Submitted:
07 October 2024
Posted:
08 October 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Sample Details
2.2. DNA Extraction, MSI Marker Panels, MIP Amplification and SEQUENCING
2.3. Sequence Analysis and MSI Classification
2.4. Comparison of Tumour Cell Content with MSI Classification
2.5. Statistical Analyses
3. Results
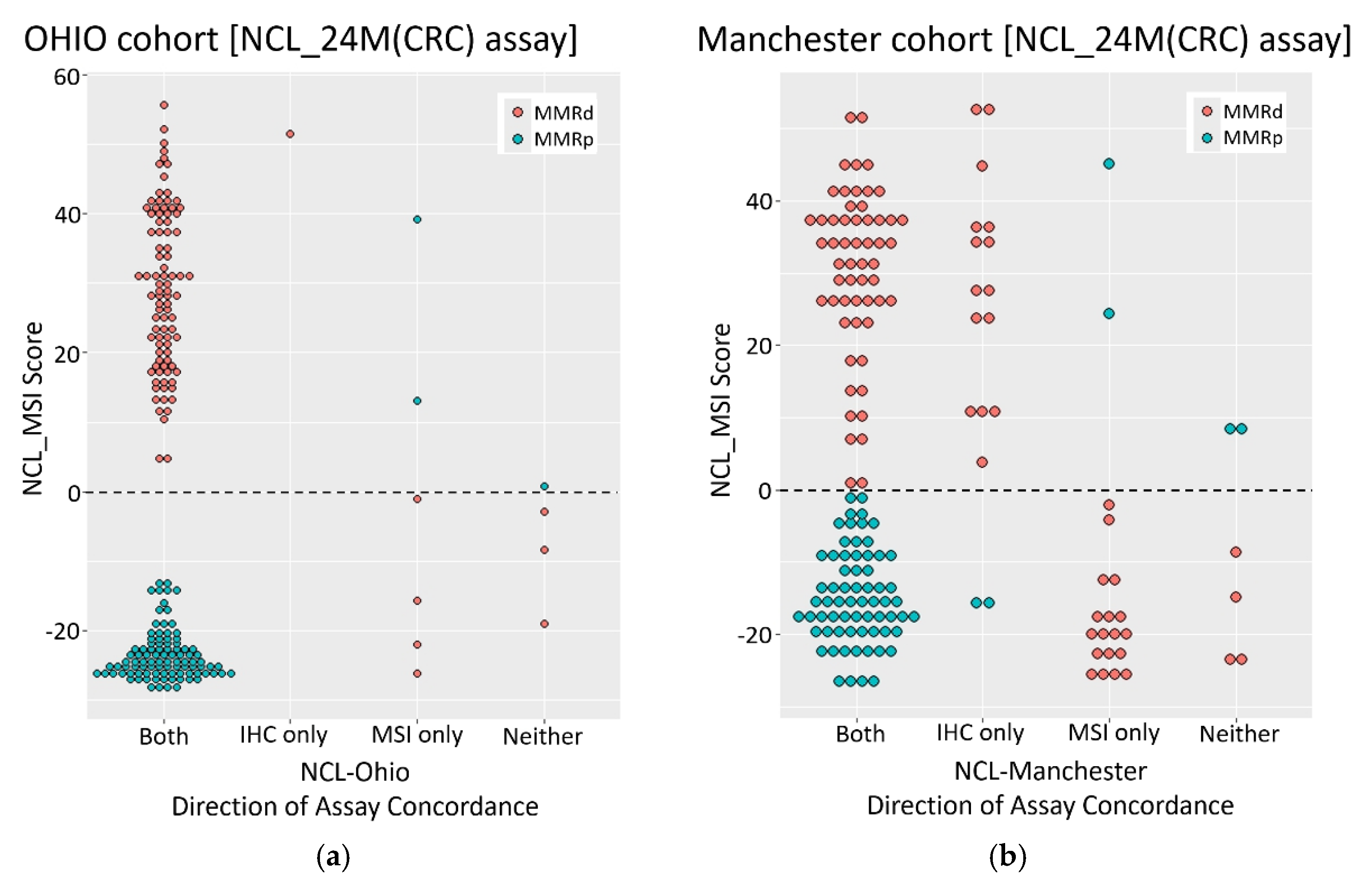
3.1. NCL_MSI Is Highly Concordant with Ohio Assay Results
3.2. NCL_MSI Increases Assay Concordance and Sensitivity in the Manchester Cohort
3.3. Increasing Marker Number and MSI Classifier Retraining Does Not Improve Assay Concordance
3.4. Improved Concordance Predominantly Affects IHC MMRd and Promega MSS Classifications
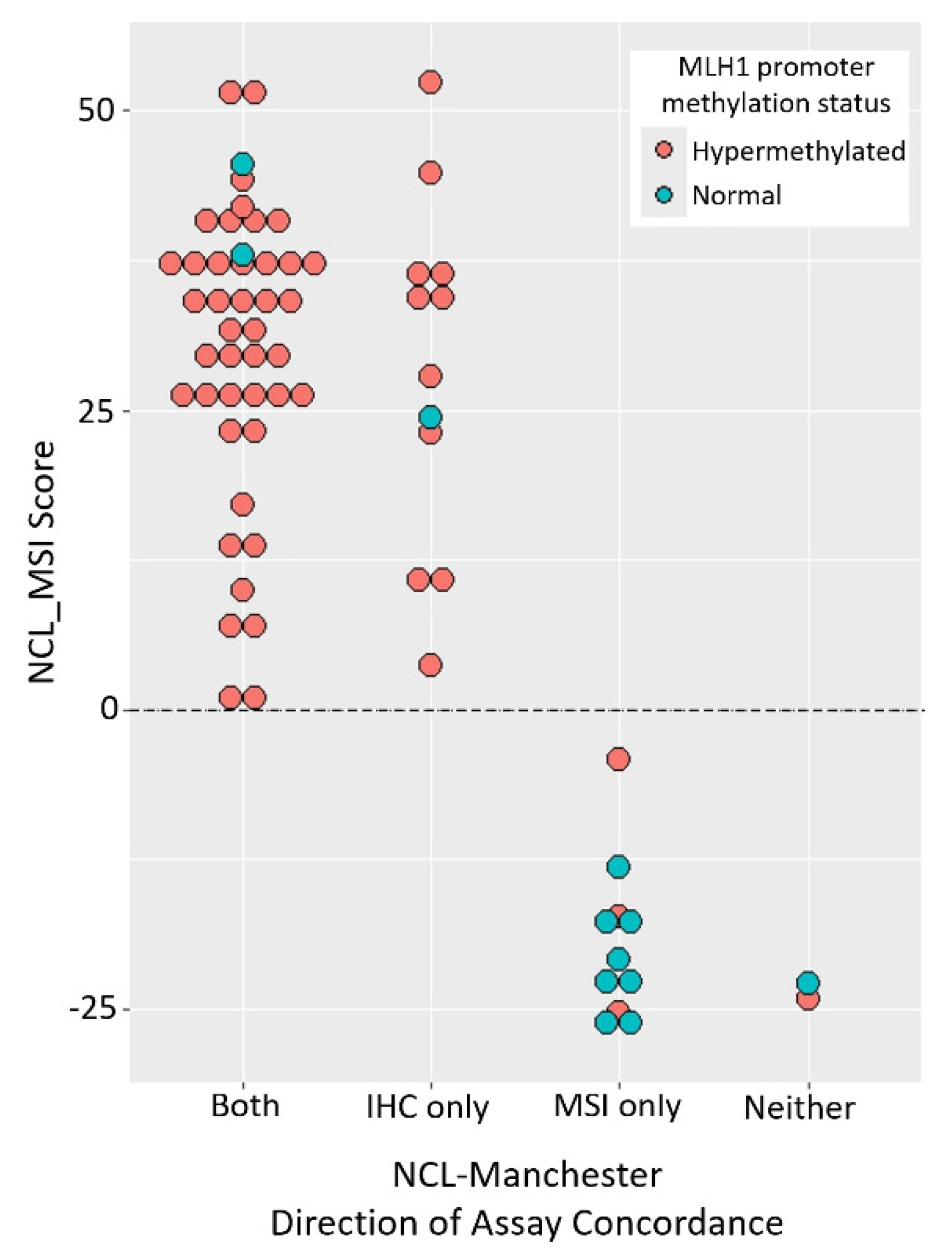
3.5. Manchester Discordance Suggests Promega MSI False Negatives and IHC False Positives
3.6. Sample Tumour Cell Content Differs between Cohorts and Impacts Upon MSI Classification
3.7. Germline Cases Are Not Identified Efficiently by Either MSI Assay in the Manchester Cohort
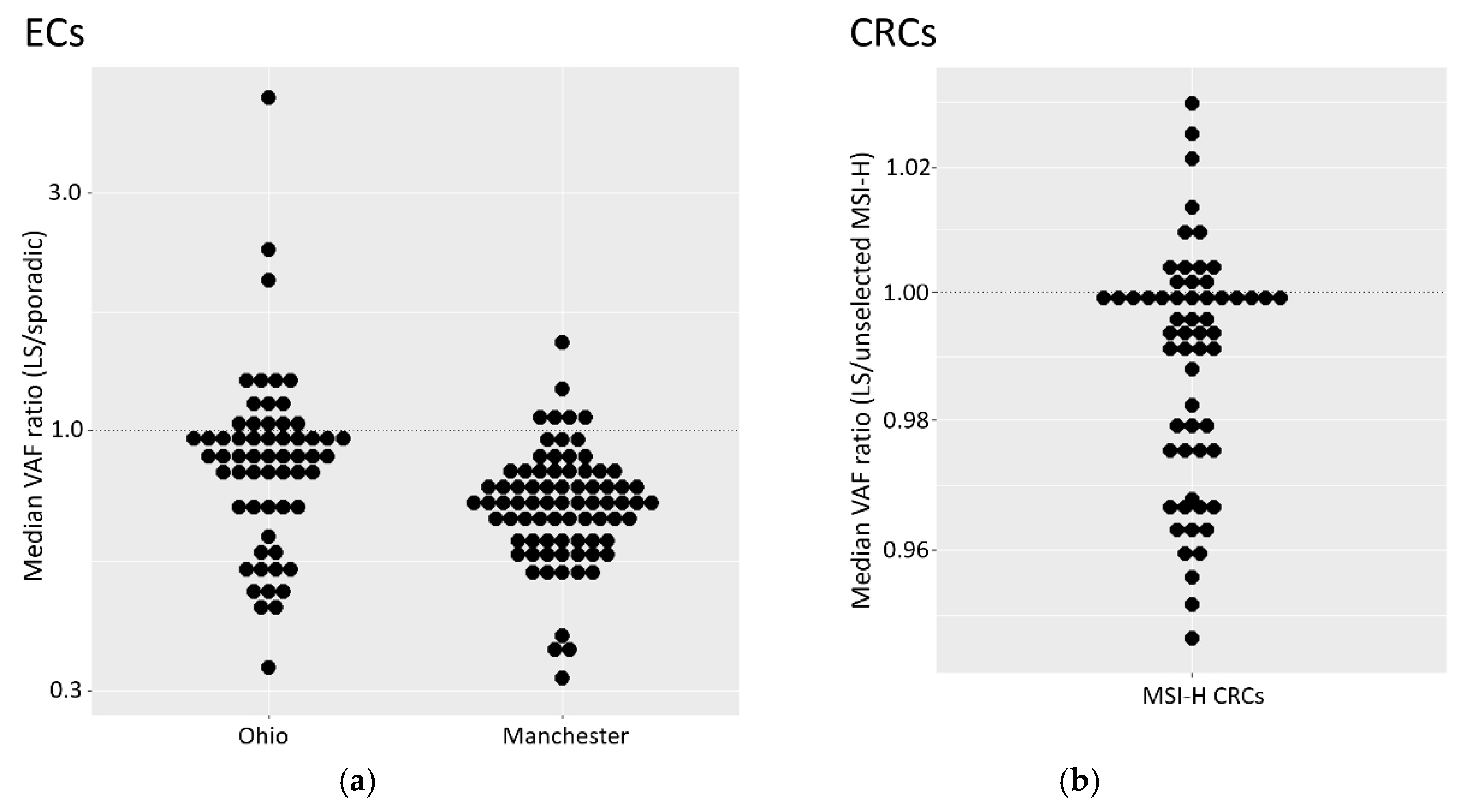
3.8. Variant Allele Frequencies Are Lower in Tumours from MMR Germline Defect Carriers
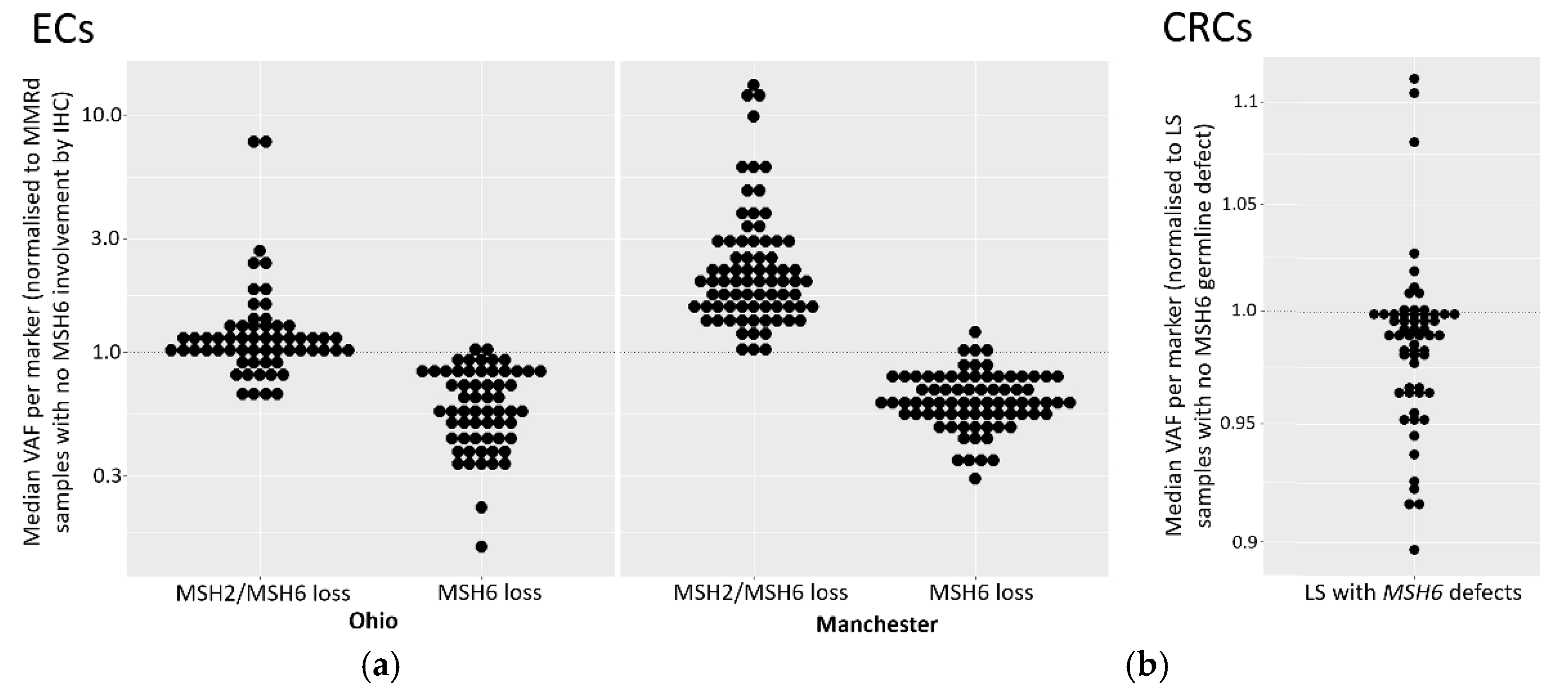
3.9. Variant Allele Frequency is Reduced in ECs with Isolated Loss of MSH6 Expression
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Le DT, Durham JN, Smith KN, Wang H, Bartlett BR, Aulakh LK, et al. Mismatch repair deficiency predicts response of solid tumors to PD-1 blockade. Science. 2017;357(6349):409-13. [CrossRef] [PubMed]
- Andre T, Shiu KK, Kim TW, Jensen BV, Jensen LH, Punt C, et al. Pembrolizumab in Microsatellite-Instability-High Advanced Colorectal Cancer. N Engl J Med. 2020;383(23):2207-18. [CrossRef] [PubMed]
- Burn J, Sheth H, Elliott F, Reed L, Macrae F, Mecklin JP, et al. Cancer prevention with aspirin in hereditary colorectal cancer (Lynch syndrome), 10-year follow-up and registry-based 20-year data in the CAPP2 study: a double-blind, randomised, placebo-controlled trial. Lancet. 2020;395(10240):1855-63. [CrossRef] [PubMed]
- Monahan KJR, N.; Monje-Garcia, L.; Armstrong, R.; Church, D. N.; Cook, J.; Elghobashy, A.; Lalloo, F.; Lane, S.; Frank D McDermott, F. D.; Miles, T.; Hardy, S. A.; Tyson, A.; Ya Wen Wang, V.; Kim, A. ; Gelinas, S; Faravelli, F.; Elmslie, F.; Shaw, A. C. The English National Lynch Syndrome transformation project: an NHS Genomic Medicine Service Alliance (GMSA) programme. BMJ Oncology. 2023;2:e000124. [CrossRef]
- Lynch HT, Lynch PM, Lanspa SJ, Snyder CL, Lynch JF, Boland CR. Review of the Lynch syndrome: history, molecular genetics, screening, differential diagnosis, and medicolegal ramifications. Clin Genet. 2009;76(1):1-18. [CrossRef] [PubMed]
- Duraturo F, Liccardo R, De Rosa M, Izzo P. Genetics, diagnosis and treatment of Lynch syndrome: Old lessons and current challenges. Oncol Lett. 2019;17(3):3048-54. [CrossRef] [PubMed]
- Battaglin F, Naseem M, Lenz HJ, Salem ME. Microsatellite instability in colorectal cancer: overview of its clinical significance and novel perspectives. Clin Adv Hematol Oncol. 2018;16(11):735-45. PubMed PubMed Central PMCID: PMCPMC7493692. [PubMed]
- Latham A, Srinivasan P, Kemel Y, Shia J, Bandlamudi C, Mandelker D, et al. Microsatellite Instability Is Associated With the Presence of Lynch Syndrome Pan-Cancer. J Clin Oncol. 2019;37(4):286-95. Epub 20181030. [CrossRef] [PubMed]
- Dominguez-Valentin M, Haupt S, Seppala TT, Sampson JR, Sunde L, Bernstein I, et al. Mortality by age, gene and gender in carriers of pathogenic mismatch repair gene variants receiving surveillance for early cancer diagnosis and treatment: a report from the prospective Lynch syndrome database. EClinicalMedicine. 2023;58:101909. Epub 20230320. [CrossRef] [PubMed]
- Moller P, Seppala TT, Ahadova A, Crosbie EJ, Holinski-Feder E, Scott R, et al. Dominantly inherited micro-satellite instable cancer - the four Lynch syndromes - an EHTG, PLSD position statement. Hered Cancer Clin Pract. 2023;21(1):19. Epub 20231011. [CrossRef] [PubMed]
- Gallon R, Gawthorpe P, Phelps RL, Hayes C, Borthwick GM, Santibanez-Koref M, et al. How Should We Test for Lynch Syndrome? A Review of Current Guidelines and Future Strategies. Cancers (Basel). 2021;13(3):406. [CrossRef] [PubMed]
- Shia, J. Immunohistochemistry versus microsatellite instability testing for screening colorectal cancer patients at risk for hereditary nonpolyposis colorectal cancer syndrome. Part I. The utility of immunohistochemistry. J Mol Diagn. 2008;10(4):293-300. Epub 20080613. [CrossRef] [PubMed]
- Zhang, L. Immunohistochemistry versus microsatellite instability testing for screening colorectal cancer patients at risk for hereditary nonpolyposis colorectal cancer syndrome. Part II. The utility of microsatellite instability testing. J Mol Diagn. 2008;10(4):301-7. Epub 20080613. [CrossRef] [PubMed]
- De’ Angelis GL, Bottarelli L, Azzoni C, De’ Angelis N, Leandro G, Di Mario F, et al. Microsatellite instability in colorectal cancer. Acta Biomed. 2018;89(9-S):97-101. [CrossRef] [PubMed]
- Berg KD, Glaser CL, Thompson RE, Hamilton SR, Griffin CA, Eshleman JR. Detection of microsatellite instability by fluorescence multiplex polymerase chain reaction. J Mol Diagn. 2000;2(1):20-8. [CrossRef] [PubMed]
- Ukkola I, Nummela P, Pasanen A, Kero M, Lepisto A, Kytola S, et al. Detection of microsatellite instability with Idylla MSI assay in colorectal and endometrial cancer. Virchows Arch. 2021;479(3):471-9. Epub 20210323. [CrossRef] [PubMed]
- Renault V, Tubacher E, How-Kit A. Assessment of Microsatellite Instability from Next-Generation Sequencing Data. Adv Exp Med Biol. 2022;1361:75-100. [CrossRef] [PubMed]
- Vanderwalde A, Spetzler D, Xiao N, Gatalica Z, Marshall J. Microsatellite instability status determined by next-generation sequencing and compared with PD-L1 and tumor mutational burden in 11,348 patients. Cancer Med. 2018;7(3):746-56. Epub 20180213. [CrossRef] [PubMed]
- Hechtman JF, Rana S, Middha S, Stadler ZK, Latham A, Benayed R, et al. Retained mismatch repair protein expression occurs in approximately 6% of microsatellite instability-high cancers and is associated with missense mutations in mismatch repair genes. Mod Pathol. 2020;33(5):871-9. Epub 20191219. [CrossRef] [PubMed]
- Snowsill T, Huxley N, Hoyle M, Jones-Hughes T, Coelho H, Cooper C, et al. A systematic review and economic evaluation of diagnostic strategies for Lynch syndrome. Health Technol Assess. 2014;18(58):1-406. [CrossRef] [PubMed]
- Snowsill TM, Ryan NAJ, Crosbie EJ. Cost-Effectiveness of the Manchester Approach to Identifying Lynch Syndrome in Women with Endometrial Cancer. Journal of Clinical Medicine. 2020;9(6). [CrossRef]
- Bartley AN, Mills AM, Konnick E, Overman M, Ventura CB, Souter L, et al. Mismatch Repair and Microsatellite Instability Testing for Immune Checkpoint Inhibitor Therapy: Guideline From the College of American Pathologists in Collaboration With the Association for Molecular Pathology and Fight Colorectal Cancer. Arch Pathol Lab Med. 2022;146(10):1194-210. [CrossRef] [PubMed]
- Luchini C, Bibeau F, Ligtenberg MJL, Singh N, Nottegar A, Bosse T, et al. ESMO recommendations on microsatellite instability testing for immunotherapy in cancer, and its relationship with PD-1/PD-L1 expression and tumour mutational burden: a systematic review-based approach. Ann Oncol. 2019;30(8):1232-43. [CrossRef] [PubMed]
- Guyot D’Asnieres De Salins A, Tachon G, Cohen R, Karayan-Tapon L, Junca A, Frouin E, et al. Discordance between immunochemistry of mismatch repair proteins and molecular testing of microsatellite instability in colorectal cancer. ESMO Open. 2021;6(3):100120. Epub 20210428. [CrossRef] [PubMed]
- Hampel H, Pearlman R, Beightol M, Zhao W, Jones D, Frankel WL, et al. Assessment of Tumor Sequencing as a Replacement for Lynch Syndrome Screening and Current Molecular Tests for Patients With Colorectal Cancer. JAMA Oncol. 2018;4(6):806-13. [CrossRef] [PubMed]
- Crosbie EJ, Ryan NAJ, Arends MJ, Bosse T, Burn J, Cornes JM, et al. The Manchester International Consensus Group recommendations for the management of gynecological cancers in Lynch syndrome. Genet Med. 2019;21(10):2390-400. [CrossRef] [PubMed]
- Berends MJ, Wu Y, Sijmons RH, van der Sluis T, Ek WB, Ligtenberg MJ, et al. Toward new strategies to select young endometrial cancer patients for mismatch repair gene mutation analysis. J Clin Oncol. 2003;21(23):4364-70. [CrossRef] [PubMed]
- Lu KH, Schorge JO, Rodabaugh KJ, Daniels MS, Sun CC, Soliman PT, et al. Prospective determination of prevalence of lynch syndrome in young women with endometrial cancer. J Clin Oncol. 2007;25(33):5158-64. Epub 20071009. [CrossRef] [PubMed]
- Rubio I, Ibanez-Feijoo E, Andres L, Aguirre E, Balmana J, Blay P, et al. Analysis of Lynch Syndrome Mismatch Repair Genes in Women with Endometrial Cancer. Oncology. 2016;91(3):171-6. Epub 20160712. [CrossRef] [PubMed]
- Wang Y, Shi C, Eisenberg R, Vnencak-Jones CL. Differences in Microsatellite Instability Profiles between Endometrioid and Colorectal Cancers: A Potential Cause for False-Negative Results? J Mol Diagn. 2017;19(1):57-64. Epub 20161101. [CrossRef] [PubMed]
- Wu X, Snir O, Rottmann D, Wong S, Buza N, Hui P. Minimal microsatellite shift in microsatellite instability high endometrial cancer: a significant pitfall in diagnostic interpretation. Mod Pathol. 2019;32(5):650-8. Epub 20181115. [CrossRef] [PubMed]
- Gallon R, Phelps R, Hayes C, Brugieres L, Guerrini-Rousseau L, Colas C, et al. Constitutional Microsatellite Instability, Genotype, and Phenotype Correlations in Constitutional Mismatch Repair Deficiency. Gastroenterology. 2023;164(4):579-92 e8. Epub 20221229. [CrossRef] [PubMed]
- Gallon R, Sheth H, Hayes C, Redford L, Alhilal G, O’Brien O, et al. Sequencing-based microsatellite instability testing using as few as six markers for high-throughput clinical diagnostics. Hum Mutat. 2020;41(1):332-41. [CrossRef] [PubMed]
- Levine MD, Pearlman R, Hampel H, Cosgrove C, Cohn D, Chassen A, et al. Up-Front Multigene Panel Testing for Cancer Susceptibility in Patients With Newly Diagnosed Endometrial Cancer: A Multicenter Prospective Study. JCO Precis Oncol. 2021;5:1588-602. [CrossRef] [PubMed]
- Pearlman R, Frankel WL, Swanson BJ, Jones D, Zhao W, Yilmaz A, et al. Prospective Statewide Study of Universal Screening for Hereditary Colorectal Cancer: The Ohio Colorectal Cancer Prevention Initiative. JCO Precis Oncol. 2021;5. Epub 20210505. [CrossRef] [PubMed]
- Ryan NAJ, McMahon R, Tobi S, Snowsill T, Esquibel S, Wallace AJ, et al. The proportion of endometrial tumours associated with Lynch syndrome (PETALS): A prospective cross-sectional study. PLoS Med. 2020;17(9):e1003263. Epub 20200917. [CrossRef] [PubMed]
- Ryan N, Wall J, Crosbie EJ, Arends M, Bosse T, Arif S, et al. Lynch syndrome screening in gynaecological cancers: results of an international survey with recommendations for uniform reporting terminology for mismatch repair immunohistochemistry results. Histopathology. 2019;75(6):813-24. Epub 20190906. [CrossRef] [PubMed]
- Halford S, Sasieni P, Rowan A, Wasan H, Bodmer W, Talbot I, et al. Low-level microsatellite instability occurs in most colorectal cancers and is a nonrandomly distributed quantitative trait. Cancer Research. 2002;62(1):53-7. PubMed PMID: WOS:000173382600012.
- Redford L, Alhilal G, Needham S, O’Brien O, Coaker J, Tyson J, et al. A novel panel of short mononucleotide repeats linked to informative polymorphisms enabling effective high volume low cost discrimination between mismatch repair deficient and proficient tumours. PLoS One. 2018;13(8):e0203052. [CrossRef] [PubMed]
- Hiatt JB, Pritchard CC, Salipante SJ, O’Roak BJ, Shendure J. Single molecule molecular inversion probes for targeted, high-accuracy detection of low-frequency variation. Genome Res. 2013;23(5):843-54. [CrossRef] [PubMed]
- Li H, Durbin R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics. 2009;25(14):1754-60. [CrossRef] [PubMed]
- McConechy MK, Talhouk A, Li-Chang HH, Leung S, Huntsman DG, Gilks CB, McAlpine JN. Detection of DNA mismatch repair (MMR) deficiencies by immunohistochemistry can effectively diagnose the microsatellite instability (MSI) phenotype in endometrial carcinomas. Gynecol Oncol. 2015;137(2):306-10. Epub 20150128. [CrossRef] [PubMed]
- Goodfellow PJ, Billingsley CC, Lankes HA, Ali S, Cohn DE, Broaddus RJ, et al. Combined Microsatellite Instability, MLH1 Methylation Analysis, and Immunohistochemistry for Lynch Syndrome Screening in Endometrial Cancers From GOG210: An NRG Oncology and Gynecologic Oncology Group Study. J Clin Oncol. 2015;33(36):4301-8. Epub 20151109. [CrossRef] [PubMed]
- Longacre TA, Broaddus R, Chuang LT, Cohen MB, Jarboe EA, Mutter GL, et al. Template for Reporting Results of Biomarker Testing of Specimens From Patients With Carcinoma of the Endometrium. Arch Pathol Lab Med. 2017;141(11):1508-12. Epub 20170316. [CrossRef] [PubMed]
- Pai RK, Plesec TP, Abdul-Karim FW, Yang B, Marquard J, Shadrach B, Roma AR. Abrupt loss of MLH1 and PMS2 expression in endometrial carcinoma: molecular and morphologic analysis of 6 cases. Am J Surg Pathol. 2015;39(7):993-9. [CrossRef] [PubMed]
- McCarthy AJ, Capo-Chichi JM, Spence T, Grenier S, Stockley T, Kamel-Reid S, et al. Heterogenous loss of mismatch repair (MMR) protein expression: a challenge for immunohistochemical interpretation and microsatellite instability (MSI) evaluation. J Pathol Clin Res. 2019;5(2):115-29. Epub 20181219. [CrossRef] [PubMed]
- Mendoza RP, Wang P, Schulte JJ, Tjota MY, Jani I, Martinez AC, et al. Endometrial Carcinomas With Subclonal Loss of Mismatch Repair Proteins: A Clinicopathologic and Genomic Study. Am J Surg Pathol. 2023;47(5):589-98. Epub 20230303. [CrossRef] [PubMed]
- Scheiderer A, Riedinger C, Kimball K, Kilgore L, Orucevic A. Reporting Subclonal Immunohistochemical Staining of Mismatch Repair Proteins in Endometrial Carcinoma in the Times of Ever-Changing Guidelines. Arch Pathol Lab Med. 2022;146(9):1114-21. [CrossRef] [PubMed]
- Riedinger CJ, Esnakula A, Haight PJ, Suarez AA, Chen W, Gillespie J, et al. Characterization of mismatch-repair (MMR)/microsatellite instability (MSI)-discordant endometrial cancers. Cancer. 2024;130(3):385-99. Epub 20230926. [CrossRef] [PubMed]
- Nagabhushana P, Kumari S, Rohilla M, Srinivasan R, Arora A, Rastogi P. Discordant immunohistochemistry in an unusual MLH1 gene variant in a case of Lynch syndrome. Gynecol Oncol Rep. 2021;37:100854. Epub 20210828. [CrossRef] [PubMed]
- Gallon R, Herrero-Belmonte P, Phelps R, Hayes C, Sollars E, Egan D, et al. A novel colorectal cancer test combining microsatellite instability and BRAF/RAS analysis: Clinical validation and impact on Lynch syndrome screening. BJC Rep. 2024;2(1):48. Epub 20240701. [CrossRef] [PubMed]
- Lin JH, Chen S, Pallavajjala A, Guedes LB, Lotan TL, Bacher JW, Eshleman JR. Validation of Long Mononucleotide Repeat Markers for Detection of Microsatellite Instability. J Mol Diagn. 2022;24(2):144-57. Epub 20211202. [CrossRef] [PubMed]
- Berends MJ, Wu Y, Sijmons RH, Mensink RG, van der Sluis T, Hordijk-Hos JM, et al. Molecular and clinical characteristics of MSH6 variants: an analysis of 25 index carriers of a germline variant. Am J Hum Genet. 2002;70(1):26-37. [CrossRef] [PubMed]
- Acharya S, Wilson T, Gradia S, Kane MF, Guerrette S, Marsischky GT, et al. hMSH2 forms specific mispair-binding complexes with hMSH3 and hMSH6. Proc Natl Acad Sci U S A. 1996;93(24):13629-34. [CrossRef] [PubMed]
- Marsischky GT, Filosi N, Kane MF, Kolodner R. Redundancy of Saccharomyces cerevisiae MSH3 and MSH6 in MSH2-dependent mismatch repair. Genes Dev. 1996;10(4):407-20. [CrossRef] [PubMed]
- Hampel H, Frankel W, Panescu J, Lockman J, Sotamaa K, Fix D, et al. Screening for Lynch syndrome (hereditary nonpolyposis colorectal cancer) among endometrial cancer patients. Cancer Res. 2006;66(15):7810-7. [CrossRef] [PubMed]
- Bennett JA, Pesci A, Morales-Oyarvide V, Da Silva A, Nardi V, Oliva E. Incidence of Mismatch Repair Protein Deficiency and Associated Clinicopathologic Features in a Cohort of 104 Ovarian Endometrioid Carcinomas. Am J Surg Pathol. 2019;43(2):235-43. [CrossRef] [PubMed]
- Gatius S, Velasco A, Varela M, Cuatrecasas M, Jares P, Setaffy L, et al. Comparison of the Idylla MSI assay with the Promega MSI Analysis System and immunohistochemistry on formalin-fixed paraffin-embedded tissue of endometrial carcinoma: results from an international, multicenter study. Virchows Arch. 2022;480(5):1031-9. Epub 20220205. [CrossRef] [PubMed]
- You JF, Buhard O, Ligtenberg MJ, Kets CM, Niessen RC, Hofstra RM, et al. Tumours with loss of MSH6 expression are MSI-H when screened with a pentaplex of five mononucleotide repeats. Br J Cancer. 2010;103(12):1840-5. Epub 20101116. [CrossRef] [PubMed]
- Helderman NC, Van Der Werf-‘t Lam AS, Msh6 Tumor G, Morreau H, Boot A, Van Wezel T, Nielsen M. Molecular Profile of MSH6-Associated Colorectal Carcinomas Shows Distinct Features From Other Lynch Syndrome-Associated Colorectal Carcinomas. Gastroenterology. 2023;165(1):271-4 e2. Epub 20230315. [CrossRef] [PubMed]
- Young J, Simms LA, Biden KG, Wynter C, Whitehall V, Karamatic R, et al. Features of colorectal cancers with high-level microsatellite instability occurring in familial and sporadic settings: parallel pathways of tumorigenesis. Am J Pathol. 2001;159(6):2107-16. [CrossRef] [PubMed]
- Bohaumilitzky L, von Knebel Doeberitz M, Kloor M, Ahadova A. Implications of Hereditary Origin on the Immune Phenotype of Mismatch Repair-Deficient Cancers: Systematic Literature Review. J Clin Med. 2020;9(6). Epub 20200604. [CrossRef] [PubMed]
- Schwitalle Y, Kloor M, Eiermann S, Linnebacher M, Kienle P, Knaebel HP, et al. Immune response against frameshift-induced neopeptides in HNPCC patients and healthy HNPCC mutation carriers. Gastroenterology. 2008;134(4):988-97. Epub 20080111. [CrossRef] [PubMed]
- Shia J, Ellis NA, Paty PB, Nash GM, Qin J, Offit K, et al. Value of histopathology in predicting microsatellite instability in hereditary nonpolyposis colorectal cancer and sporadic colorectal cancer. Am J Surg Pathol. 2003;27(11):1407-17. [CrossRef] [PubMed]
- Janikovits J, Muller M, Krzykalla J, Korner S, Echterdiek F, Lahrmann B, et al. High numbers of PDCD1 (PD-1)-positive T cells and B2M mutations in microsatellite-unstable colorectal cancer. Oncoimmunology. 2018;7(2):e1390640. Epub 20171106. [CrossRef] [PubMed]
- Koornstra JJ, de Jong S, Boersma-van Eck W, Zwart N, Hollema H, de Vries EG, Kleibeuker JH. Fas ligand expression in lynch syndrome-associated colorectal tumours. Pathol Oncol Res. 2009;15(3):399-406. [CrossRef] [PubMed]
- Pakish JB, Zhang Q, Chen Z, Liang H, Chisholm GB, Yuan Y, et al. Immune Microenvironment in Microsatellite-Instable Endometrial Cancers: Hereditary or Sporadic Origin Matters. Clin Cancer Res. 2017;23(15):4473-81. Epub 20170306. [CrossRef] [PubMed]
- Ramchander NC, Ryan NAJ, Walker TDJ, Harries L, Bolton J, Bosse T, et al. Distinct Immunological Landscapes Characterize Inherited and Sporadic Mismatch Repair Deficient Endometrial Cancer. Front Immunol. 2019;10:3023. Epub 20200109. [CrossRef] [PubMed]
- Staffa L, Echterdiek F, Nelius N, Benner A, Werft W, Lahrmann B, et al. Mismatch repair-deficient crypt foci in Lynch syndrome--molecular alterations and association with clinical parameters. PLoS One. 2015;10(3):e0121980. Epub 20150327. [CrossRef] [PubMed]
- Weyand CM, Goronzy JJ. Aging of the Immune System. Mechanisms and Therapeutic Targets. Ann Am Thorac Soc. 2016;13 Suppl 5(Suppl 5):S422-S8. [CrossRef] [PubMed]
- Stelloo E, Jansen AML, Osse EM, Nout RA, Creutzberg CL, Ruano D, et al. Practical guidance for mismatch repair-deficiency testing in endometrial cancer. Ann Oncol. 2017;28(1):96-102. [CrossRef] [PubMed]
- Li X, Zhang S, Zeng J, Song SS, Liu X, Kang W, et al. Heterogeneous expression of mismatch repair proteins and interpretation of immunohistochemical results in colorectal cancer and endometrial cancer. Pathol Res Pract. 2023;248:154647. Epub 20230702. [CrossRef] [PubMed]
- Committee HaSC. Clearing the Backlog Caused by the Pandemic. HC; 2021. p. 43.
- Aldridge J. obot RCoP. Written evidence submitted by the Royal College of Pathologists (CBP0011) 2021. Available from: https://committees.parliament.uk/writtenevidence/38399/html/.
| Cohort | Ohio | Manchester | ||||
|---|---|---|---|---|---|---|
| Dataset | Train | Val | Merged | Train | Val | Merged |
| IHC Result (MMRd/MMRp) | (45/51) | (52/48) | (97/99) | (42/38) | (51/34) | (93/72) |
| A. Original Analyses | ||||||
| Concordance MSI v IHC | 95% | 97% | 96% | 73% | 82% | 78% |
| Sensitivity MSI v IHC | 96% | 94% | 95% | 57% | 71% | 65% |
| Specificity MSI v IHC | 94% | 100% | 97% | 89% | 100% | 94% |
| B. NCL 24 Marker Panel | ||||||
| Concordance NCL_MSI v MSI MSMSIMSI_MSI) | 97% | 98% | 97% | 83% | 89% | 86% |
| Concordance NCL_MSI v IHC | 94% | 95% | 94% | 85% | 84% | 84% |
| Sensitivity NCL_MSI v IHC | 93% | 92% | 93% | 76% | 76% | 76% |
| Specificity NCL_MSI v IHC | 94% | 98% | 96% | 95% | 94% | 94% |
| % Ohio (n/total) | % Man (n/total) | |||
|---|---|---|---|---|
| MMRd | 49% | (97/196) | 56% | (93/165) |
| MSI-H | 48% | (95/196) | 39% | (64/165) |
| Concordance | 96% | (188/196) | 78% | (129/165) |
| Sample | Cohort | IHC Loss | Promega MSI | NCL_MSI Score | NCL_MSI Status | Indicates Germline Testing IHC Promega NCL | ||
|---|---|---|---|---|---|---|---|---|
| ECT176 | OT | MSH6 | MSI-H | 13.1 | MSI-H | Y | Y | Y |
| ECT184 | OT | MSH2/MSH6 | MSI-H | 15.7 | MSI-H | Y | Y | Y |
| ECV69 | OV | MSH6 | MSI-H | 17.7 | MSI-H | Y | Y | Y |
| ECV100 | OV | MSH2/MSH6 | MSI-H | 16.9 | MSI-H | Y | Y | Y |
| ECT192 | OV | No Loss | MSI-H | 39.2 | MSI-H | N | Y | Y |
| ECT102 | OV | MSH6 | MSS | -1.1 | MSS | Y | N | N |
| PET256 | MV | MLH1/PMS2 | MSI-H | 10.2 | MSI-H | Y | Y | Y |
| PET16 | MT | MLH1/PMS2 | MSI-H | 37.2 | MSI-H | Y | Y | Y |
| PET61 | MV | MSH6/MSH2 | MSI-H | 45.7 | MSI-H | Y | Y | Y |
| PET215 | MV | MSH6 | MSI-H | 26.2 | MSI-H | Y | Y | Y |
| PET173 | MV | PMS2 | MSI-H | 28.2 | MSI-H | Y | Y | Y |
| PET213 | MV | MSH6 | MSI-H | 30.3 | MSI-H | Y | Y | Y |
| PET255 | MT | MSH6 | MSS | 11.4 | MSI-H | Y | N | Y |
| PET31 | MT | MSH6 | MSS | -19.1 | MSS | Y | N | N |
| PET128 | MT | MSH6 | MSI-L | -24.6 | MSS | Y | N | N |
| 165BRC | MT | MSH6/MSH2 | MSS | 53.0 | MSI-H | Y | N | Y |
| 882BRC | MT | PMS2 | MSS | -15.6 | MSS | Y | N | N |
| PET241 | MV | MSH6 | MSI-H | -14.8 | MSS | Y | Y | N |
| PRE011 | MV | MSH6 | MSS | -21.0 | MSS | Y | N | N |
| PET96 | MV | MSH6 | MSS | -20.0 | MSS | Y | N | N |
| Sample | Cohort | IHC Loss | Promega MSI | NCL_MSI Score | NCL_MSI Status | Indicates Germline Testing IHC Promega NCL | ||
|---|---|---|---|---|---|---|---|---|
| ECT176 | OT | MSH6 | MSI-H | 13.1 | MSI-H | Y | Y | Y |
| ECT184 | OT | MSH2/MSH6 | MSI-H | 15.7 | MSI-H | Y | Y | Y |
| ECV69 | OV | MSH6 | MSI-H | 17.7 | MSI-H | Y | Y | Y |
| ECV100 | OV | MSH2/MSH6 | MSI-H | 16.9 | MSI-H | Y | Y | Y |
| ECT192 | OV | No Loss | MSI-H | 39.2 | MSI-H | N | Y | Y |
| ECT102 | OV | MSH6 | MSS | -1.1 | MSS | Y | N | N |
| PET256 | MV | MLH1/PMS2 | MSI-H | 10.2 | MSI-H | Y | Y | Y |
| PET16 | MT | MLH1/PMS2 | MSI-H | 37.2 | MSI-H | Y | Y | Y |
| PET61 | MV | MSH6/MSH2 | MSI-H | 45.7 | MSI-H | Y | Y | Y |
| PET215 | MV | MSH6 | MSI-H | 26.2 | MSI-H | Y | Y | Y |
| PET173 | MV | PMS2 | MSI-H | 28.2 | MSI-H | Y | Y | Y |
| PET213 | MV | MSH6 | MSI-H | 30.3 | MSI-H | Y | Y | Y |
| PET255 | MT | MSH6 | MSS | 11.4 | MSI-H | Y | N | Y |
| PET31 | MT | MSH6 | MSS | -19.1 | MSS | Y | N | N |
| PET128 | MT | MSH6 | MSI-L | -24.6 | MSS | Y | N | N |
| 165BRC | MT | MSH6/MSH2 | MSS | 53.0 | MSI-H | Y | N | Y |
| 882BRC | MT | PMS2 | MSS | -15.6 | MSS | Y | N | N |
| PET241 | MV | MSH6 | MSI-H | -14.8 | MSS | Y | Y | N |
| PRE011 | MV | MSH6 | MSS | -21.0 | MSS | Y | N | N |
| PET96 | MV | MSH6 | MSS | -20.0 | MSS | Y | N | N |
| % Ohio (n/total) | % Man (n/total) | |||
|---|---|---|---|---|
| MMRd | 49% | (97/196) | 56% | (93/165) |
| MSI-H | 48% | (95/196) | 39% | (64/165) |
| Concordance | 96% | (188/196) | 78% | (129/165) |
| Cohort | Ohio | Manchester | ||||
|---|---|---|---|---|---|---|
| Dataset | Train | Val | Merged | Train | Val | Merged |
| IHC Result (MMRd/MMRp) | (45/51) | (52/48) | (97/99) | (42/38) | (51/34) | (93/72) |
| A. Original Analyses | ||||||
| Concordance MSI v IHC | 95% | 97% | 96% | 73% | 82% | 78% |
| Sensitivity MSI v IHC | 96% | 94% | 95% | 57% | 71% | 65% |
| Specificity MSI v IHC | 94% | 100% | 97% | 89% | 100% | 94% |
| B. NCL 24 Marker Panel | ||||||
| Concordance NCL_MSI v MSI MSMSIMSI_MSI) | 97% | 98% | 97% | 83% | 89% | 86% |
| Concordance NCL_MSI v IHC | 94% | 95% | 94% | 85% | 84% | 84% |
| Sensitivity NCL_MSI v IHC | 93% | 92% | 93% | 76% | 76% | 76% |
| Specificity NCL_MSI v IHC | 94% | 98% | 96% | 95% | 94% | 94% |
| Sample | Cohort | IHC Loss | Promega MSI | NCL_MSI Score | NCL_MSI Status | Indicates Germline Testing IHC Promega NCL | ||
|---|---|---|---|---|---|---|---|---|
| ECT176 | OT | MSH6 | MSI-H | 13.1 | MSI-H | Y | Y | Y |
| ECT184 | OT | MSH2/MSH6 | MSI-H | 15.7 | MSI-H | Y | Y | Y |
| ECV69 | OV | MSH6 | MSI-H | 17.7 | MSI-H | Y | Y | Y |
| ECV100 | OV | MSH2/MSH6 | MSI-H | 16.9 | MSI-H | Y | Y | Y |
| ECT192 | OV | No Loss | MSI-H | 39.2 | MSI-H | N | Y | Y |
| ECT102 | OV | MSH6 | MSS | -1.1 | MSS | Y | N | N |
| PET256 | MV | MLH1/PMS2 | MSI-H | 10.2 | MSI-H | Y | Y | Y |
| PET16 | MT | MLH1/PMS2 | MSI-H | 37.2 | MSI-H | Y | Y | Y |
| PET61 | MV | MSH6/MSH2 | MSI-H | 45.7 | MSI-H | Y | Y | Y |
| PET215 | MV | MSH6 | MSI-H | 26.2 | MSI-H | Y | Y | Y |
| PET173 | MV | PMS2 | MSI-H | 28.2 | MSI-H | Y | Y | Y |
| PET213 | MV | MSH6 | MSI-H | 30.3 | MSI-H | Y | Y | Y |
| PET255 | MT | MSH6 | MSS | 11.4 | MSI-H | Y | N | Y |
| PET31 | MT | MSH6 | MSS | -19.1 | MSS | Y | N | N |
| PET128 | MT | MSH6 | MSI-L | -24.6 | MSS | Y | N | N |
| 165BRC | MT | MSH6/MSH2 | MSS | 53.0 | MSI-H | Y | N | Y |
| 882BRC | MT | PMS2 | MSS | -15.6 | MSS | Y | N | N |
| PET241 | MV | MSH6 | MSI-H | -14.8 | MSS | Y | Y | N |
| PRE011 | MV | MSH6 | MSS | -21.0 | MSS | Y | N | N |
| PET96 | MV | MSH6 | MSS | -20.0 | MSS | Y | N | N |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




