Submitted:
25 October 2024
Posted:
25 October 2024
Read the latest preprint version here
Abstract
Keywords:
1. Introduction
2. Literature Review
3. Data and Methods
3.1. Data Introduction
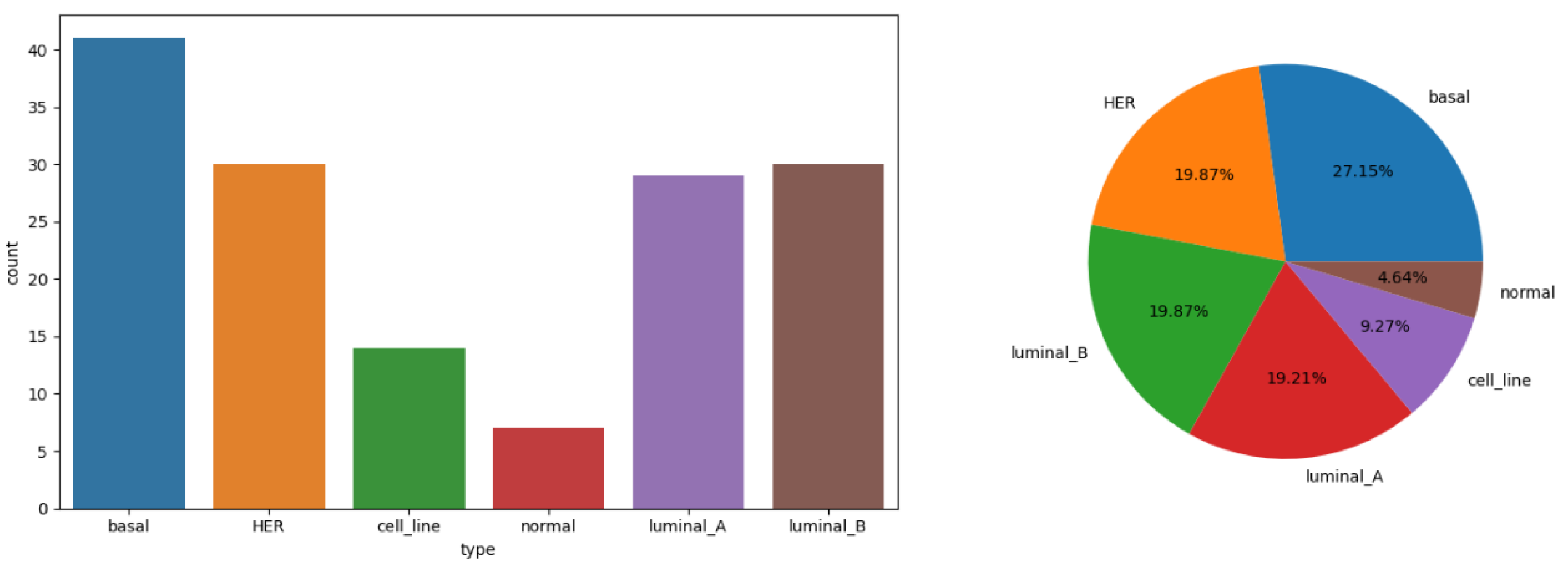
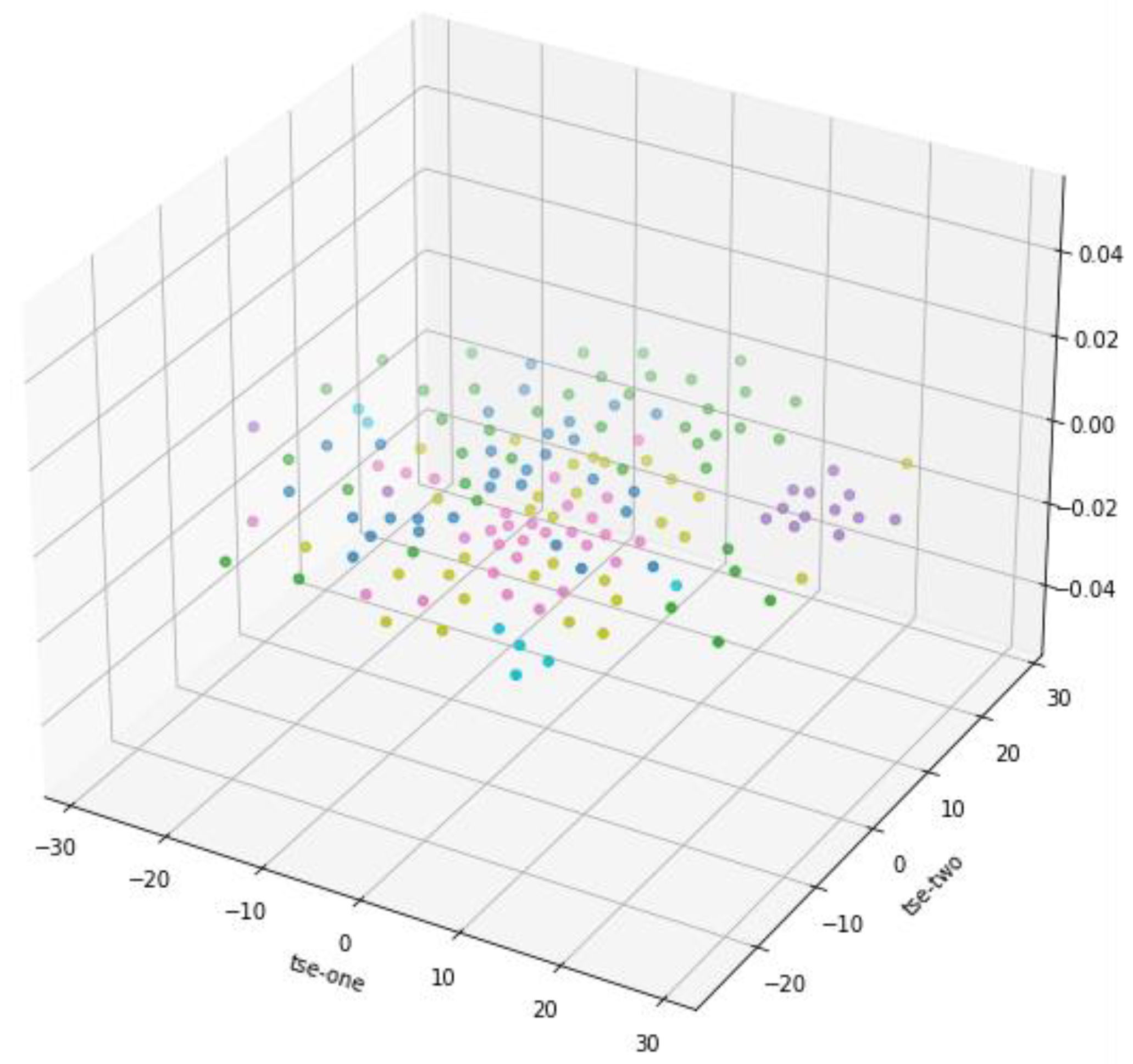
3.2. T-Distributed Stochastic Neighbor Embedding
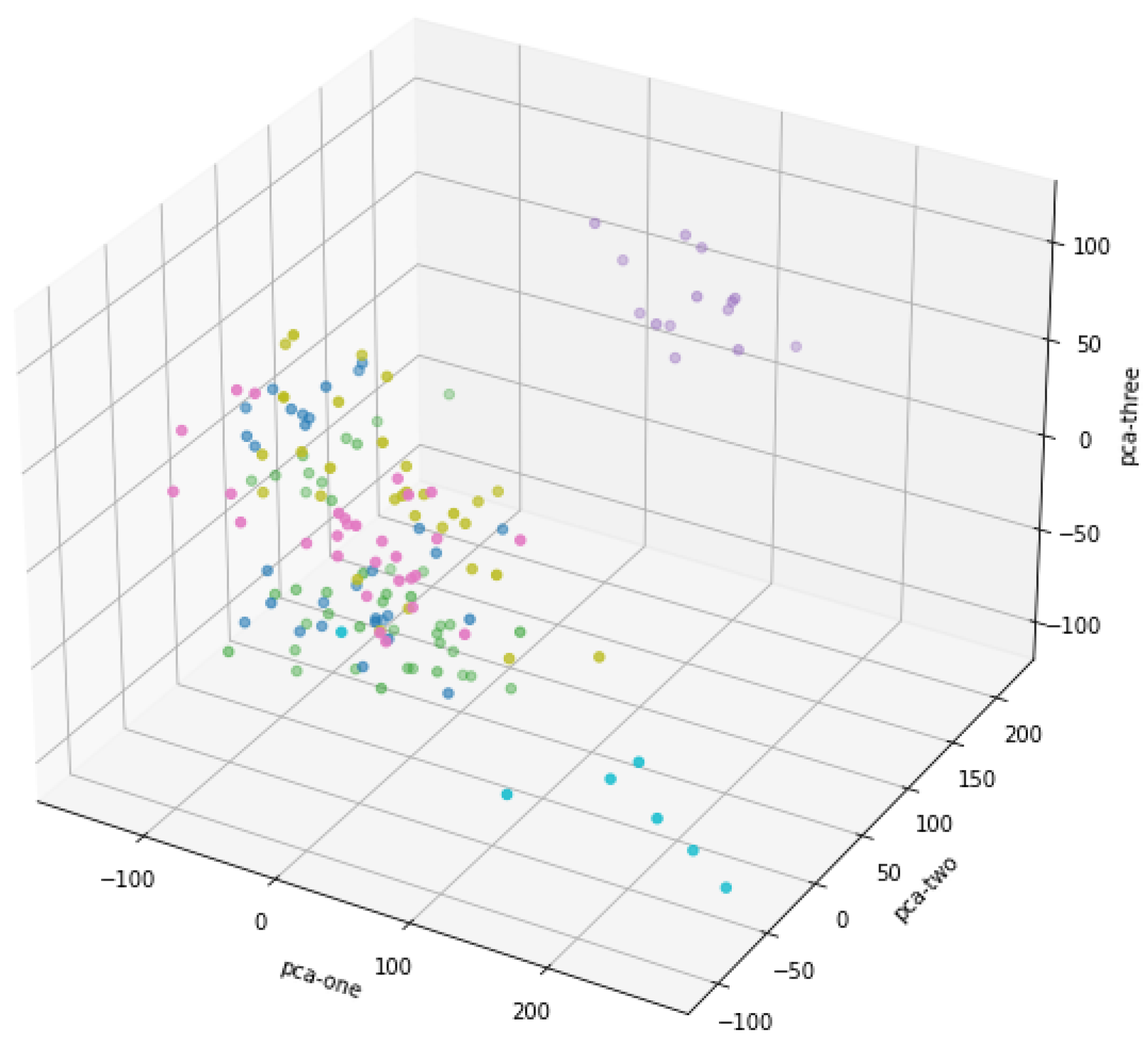
3.3. Principal Component Analysis
3.4. Extreme Gradient Boosting
4. Model Analysis
5. Conclusions
References
- Zelli, V.; Manno, A.; Compagnoni, C. , et al. Classification of tumor types using XGBoost machine learning model: a vector space transformation of genomic alterations. Journal of Translational Medicine 2023, 21, 836. [Google Scholar] [CrossRef] [PubMed]
- Hoque, R.; Das, S.; Hoque, M. , et al. Breast Cancer Classification using XGBoost. World Journal of Advanced Research and Reviews 2024, 21, 1985–1994. [Google Scholar] [CrossRef]
- Song, Y.; Westerhuis, J.A.; Aben, N. , et al. Principal component analysis of binary genomics data. Briefings in bioinformatics 2019, 20, 317–329. [Google Scholar] [CrossRef] [PubMed]
- Laghmati, S.; Hamida, S.; Hicham, K. , et al. An improved breast cancer disease prediction system using ML and PCA. Multimedia Tools and Applications 2024, 83, 33785–33821. [Google Scholar] [CrossRef]
- Sharma, A.; Mishra, P.K. Performance analysis of machine learning based optimized feature selection approaches for breast cancer diagnosis. International Journal of Information Technology 2022, 14, 1949–1960. [Google Scholar] [CrossRef]
- Nguyen, Q.H.; Do, T.T.T.; Wang, Y. , et al. In Breast cancer prediction using feature selection and ensemble voting. In Proceedings of the 2019 International Conference on System Science and Engineering (ICSSE). IEEE; 2019; pp. 250–254. [Google Scholar]
- Song, X.; Zhu, J.; Tan, X. , et al. XGBoost-based feature learning method for mining COVID-19 novel diagnostic markers. Frontiers in Public Health 2022, 10, 926069. [Google Scholar] [CrossRef] [PubMed]
- Liew, X.Y.; Hameed, N.; Clos, J. An investigation of XGBoost-based algorithm for breast cancer classification. Machine Learning with Applications 2021, 6, 100154. [Google Scholar] [CrossRef]
- Meng, C.; Zeleznik, O.A.; Thallinger, G.G. , et al. Dimension reduction techniques for the integrative analysis of multi-omics data. Briefings in bioinformatics 2016, 17, 628–641. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.; Liu, C.C.; Li, W. , et al. Discovery of multi-dimensional modules by integrative analysis of cancer genomic data. Nucleic acids research 2012, 40, 9379–9391. [Google Scholar] [CrossRef] [PubMed]
- Chiu PK, F.; Shen, X.; Wang, G. , et al. Enhancement of prostate cancer diagnosis by machine learning techniques: an algorithm development and validation study. Prostate cancer and prostatic diseases 2022, 25, 672–676. [Google Scholar] [CrossRef] [PubMed]
| Precision | Recall | f1-Score | Support | |
|---|---|---|---|---|
| 0 | 0.50 | 0.67 | 0.57 | 9 |
| 1 | 0.93 | 0.78 | 0.85 | 18 |
| 2 | 0.83 | 1.00 | 0.91 | 5 |
| 3 | 1.00 | 0.77 | 0.87 | 13 |
| 4 | 1.00 | 0.50 | 0.67 | 14 |
| 5 | 1.00 | 1.00 | 1.00 | 2 |
| micro avg | 0.85 | 0.72 | 0.78 | 61 |
| macro avg | 0.88 | 0.79 | 0.81 | 61 |
| weighted avg | 0.89 | 0.72 | 0.78 | 61 |
| samples avg | 0.69 | 0.72 | 0.70 | 61 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




