Submitted:
27 October 2024
Posted:
28 October 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results
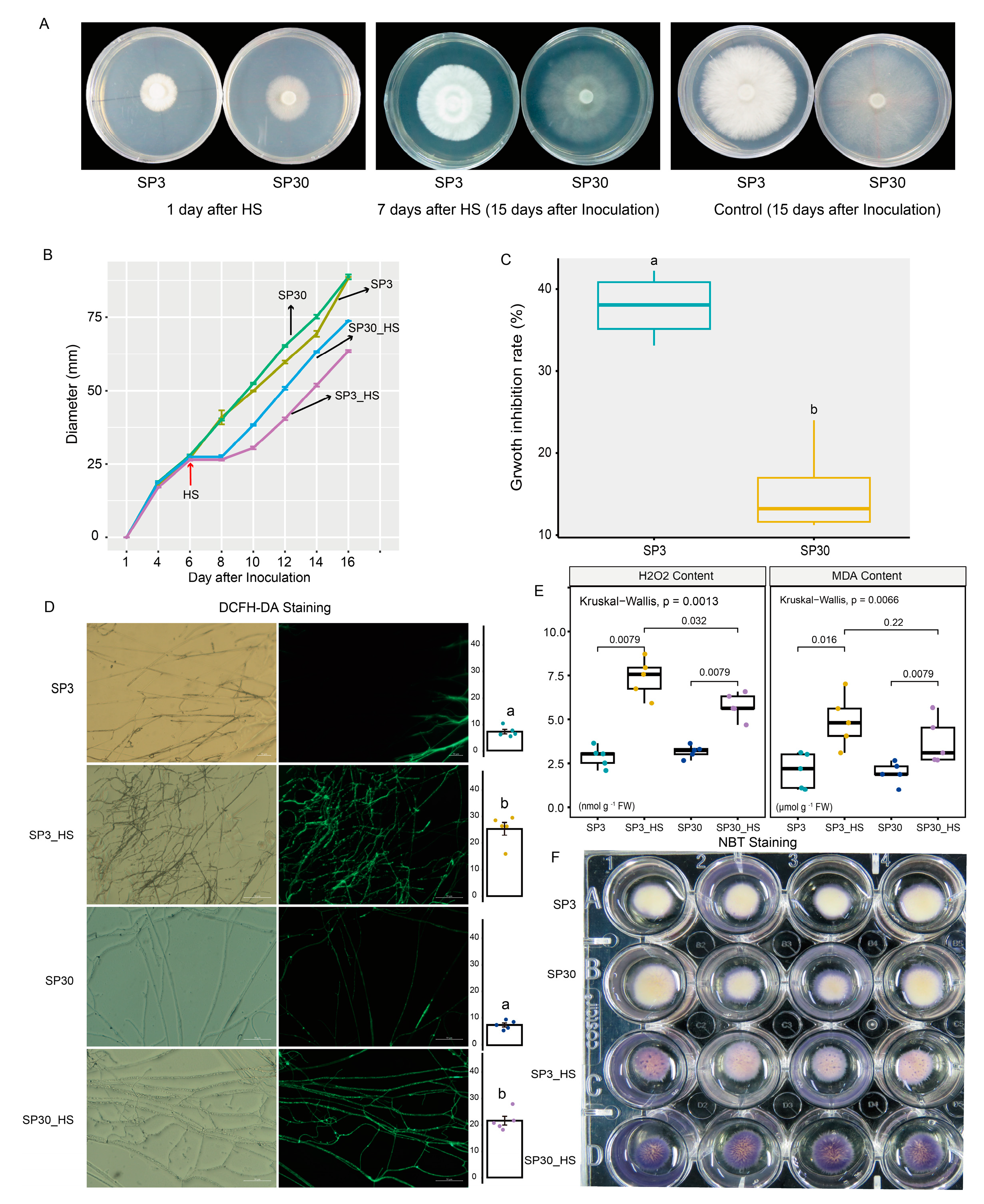
2.1. Thermostress Triggered ROS Production and Inhibited Mycelium Growth of the Two Monokaryons
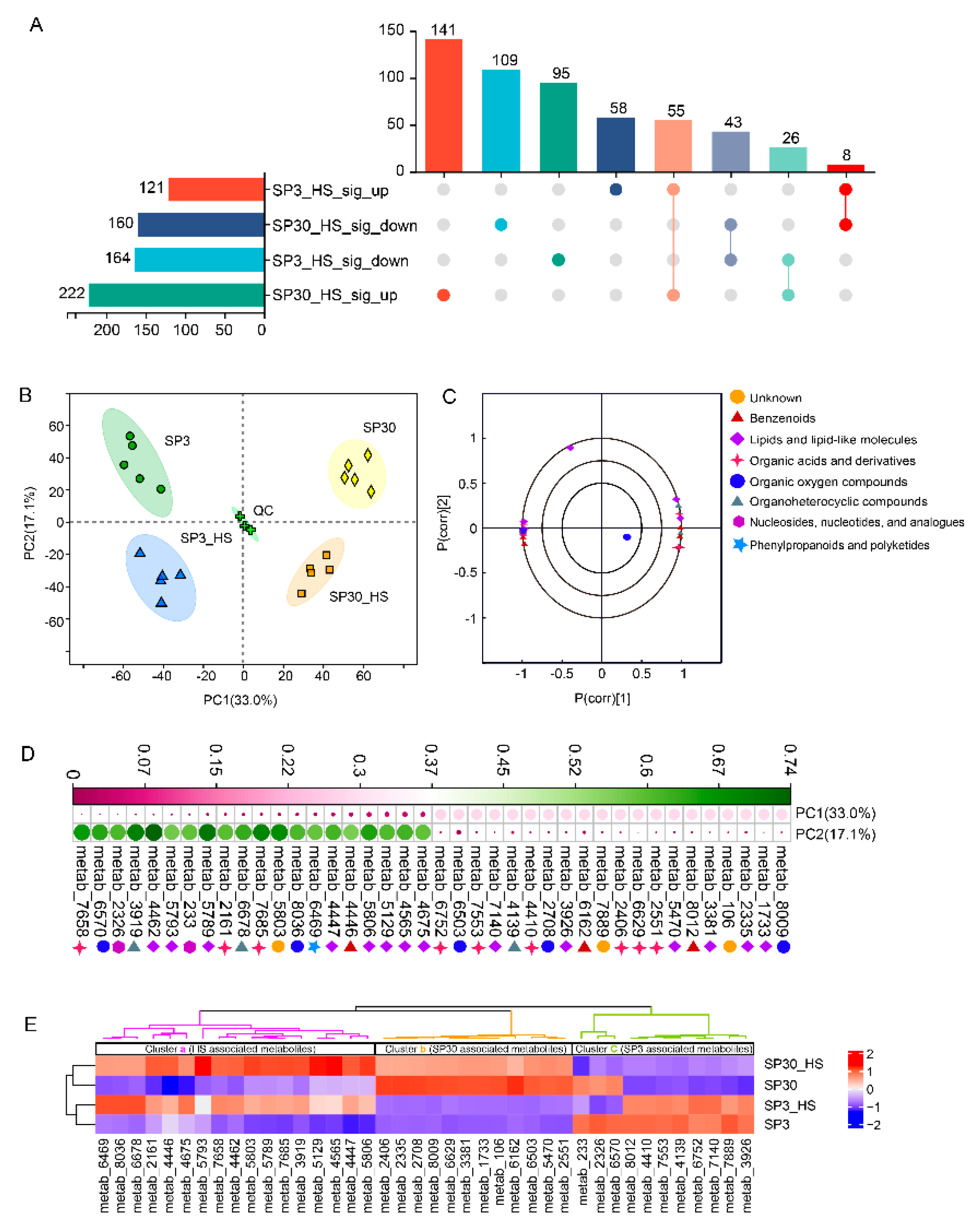
2.2. The Two Monokaryons Showed Distinct Metabolic Profiles under Normal and HS Conditions
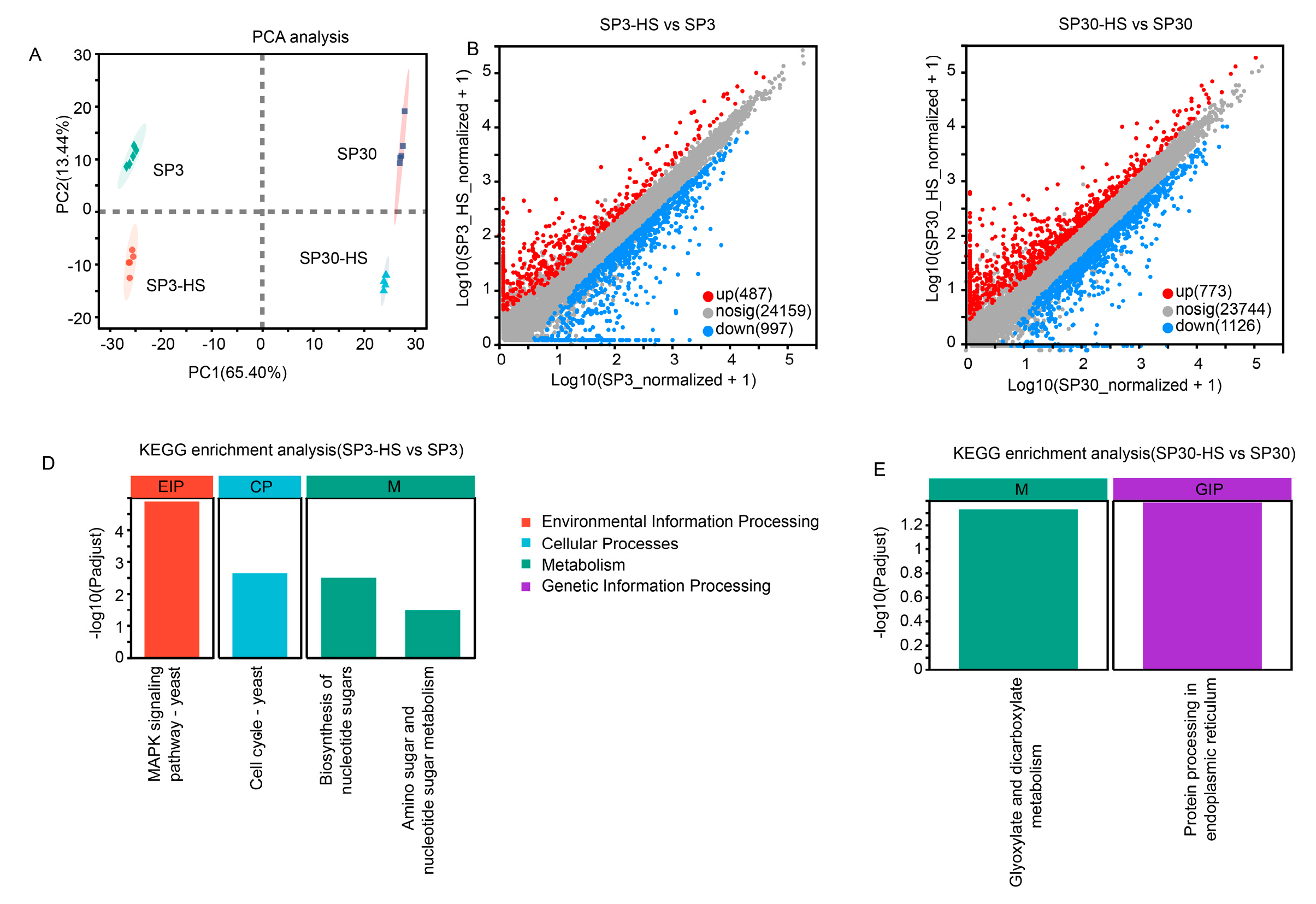
2.3. The Two Monokaryons Showed Distinct Transcriptomic Profiles under Normal and HS Conditions
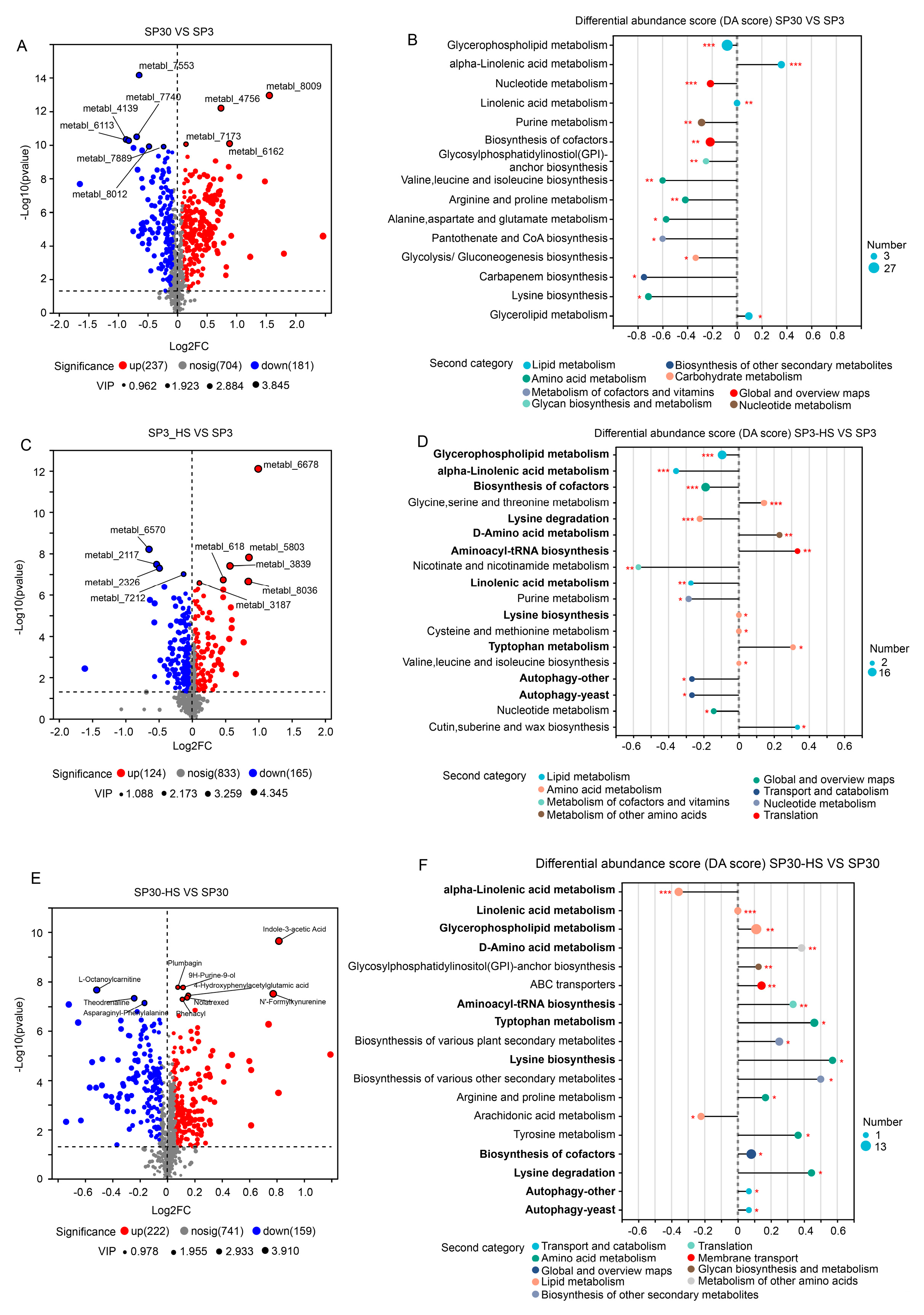
2.4. Identification of the Key Metabolic Pathway of the Two Monokaryons in Response to HS
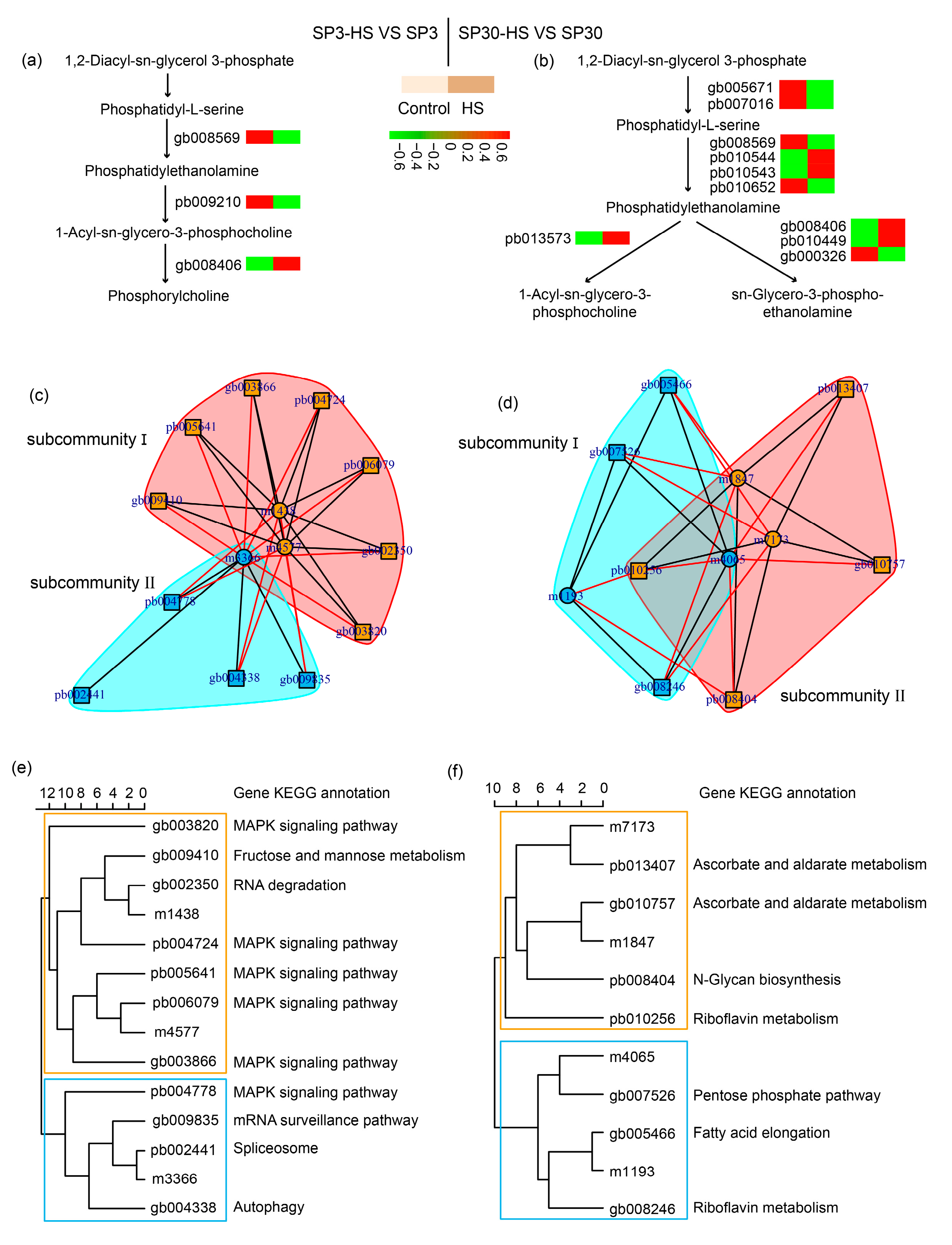
2.5. Glycerophospholipid Metabolism Pathway Analysis
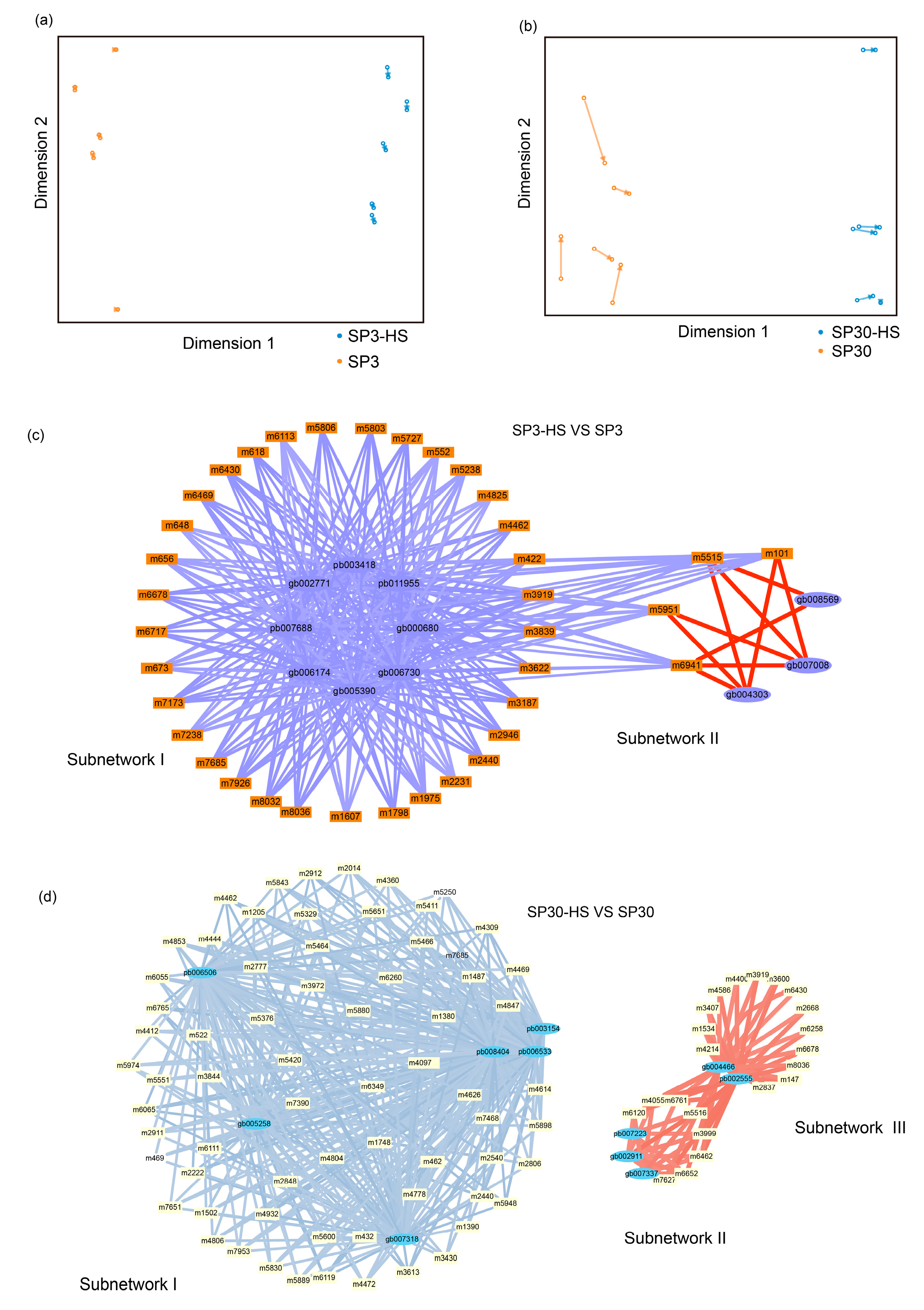
2.6. Integrative Analysis of Upregulated Metabolites with Transcriptome
3. Discussion
4. Materials and Methods
4.1. Fungal Strains and Cultivation
4.2. Measurement of Growth Rate
4.3. Detection of Reactive Oxygen Species and Malondialdehyde Content
4.4. RNA Isolation, Sequencing, Differential Expression Analysis and Functional Annotation
4.5. Non-Target Metabolomics Analysis
4.6. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Singh, M.; Kamal, S.; Sharma, V.P. Status and trends in world mushroom production-III-world production of different mushroom species in 21st century. Mushroom Res. 2020, 29, 75–111. [Google Scholar] [CrossRef]
- Ahmad, I.; Arif, M.; Xu, M.; Zhang, J.; Ding, Y.; Lyu, F. Therapeutic values and nutraceutical properties of shiitake mushroom (Lentinula edodes): A review. Trends Food Sci. Technol. 2023, 134, 123–135. [Google Scholar] [CrossRef]
- Xu, X.; Yu, C.; Liu, Z.; Cui, X.; Guo, X.; Wang, H. Chemical composition, antioxidant and anti-inflammatory activity of shiitake mushrooms (lentinus edodes). J. Fungi 2024, 10, 552. [Google Scholar] [CrossRef]
- Zhang, Q.; Feng, R.; Miao, R.; Lin, J.; Cao, L.; Ni, Y.; Li, W.; Zhao, X. Combined transcriptomics and metabolomics analysis reveals the molecular mechanism of heat tolerance of Le023M, a mutant in Lentinula edodes. Heliyon 2023, 9. [Google Scholar] [CrossRef]
- Sierra-Patev, S.; Min, B.; Naranjo-Ortiz, M.; Looney, B.; Konkel, Z.; Slot, J.C.; Sakamoto, Y.; Steenwyk, J.L.; Rokas, A.; Carro, J. A global phylogenomic analysis of the shiitake genus Lentinula. Proc. Natl. Acad. Sci. U. S. A 2023, 120, e2214076120. [Google Scholar] [CrossRef]
- Menolli Jr, N.; Sanchez-Ramirez, S.; Sanchez-Garcia, M.; Wang, C.; Patev, S.; Ishikawa, N.K.; Mata, J.L.; Lenz, A.R.; Vargas-Isla, R.; Liderman, L. Global phylogeny of the shiitake mushroom and related Lentinula species uncovers novel diversity and suggests an origin in the Neotropics. Mol. Phylogenet. and Evo. 2022, 173, 107494. [Google Scholar] [CrossRef] [PubMed]
- Zhao, X.; Chen, M.; Zhao, Y.; Zha, L.; Yang, H.; Wu, Y. GC–MS-based nontargeted and targeted metabolic profiling identifies changes in the Lentinula edodes mycelial metabolome under high-temperature stress. Int. J. Mol. Sci. 2019, 20, 2330. [Google Scholar] [CrossRef]
- Yang, H.; Jiang, J.; Chen, M.; Song, X.; Yu, C.; Chen, H.; Zhao, Y. Homologous delta-12 fatty acid desaturase (fad2) genes affect gene expression and linoleic acid levels in Lentinula edodes under heat stress. J. Fungi 2024, 10, 496. [Google Scholar] [CrossRef]
- Shen, Y.; Cai, W.; Zhou, S.; Jin, Q.; Feng, W.; Fan, L.; Song, T.; Li, L. Phenotype analysis of mycelium growth regeneration after heat stress in a Lentinula edodes F2 population. J. Hortic. Sci. Biotechnol 2017, 92, 397–403. [Google Scholar] [CrossRef]
- González-García, M.P.; Conesa, C.M.; Lozano-Enguita, A.; Baca-González, V.; Simancas, B.; Navarro-Neila, S.; Sánchez-Bermúdez, M.; Salas-González, I.; Caro, E.; Castrillo, G. Temperature changes in the root ecosystem affect plant functionality. Plant Commun. 2023, 4. [Google Scholar] [CrossRef]
- Guo, Y.; Gao, Q.; Fan, Y.; Song, S.; Yan, D.; Zhao, J.; Chen, Y.; Liu, Y.; Wang, S. Two strains of Lentinula edodes differ in their transcriptional and metabolic patterns and respond differently to thermostress. J. Fungi 2023, 9, 179. [Google Scholar] [CrossRef] [PubMed]
- Wang, G.-Z.; Ma, C.-J.; Luo, Y.; Zhou, S.-S.; Zhou, Y.; Ma, X.-L.; Cai, Y.-L.; Yu, J.-J.; Bian, Y.-B.; Gong, Y.-H. Proteome and transcriptome reveal involvement of heat shock proteins and indoleacetic acid metabolism process in Lentinula edodes thermotolerance. Cell. Physiol. Biochem. 2018, 50, 1617–1637. [Google Scholar] [CrossRef] [PubMed]
- Luo, L.; Zhang, S.; Wu, J.; Sun, X.; Ma, A. Heat stress in macrofungi: effects and response mechanisms. Appl. Microbiol. Biotechnol. 2021, 105, 7567–7576. [Google Scholar] [CrossRef] [PubMed]
- Wang, G.; Luo, Y.; Wang, C.; Zhou, Y.; Mou, C.; Kang, H.; Xiao, Y.; Bian, Y.; Gong, Y.H. Hsp40 protein LeDnaJ07 enhances the thermotolerance of Lentinula edodes and regulates IAA biosynthesis by interacting LetrpE. Front. Microbiol. 2020, 11, 707. [Google Scholar] [CrossRef] [PubMed]
- Xu, R.; Zhou, S.; Song, J.; Zhong, H.; Zhu, T.; Gong, Y.; Zhou, Y.; Bian, Y. Comparative transcriptome analysis provides insights into the mechanism by which 2,4-dichlorophenoxyacetic acid improves thermotolerance in Lentinula edodes. Front. Microbiol. 2022, 13, 910255. [Google Scholar] [CrossRef] [PubMed]
- Shen, N.; Xie, H.; Liu, K.; Li, X.; Wang, L.; Deng, Y.; Chen, L.; Bian, Y.; Xiao, Y. Near-gapless genome and transcriptome analyses provide insights into fruiting body development in Lentinula edodes. Int. J. Biol. Macromol. 2024, 263, 130610. [Google Scholar] [CrossRef]
- Gao, Q.; Yan, D.; Song, S.; Fan, Y.; Wang, S.; Liu, Y.; Huang, Y.; Rong, C.; Guo, Y.; Zhao, S. Haplotype-resolved genome analyses reveal genetically distinct nuclei within a commercial cultivar of Lentinula edodes. J. Fungi 2022, 8, 167. [Google Scholar] [CrossRef]
- Yoo, S.-i.; Moon, S.; Hong, C.P.; Park, S.-G.; Shim, D.; Ryu, H. Genome sequencing of Lentinula edodes revealed a genomic variant block associated with a thermo-tolerant trait in fruit body formation. J. Fungi 2024, 10, 628. [Google Scholar] [CrossRef]
- Feng, B.; Xiong, J.; Tao, L. How rice plants response to abiotic stresses. Int. J. Mol. Sci. 2023, 24, 12806. [Google Scholar] [CrossRef]
- Zhou, J.; Tang, X.; Li, J.; Dang, S.; Ma, H.; Zhang, Y. Comparative transcriptomic and metabolomic analyses provide insights into the responses to high temperature stress in alfalfa (Medicago sativa L.). BMC Plant Bio. 2024, 24, 776. [Google Scholar] [CrossRef]
- Sun, M.; Huang, D.; Zhang, A.; Khan, I.; Yan, H.; Wang, X.; Zhang, X.; Zhang, J.; Huang, L. Transcriptome analysis of heat stress and drought stress in pearl millet based on Pacbio full-length transcriptome sequencing. BMC Plant Bio. 2020, 20, 323. [Google Scholar] [CrossRef] [PubMed]
- Jiang, C.; Ge, J.; He, B.; Zhang, Z.; Hu, Z.; Li, Y.; Zeng, B. Transcriptomic analysis reveals Aspergillus oryzae responds to temperature stress by regulating sugar metabolism and lipid metabolism. Plos one 2022, 17, e0274394. [Google Scholar] [CrossRef] [PubMed]
- Yan, Z.; Zhao, M.; Wu, X.; Zhang, J. Metabolic response of Pleurotus ostreatus to continuous heat stress. Front. Microbiol 2020, 10, 3148. [Google Scholar] [CrossRef] [PubMed]
- Nieuwenhuis, B.P.; James, T.Y. The frequency of sex in fungi. Philos. T. R. Soc. B. 2016, 371, 20150540. [Google Scholar] [CrossRef] [PubMed]
- Sperschneider, J.; Yildirir, G.; Rizzi, Y.S.; Malar C, M.; Mayrand Nicol, A.; Sorwar, E.; Villeneuve-Laroche, M.; Chen, E.C.H.; Iwasaki, W.; Brauer, E.K.; et al. Arbuscular mycorrhizal fungi heterokaryons have two nuclear populations with distinct roles in host–plant interactions. Nat. Microbiol. 2023, 8, 2142–2153. [Google Scholar] [CrossRef] [PubMed]
- Gehrmann, T.; Pelkmans, J.F.; Ohm, R.A.; Vos, A.M.; Sonnenberg, A.S.M.; Baars, J.J.P.; Wösten, H.A.B.; Reinders, M.J.T.; Abeel, T. Nucleus-specific expression in the multinuclear mushroom-forming fungus <i>Agaricus bisporus</i> reveals different nuclear regulatory programs. Proc. Natl. Acad. Sci. U. S. A. 2018, 115, 4429–4434. [Google Scholar] [PubMed]
- Ling, Y.-Y.; Ling, Z.-L.; Zhao, R.-L. Construction of a heat-resistant strain of Lentinus edodes by fungal Hsp20 protein overexpression and genetic transformation. Front. Microbiol. 2022, 13, 1009885. [Google Scholar] [CrossRef] [PubMed]
- Ren, A.; Liu, R.; Miao, Z.G.; Zhang, X.; Cao, P.F.; Chen, T.X.; Li, C.Y.; Shi, L.; Jiang, A.L.; Zhao, M.W. Hydrogen-rich water regulates effects of ROS balance on morphology, growth and secondary metabolism via glutathione peroxidase in Ganoderma lucidum. Environ. Microbiol. 2017, 19, 566–583. [Google Scholar] [CrossRef]
- Kim, D.; Langmead, B.; Salzberg, S.L. HISAT: a fast spliced aligner with low memory requirements. Nat. Methods 2015, 12, 357–360. [Google Scholar] [CrossRef]
- Pertea, M.; Pertea, G.M.; Antonescu, C.M.; Chang, T.-C.; Mendell, J.T.; Salzberg, S.L. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat. Biotechnol. 2015, 33, 290–295. [Google Scholar] [CrossRef]
- Li, B.; Dewey, C.N. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC bioinf. 2011, 12, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome biol. 2014, 15, 1–21. [Google Scholar] [CrossRef]
- Benjamini, Y.; Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc., Ser. B, Methodol. 1995, 57, 289–300. [Google Scholar] [CrossRef]
- Zhang, J.; Ye, L.; Chen, Q.; Wang, F. Response analysis of Pinus sibirica to pine wood nematode infection through transcriptomics and metabolomics study. Front. Plant Sci. 2024, 15, 1383018. [Google Scholar] [CrossRef]
- Lê, S.; Josse, J.; Husson, F. FactoMineR: an R package for multivariate analysis. J. Stat. Softw. 2008, 25, 1–18. [Google Scholar] [CrossRef]
- Kassambara, A.; Mundt, F. Package ‘factoextra’. Extract and visualize the results of multivariate data analyses. Available online: https://cran.r-project.org/web/packages/factoextra/index.html (accessed on 25.10.2024.
- Dixon, P. VEGAN, a package of R functions for community ecology. J. Veg. Sci. 2003, 14, 927–930. [Google Scholar] [CrossRef]
- Lê Cao, K.-A.; Rossouw, D.; Robert-Granié, C.; Besse, P. A sparse PLS for variable selection when integrating omics data. Stat. Appl. Genet. Mol. Biol. 2008, 7. [Google Scholar] [CrossRef]
- Rohart, F.; Gautier, B.; Singh, A.; Lê Cao, K.-A. mixOmics: An R package for ‘omics feature selection and multiple data integration. PLoS Comput. Biol. 2017, 13, e1005752. [Google Scholar] [CrossRef]
| HSSP3 VS SP3 | ||||
| Gene ID | Log2FC | Padjust | KEGG | Swiss-Prot description |
| pb011955 | 3.015065 | 8.11E-205 | Protein processing in endoplasmic reticulum | Heat shock protein 16 |
| pb003418 | 2.197605 | 9.04E-178 | Longevity regulating pathway | Heat shock protein 104 |
| gb002771 | 3.061065 | 2.23E-101 | Longevity regulating pathway | Heat shock protein 104 |
| pb007688 | 1.614501 | 1.09E-134 | Biotin metabolism | Biotin--protein ligase |
| gb006174 | 1.614501 | 1.09E-134 | Biotin metabolism | Biotin--protein ligase |
| HSSP30VS SP30 | ||||
| pb006533 | -5.35584 | 2.07E-241 | - | |
| pb003154 | -1.10127 | 1.19E-67 | - | Uncharacterized secreted protein ARB_0804 |
| gb005258 | -6.18295 | 0 | - | - |
| gb007318 | -1.38438 | 1.03E-70 | - | gmc oxidoreductase |
| pb008404 | -1.24555 | 9.01E-170 | Protein processing in endoplasmic reticulum | Probable mannosyl-oligosaccharide alpha-1,2-mannosidase 1B |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




