Submitted:
06 October 2024
Posted:
07 October 2024
You are already at the latest version
Abstract
Keywords:
Introduction
Methods

Results

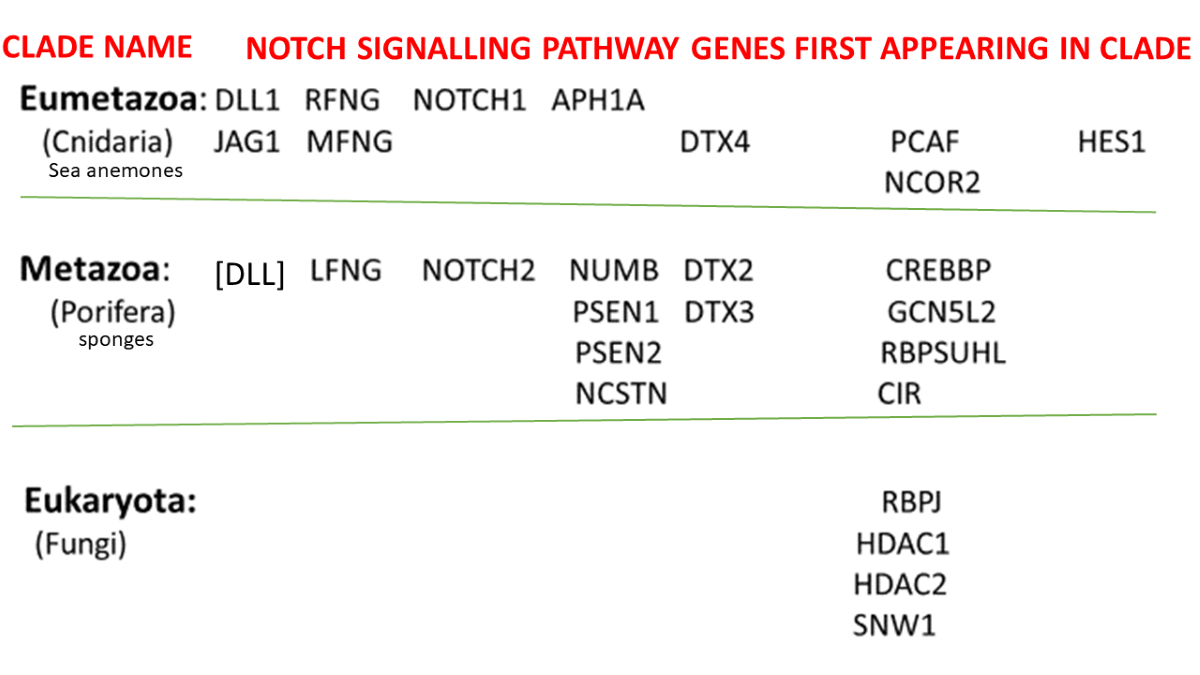
Exploring the Deep Evolutionary Origins of the Notch, DLL and JAG Proteins
Discussion
Conclusions
References
- Jin, M.; Zhang, H.; Xu, B.; Li, Y.; Qin, H.; Yu, S.; He, J. Jag2b-Notch3/1b-mediated Neuron-to-glia Crosstalk Controls Retinal Gliogenesis. EMBO Rep. 2022, 23, e54922. [Google Scholar] [CrossRef] [PubMed]
- Eng, J.W.-L.; Kato, Y.; Adachi, Y.; Swaminathan, B.; Naiche, L.A.; Vadakath, R.; Sakamoto, Y.; Nakazawa, Y.; Tachino, S.; Ito, K.; et al. Inhibition of Notch4 Using Novel Neutralizing Antibodies Reduces Tumor Growth in Murine Cancer Models by Targeting the Tumor Endothelium. Cancer Res. Commun. 2024, 4, 1881–1893. [Google Scholar] [CrossRef] [PubMed]
- MacGrogan, D.; Münch, J.; Pompa, J.L. de la Notch and Interacting Signalling Pathways in Cardiac Development, Disease, and Regeneration. Nat. Rev. Cardiol. 2018, 15, 685–704. [Google Scholar] [CrossRef] [PubMed]
- Kuriyama, K.; Kodama, Y.; Shiokawa, M.; Nishikawa, Y.; Marui, S.; Kuwada, T.; Sogabe, Y.; Kakiuchi, N.; Tomono, T.; Matsumori, T.; et al. Essential Role of Notch/Hes1 Signaling in Postnatal Pancreatic Exocrine Development. J. Gastroenterol. 2021, 56, 673–687. [Google Scholar] [CrossRef]
- Demitrack, E.S.; Samuelson, L.C. Notch Regulation of Gastrointestinal Stem Cells. J. Physiol. 2016, 594, 4791–4803. [Google Scholar] [CrossRef]
- Chen, W.; Wei, W.; Yu, L.; Ye, Z.; Huang, F.; Zhang, L.; Hu, S.; Cai, C. Mammary Development and Breast Cancer: A Notch Perspective. J. Mammary Gland Biol. Neoplasia 2021, 26, 309–320. [Google Scholar] [CrossRef]
- Kiyokawa, H.; Morimoto, M. Notch Signaling in the Mammalian Respiratory System, Specifically the Trachea and Lungs, in Development, Homeostasis, Regeneration, and Disease. Dev., Growth Differ. 2020, 62, 67–79. [Google Scholar] [CrossRef]
- Ehebauer, M.; Hayward, P.; Arias, A.M. Notch, a Universal Arbiter of Cell Fate Decisions. Science 2006, 314, 1414–1415. [Google Scholar] [CrossRef]
- Gazave, E.; Lapébie, P.; Richards, G.S.; Brunet, F.; Ereskovsky, A.V.; Degnan, B.M.; Borchiellini, C.; Vervoort, M.; Renard, E. Origin and Evolution of the Notch Signalling Pathway: An Overview from Eukaryotic Genomes. BMC Evol. Biol. 2009, 9, 249. [Google Scholar] [CrossRef]
- Lv, Y.; Pang, X.; Cao, Z.; Song, C.; Liu, B.; Wu, W.; Pang, Q. Evolution and Function of the Notch Signaling Pathway: An Invertebrate Perspective. Int. J. Mol. Sci. 2024, 25, 3322. [Google Scholar] [CrossRef]
- Babonis, L.S.; Martindale, M.Q. Phylogenetic Evidence for the Modular Evolution of Metazoan Signalling Pathways. Philos. Trans. R. Soc. B: Biol. Sci. 2017, 372, 20150477. [Google Scholar] [CrossRef] [PubMed]
- Domazet-Lošo, T.; Tautz, D. Phylostratigraphic Tracking of Cancer Genes Suggests a Link to the Emergence of Multicellularity in Metazoa. BMC Biol. 2010, 8, 66. [Google Scholar] [CrossRef] [PubMed]
- Fiddes, I.T.; Lodewijk, G.A.; Mooring, M.; Bosworth, C.M.; Ewing, A.D.; Mantalas, G.L.; Novak, A.M.; Bout, A. van den; Bishara, A.; Rosenkrantz, J.L.; et al. Human-Specific NOTCH2NL Genes Affect Notch Signaling and Cortical Neurogenesis. Cell 2018, 173, 1356–1369. [Google Scholar] [CrossRef] [PubMed]
- Bertrand, S.; Petillon, Y.L.; Somorjai, I.M.L.; Escriva, H. Developmental Cell-Cell Communication Pathways in the Cephalochordate Amphioxus: Actors and Functions. Int. J. Dev. Biol. 2017, 61, 697–722. [Google Scholar] [CrossRef]
- Sánchez-Iranzo, H.; Halavatyi, A.; Diz-Muñoz, A. Strength of Interactions in the Notch Gene Regulatory Network Determines Patterning and Fate in the Notochord. eLife 2022, 11, e75429. [Google Scholar] [CrossRef]
- Materna, M.; Delmonte, O.M.; Bosticardo, M.; Momenilandi, M.; Conrey, P.E.; Muylder, B.C.-D.; Bravetti, C.; Bellworthy, R.; Cederholm, A.; Staels, F.; et al. The Immunopathological Landscape of Human Pre-TCRα Deficiency: From Rare to Common Variants. Science 2024, 383, eadh4059–eadh4059. [Google Scholar] [CrossRef]
- Zuma, A.A.; Souza, W. de Histone Deacetylases as Targets for Antitrypanosomal Drugs. Futur. Sci. OA 2018, 4, FSO325. [Google Scholar] [CrossRef]
- Převorovský, M.; Půta, F.; Folk, P. Fungal CSL Transcription Factors. BMC Genom. 2007, 8, 233. [Google Scholar] [CrossRef]
- Oravcová, M.; Teska, M.; Půta, F.; Folk, P.; Převorovský, M. Fission Yeast CSL Proteins Function as Transcription Factors. PLoS ONE 2013, 8, e59435. [Google Scholar] [CrossRef]
- Hálová, M.; Gahura, O.; Převorovský, M.; Cit, Z.; Novotný, M.; Valentová, A.; Abrhámová, K.; Půta, F.; Folk, P. Nineteen Complex–Related Factor Prp45 Is Required for the Early Stages of Cotranscriptional Spliceosome Assembly. RNA 2017, 23, 1512–1524. [Google Scholar] [CrossRef]
- Schenkelaars, Q.; Pratlong, M.; Kodjabachian, L.; Fierro-Constain, L.; Vacelet, J.; Bivic, A.L.; Renard, E.; Borchiellini, C. Animal Multicellularity and Polarity without Wnt Signaling. Sci. Rep. 2017, 7, 15383. [Google Scholar] [CrossRef] [PubMed]
- Richards, G.S.; Degnan, B.M. The Dawn of Developmental Signaling in the Metazoa. Cold Spring Harb. Symp. Quant. Biol. 2009, 74, 81–90. [Google Scholar] [CrossRef] [PubMed]
- Richards, G.S.; Degnan, B.M. The Expression of Delta Ligands in the Sponge Amphimedon Queenslandica Suggests an Ancient Role for Notch Signaling in Metazoan Development. EvoDevo 2012, 3, 15. [Google Scholar] [CrossRef] [PubMed]
- Münder, S.; Tischer, S.; Grundhuber, M.; Büchels, N.; Bruckmeier, N.; Eckert, S.; Seefeldt, C.A.; Prexl, A.; Käsbauer, T.; Böttger, A. Notch-Signalling Is Required for Head Regeneration and Tentacle Patterning in Hydra. Dev. Biol. 2013, 383, 146–157. [Google Scholar] [CrossRef]
- Pan, Q.; Mercker, M.; Klimovich, A.; Wittlieb, J.; Marciniak-Czochra, A.; Böttger, A. Genetic Interference with HvNotch Provides New Insights into the Role of the Notch-Signalling Pathway for Developmental Pattern Formation in Hydra. Sci. Rep. 2024, 14, 8553. [Google Scholar] [CrossRef]
- Appella, E.; Weber, I.T.; Blasi, F. Structure and Function of Epidermal Growth Factor-like Regions in Proteins. FEBS Lett. 1988, 231, 1–4. [Google Scholar] [CrossRef]
- Suga, H.; Dacre, M.; Mendoza, A. de; Shalchian-Tabrizi, K.; Manning, G.; Ruiz-Trillo, I. Genomic Survey of Premetazoans Shows Deep Conservation of Cytoplasmic Tyrosine Kinases and Multiple Radiations of Receptor Tyrosine Kinases. Sci. Signal. 2012, 5, ra35. [Google Scholar] [CrossRef]
- Li E, Hristova K. Receptor tyrosine kinase transmembrane domains: Function, dimer structure and dimerization energetics. Cell Adh Migr. [CrossRef]
- Newman, S.A. ‘Biogeneric’ Developmental Processes: Drivers of Major Transitions in Animal Evolution. Philos. Trans. R. Soc. B: Biol. Sci. 2016, 371, 20150443. [Google Scholar] [CrossRef]
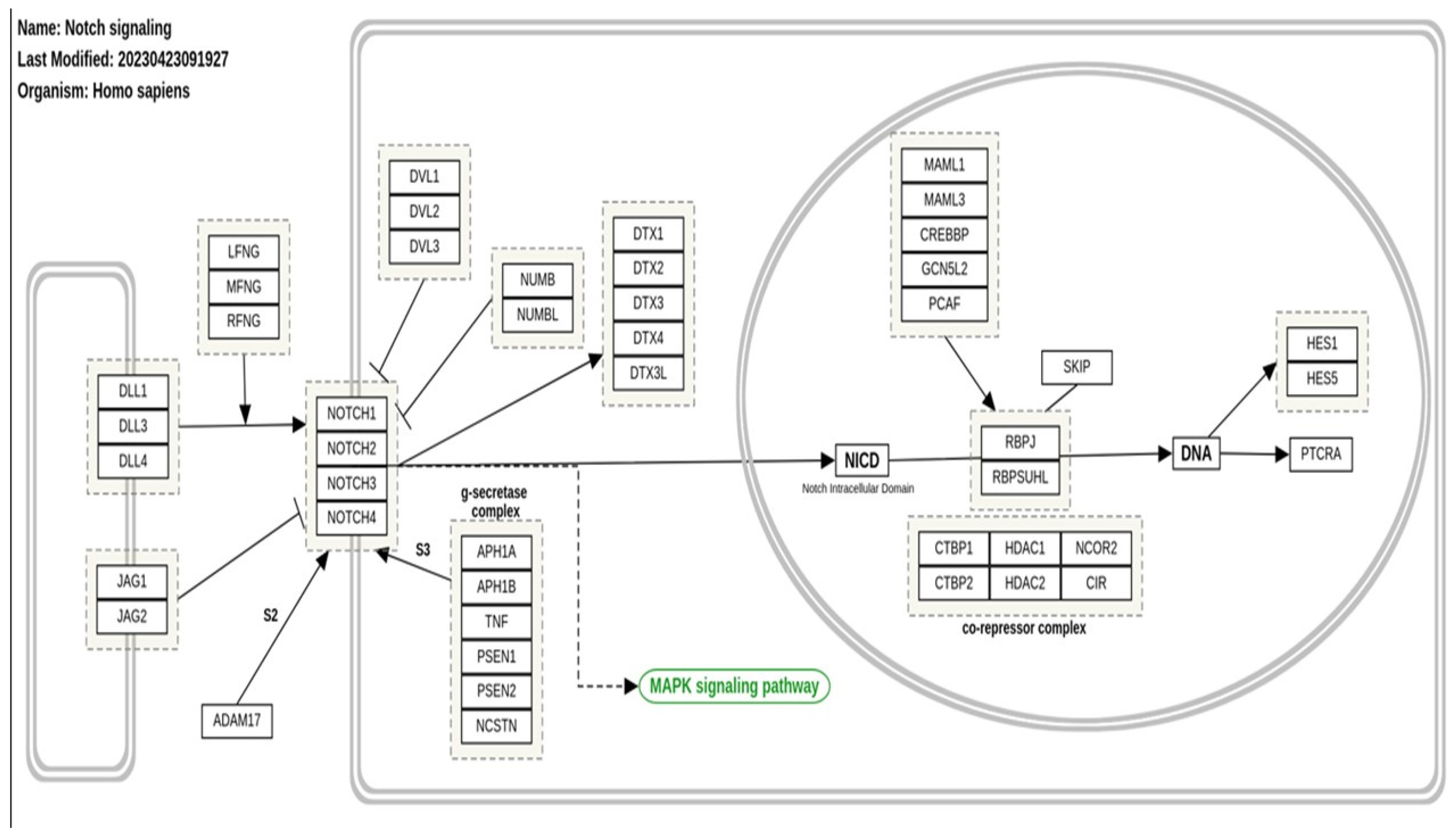
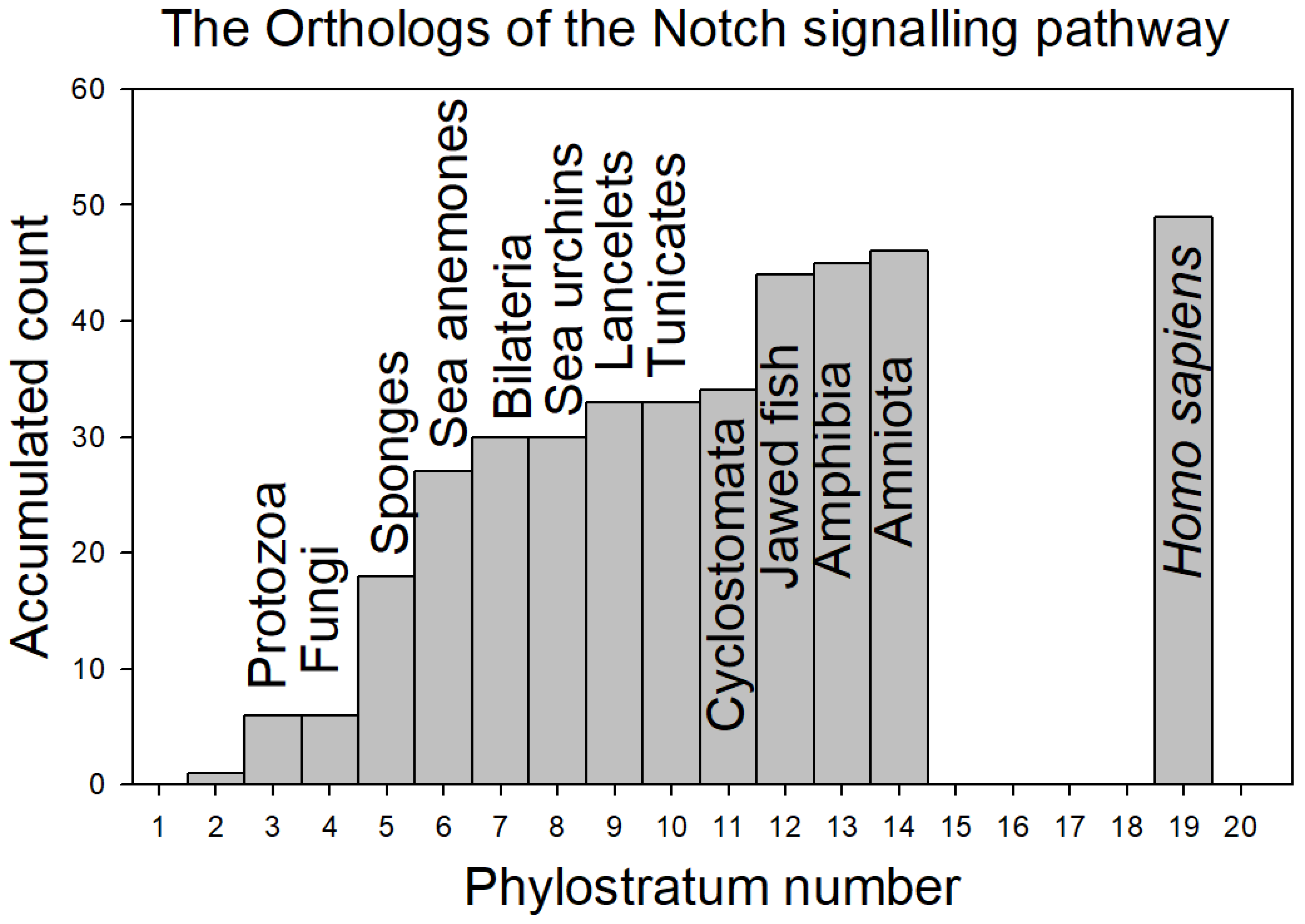
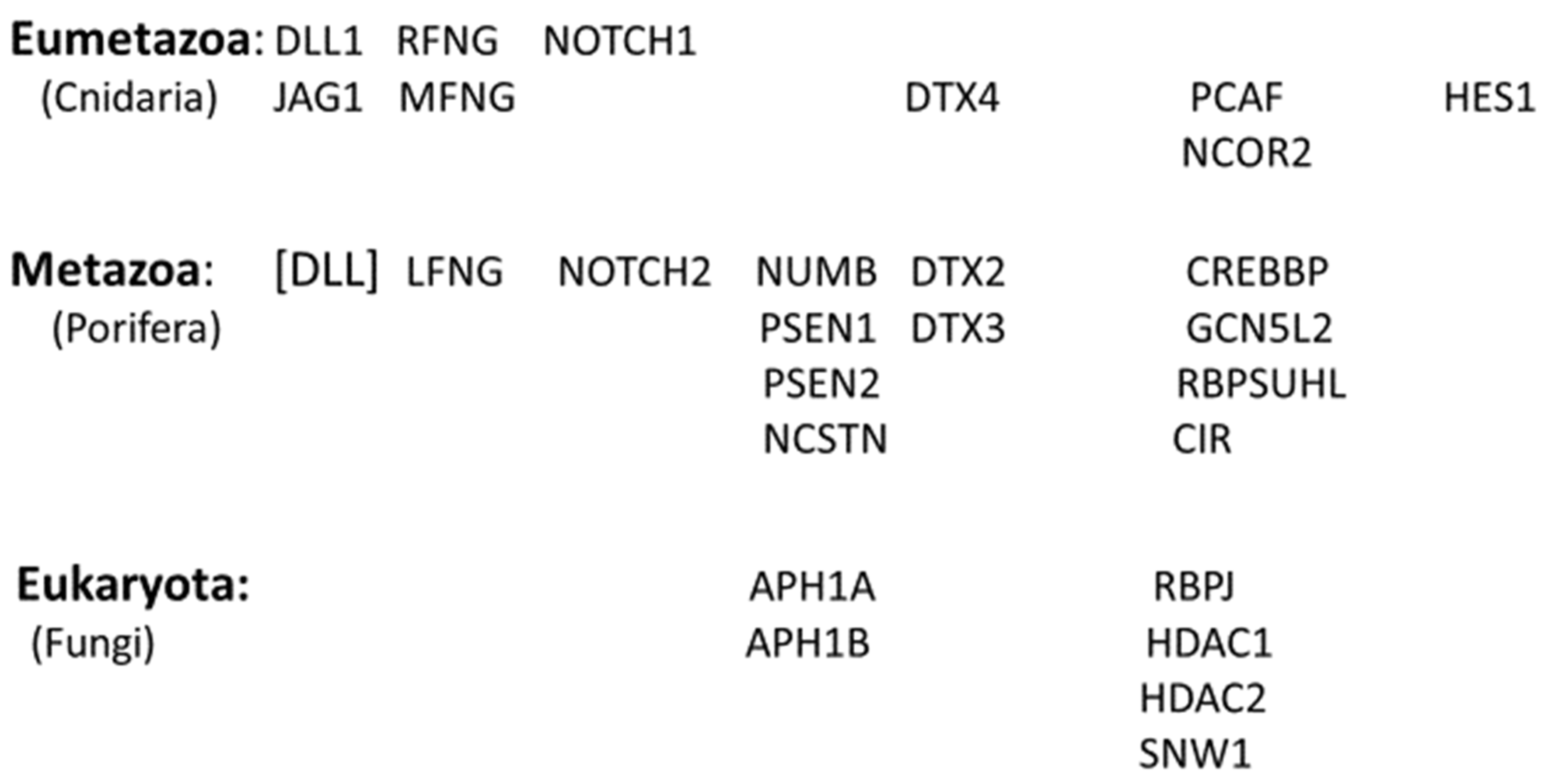




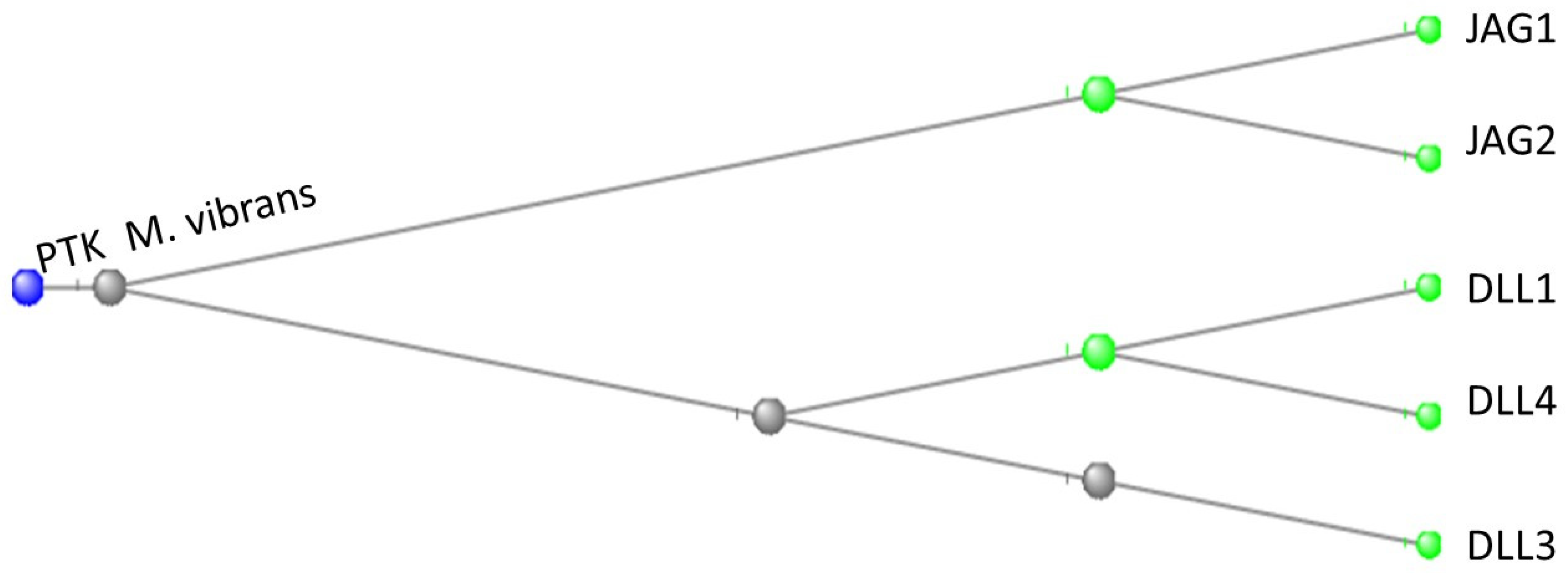
| TABLE 1a 0RTHOLOGS OF THE NOTCH SIGNALLING GENES (A to H) | |||||||
| HGNC (or name on WP268) | Uniprot | Ortholog found in: | PSn* | E value** | |||
| ADAM17 | P78536 | Branchiostomatidae | 9 | 6.00E-178 | |||
| APH1A | Q96BI3 | Fungi | 3 | 1.00E-34 | |||
| APH1B | Q8WW43 | Fungi | 3 | 3.00E-40 | |||
| CIR1 (CIR) | Q86X95 | Porifera | 5 | 1.00E-63 | |||
| CREBBP | Q92793 | Porifera | 5 | 0.00E+00 | |||
| CTBP1 | Q13363 | Bilateria | 7 | 0.00E+00 | |||
| CTBP2 | P56545 | Branchiostomatidae | 9 | 0.00E+00 | |||
| DLL1 | O00548 | Cnidaria | 6 | 2.00E-110 | |||
| DLL3 | Q9NYJ7 | Elasmobranchi sharks and rays | 12 | 1.00E-111 | |||
| DLL4 | Q9NR61 | Branchiostomatidae | 9 | 3.00E-180 | |||
| DTX1 | Q86Y01 | Bilateria | 7 | 5.00E-100 | |||
| DTX2 | Q86UW9 | Porifera | 5 | 6.00E-83 | |||
| DTX3 | Q8N9I9 | Porifera | 5 | 8.00E-63 | |||
| DTX3L | Q8TDB6 | Elasmobranchi sharks and rays | 12 | 9.00E-59 | |||
| DTX4 | Q9Y2E6 | Cnidaria | 6 | 2.00E-87 | |||
| DVL1 | O14640 | Elasmobranchi sharks and rays | 12 | 0.00E+00 | |||
| DVL2 | O14641 | Elasmobranchi sharks and rays | 12 | 0.00E+00 | |||
| DVL3 | Q92997 | Cyclostomata | 11 | 0.00E+00 | |||
| KAT2A (GCN5L2) | Q92830 | Porifera | 5 | 0.00E+00 | |||
| HDAC1 | Q13547 | Fungi | 3 | 0.00E+00 | |||
| HDAC2 | Q92769 | Protista | 2 | 0.00E+00 | |||
| HES1 | Q14469 | Cnidaria | 6 | 6.00E-50 | |||
| HES5 | Q5TA89 | Elasmobranchi sharks and rays | 12 | 5.00E-43 | |||
| Footnotes: *Phylostratum number ** Expect value in a BLAST 2-sequence comparison | |||||||
| TABLE 1b 0RTHOLOGS OF THE NOTCH SIGNALLING GENES (J to T) | |||||||
| HGNC (and name on WP268) | Uniprot | Ortholog found in: | PSn* | E value** | |||
| JAG1 | P78504 | Cnidaria | 6 | 0.00E+00 | |||
| JAG2 | Q9Y219 | Elasmobranchi sharks and rays | 12 | 0.00E+00 | |||
| LFNG | Q8NES3 | Porifera | 5 | 8.00E-51 | |||
| MAML1 | Q92585 | Elasmobranchi sharks and rays | 12 | 0.00E+00 | |||
| MAML3 | Q96JK9 | Elasmobranchi sharks and rays | 12 | 0.00E+00 | |||
| MFNG | O00587 | Cnidaria | 6 | 3.00E-91 | |||
| NCOR2 | Q9Y618 | Cnidaria | 6 | 2.20E-117 | |||
| NCSTN | Q92542 | Porifera | 5 | 1.00E-106 | |||
| NOTCH1 | P46531 | Cnidaria | 6 | 0.00E+00 | |||
| NOTCH2 | Q04721 | Porifera | 5 | 0.00E+00 | |||
| NOTCH2NLA | Q7Z3S9 | Humans | 19 | Identical | |||
| NOTCH2NLB | P0DPK3 | Humans | 19 | Identical | |||
| NOTCH2NLC | P0DPK4 | Humans | 19 | Identical | |||
| NOTCH3 | Q9UM47 | Elasmobranchi sharks and rays | 12 | 0.00E+00 | |||
| NOTCH4 | Q99466 | Amphibia | 13 | 0.00E+00 | |||
| NUMB | P49757 | Porifera | 5 | 2.00E-47 | |||
| NUMBL | Q9Y6R0 | Bilateria | 7 | 4.00E-111 | |||
| KAT2B (PCAF) | Q92831 | Cnidaria | 6 | 0 | |||
| PSEN1 | P49768 | Porifera | 5 | 2E-132 as PSN | |||
| PSEN2 | P49810 | Porifera | 5 | 3E-122 | |||
| PTCRA | Q6ISU1 | Amniota | 14 | 1.00E-34 | |||
| RBPJ | Q06330 | Fungus | 3 | 9.00E-53 | |||
| RBPJL (RBPSUHL) | Q9UBG7 | Porifera | 5 | 2.00E-150 | |||
| RFNG | Q9Y644 | Cnidaria | 6 | 7.00E-88 | |||
| SNW1 (SKIP) | Q13573 | Fungus | 3 | 2E-170 | |||
| TNF | P01375 | Elasmobranchi sharks and rays | 12 | 5.00E-34 | |||
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




