Submitted:
27 October 2024
Posted:
28 October 2024
You are already at the latest version
Abstract
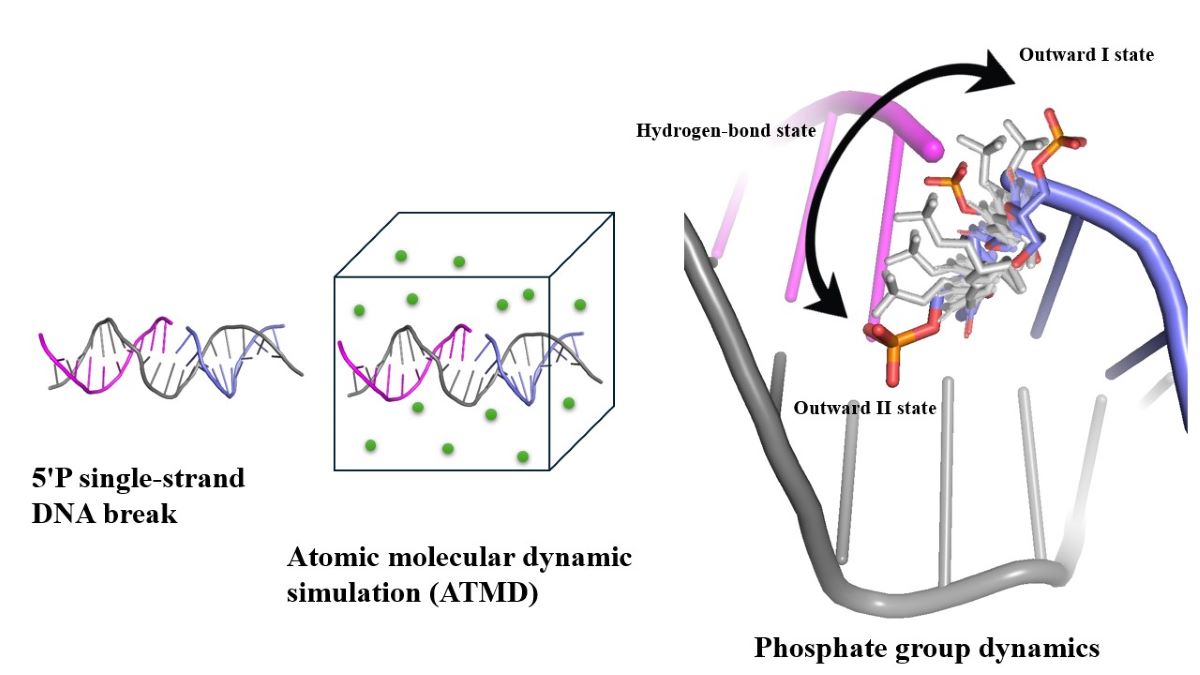
Keywords:
Introduction
Methods
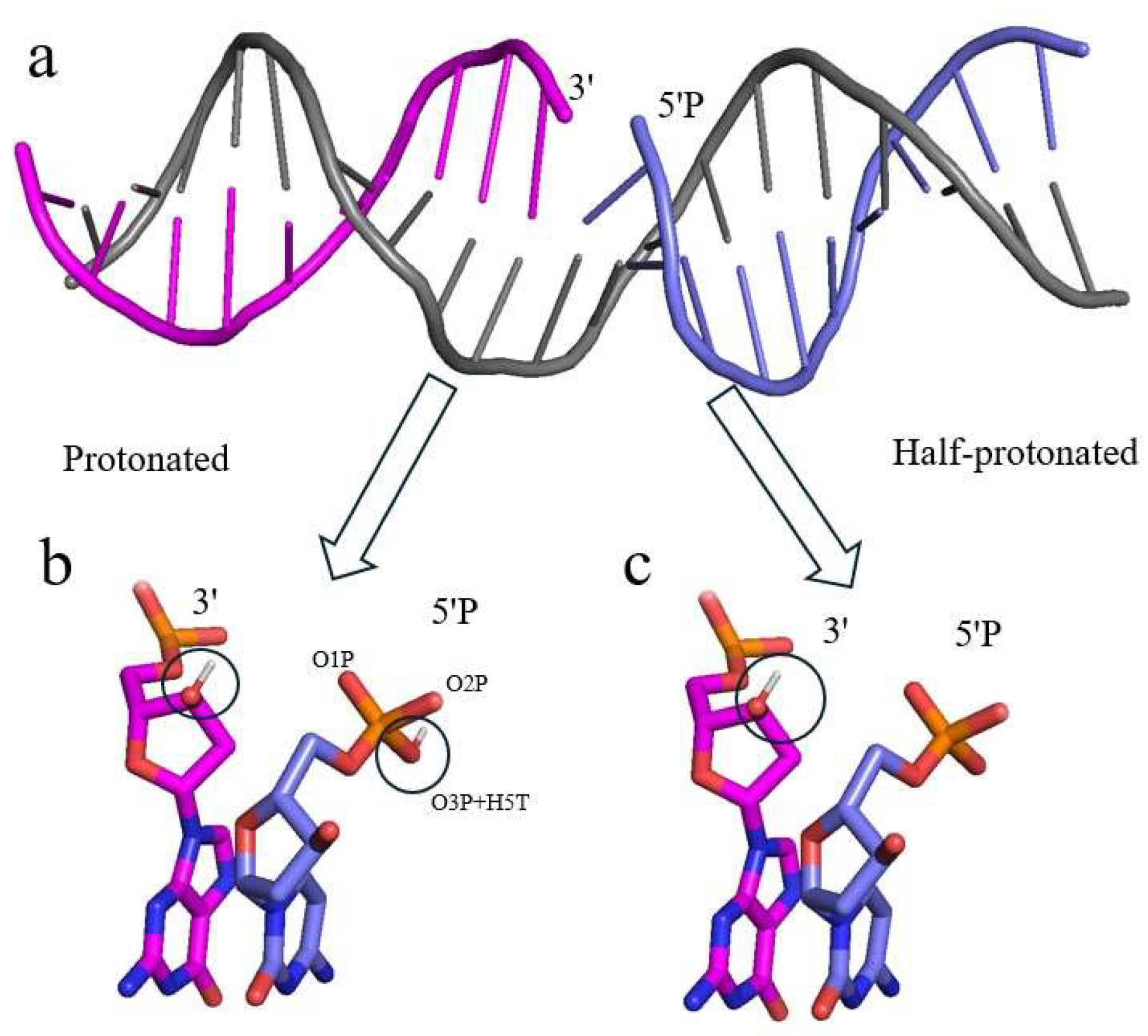
Damaged DNA Templates
Charge Distributions of the 5’ Phosphate Group
| Atom name | Standard AMBER | Bunta group | Acpype | Charge set1 | Charge set2 | Charge set3 | Charge set4 | Charge set5 | Half-pro | |
| Gaussian ESP | RESP | |||||||||
| O3’ | -0.5232 | -0.7894 | -0.5991 | -0.5232 | -0.5232 | -0.5232 | -0.5232 | -0.5232 | -0.5232 | -0.5232 |
| H3T | 0.4396 | -0.4529 | 0.3541 | 0.4396 | 0.4396 | 0.4396 | 0.4396 | 0.4396 | 0.4396 | 0.4396 |
| P | 1.1659 | 1.3139 | 1.1436 | 1.4248 | 1.1559 | 1.1659 | 1.1659 | 1.1659 | 1.1659 | 1.1659 |
| O3P | - | -0.6602 | -0.5907 | -0.7784 | -0.5054 | -0.4954 | -0.4954 | -0.4954 | -0.4954 | -0.6226 |
| O2P | -0.7761 | - | - | -0.8395 | -0.7761 | -0.7761 | -0.7761 | -0.7761 | -0.7761 | -0.6226 |
| O1P | -0.7761 | - | - | -0.8395 | -0.7761 | -0.7761 | -0.7761 | -0.7761 | -0.7761 | -0.6226 |
| H5T | 0.4422 | 0.4563 | 0.4334 | 0.4280 | 0.2097 | 0.1897 | 0.1797 | 0.1697 | 0.1597 | - |
| O5’ | -0.6318 | - | - | -0.5902 | -0.5043 | -0.4943 | -0.4843 | -0.4743 | -0.4643 | -0.4943 |
| C5’ | -0.0069 | - | - | 0.1594 | 0.2064 | 0.2064 | 0.2064 | 0.2064 | 0.2064 | 0.2064 |
| H5’1 | 0.0754 | - | - | 0.0757 | 0.0021 | 0.0021 | 0.0021 | 0.0021 | 0.0021 | 0.0021 |
| H5’2 | 0.0754 | - | - | 0.0757 | 0.0021 | 0.0021 | 0.0021 | 0.0021 | 0.0021 | 0.0021 |
| Total | -1 | -2.9121 | -2.0378 | -2 | -2 | -2 | -2 | -2 | -2 | -3 |
Molecular Dynamics Protocol
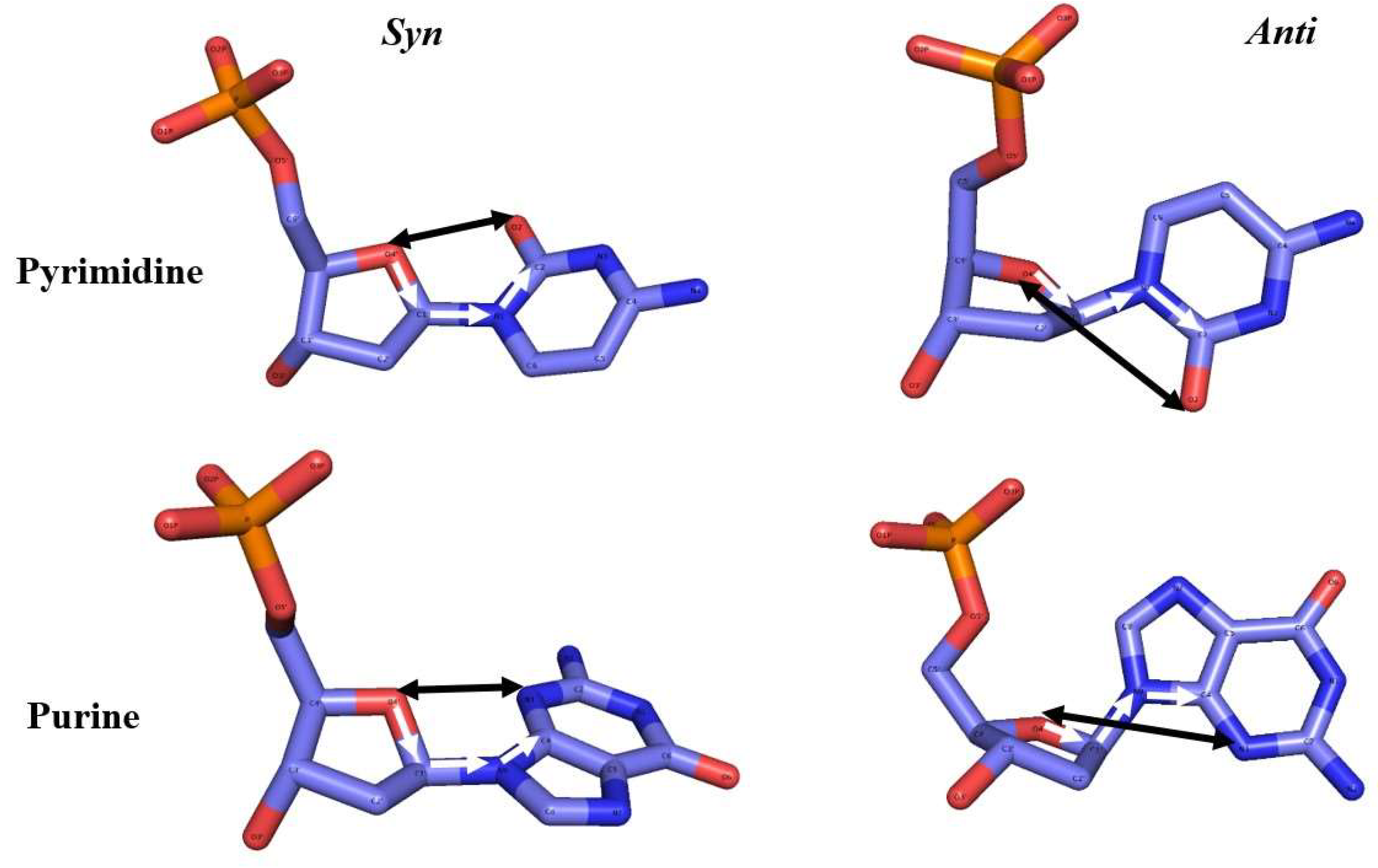
Conformational and Structural Analysis
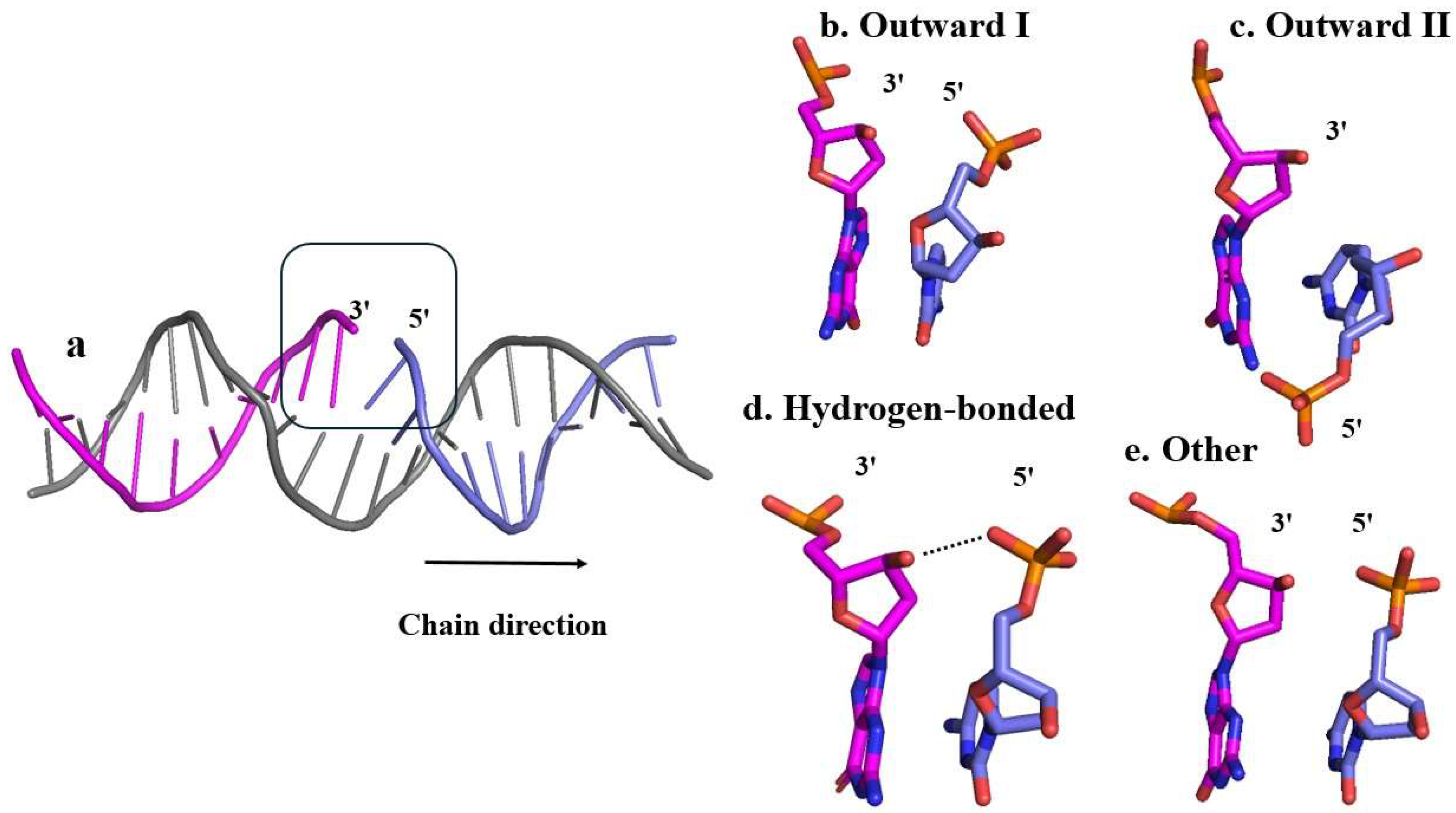
5’-Phosphate Group Dynamics
Results
Charge Sets
DNA Form Analysis
| % | Intact | Set3 | Set4 | |||
| BI | BII | BI | BII | BI | BII | |
| C-break | 83.03 ± 0.50 | 16.97 ± 0.66 | 84.02 ± 0.50 | 15.98 ± 0.51 | 85.94 ± 0.51 | 14.06 ± 0.85 |
| T-break | 84.45 ± 0.51 | 15.55 ± 0.78 | 84.73 ± 0.50 | 15.27 ± 0.70 | 84.47 ± 0.51 | 15.53 ± 0.73 |
| G-break | 81.16 ± 0.50 | 18.84 ± 0.75 | 84.71 ± 0.50 | 15.29 ± 0.63 | 85.52 ± 0.49 | 14.48 ± 0.91 |
| A-break | 81.79 ± 0.53 | 18.21 ± 0.71 | 81.94 ± 0.51 | 18.06 ± 0.82 | 86.11 ± 0.48 | 13.89 ± 0.77 |
| % | Set5 | Half-pro | Non-P | |||
| BI | BII | BI | BII | BI | BII | |
| C-break | 86.06 ± 0.51 | 13.94 ± 0.83 | 85.52 ± 0.49 | 14.48 ± 0.80 | 84.81 ± 0.50 | 15.19 ± 0.68 |
| T-break | 85.59 ± 0.49 | 14.41 ±0.74 | 84.61 ± 0.49 | 15.39 ± 0.77 | 85.53 ± 0.49 | 14.47 ± 0.68 |
| G-break | 82.71 ± 0.48 | 17.29 ± 0.68 | 85.48 ± 0.48 | 14.52 ± 0.85 | 86.25 ± 0.46 | 13.75 ± 0.72 |
| A-break | 83.94 ± 0.48 | 16.06 ± 0.66 | 81.99 ± 0.50 | 18.01 ± 0.67 | 84.92 ± 0.50 | 15.08 ± 0.74 |
Helical Parameters
Intra-Base Pair
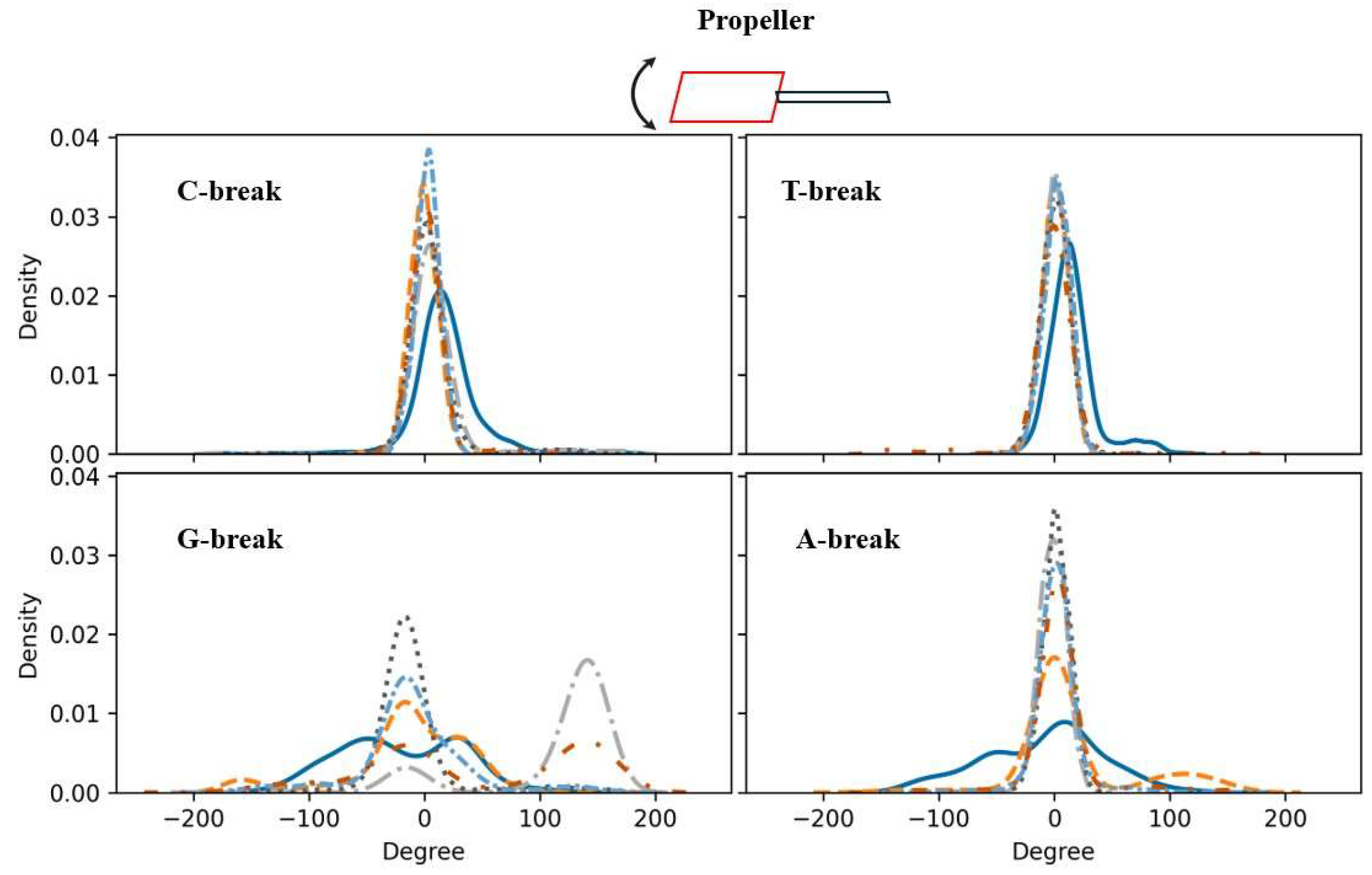
Inter-Base Pair Step
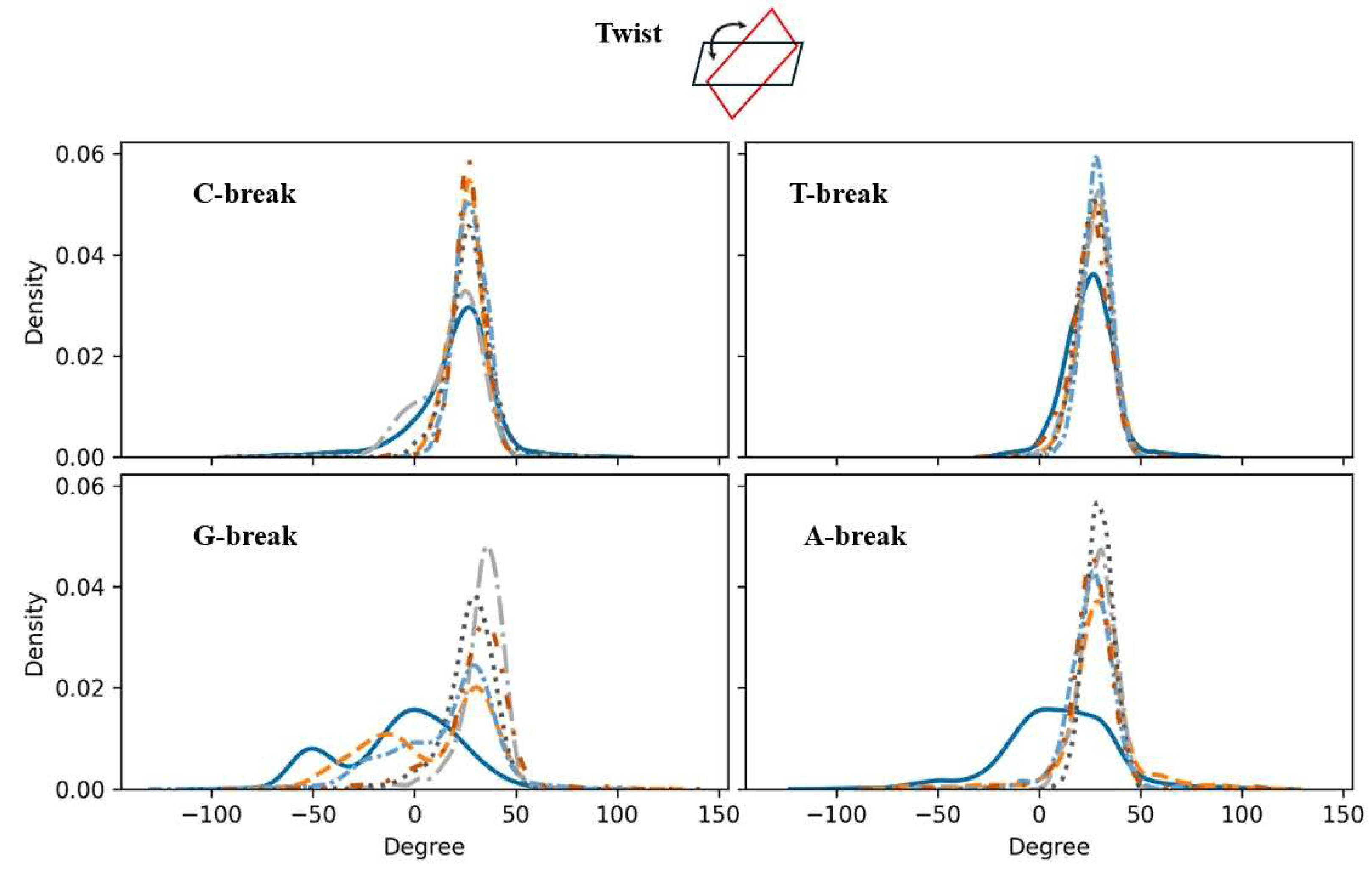
Local Bending Angles
| Intact | Set3 | Set4 | Set5 | Half-pro | Non-P | |
| C-break | 118.25 ± 11.32 | 107.69 ± 11.82 | 110.38 ± 12.12 | 110.64 ±15.25 | 106.79 ± 14.95 | 132.87 ± 18.98 |
| T-break | 121.66 ± 11.62 | 118.97 ± 17.80 | 125.48 ± 17.38 | 135.01 ± 15.75 | 119.98 ± 12.97 | 138.89 ± 18.81 |
| G-break | 113.61 ± 9.62 | 115. 48 ± 14.64 | 112.60 ± 11.68 | 113.62 ± 12.11 | 116.40 ± 12.92 | 125.66 ± 13.78 |
| A-break | 115.19 ± 11.42 | 107.77 ± 11.04 | 107.54 ± 10.37 | 112.71 ± 11.17 | 112.61 ± 11.09 | 122.95 ± 12.75 |
Local Stiffness
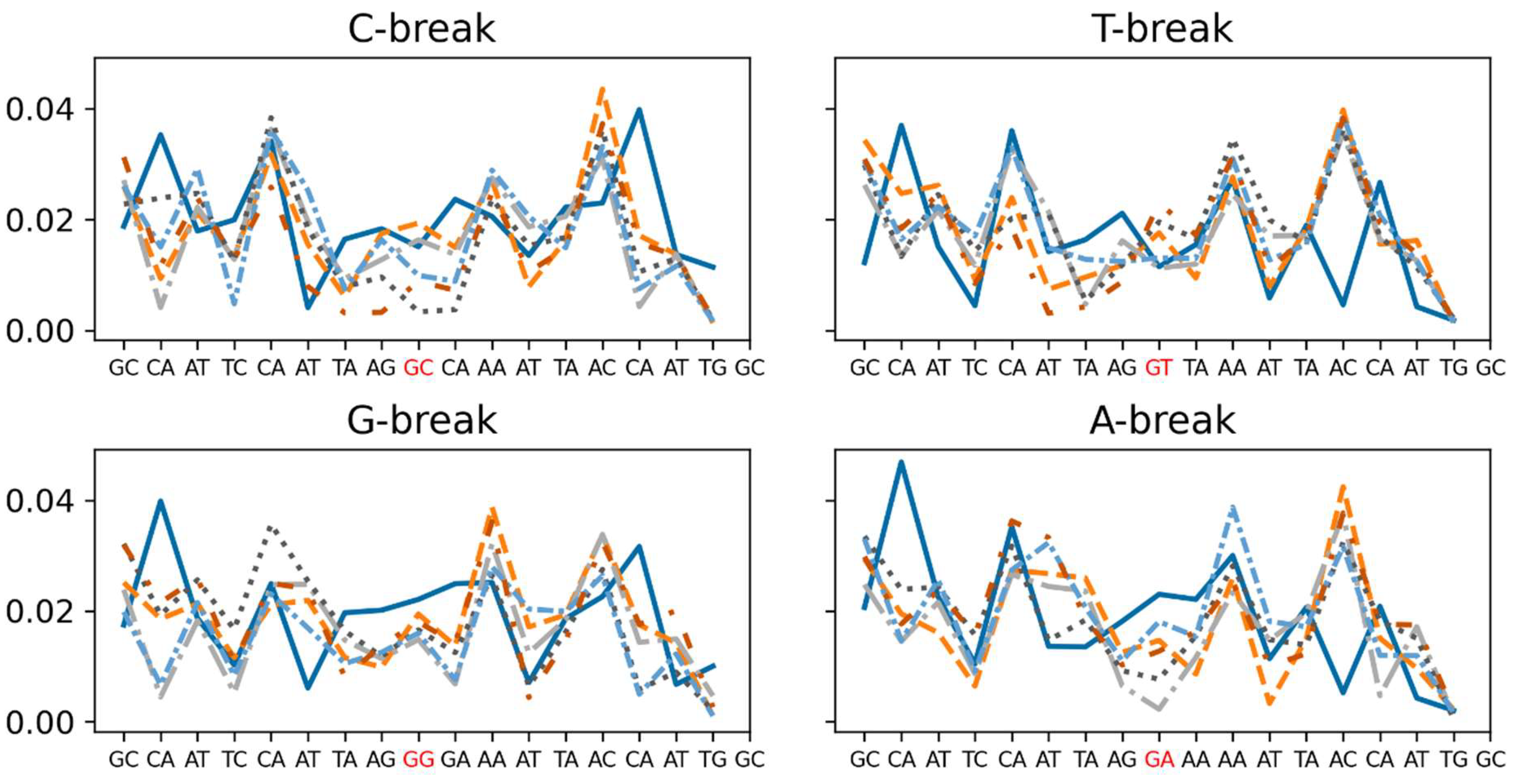
5’. Phosphate Group Dynamics
| A | ||||||
|---|---|---|---|---|---|---|
| C-break | Set3 | Set4 | Set5 | Half-pro | ||
| H-bond | 0.50 ± 0.02 | 0.57 ± 0.02 | 0.50 ± 0.02 | 0.47± 0.02 | ||
| Outward I | 0.18 ± 0.01 | 0.28 ± 0.01 | 0.31 ± 0.01 | 0.35 ± 0.02 | ||
| Outward II | 0.20 ± 0.01 | 0.01 ± 0.00 | 0.02 ± 0.00 | 0.01 ± 0.00 | ||
| Other | 0.11 ± 0.01 | 0.14 ± 0.01 | 0.17 ± 0.01 | 0.17 ± 0.01 | ||
| B | ||||||
| T-break | Set3 | Set4 | Set5 | Half-pro | ||
| H-bond | 0.46 ± 0.02 | 0.50 ± 0.02 | 0.43 ± 0.02 | 0.31± 0.01 | ||
| Outward I | 0.30 ± 0.01 | 0.31 ± 0.01 | 0.38 ± 0.02 | 0.40 ± 0.02 | ||
| Outward II | 0.01 ± 0.00 | 0.03 ± 0.01 | 0.01 ± 0.00 | 0.07 ± 0.01 | ||
| Other | 0.23 ± 0.01 | 0.16 ± 0.01 | 0.19 ± 0.01 | 0.22 ± 0.01 | ||
| C | ||||||
| G-break | Set3 | Set4 | Set5 | Half-pro | ||
| H-bond | 0.39 ± 0.02 | 0.38 ± 0.02 | 0.38 ± 0.02 | 0.25 ± 0.01 | ||
| Outward I | 0.37 ± 0.02 | 0.45 ± 0.02 | 0.48 ± 0.01 | 0.62 ± 0.02 | ||
| Outward II | 0.21 ± 0.01 | 0.12 ± 0.01 | 0.12 ± 0.01 | 0.10 ± 0.01 | ||
| Other | 0.03 ± 0.01 | 0.05 ± 0.01 | 0.02 ± 0.00 | 0.03 ± 0.01 | ||
| D | ||||||
| A-break | Set3 | Set4 | Set5 | Half-pro | ||
| H-bond | 0.50 ± 0.02 | 0.57 ± 0.02 | 0.37 ± 0.02 | 0.30 ± 0.01 | ||
| Outward I | 0.41 ± 0.02 | 0.35 ± 0.02 | 0.36 ± 0.02 | 0.42 ± 0.02 | ||
| Outward II | 0.06 ± 0.01 | 0.03 ± 0.01 | 0.19 ± 0.01 | 0.23 ± 0.01 | ||
| Other | 0.03 ± 0.01 | 0.04 ± 0.01 | 0.08 ± 0.01 | 0.05 ± 0.01 | ||
Radial Distribution Function
Discussion
Conclusions
Supplementary Materials
Acknowledgements
References
- Abbotts, R., & DM, W. (2017). Coordination of DNA single strand break repair. Free Radical Biology and Medicine, 228-244. doi:https://doi.org/10.1016/j.freeradbiomed.2016.11.039.
- Afonin, K., Kasprzak, W., Bindewald, E., Puppala, P., Diehl, A., Hall, K., . . . Shapiro, B. (2021). Computational and experimental characterization of RNA cubic nanoscaffolds. In Therapeutic RNA Nanotechnology (pp. 121-149). New York: Jenny Standford Puhlishing. doi:https://doi.org/10.1201/9781003122005.
- Alberts, B., Johnson, A., Lewis, J., Raff, M., Roberts, K., & Walter, P. (2003). The Structure and Function of DNA. In Molecular Biology of the Cell. New York: Garland Science. doi:https://doi.org/10.1093/aob/mcg023.
- Alhmoud, J., Woolley, J., Moustafa, A.-E., & Mallei, M. (2021). DNA damage/repair management in cancers. In Advances in Medical Biochemistry, Genomics, Physiology, and Pathology (pp. 309-339). New York: Jenny Standford Publishing. doi:https://doi.org/10.1201/9781003180449.
- Andrio, P., Hospital, A., Conejero, J., Jordá, L., Del Pino, M., Codo, L., & Soiland-Reyes, S. (2019). BioExcel Building Blocks, a software library for interoperable biomolecular simulation workflows. Nature scientific data. doi:https://doi.org/10.1038/s41597-019-0177-4.
- Bayly, C., Cieplak, P., Cornell, W., & Kollman, P. (1993). A well-behaved electrostatic potential based method using charge restraints for deriving atomic charges: the RESP model. The Journal of Physical Chemistry, 10269-10280. doi:https://doi.org/10.1021/j100142a004.
- Bekker, H., Berendsen, H., Dijkstra, E., Achterop, S., Vondrumen, R., Vanderspoel, D., . . . Renardus, M. (1993). Gromacs-a parallel computer for molecular-dynamics simulations. 4th international conference on computational physics (pp. 252-256). World Scientific Publishing. https://doi.org/10.1074/jbc.m116.736728.
- Berendsen, H., Grigera, J.-R., & Straatsma, T. (1987). The missing term in effective pair potentials. Journal of Physical Chemistry, 6269-6271.https://doi.org/10.1021/ja963909j.
- Bernstein, C., Prasad, A., Nfonsam, V., & Bernstein, H. (2013). DNA damage, DNA repair and cancer. InTech Rijeka, Croatia. https://doi.org/10.1093/nar/gkad013.
- Bhattacharyya, D., & Bansal, M. (1990). Local variability and base sequence effects in DNA crystal structures. Journal of Biomolecular Structure and Dynamics. doi:https://doi.org/10.1080/07391102.1990.10507828.
- Bilokapic, S., Suskiewicz, M., Ahel, I., & Halic, M. (2020). Bridging of DNA breaks activates PARP2–HPF1 to modify chromatin. Nature, 609-613. doi:https://doi.org/10.1038/s41586-020-2725-7.
- Bishop, T. (2008). Geometry of the nucleosomal DNA superhelix. Biophysical journal. doi:https://doi.org/10.1529/biophysj.107.122853.
- Bowman, J., & Willams, L. D. (2023). Nucleic acids. In Encyclopedia of astrobiology (pp. 2096-2103). Berlin: Springer Berlin Heidelberg. doi:https://doi.org/10.1007/978-3-662-65093-6_1079.
- Brown, I. (1976). On the geometry of O–H⋯ O hydrogen bond. s Acta Crystallographica Section A: Crystal Physics, Diffraction, Theoretical and General Crystallography. doi:https://doi.org/10.1107/S0567739476000041.
- Brown, J., O’Carrigan, B., Jackson, S., & Yap, T. (2017). Targeting DNA repair in cancer: beyond PARP inhibitors. Cancer discovery, 20-37. doi:https://doi.org/10.1158/2159-8290.CD-16-0860.
- Bunta, J., Laaksonen, A., Pinak, M., & Nemoto, T. (2006). DNA strand break: Structural and electrostatic properties studied by molecular dynamics simulation. Computational Biology and Chemistry, 112-119. doi:https://doi.org/10.1016/j.compbiolchem.2005.12.001.
- Bussi, G., Donadio, D., & Parrinello, M. (2007). Canonical sampling through velocity rescaling . The Journal of chemical physics . doi:https://doi.org/10.1063/1.2408420.
- Caldecott, K. (2008). Single-strand break repair and genetic disease. Nature Reviews Genetics, 619-631. doi:https://doi.org/10.1038/nrg2380.
- Caldecott, K. (2022). DNA single-strand break repair and human genetic disease. Trends in cell biology, 733-745. doi:https://doi.org/10.1016/j.tcb.2022.04.010.
- Caldecott, K. (2024). Causes and consequences of DNA single-strand breaks. Trends in Biochemical Sciences. doi:https://doi.org/10.1016/j.tibs.2023.11.001.
- Campo, M. (2010). Structural and dynamic properties of SPC/E water. Papers in physics.
- Cannan, W., Rashid, I., Tomkinson, A., Wallace, S., & Pederson, D. (2017). The human ligase IIIα-XRCC1 protein complex performs DNA nick repair after transient unwrapping of nucleosomal DNA. Journal of Biological Chemistry, 5227-5238. doi:https://doi.org/10.1074/jbc.M116.736728.
- Carusillo, A., & Mussolino, C. (2020). DNA Damage: From Threat to Treatment. Cells, 1665. doi:https://doi.org/10.3390/cells9071665.
- Case, D., Darden, T., Cheatham III, T., Simmerling, C., Wang, J., Duke, R., . . . Kollman, P. (2006). AMBER 9. San Francisco.
- Cherepanov, A., & de Vries, S. (2002). Dynamic mechanism of nick recognition by DNA ligase. European Journal of Biochemistry, 5993-5999. doi:https://doi.org/10.1046/j.1432-1033.2002.03309.x.
- Cieplak, P., Cheatham, T., & Kollman, P. (1997). Cieplak, P., Cheatham, T. E., & Kollman, P. A. (1997). Molecular Dynamics Simulations Find That 3 ‘Phosphoramidate Modified DNA Duplexes Undergo a B to A Transition and Normal DNA Duplexes an A to B Transition. Journal of the American Chemical Society, 6722-6730. doi:https://doi.org/10.1021/ja963909j.
- Clayden, J., Greeves, N., & Warren, S. (2012). Organic chemistry. Oxford University Press.
- Dans, P., Zeida, A., Machado, M., & Pantano, S. (2010). A coarse grained model for atomic-detailed DNA simulations with explicit electrostatics. Journal of chemical theory and computation, 1711-1725. doi:https://doi.org/10.1021/ct900653p.
- Darden, T., York, D., & Pedersen, L. (1993). Particle mesh Ewald: An N⋅ log (N) method for Ewald sums in large systems. The Journal of chemical physics, 10089-10092.
- Dexheimer, T. (2013). DNA repair pathways and mechanisms. DNA repair of cancer stem cells, 19-32. doi:https://doi.org/10.1007/978-94-007-4590-2_2.
- Djuranovic, D., & Hartmann, B. (2004). DNA fine structure and dynamics in crystals and in solution: the impact of BI/BII backbone conformations. Biopolymers: Original Research on Biomolecules. doi: https://doi.org/10.1002/bip.10528.
- Esmaeeli, R., Bauzá, A., & Perez, A. (2023). Structural predictions of protein–DNA binding: MELD-DNA. Nucleic Acids Research, 1625-1636. doi:https://doi.org/10.1093/nar/gkad013.
- Friedberg, E. (2008). A brief history of the DNA repair field. Cell research, 3-7.
- Frisch, M., Trucks, G., Schlegal, H., Scuseria, G., Robb, M., Cheeseman, J., . . . Pople, J. (2004). Gaussian 03, Revision C.02. Gaussian, Inc., Wallingford CT.
- Furse, K., & Corcelli, S. (2010). Molecular dynamics simulations of DNA solvation dynamics. The Journal of Physical Chemistry Letters, 1813-1820. doi:https://doi.org/10.1021/jz100485e.
- Giglia-Mari, G., Zotter, A., & Vermeulen, W. (2011). DNA damage response. Cold Spring Harbor perspectives in biology, a000745. doi:DOI: 10.1101/cshperspect.a000745.
- González, M. (2011). Force fields and molecular dynamics simulations. École thématique de la Société Française de la Neutronique, 169-200. doi:https://doi.org/10.1051/sfn/201112009.
- Heddi, B., Oguey, C., Lavelle, C., Foloppe, N., & Hartmann, B. (2010). Intrinsic flexibility of B-DNA: the experimental TRX scale. Nucleic acids research. doi:https://doi.org/10.1093/nar/gkp962.
- Hess, B., Bekker, H., Berendsen, H., & Fraaije, J. (1997). LINCS: a linear constraint solver for molecular simulations. Journal of computational chemistry 18, 1463-1472. doi:https://doi.org/10.1002/(SICI)1096-987X(199709)18:12<1463::AID-JCC4>3.0.CO;2-H.
- Howes, T., & Tomkinson, A. (2012). DNA Ligase I, the Replicative DNA Ligase. The eukaryotic replisome: A guide to protein structure and function, 327-341. doi:https://doi.org/10.1007/978-94-007-4572-8_17.
- Huang, R., & Zhou, P.-K. (2021). DNA damage repair: historical perspectives, mechanistic pathways and clinical translation for targeted cancer therapy. Nature, 254. doi:https://doi.org/10.1038/s41392-021-00648-7.
- Jackson, S., & Bartek, J. (2009). The DNA-damage response in human biology and disease. Nature 461, 1071-1078. doi:https://doi.org/10.1038/nature08467.
- Jeggo, P. (1998). DNA Breakage and Repair. In Advances in Genetics 38 (pp. 185-218). Academic Press. doi:https://doi.org/10.1016/S0065-2660(08)60144-3.
- Kagami, L., Wilter, A., Diaz, A., & Vranken, W. (2023). The ACPYPE web server for small-molecule MD topology generation. Bioinformatics. doi:https://doi.org/10.1093/bioinformatics/btad350.
- Karagiannias, T., & El-Osta, A. (2004). DNA damage repair and transcription: Double-strand breaks: signaling pathways and repair mechanisms. Cellular and Molecular Life Sciences CMLS, 2137-2147. doi:https://doi.org/10.1007/s00018-004-4174-0.
- Karakaidos, P., Karagiannis, D., & Rampias, T. (2020). Resolving DNA damage: epigenetic regulation of DNA repair. Molecules 25(11), 2496. doi:https://doi.org/10.3390/molecules25112496.
- Krupa, P., Wales, D., & Sieradzan, A. (2018). Computational Studies of the Mechanical Stability for Single-Strand Break DNA. The Journal of Physical Chemistry B, 8166-8173. doi:https://doi.org/10.1021/acs.jpcb.8b05417.
- Landau, L., Lifshitz, E., & Reichl, L. (1981). Statistical Physics, Part 1 (Third Edition). Physics today. doi:https://doi.org/10.1063/1.2889978.
- Langelier, M.-F., Riccio, A., & Pascal, J. (2014). PARP-2 and PARP-3 are selectively activated by 5′ phosphorylated DNA breaks through an allosteric regulatory mechanism shared with PARP-1. Nucleic acids research, 7762-7775. doi:https://doi.org/10.1093/nar/gku474.
- Lankas, F., Langowski, J., Šponer, J., & Cheatham, T. (2003). DNA basepair step deformability inferred from molecular dynamics simulations. Biophysical journal. doi:https://doi.org/10.1016/S0006-3495(03)74710-9.
- Lans, H., Marteijn, J., & Vermeulen, W. (2012). ATP-dependent chromatin remodeling in the DNA-damage response. Epigenetics & Chromatin 5, 4. doi: https://doi.org/10.1186/1756-8935-5-4.
- Laughton, C., & Harris, S. (2011). The atomistic simulation of DNA. Wiley Interdisciplinary Reviews: Computational Molecular Science, 590-600. doi:https://doi.org/10.1002/wcms.46.
- Lavery, R., Moakher, M., Maddocks, J., Petkeviciute, D., & Zakrzewska, K. (2009). Conformational analysis of nucleic acids revisited: Curves+. Nucleic acids research. doi:https://doi.org/10.1093/nar/gkp608.
- Lindahl, T. (1993). Instability and decay of the primary structure of DNA. Nature 362, 709-715. doi:https://doi.org/10.1038/362709a0.
- Lindahl, T., & Nyberg, B. (1972). Rate of depurination of native deoxyribonucleic acid. In Biochemistry (pp. 3610-3618).
- Lukas, J., Lukas, C., & Bartek, J. (2011). More than just a focus: The chromatin response to DNA damage and its role in genome integrity maintenance. Nature cell biology, 1161-1169. doi:https://doi.org/10.1038/ncb2344.
- Lukin, M., & de Los Santos, C. (2006). NMR Structures of Damaged DNA. Chemical Reviews, 607-686. doi:https://doi.org/10.1021/cr0404646.
- Mackerell Jr., A. (2004). Empirical force fields for biological macromolecules: overview and issues. Journal of computational chemistry, 1584-1604. doi:https://doi.org/10.1002/jcc.20082.
- Maginn, E., & Elliott, J. (2010). Historical perspective and current outlook for molecular dynamics as a chemical engineering tool. Industrial & engineering chemistry research, 3059-3078. doi:https://doi.org/10.1021/ie901898k.
- Mark, P., & Nilsson, L. (2011). Structure and Dynamics of the TIP3P, SPC, and SPC/E Water Models at 298 K. The Journal of Physical Chemistry A, 9954-9960. doi:https://doi.org/10.1021/jp003020w.
- McKinnon, P., & Caldecott, K. (2007). DNA Strand Break Repair and Human Genetic Disease. Annu. Rev. Genomics Hum. Genet. 8, no. 1 , 37-55. doi:https://doi.org/10.1146/annurev.genom.7.080505.115648.
- Menoni, H., Mascio, P., Cadet, J., Dimitrov, S., & Angelov, D. (2017). Chromatin associated mechanisms in base excision repair - nucleosome remodeling and DNA transcription, two key players. Free Radical Biology and Medicine 107, 159-169. doi:https://doi.org/10.1016/j.freeradbiomed.2016.12.026.
- Michaud-Agrawal, N., Denning, E., Woolf, T., & Beckstein, O. (2011). MDAnalysis: a toolkit for the analysis of molecular dynamics simulations. Journal of computational chemistry. doi: https://doi.org/10.1002/jcc.21787.
- Milano, L., Gautam, A., & Caldecott, K. (2024). DNA damage and transcription stress. Molecular Cell, 70-79. doi:https://doi.org/10.1016/j.molcel.2023.11.014.
- Minchin, S., & Lodge, J. (2019). Understanding biochemistry: structure and function of nucleic acids. Essays in biochemistry 63(4), 433-456. doi:https://doi.org/10.1042/EBC20180038.
- Misra, M., & Yadav, A. (2022). Assessment of Available AMBER Force Fields to Model DNA-Ligand Interactions. Biointerface Res. Appl. Chem. 13, 156. doi:https://doi.org/10.33263/BRIAC132.156.
- Molinaro, C., Martoriati, A., & Cailliau, K. (2021). Proteins from the DNA damage response: Regulation, dysfunction, and anticancer strategies. Cancers, 3819. doi:https://doi.org/10.3390/cancers13153819.
- Nair, N., Shoaib, M., & Sørensen, C. (2017). Chromatin Dynamics in Genome Stability: Roles in Suppressing Endogenous DNA Damage and Facilitating DNA Repair. International Journal of Molecular Sciences, 1486. doi:https://doi.org/10.3390/ijms18071486.
- Nosé, S., & Klein, M. (1983). Constant pressure molecular dynamics for molecular systems. Molecular Physics, 1055-1076. doi:https://doi.org/10.1080/00268978300102851.
- O’Connor, M. (2015). Targeting the DNA damage response in cancer. Molecular cell, 547-560. doi:https://doi.org/10.1016/j.molcel.2015.10.040.
- Obaji, E., Haikarainen, T., & Lehtiö, L. (2018). Structural basis for DNA break recognition by ARTD2/PARP2. Nucleic acids research, 12154-12165. doi:https://doi.org/10.1093/nar/gky927.
- Obaji, E., Maksimainen, M., Galera-Prat, A., & Lehtiö, L. (2021). Activation of PARP2/ARTD2 by DNA damage induces conformational changes relieving enzyme autoinhibition. Nature communications, 3479. doi:https://doi.org/10.1038/s41467-021-23800-x.
- Ogawa, T., & Okazaki, T. (1980). Discontinuous DNA replication. Annual review of biochemistry, 421-457. doi:https://doi.org/10.1146/annurev.bi.49.070180.002225.
- Ohyama, T., Vladimir N., P., & Sinden, R. (2005). DNA: alternative conformations and biology. DNA conformation and transcription, 3-17.
- Parrinello, M., & Aneesur, R. (1981, 7182-7190). Polymorphic transitions in single crystals: A new molecular dynamics method. Journal of Applied physics 52. doi:https://doi.org/10.1063/1.328693.
- Pérez, A., Luque, F., & Orozco, M. (2012). Frontiers in molecular dynamics simulations of DNA. Accounts of chemical research, 196-205. doi:https://doi.org/10.1021/ar2001217.
- Petrová, P., Koča, J., & Imberty, A. (1999). Potential Energy Hypersurfaces of Nucleotide Sugars: Ab Initio Calculations, Force-Field Parametrization, and Exploration of the Flexibility. Journal of the American Chemical Society, 5535-5547. doi:https://doi.org/10.1021/ja983854g.
- Prasad, R., Beard, W., & Wilson, S. (1994). Studies of gapped DNA substrate binding by mammalian DNA polymerase beta. Dependence on 5’-phosphate group. Journal of Biological Chemistry, 18096-18101. doi:https://doi.org/10.1016/S0021-9258(17)32422-5.
- Rohs, R., Jin, X., West, S., Joshi, R., Honig, B., & Mann, R. (2010). Origins of specificity in protein-DNA recognition. Annual review of biochemistry 79, 233-269. doi:https://doi.org/10.1146/annurev-biochem-060408-091030.
- Saenger, W. (2013). Principles of nucleic acid strucure. Springer Science & Business Media.
- Sallmyr, A., Rashid, I., Bhandari, S., Naila, T., & Tomkinson, A. (2020). Human DNA ligases in replication and repair. DNA repair, 102908. doi:https://doi.org/10.1016/j.dnarep.2020.102908.
- Salo-Ahen, O., Alanko, I., Bhadane, R., Bonvin, A., Honorato, R., Hossain, S., . . . Larsen, A. (2020). Molecular dynamics simulations in drug discovery and pharmaceutical development. Processes, 71. doi:https://doi.org/10.3390/pr9010071.
- Salomon-Ferrer, R., Case, D., & Walker, R. (2013). An overview of the Amber biomolecular simulation package. Computational Molecular Science. doi: https://doi.org/10.1002/wcms.1121.
- Samanta, S., Mukherjee, S., Chakrabarti, J., & Bhattacharyya, D. (2009). Structural properties of polymeric DNA from molecular dynamics simulations. The Journal of chemical physics. doi:https://doi.org/10.1063/1.3078797.
- Schrödinger, L., & DeLano, W. (2020). The PyMOL Molecular Graphics System.
- Sirbu, B., & Cortez, D. (2013). DNA Damage Response: Three Levels of DNA Repair Regulation. Cold Spring Harbor perspectives in biology, 5(8), a012724.
- Song, H.-Y., Shen, R., Mahasin, H., Guo, Y.-N., & Wang, D.-G. (2023). DNA replication: Mechanisms and therapeutic interventions for diseases. MedComm, e210. doi: https://doi.org/10.1002/mco2.210.
- Sousa da Silva, A., & Vranken, W. (2012). ACPYPE - AnteChamber PYthon Parser interfacE. BMC research notes, 1-8. doi:https://doi.org/10.1186/1756-0500-5-367.
- Spenser, M. (1959). The stereochemistry of deoxyribonucleic acid. II. Hydrogen-bonded pairs of bases. Acta Crystallographica. doi:https://doi.org/10.1107/S0365110X59000160.
- Steinbrecher, T., Latzer, J., & Case, D. (2012). Revised AMBER Parameters for Bioorganic Phosphates. Journal of Chemical Theory and Computation, 4405-4412. doi:https://doi.org/10.1021/ct300613v.
- Taylor, R., Kennard, O., & Versichel, W. (1980). The geometry of the N–H⋯ O= C hydrogen bond. 3. Hydrogen-bond distances and angles. Acta Crystallographica Section B: Structural Science 40. doi:https://doi.org/10.1107/S010876818400210X.
- Teletchéa, S., Hartmann, B., & Kozelka, J. (2004). Discrimination between BI and BII conformational substates of B-DNA based on sugar-base interproton distances. Journal of Biomolecular Structure and Dynamics. doi:https://doi.org/10.1080/07391102.2004.10506942.
- Thaplyal, P., & Bevilacqua, P. (2014). Experimental approaches for measuring pKa’s in RNA and DNA. Methods in enzymology, 189-219. doi:https://doi.org/10.1016/B978-0-12-801122-5.00009-X.
- Timson, D., Singleton, M., & Wigley, D. (2000). DNA ligases in the repair and replication of DNA. Mutation Research/DNA Repair, 301-318. doi:https://doi.org/10.1016/S0921-8777(00)00033-1.
- Toukan, K., & Rahman, A. (1985). Molecular-dynamics study of atomic motions in water. Physical Review B, 2643-2648. doi:https://org/doi/10.1103/PhysRevB.31.2643.
- Tucker, M., Piana, S., Tan, D., LeVine, M., & Shaw, D. (2022). Development of Force Field Parameters for the Simulation of Single- and Double-Stranded DNA Molecules and DNA–Protein Complexes. The Journal of Physical Chemistry B, 4442-4457. doi:https://doi.org/10.1021/acs.jpcb.1c10971.
- Urrutia, K., Xu, W., & Zhao, L. (2022). The 5′-phosphate enhances the DNA-binding and exonuclease activities of human mitochondrial genome maintenance exonuclease 1 (MGME1). Journal of Biological Chemistry. doi:https://doi.org/10.1016/j.jbc.2022.102306.
- Waters, K., & Spratt, D. (2024). New Discoveries on Protein Recruitment and Regulation during the Early Stages of the DNA Damage Response Pathways. International Journal of Molecular Sciences, 1676. doi:https://doi.org/10.3390/ijms25031676.
- Watson, J., & Crick, F. (1953). Molecular structure of nucleic acids: a structure for deoxyribose nucleic acid. Nature 171(4356), 737/738. doi:https://doi.org/10.1038/171737a0.
- Wolffe, A. (1998). Chromatin: structure and function. Academic Press.
- Yamaguchi, H., Siebers, J.-G., Furukawa, A., Otagiri, N., & Osman, R. (2002). Molecular dynamics simulation of a DNA containing a single strand break. Radiation protection dosimetry, 103-108. doi:https://doi.org/10.1093/oxfordjournals.rpd.a006737.
- Yoo, J., Winogradoff, D., & Aksimentiev, A. (2020). Molecular dynamics simulations of DNA–DNA and DNA–protein interactions. Current Opinion in Structural Biology, 88-96. doi:https://doi.org/10.1016/j.sbi.2020.06.007.
- Zenkova, M. (2004). Artificial nucleases (Vol. 13). Springer Science & Business Media.
- Zgarbová, M., Otyepka, M., Šponer, J., Mládek, A., Banáš, P., Cheatham, T., & Jurečka, P. (2011). Refinement of the Cornell et al. Nucleic Acids Force Field Based on Reference Quantum Chemical Calculations of Glycosidic Torsion Profiles. (2902, Ed.) Journal of Chemical Theory and Computation, 2886. doi:https://doi.org/10.1021/ct200162x.
- Zhang, F., & Collins, M. (1995). Model simulations of DNA dynamics. Physical Review E, 4217. doi:https://doi.org/10.1103/PhysRevE.52.4217.
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




