Gene and a near Complete Return of Chloroquine Susceptible Falciparum Malaria in Southeast of Tanzania. Sci Rep 2020, 10, 3500. [CrossRef]
- Brokhattingen, N.; Matambisso, G.; da Silva, C.; Neubauer Vickers, E.; Pujol, A.; Mbeve, H.; Cisteró, P.; Maculuve, S.; Cuna, B.; Melembe, C.; et al. Genomic Malaria Surveillance of Antenatal Care Users Detects Reduced Transmission Following Elimination Interventions in Mozambique. Nature Communications 2024 15:1 2024, 15, 1–12. [CrossRef]
- Huijben, S.; Macete, E.; Mombo-Ngoma, G.; Ramharter, M.; Kariuki, S.; Desai, M.; Shi, Y.P.; Mwangoka, G.; Massougbodji, A.; Cot, M.; et al. Counter-Selection of Antimalarial Resistance Polymorphisms by Intermittent Preventive Treatment in Pregnancy. J Infect Dis 2020, 221, 293–303. [CrossRef]
- Arnaldo, P. Evaluation of Intermittent Preventive Treatment during Pregnancy (IPTp) in Chókwè District, Southern Mozambique: Coverage and Effect on Pregnancy and Parasitological Outcomes 2019.
- Gupta, H.; Macete, E.; Bulo, H.; Salvador, C.; Warsame, M.; Carvalho, E.; Ménard, D.; Ringwald, P.; Bassat, Q.; Enosse, S.; et al. Drug-Resistant Polymorphisms and Copy Numbers in Plasmodium Falciparum, Mozambique, 2015. Emerg Infect Dis 2018, 24, 40. [CrossRef]
- Gupta, H.; Galatas, B.; Chidimatembue, A.; Huijben, S.; Cisteró, P.; Matambisso, G.; Nhamussua, L.; Simone, W.; Bassat, Q.; Ménard, D.; et al. Effect of Mass Dihydroartemisinin–Piperaquine Administration in Southern Mozambique on the Carriage of Molecular Markers of Antimalarial Resistance. PLoS One 2020, 15, e0240174. [CrossRef]
- Raman, J.; Mauff, K.; Muianga, P.; Mussa, A.; Maharaj, R.; Barnes, K.I. Five Years of Antimalarial Resistance Marker Surveillance in Gaza Province, Mozambique, Following Artemisinin-Based Combination Therapy Roll Out. PLoS One 2011, 6. [CrossRef]
- Mayor, A.; Serra-Casas, E.; Sanz, S.; Aponte, J.J.; Macete, E.; Mandomando, I.; Puyol, L.; Berzosa, P.; Dobaño, C.; Aide, P.; et al. Molecular Markers of Resistance to Sulfadoxine-Pyrimethamine during Intermittent Preventive Treatment for Malaria in Mozambican Infants. J Infect Dis 2008, 197, 1737–1742. [CrossRef]
- Chidimatembue, A.; Svigel, S.S.; Mayor, A.; Aíde, P.; Nhama, A.; Nhamussua, L.; Nhacolo, A.; Bassat, Q.; Salvador, C.; Enosse, S.; et al. Molecular Surveillance for Polymorphisms Associated with Artemisinin-Based Combination Therapy Resistance in Plasmodium Falciparum Isolates Collected in Mozambique, 2018. Malar J 2021, 20, 1–9. [CrossRef]
- Brokhattingen, N.; Matambisso, G.; da Silva, C.; Neubauer Vickers, E.; Pujol, A.; Mbeve, H.; Cisteró, P.; Maculuve, S.; Cuna, B.; Melembe, C.; et al. Genomic Malaria Surveillance of Antenatal Care Users Detects Reduced Transmission Following Elimination Interventions in Mozambique. Nat Commun 2024, 15, 2402. [CrossRef]
- Huijben, S.; Macete, E.; Mombo-Ngoma, G.; Ramharter, M.; Kariuki, S.; Desai, M.; Shi, Y.P.; Mwangoka, G.; Massougbodji, A.; Cot, M.; et al. Counter-Selection of Antimalarial Resistance Polymorphisms by Intermittent Preventive Treatment in Pregnancy. J Infect Dis 2020, 221, 293–303. [CrossRef]
- Raman, J.; Little, F.; Roper, C.; Kleinschmidt, I.; Cassam, Y.; Maharaj, R.; Barnes, K.I. Five Years of Large-Scale Dhfr and Dhps Mutation Surveillance Following the Phased Implementation of Artesunate Plus Sulfadoxine-Pyrimethamine in Maputo Province, Southern Mozambique. Am J Trop Med Hyg 2010, 82, 788. [CrossRef]
- Enosse, S.; Magnussen, P.; Abacassamo, F.; Gómez-Olivé, X.; Rønn, A.M.; Thompson, R.; Alifrangis, M. Rapid Increase of Plasmodium Falciparum Dhfr/Dhps Resistant Haplotypes, after the Adoption of Sulphadoxine-Pyrimethamine as First Line Treatment in 2002, in Southern Mozambique. Malar J 2008, 7. [CrossRef]
- Matambisso, G.; Brokhattingen, N.; Maculuve, S.; Cístero, P.; Mbeve, H.; Escoda, A.; Bambo, G.; Cuna, B.; Melembe, C.; Ndimande, N.; et al. Sustained Clinical Benefit of Malaria Chemoprevention with Sulfadoxine-Pyrimethamine (SP) in Pregnant Women in a Region with High SP Resistance Markers. J Infect 2024, 88. [CrossRef]
- Figueroa-Romero, A.; Bissombolo, D.; Meremikwu, M.; Ratsimbasoa, A.; Sacoor, C.; Arikpo, I.; Lemba, E.; Nhama, A.; Rakotosaona, R.; Llach, M.; et al. Prevalence of Molecular Markers of Resistance to Sulfadoxine–Pyrimethamine before and after Community Delivery of Intermittent Preventive Treatment of Malaria in Pregnancy in Sub-Saharan Africa: A Multi-Country Evaluation. Lancet Glob Health 2023, 11, e1765–e1774. [CrossRef]
- Lobo, E.; de Sousa, B.; Rosa, S.; Figueiredo, P.; Lobo, L.; Pateira, S.; Fernandes, N.; Nogueira, F. Prevalence of Pfmdr1 Alleles Associated with Artemether-Lumefantrine Tolerance/Resistance in Maputo before and after the Implementation of Artemisinin-Based Combination Therapy. Malar J 2014, 13, 300. [CrossRef]
- Brown, N.; Silva, C. da; Webb, C.; Matias, D.; Dias, B.; Cancio, B.; Silva, M.; Viegas, R.; Salvador, C.; Chivale, N.; et al. Antimalarial Resistance Risk in Mozambique Detected by a Novel Quadruplex Droplet Digital PCR Assay. Antimicrob Agents Chemother 2024. [CrossRef]
- Ross, L.S.; Dhingra, S.K.; Mok, S.; Yeo, T.; Wicht, K.J.; Kümpornsin, K.; Takala-Harrison, S.; Witkowski, B.; Fairhurst, R.M.; Ariey, F.; et al. Emerging Southeast Asian PfCRT Mutations Confer Plasmodium Falciparum Resistance to the First-Line Antimalarial Piperaquine. Nat Commun 2018, 9, 3314. [CrossRef]
- Matambisso, G.; Brokhattingen, N.; Maculuve, S.; Cístero, P.; Mbeve, H.; Escoda, A.; Bambo, G.; Cuna, B.; Melembe, C.; Ndimande, N.; et al. Sustained Clinical Benefit of Malaria Chemoprevention with Sulfadoxine-Pyrimethamine (SP) in Pregnant Women in a Region with High SP Resistance Markers. Journal of Infection 2024, 88, 106144. [CrossRef]
- Orlando, P. Evaluation of Intermittent Preventive Treatment during Pregnancy (IPTp) in Chókwè District, Southern Mozambique: Coverage and Effect on Pregnancy and Parasitological Outcomes, University of Antwerp to be defended: Antwerpen, 2019.
- Arnaldo, P.; Rovira-Vallbona, E.; Langa, J.S.; Salvador, C.; Guetens, P.; Chiheb, D.; Xavier, B.; Kestens, L.; Enosse, S.M.; Rosanas-Urgell, A. Uptake of Intermittent Preventive Treatment and Pregnancy Outcomes: Health Facilities and Community Surveys in Chókwè District, Southern Mozambique. Malar J 2018, 17, 109. [CrossRef]
- Kayode, A.T.; Ajogbasile, F. V.; Akano, K.; Uwanibe, J.N.; Oluniyi, P.E.; Eromon, P.J.; Folarin, O.A.; Sowunmi, A.; Wirth, D.F.; Happi, C.T. Polymorphisms in Plasmodium Falciparum Dihydropteroate Synthetase and Dihydrofolate Reductase Genes in Nigerian Children with Uncomplicated Malaria Using High-Resolution Melting Technique. Sci Rep 2021, 11, 471. [CrossRef]
- Duah, N.O.; Quashie, N.B.; Abuaku, B.K.; Sebeny, P.J.; Kronmann, K.C.; Koram, K.A. Surveillance of Molecular Markers of Plasmodium Falciparum Resistance to Sulphadoxine-Pyrimethamine 5 Years after the Change of Malaria Treatment Policy in Ghana. The American Society of Tropical Medicine and Hygiene 2012, 87, 996–1003. [CrossRef]
- Duah, N.O.; Matrevi, S.A.; de Souza, D.K.; Binnah, D.D.; Tamakloe, M.M.; Opoku, V.S.; Onwona, C.O.; Narh, C.A.; Quashie, N.B.; Abuaku, B.; et al. Increased Pfmdr1 Gene Copy Number and the Decline in Pfcrt and Pfmdr1 Resistance Alleles in Ghanaian Plasmodium Falciparum Isolates after the Change of Anti-Malarial Drug Treatment Policy. Malar J 2013, 12, 377. [CrossRef]
- Njiro, B.J.; Mutagonda, R.F.; Chamani, A.T.; Mwakyandile, T.; Sabas, D.; Bwire, G.M. Molecular Surveillance of Chloroquine-Resistant Plasmodium Falciparum in Sub-Saharan African Countries after Withdrawal of Chloroquine for Treatment of Uncomplicated Malaria: A Systematic Review. J Infect Public Health 2022, 15, 550–557. [CrossRef]
- Niba, P.T.N.; Nji, A.M.; Evehe, M.-S.; Ali, I.M.; Netongo, P.M.; Ngwafor, R.; Moyeh, M.N.; Ngum, L.N.; Ndum, O.E.; Acho, F.A.; et al. Drug Resistance Markers within an Evolving Efficacy of Anti-Malarial Drugs in Cameroon: A Systematic Review and Meta-Analysis (1998–2020). Malar J 2021, 20, 32. [CrossRef]
- Okell, L.C.; Reiter, L.M.; Ebbe, L.S.; Baraka, V.; Bisanzio, D.; Watson, O.J.; Bennett, A.; Verity, R.; Gething, P.; Roper, C.; et al. Emerging Implications of Policies on Malaria Treatment: Genetic Changes in the Pfmdr-1 Gene Affecting Susceptibility to Artemether–Lumefantrine and Artesunate–Amodiaquine in Africa. BMJ Glob Health 2018, 3, e000999. [CrossRef]
- Groger, M.; Veletzky, L.; Lalremruata, A.; Cattaneo, C.; Mischlinger, J.; Zoleko-Manego, R.; Endamne, L.; Klicpera, A.; Kim, J.; Nguyen, T.; et al. Prospective Clinical Trial Assessing Species-Specific Efficacy of Artemether-Lumefantrine for the Treatment of Plasmodium Malariae, Plasmodium Ovale, and Mixed Plasmodium Malaria in Gabon. Antimicrob Agents Chemother 2018, 62. [CrossRef]
- Gaye, A.; Sy, M.; Ndiaye, T.; Siddle, K.J.; Park, D.J.; Deme, A.B.; Mbaye, A.; Dieye, B.; Ndiaye, Y.D.; Neafsey, D.E.; et al. Amplicon Deep Sequencing of Kelch13 in Plasmodium Falciparum Isolates from Senegal. Malar J 2020, 19, 134. [CrossRef]
- Jeang, B.; Zhong, D.; Lee, M.-C.; Atieli, H.; Yewhalaw, D.; Yan, G. Molecular Surveillance of Kelch 13 Polymorphisms in Plasmodium Falciparum Isolates from Kenya and Ethiopia. Malar J 2024, 23, 36. [CrossRef]
- da Silva, C.; Boene, S.; Datta, D.; Rovira-Vallbona, E.; Aranda-Díaz, A.; Cisteró, P.; Hathaway, N.; Tessema, S.; Chidimatembue, A.; Matambisso, G.; et al. Targeted and Whole-Genome Sequencing Reveal a North-South Divide in P. Falciparum Drug Resistance Markers and Genetic Structure in Mozambique. Commun Biol 2023, 6, 619. [CrossRef]
- Leroy, D.; Macintyre, F.; Adoke, Y.; Ouoba, S.; Barry, A.; Mombo-Ngoma, G.; Ndong Ngomo, J.M.; Varo, R.; Dossou, Y.; Tshefu, A.K.; et al. African Isolates Show a High Proportion of Multiple Copies of the Plasmodium Falciparum Plasmepsin-2 Gene, a Piperaquine Resistance Marker. Malar J 2019, 18, 126. [CrossRef]
- Uwimana, A.; Umulisa, N.; Venkatesan, M.; Svigel, S.S.; Zhou, Z.; Munyaneza, T.; Habimana, R.M.; Rucogoza, A.; Moriarty, L.F.; Sandford, R.; et al. Association of Plasmodium Falciparum Kelch13 R561H Genotypes with Delayed Parasite Clearance in Rwanda: An Open-Label, Single-Arm, Multicentre, Therapeutic Efficacy Study. Lancet Infect Dis 2021, 21, 1120–1128. [CrossRef]
- Asua, V.; Conrad, M.D.; Aydemir, O.; Duvalsaint, M.; Legac, J.; Duarte, E.; Tumwebaze, P.; Chin, D.M.; Cooper, R.A.; Yeka, A.; et al. Changing Prevalence of Potential Mediators of Aminoquinoline, Antifolate, and Artemisinin Resistance Across Uganda. J Infect Dis 2021, 223, 985–994. [CrossRef]
- Conrad, M.D.; Asua, V.; Garg, S.; Giesbrecht, D.; Niaré, K.; Smith, S.; Namuganga, J.F.; Katairo, T.; Legac, J.; Crudale, R.M.; et al. Evolution of Partial Resistance to Artemisinins in Malaria Parasites in Uganda. New England Journal of Medicine 2023, 389, 722–732. [CrossRef]
- Greenwood, B. Artemisinin-Resistant and HRP-Negative Malaria Parasites in Africa. New England Journal of Medicine 2023, 389, 1162–1164. [CrossRef]
- Health Organization, W. WHO Guidelines for Malaria - 14 March 2023; 2023;
- Ndwiga, L.; Kimenyi, K.M.; Wamae, K.; Osoti, V.; Akinyi, M.; Omedo, I.; Ishengoma, D.S.; Duah-Quashie, N.; Andagalu, B.; Ghansah, A.; et al. A Review of the Frequencies of Plasmodium Falciparum Kelch 13 Artemisinin Resistance Mutations in Africa. Int J Parasitol Drugs Drug Resist 2021, 16, 155–161. [CrossRef]
- Von Wowern, F.; Makenga, G.; Wellmann Thomsen, S.; Wellmann Thomsen, L.; Filtenborg Hocke, E.; Baraka, V.; Opot, B.H.; Minja, D.T.R.; Lusingu, J.P.A.; Van-geertruyden, J.-P.; et al. Lack of Selection of Antimalarial Drug Resistance Markers after Intermittent Preventive Treatment of Schoolchildren (IPTsc) against Malaria in Northeastern Tanzania. International Journal of Infectious Diseases 2024, 146, 107102. [CrossRef]
- Tajebe, A.; Aemero, M.; Francis, K.; Magoma, G. Identification of Chloroquine Resistance Pfcrt-K76T and Determination of Pfmdr1-N86Y Copy Number by SYBR Green I QPCR. Asian Pac J Trop Biomed 2015, 5, 208–220. [CrossRef]
- Ngalah, B.S.; Ingasia, L.A.; Cheruiyot, A.C.; Chebon, L.J.; Juma, D.W.; Muiruri, P.; Onyango, I.; Ogony, J.; Yeda, R.A.; Cheruiyot, J.; et al. Analysis of Major Genome Loci Underlying Artemisinin Resistance and Pfmdr1 Copy Number in Pre- and Post-ACTs in Western Kenya. Sci Rep 2015, 5, 8308. [CrossRef]
- Tadele, G.; Jawara, A.; Oboh, M.; Oriero, E.; Dugassa, S.; Amambua-Ngwa, A.; Golassa, L. Clinical Isolates of Uncomplicated Falciparum Malaria from High and Low Malaria Transmission Areas Show Distinct Pfcrt and Pfmdr1 Polymorphisms in Western Ethiopia. Malar J 2023, 22, 171. [CrossRef]
- Agomo, C.O.; Mishra, N.; Olukosi, Y.A.; Gupta, R.; Kamlesh, K.; Aina, O.O.; Awolola, S.T. Mutations in Pfcrt and Pfmdr1 Genes of Plasmodium Falciparum Isolates from Two Sites in Northcentral and Southwest Nigeria. Infection, Genetics and Evolution 2021, 95, 105042. [CrossRef]
- Yobi, D.M.; Kayiba, N.K.; Mvumbi, D.M.; Boreux, R.; Kabututu, P.Z.; Situakibanza, H.N.T.; Umesumbu, S.E.; De Mol, P.; Speybroeck, N.; Mvumbi, G.L.; et al. Assessment of Plasmodium Falciparum Anti-Malarial Drug Resistance Markers in Pfk13-Propeller, Pfcrt and Pfmdr1 Genes in Isolates from Treatment Failure Patients in Democratic Republic of Congo, 2018–2019. Malar J 2021, 20, 144. [CrossRef]
- Liu, Y.; Liang, X.; Li, J.; Chen, J.; Huang, H.; Zheng, Y.; He, J.; Ehapo, C.S.; Eyi, U.M.; Yang, P.; et al. Molecular Surveillance of Artemisinin-Based Combination Therapies Resistance in Plasmodium Falciparum Parasites from Bioko Island, Equatorial Guinea. Microbiol Spectr 2022, 10. [CrossRef]
- Ebong, C.; Sserwanga, A.; Namuganga, J.F.; Kapisi, J.; Mpimbaza, A.; Gonahasa, S.; Asua, V.; Gudoi, S.; Kigozi, R.; Tibenderana, J.; et al. Efficacy and Safety of Artemether-Lumefantrine and Dihydroartemisinin-Piperaquine for the Treatment of Uncomplicated Plasmodium Falciparum Malaria and Prevalence of Molecular Markers Associated with Artemisinin and Partner Drug Resistance in Uganda. Malar J 2021, 20, 484. [CrossRef]
- Koko, V.S.; Warsame, M.; Vonhm, B.; Jeuronlon, M.K.; Menard, D.; Ma, L.; Taweh, F.; Tehmeh, L.; Nyansaiye, P.; Pratt, O.J.; et al. Artesunate–Amodiaquine and Artemether–Lumefantrine for the Treatment of Uncomplicated Falciparum Malaria in Liberia: In Vivo Efficacy and Frequency of Molecular Markers. Malar J 2022, 21, 134. [CrossRef]
- Amaratunga, C.; Lim, P.; Suon, S.; Sreng, S.; Mao, S.; Sopha, C.; Sam, B.; Dek, D.; Try, V.; Amato, R.; et al. Dihydroartemisinin–Piperaquine Resistance in Plasmodium Falciparum Malaria in Cambodia: A Multisite Prospective Cohort Study. Lancet Infect Dis 2016, 16, 357–365. [CrossRef]
- Amato, R.; Lim, P.; Miotto, O.; Amaratunga, C.; Dek, D.; Pearson, R.D.; Almagro-Garcia, J.; Neal, A.T.; Sreng, S.; Suon, S.; et al. Genetic Markers Associated with Dihydroartemisinin–Piperaquine Failure in Plasmodium Falciparum Malaria in Cambodia: A Genotype–Phenotype Association Study. Lancet Infect Dis 2017, 17, 164–173. [CrossRef]
- Witkowski, B.; Duru, V.; Khim, N.; Ross, L.S.; Saintpierre, B.; Beghain, J.; Chy, S.; Kim, S.; Ke, S.; Kloeung, N.; et al. A Surrogate Marker of Piperaquine-Resistant Plasmodium Falciparum Malaria: A Phenotype–Genotype Association Study. Lancet Infect Dis 2017, 17, 174–183. [CrossRef]
- Arroz, J.A.H. Increase in Cases of Malaria in Mozambique, 2014: Epidemic or New Endemic Pattern? Rev Saude Publica 2016, 50. [CrossRef]
| Mutation | Gene | Province | Nº of studies (2008-2024) |
References | Year of sample collection* |
|---|---|---|---|---|---|
| SNP | pfcrt | Maputo | 6 | [10,50,53,54,55,56,63,69,70] | 2015 - 2019 [69] |
| Gaza | 2 | 2015 [56] | |||
| Inhambane | 1 | 2018 [54] | |||
| Zambezia | |||||
| Tete | |||||
| Cabo Delgado | |||||
| pfdhfr, pfdhps | Maputo | 10 | [10,45,46,51,53,56,63,69,70,71,72,73,74] | 2015 - 2019 [69] | |
| Gaza | 3 | 2014 - 2015 [63] | |||
| Tete | 1 | 2015 [56] | |||
| Sofala | 2 | ||||
| Cabo Delgado | 1 | 2015 [56] | |||
| pfmdr1 | Maputo | 5 | [10,53,54,55,56,63,69,70,75] | 2015 - 2019 [69] | |
| Gaza | 3 | 2014 - 2015 [63] | |||
| Inhambane | 1 | 2018 [54] | |||
| Zambezia | |||||
| Tete | 2 | ||||
| Sofala | 1 | ||||
| Cabo Delgado | 2 | ||||
| pfk13 | Maputo | 4 | [47,48,54,55,69] | 2021 [47] | |
| Gaza | 1 | ||||
| Zambezia | |||||
| Tete | |||||
| Sofala | |||||
| Manica | |||||
| Cabo Delgado | |||||
| Niassa | |||||
| CNV |
pfpm2/ pfpm3/ pfmdr1 |
Maputo | 2 | [55,56,76] | 2021 [76] |
| Gaza | 1 | 2015 [56] | |||
| Tete | |||||
| Sofala | |||||
| Manica | 2021 [76] | ||||
| Niassa |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




