Submitted:
26 October 2024
Posted:
29 October 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Datasets and Symptom Analysis
2.1.1. Image Data
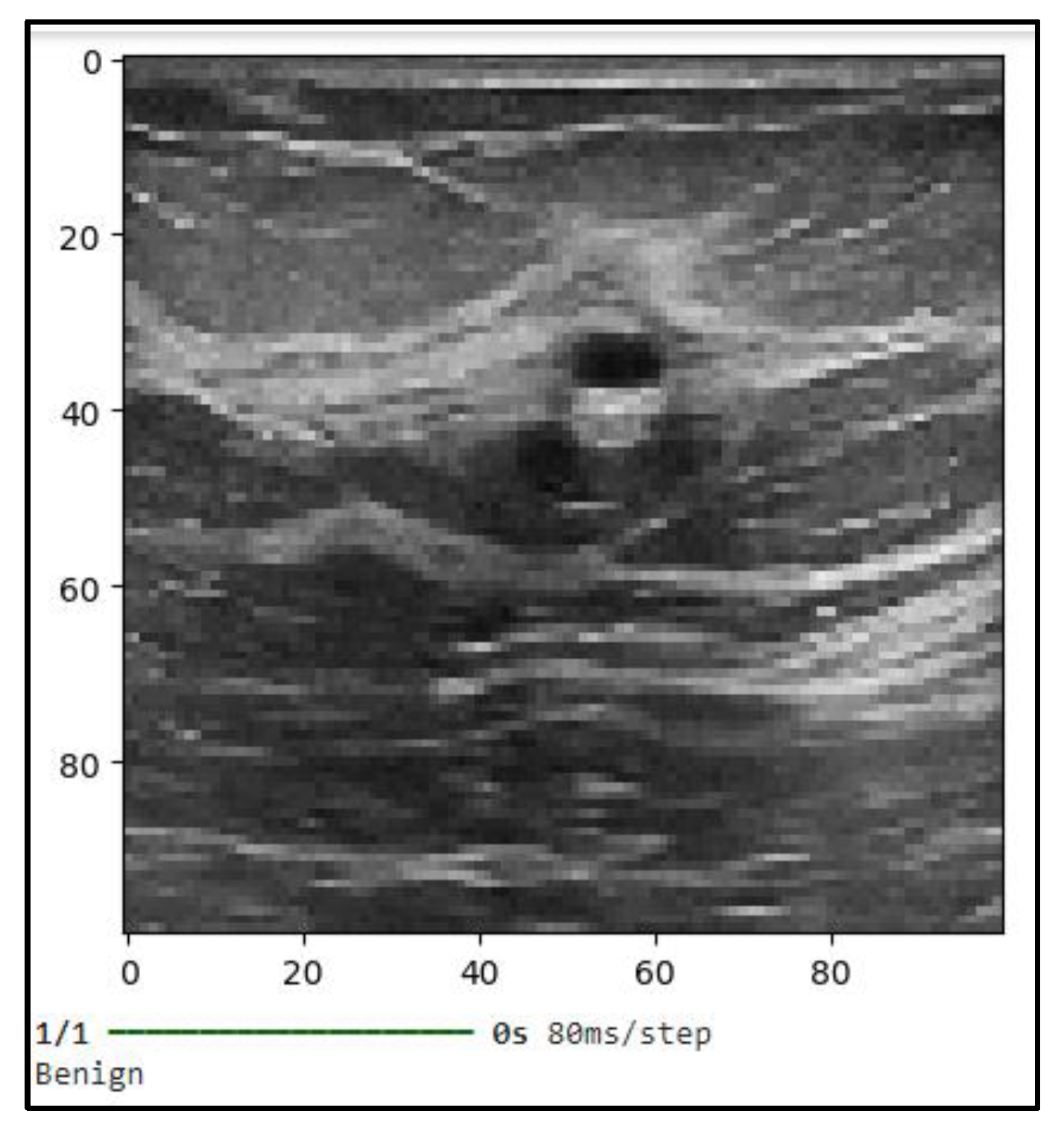
- Ultrasound Images: A total of 100 breast ultrasound images were collected. Ultrasound imaging is commonly used in breast cancer detection due to its non-invasive nature and ability to distinguish between solid masses and cysts. These images typically help in identifying the presence of lumps or abnormalities in breast tissue. The dataset includes both benign and malignant cases, which were pre-labelled.
- Pathology Images: Additionally, 100 pathology images were included in the dataset. Pathological images provide a microscopic view of breast tissue and are critical for confirming cancer diagnoses. These images are highly detailed and allow for the observation of cellular structures, which are indicative of cancerous changes in the tissue. The pathology dataset also contains both benign and malignant cases, providing an opportunity for the model to learn from diverse patterns.
2.1.2. Clinical Data
- ID: A unique identifier assigned to each patient in the dataset. This feature does not contribute to the predictive model but is used to track and distinguish patient records.
- Radius Mean: The average distance from the center to the perimeter of the tumor. This feature reflects the size of the tumor, where larger radius may indicate a higher likelihood of malignancy.
- Texture Mean: Represents the variation in gray levels within the tumor, which corresponds to the roughness of the tumor surface. Tumors with uneven textures are more likely to be cancerous.
- Perimeter Mean: The average length around the boundary of the tumor. A longer perimeter often indicates an irregular tumor shape, which can be associated with malignant growth.
- Area Mean: The mean area occupied by the tumor cells. Larger areas tend to correlate with more aggressive tumor types and higher chances of malignancy.
- Smoothness Mean: Measures the regularity of the tumor's borders. Benign tumors generally have smoother edges, while malignant tumors tend to have more jagged or irregular borders.
- Compactness Mean: A measure of how closely packed the tumor cells are, calculated using the perimeter and area. Higher compactness can indicate malignancy, as cancerous cells often exhibit irregular, tight clustering.
- Concavity Mean: The degree to which parts of the tumor contour are indented. More concavities suggest irregular shapes, often associated with malignant tumors.
- Concave Points Mean: The number of distinct concave areas or "inward dents" in the tumor's boundary. More concave points typically indicate a higher risk of cancer.
- Symmetry Mean: Describes how symmetrical the tumor is. Benign tumors are generally more symmetrical, while malignant tumors tend to show asymmetry.
- Fractal Dimension Mean: A measure of the complexity of the tumor’s boundary, quantifying how detailed or rough the edges are. Higher fractal dimensions can indicate a more irregular, and potentially cancerous, tumor shape.
- Radius SE: The standard deviation of the radius, indicating variability in tumor size within the sample. Higher variability can be a sign of aggressive, heterogeneous tumors.
- Texture SE: Measures how much the texture varies across different sections of the tumor. High texture variability may indicate uneven growth, often seen in malignant tumors.
- Perimeter SE: Describes how much the perimeter measurements differ across the tumor. Greater deviations in perimeter can suggest irregular tumor growth, common in cancerous tissues.
- Area SE: Shows the variability in the tumor's area measurements, with greater variation possibly indicating more aggressive or invasive cancer.
- Smoothness SE: Indicates how much the smoothness of the tumor’s borders varies. Large fluctuations in smoothness can suggest inconsistent cell growth, often linked to malignancy.
- Compactness SE: Shows the degree of variation in how tightly packed the tumor cells are. High variation may indicate that the tumor is cancerous, as malignant cells often exhibit abnormal packing.
- Concavity SE: The standard deviation of the concavities, showing the variability in the indentations along the tumor’s contour. Malignant tumors often exhibit more irregular concavity.
- Concave Points SE: Measures the variability in the number of concave points. A higher standard error can signal irregular tumor shapes, commonly associated with malignancy.
- Symmetry SE: The standard deviation of the symmetry values across the tumor. Greater variation in symmetry may be a red flag for cancerous growth.
- Fractal Dimension SE: Measures the variability in the fractal dimension, which can indicate how much the complexity of the tumor’s boundary fluctuates. Higher values often correlate with malignancy.
- Radius Worst: The largest radius observed in the tumor sample. Larger radii can suggest more aggressive or advanced cancer.
- Texture Worst: The highest variation in texture observed in any part of the tumor. Irregular texture at its worst often indicates malignancy.
- Perimeter Worst: The longest perimeter measurement, representing the largest observed boundary. Larger, irregular boundaries often point to malignancy.
- Area Worst: The maximum area occupied by tumor cells. Larger areas often correlate with more advanced stages of cancer.
- Smoothness Worst: The most irregular borders observed. High smoothness variability can signal more invasive cancer.
- Compactness Worst: The highest level of cell compactness observed. Extremely compact cell formations are more commonly seen in cancerous tumors.
- Concavity Worst: The deepest concavity or inward curve along the tumor's boundary. A more pronounced concavity often signals malignancy.
- Concave Points Worst: The maximum number of concave points identified. More concave points are a red flag for irregular, cancerous growth.
- Symmetry Worst: The most asymmetrical shape of the tumor. Malignant tumors tend to have more irregular, asymmetrical growth patterns.
- Fractal Dimension Worst: The most complex boundary found in the tumor. Higher fractal dimension values are associated with irregular and invasive cancer growth.
2.2. Machine Learning Model
3. Result
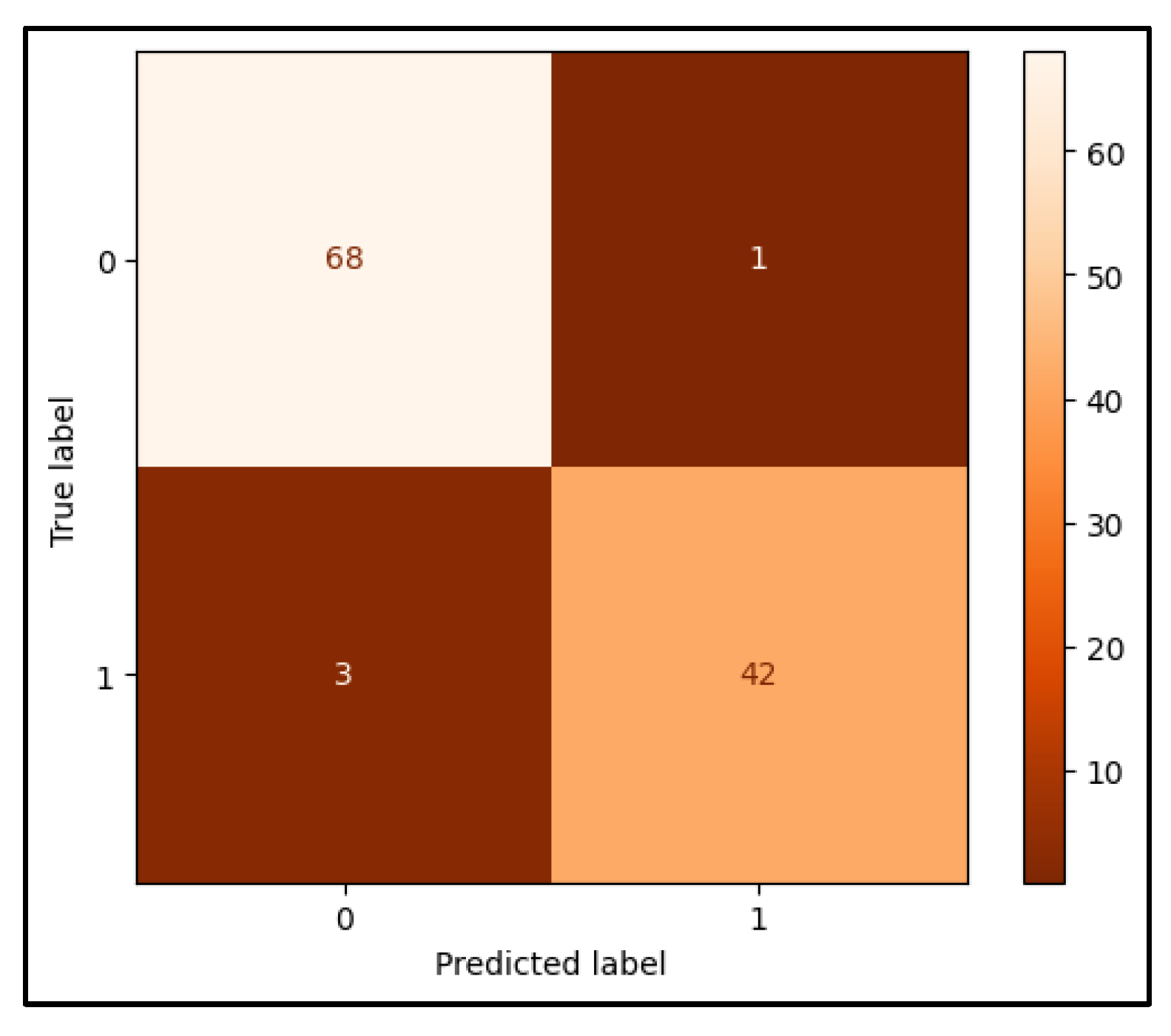
3.1. CNN-Based Classification of Ultrasound Images for Breast Cancer Detection
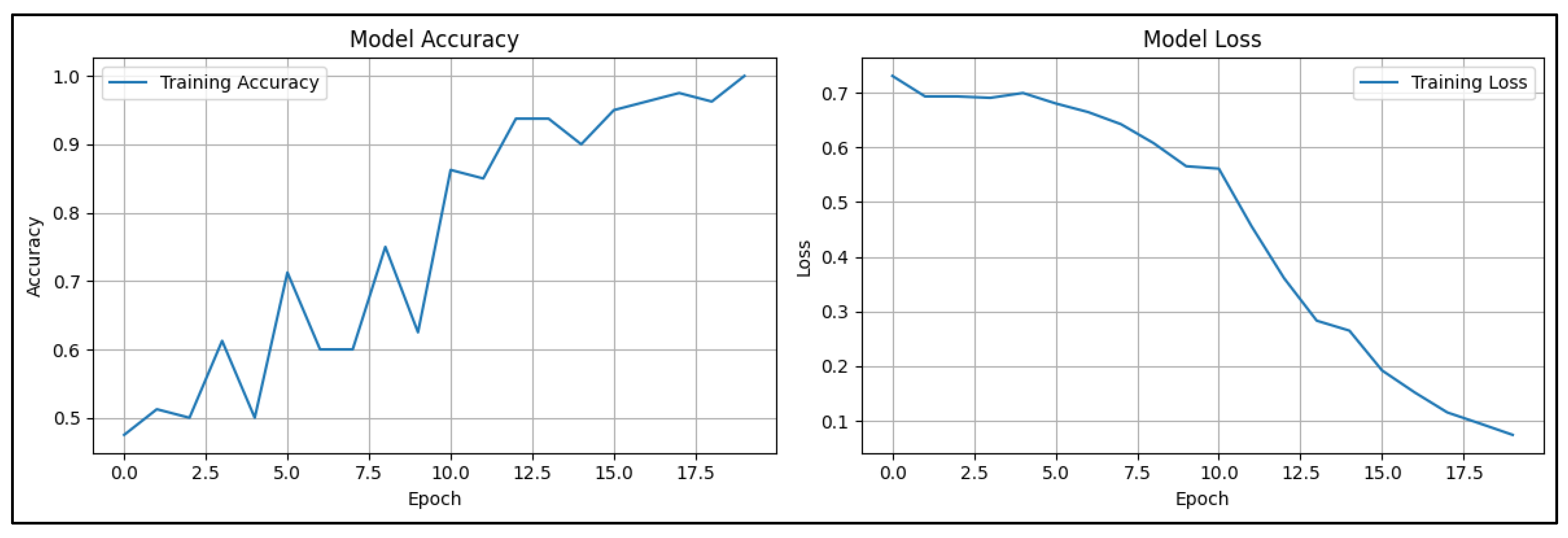
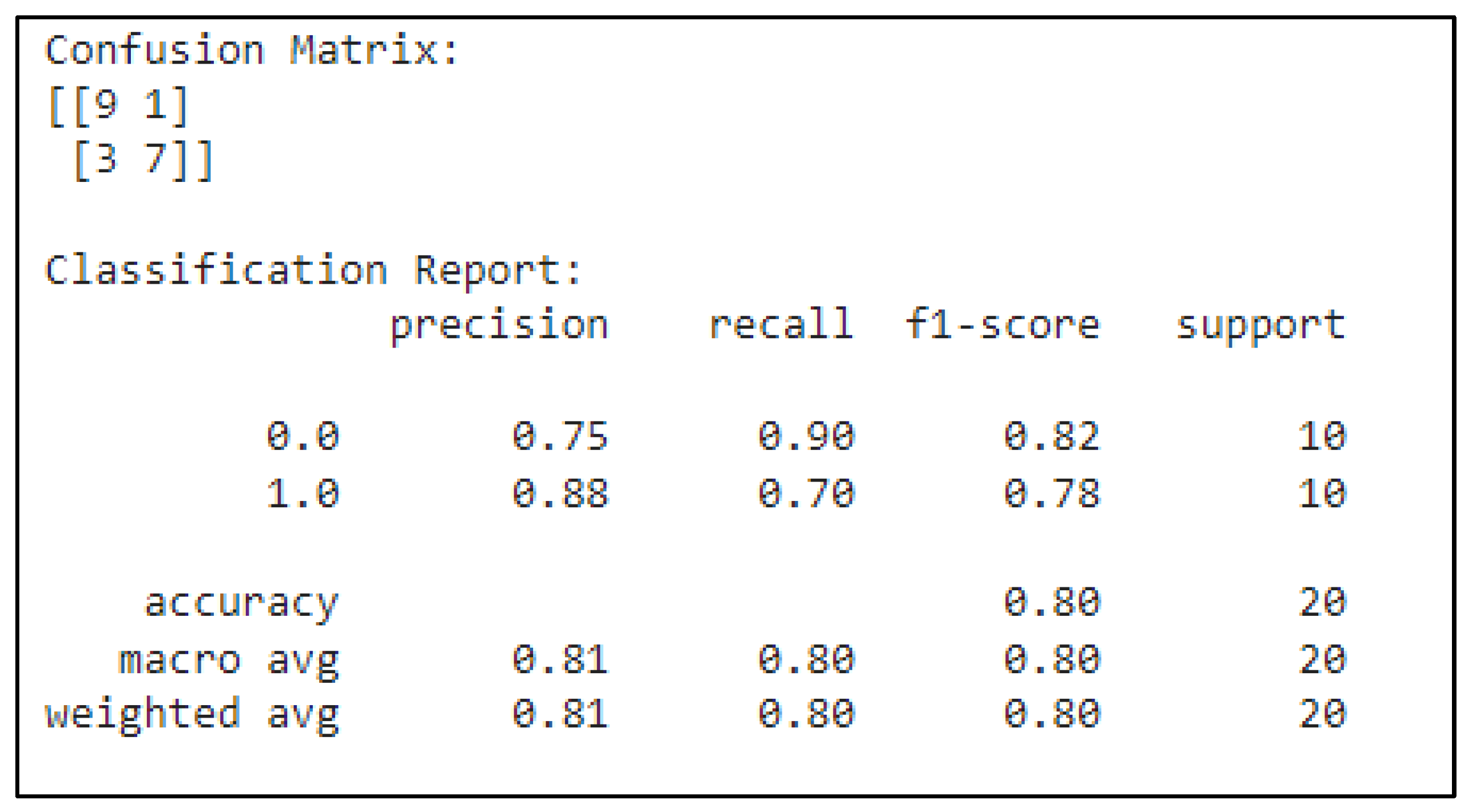
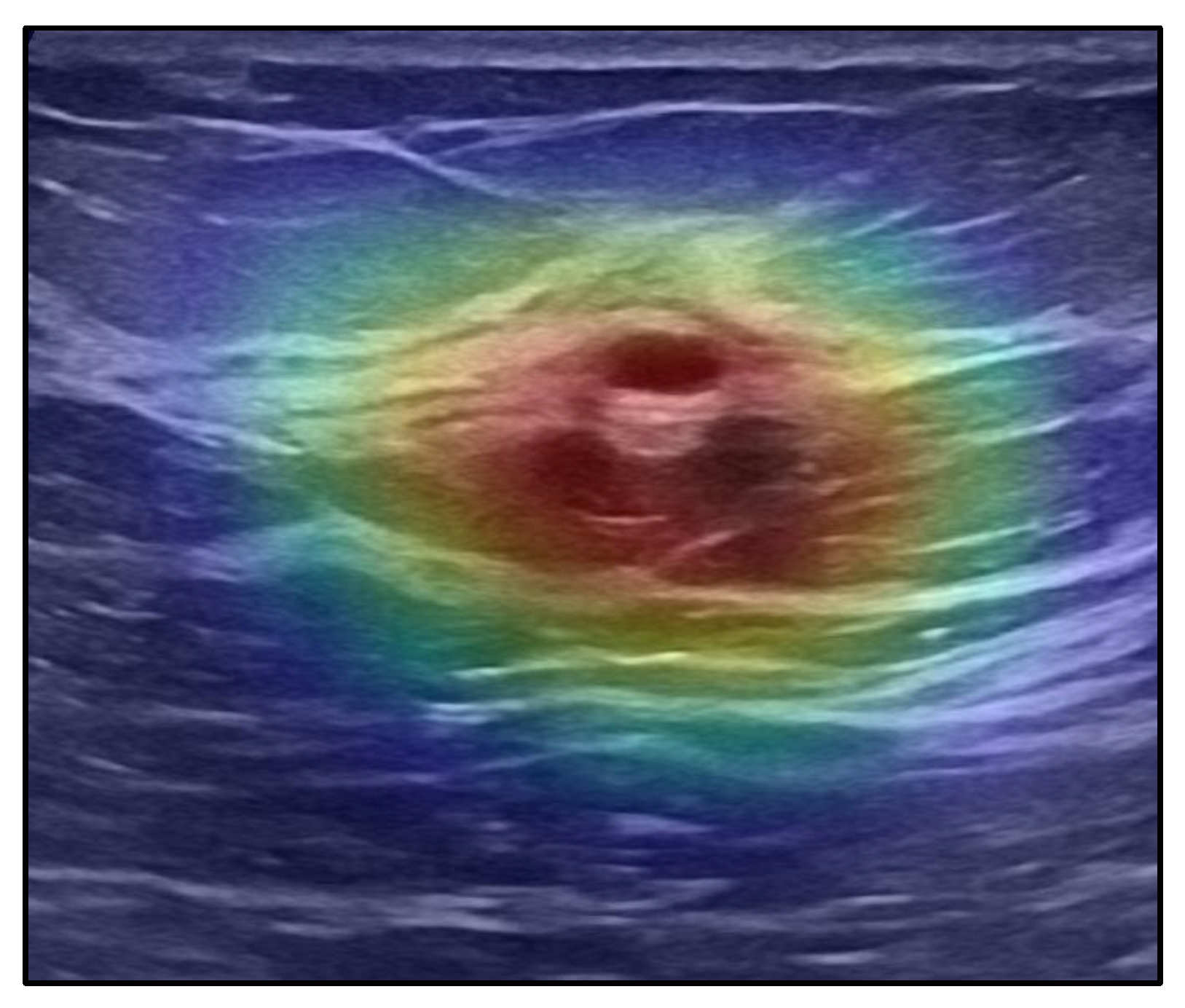
3.2. CNN-Based Classification of Pathology Images for Breast Cancer Detection
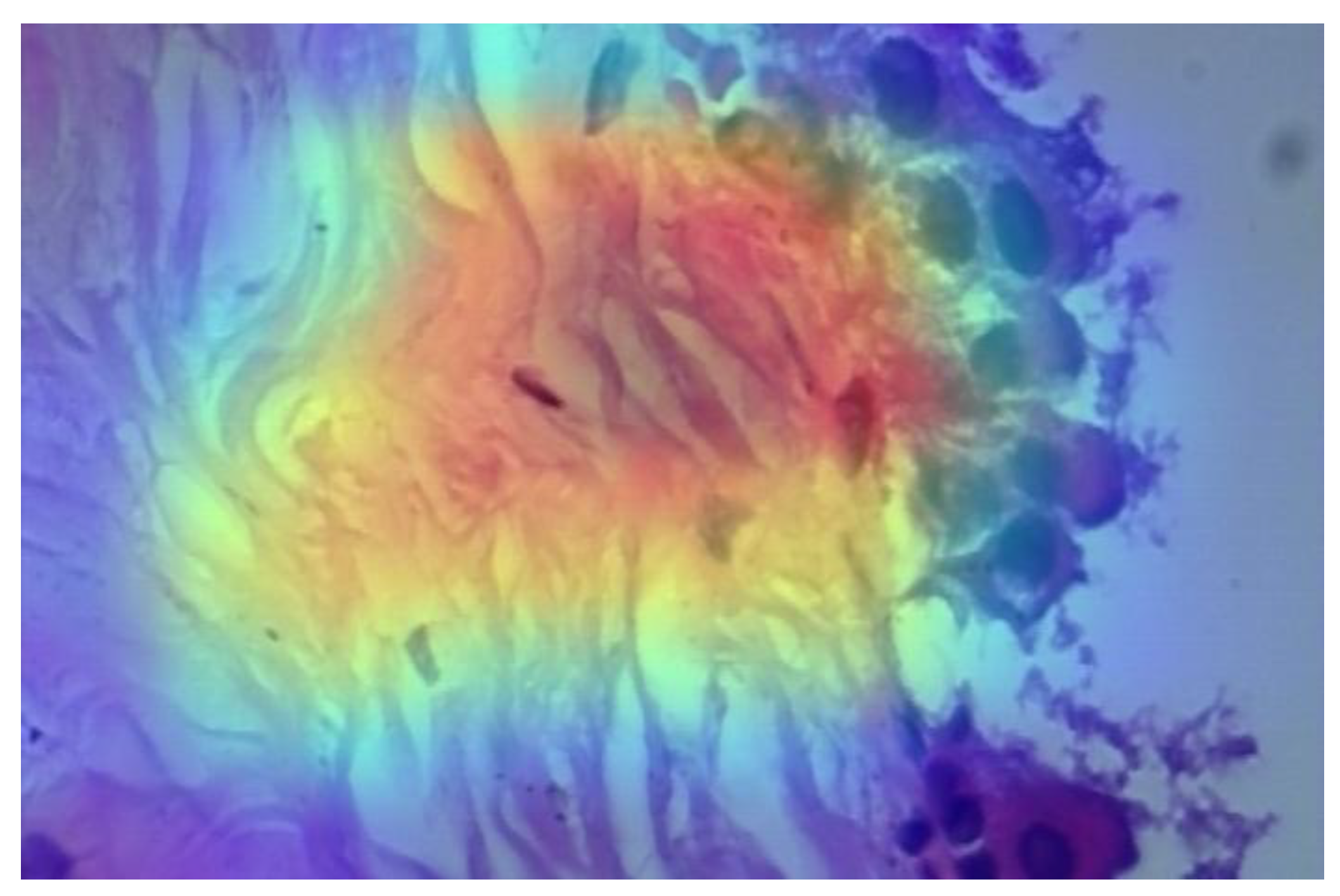
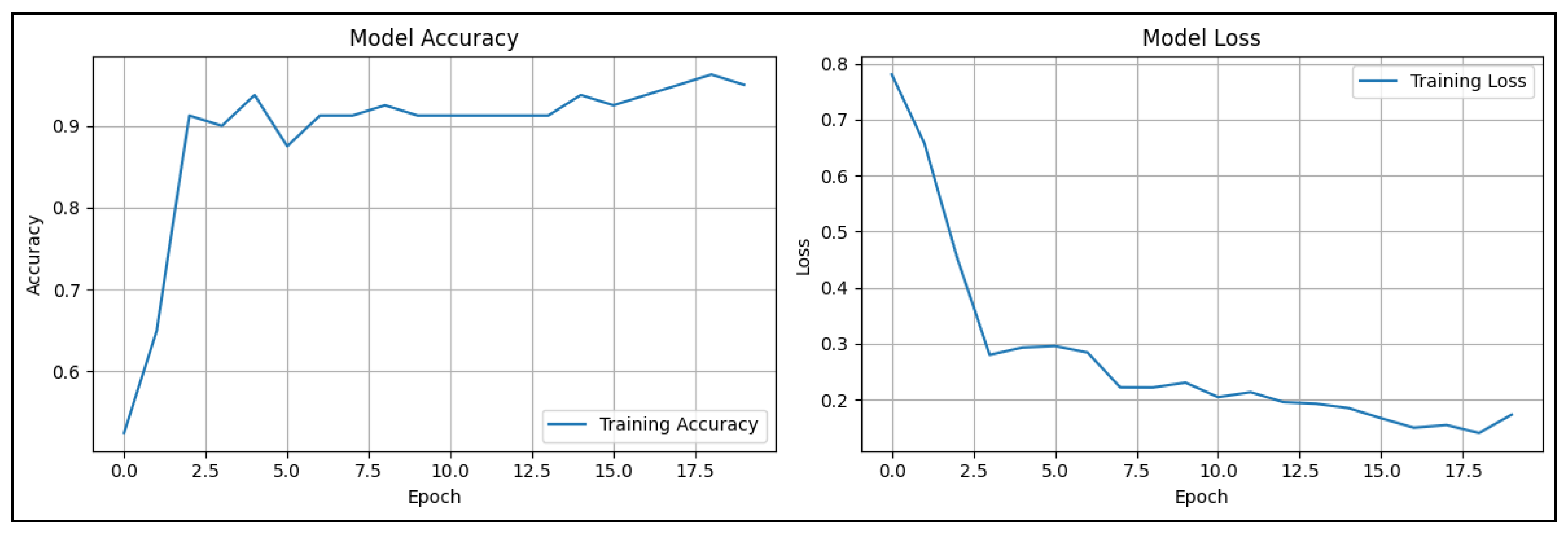
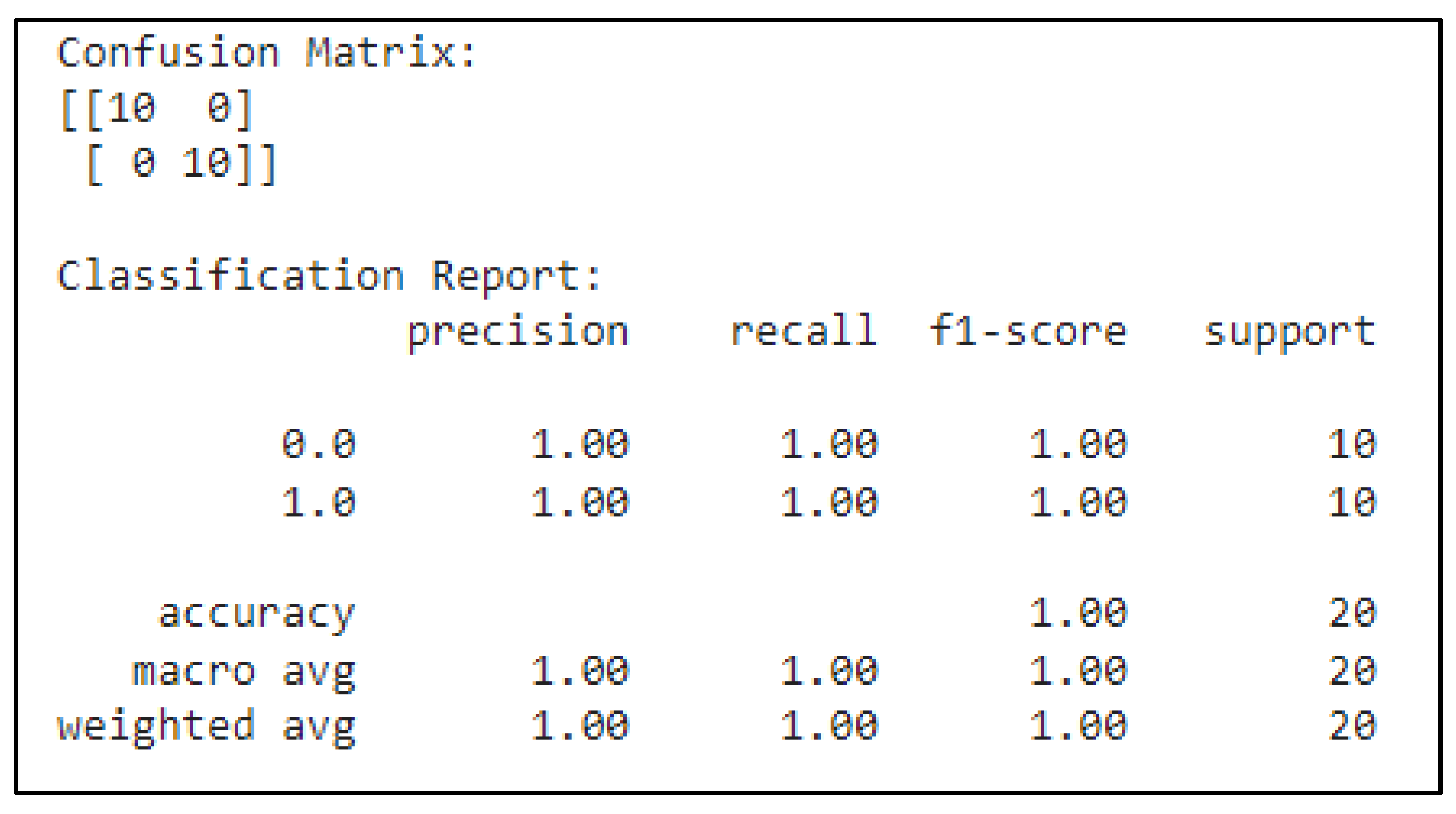
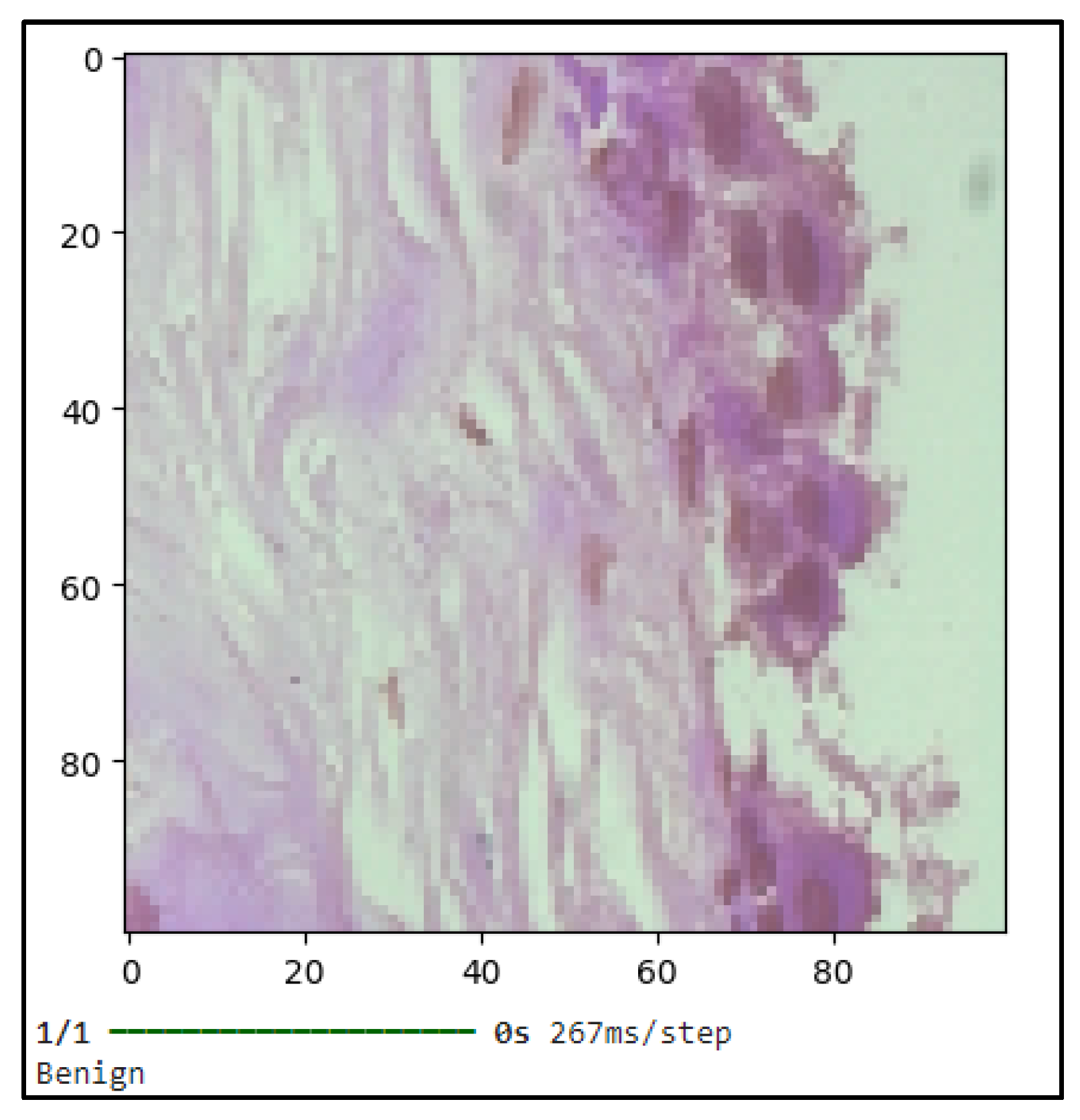
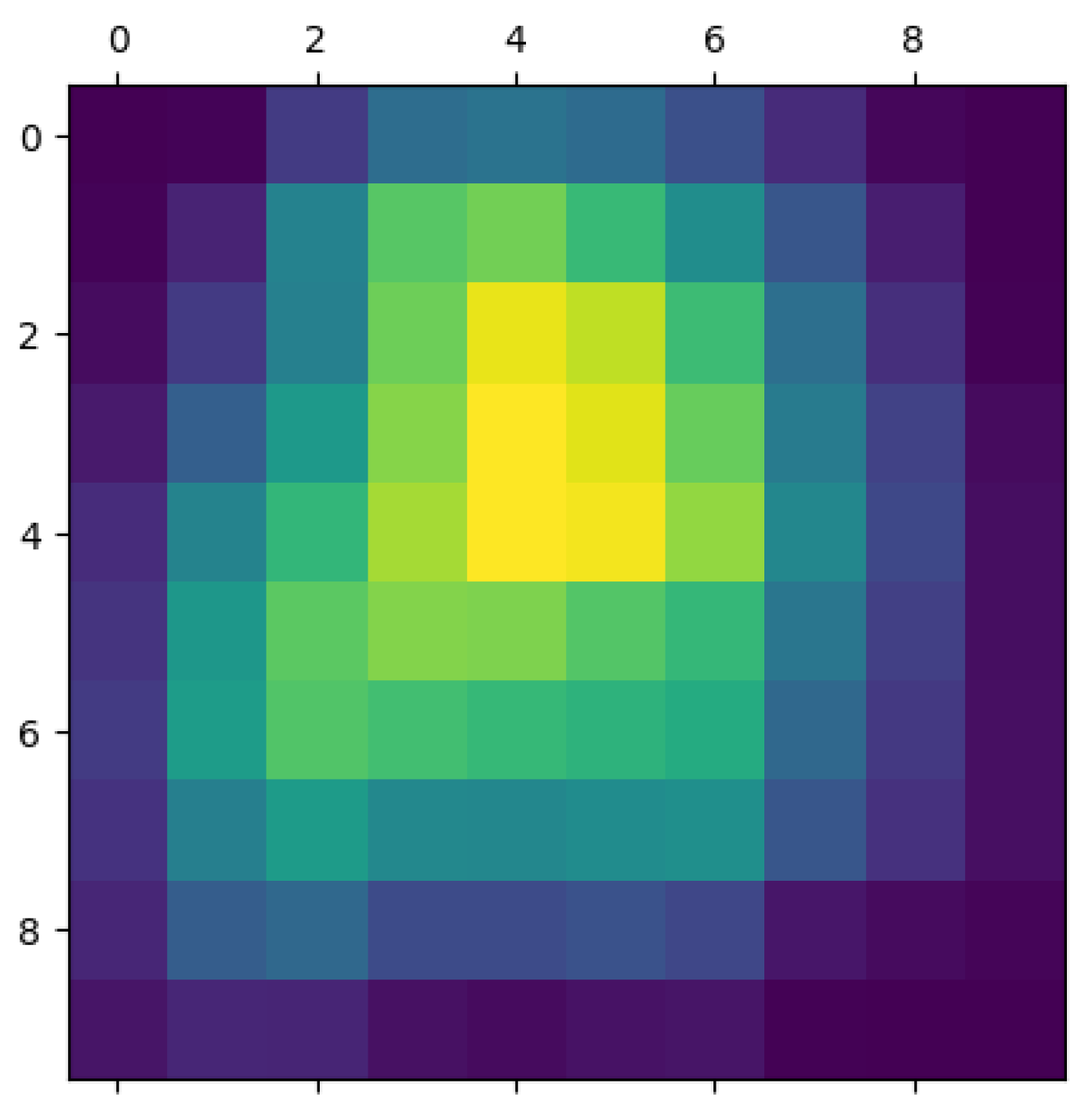

3.3. ANN-Based Classification of Patients' Data for Breast Cancer Detection
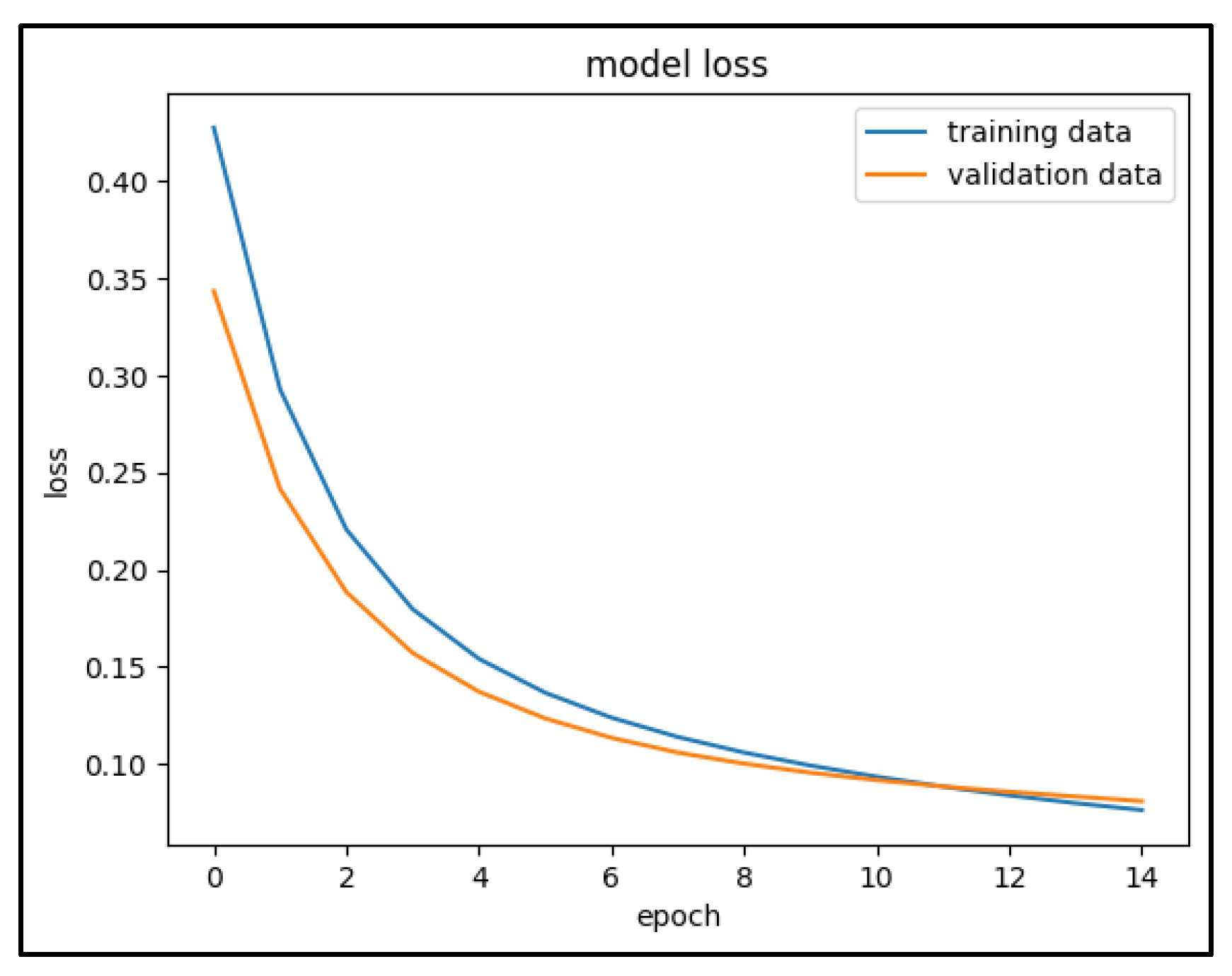
3. Discussion
4. Conclusion
Author Contributions
Data Availability Statement
Acknowledgements
Conflicts of Interest
References
- J. S. Ahn et al., “Artificial Intelligence in Breast Cancer Diagnosis and Personalized Medicine,” Oct. 01, 2023, Korean Breast Cancer Society. [CrossRef]
- H. Sung et al., “Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries,” CA Cancer J Clin, vol. 71, no. 3, pp. 209–249, May 2021. [CrossRef]
- Z. S. Lima, M. R. Z. S. Lima, M. R. Ebadi, G. Amjad, and L. Younesi, “Application of imaging technologies in breast cancer detection: A review article,” Mar. 15, 2019, Open Access Macedonian Journal of Medical Sciences. [CrossRef]
- L. Fass, “Imaging and cancer: A review,” 2008, Elsevier. [CrossRef]
- W. A. Berg et al., “Combined screening with ultrasound and mammography vs mammography alone in women at elevated risk of breast cancer,” JAMA, vol. 299, no. 18, pp. 2151–2163, May 2008. [CrossRef]
- Gheonea, *!!! REPLACE !!!*; et al. , “Review The Role of Imaging Techniques in Diagnosis of Breast Cancer,” 2011.
- R. Gerami et al., “Review Article A literature review on the imaging methods for breast cancer,” 2022. [Online]. Available: www.ijppp.org.
- Hosny, C. Parmar, J. Quackenbush, L. H. Schwartz, and H. J. W. L. Aerts, “Artificial intelligence in radiology,” Aug. 01, 2018, Nature Publishing Group. [CrossRef]
- F. Sadoughi, Z. F. Sadoughi, Z. Kazemy, F. Hamedan, L. Owji, M. Rahmanikatigari, and T. T. Azadboni, “Artificial intelligence methods for the diagnosis of breast cancer by image processing: A review,” 2018, Dove Medical Press Ltd. [CrossRef]
- W. T. Tran et al., “Personalized Breast Cancer Treatments Using Artificial Intelligence in Radiomics and Pathomics,” Dec. 01, 2019, Elsevier Inc. [CrossRef]
- F. Boniolo, E. F. Boniolo, E. Dorigatti, A. J. Ohnmacht, D. Saur, B. Schubert, and M. P. Menden, “Artificial intelligence in early drug discovery enabling precision medicine,” 2021, Taylor and Francis Ltd. [CrossRef]
- S. M. McKinney et al., “International evaluation of an AI system for breast cancer screening,” Nature, vol. 577, no. 7788, pp. 89–94, Jan. 2020. [CrossRef]
- K. Dembrower, A. K. Dembrower, A. Crippa, E. Colón, M. Eklund, and F. Strand, “Artificial intelligence for breast cancer detection in screening mammography in Sweden: a prospective, population-based, paired-reader, non-inferiority study,” Lancet Digit Health, vol. 5, no. 10, pp. e703–e711, Oct. 2023. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).



