Submitted:
03 October 2024
Posted:
04 October 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results
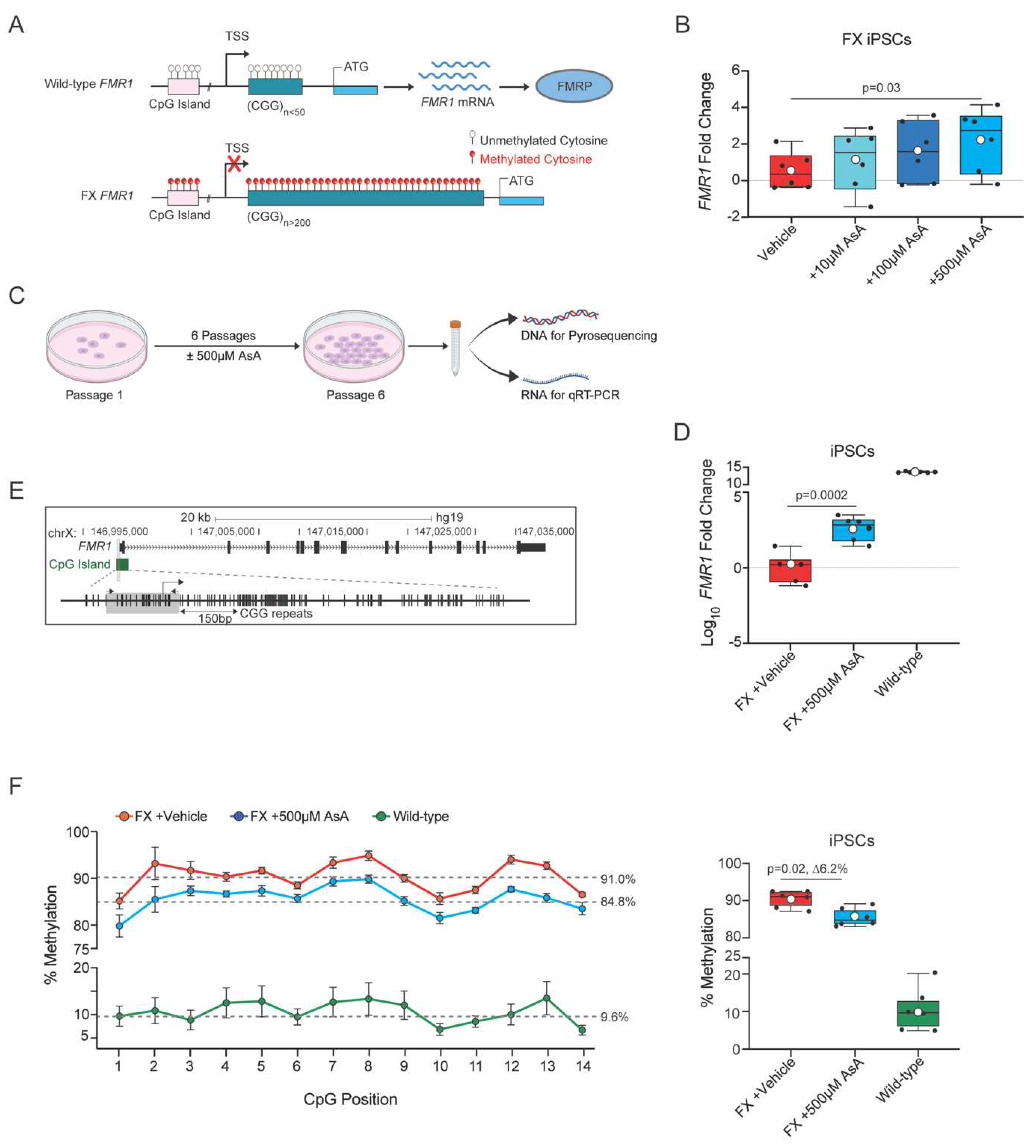
2.1. Ascorbic Acid Restores FMR1 Expression by Reversing Hypermethylation in FX iPSCs
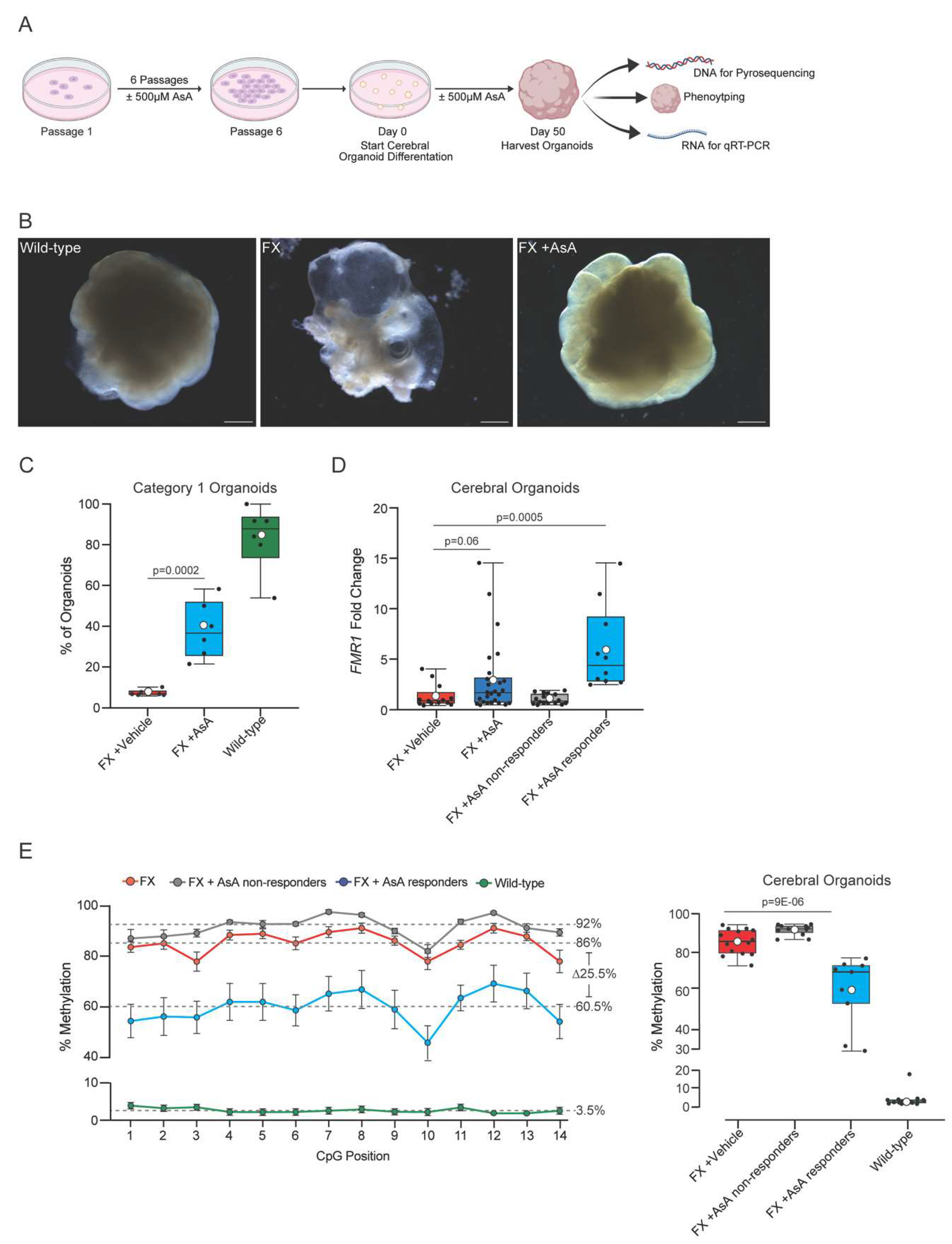
2.2. Ascorbic Acid Reactivates FMR1 and Reduces Methylation in FX Cerebral Organoids
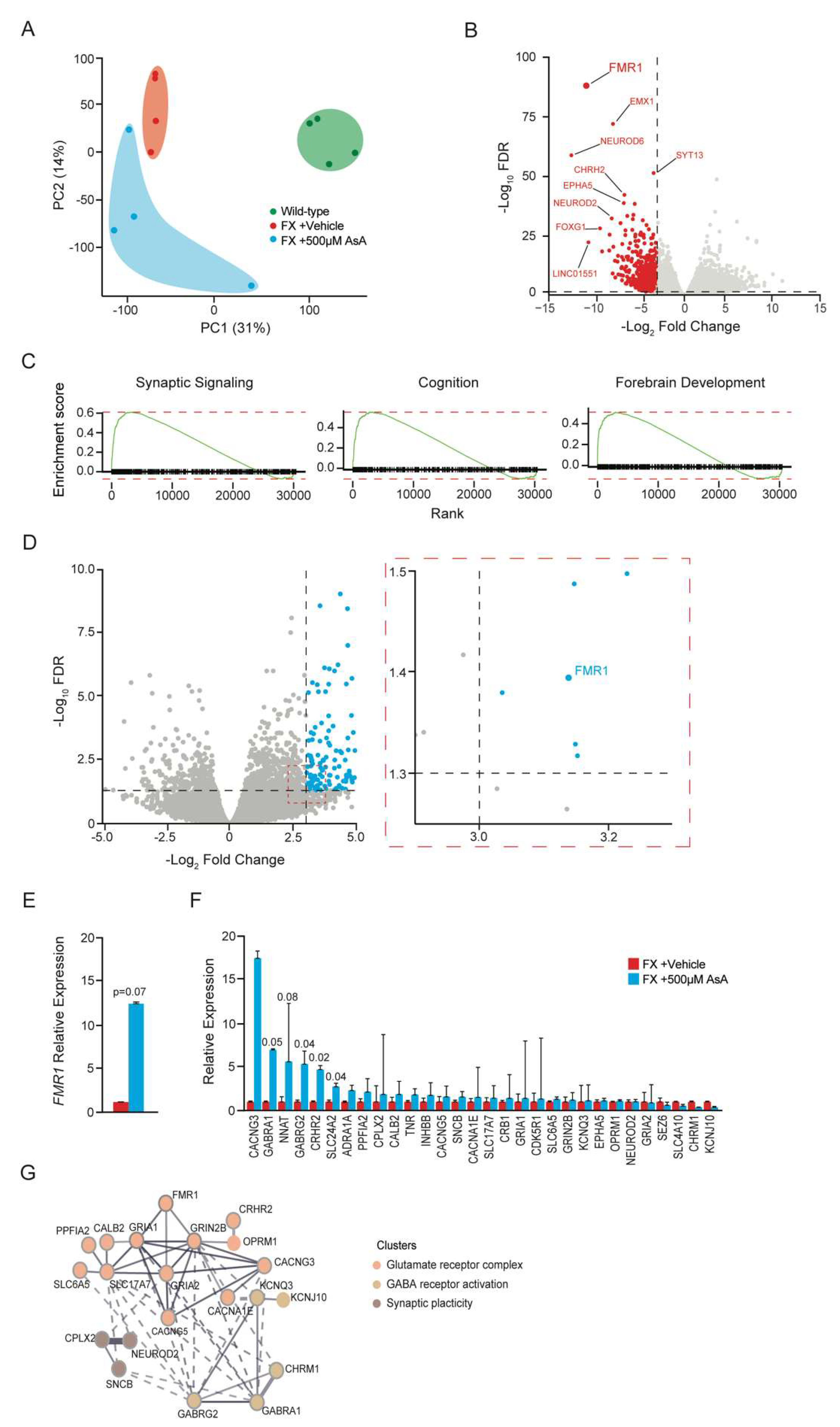
2.3. Ascorbic Acid Treatment Modulates Gene Expression in FX Cerebral Organoids Related to Neurodevelopment
3. Discussion
4. Materials and Methods
4.2. Cerebral Organoid Differentiation
4.3. DNA and RNA Extraction
4.4. RT-PCR
4.4. Pyrosequencing
4.5. RNASeq and Bioinformatic Analysis
4.6. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgements
Conflicts of Interest
References
- Crawford, D. C., J. M. Acuna, and S. L. Sherman. “Fmr1 and the Fragile X Syndrome: Human Genome Epidemiology Review.”. Genet Med 2001, 3, 359–71. [Google Scholar] [CrossRef]
- Bhakar, A. L., G. Dolen, and M. F. Bear. “The Pathophysiology of Fragile X (and What It Teaches Us About Synapses).”. Annu Rev Neurosci 2012, 35, 417–43. [Google Scholar] [CrossRef] [PubMed]
- Davidson, M., S. A. Sebastian, Y. Benitez, S. Desai, J. Quinonez, S. Ruxmohan, J. D. Stein, and W. Cueva. “Behavioral Problems in Fragile X Syndrome: A Review of Clinical Management.”. Cureus 2022, 14, e21840. [Google Scholar]
- Verkerk, A. J., M. Pieretti, J. S. Sutcliffe, Y. H. Fu, D. P. Kuhl, A. Pizzuti, O. Reiner, S. Richards, M. F. Victoria, F. P. Zhang, and et al. “Identification of a Gene (Fmr-1) Containing a Cgg Repeat Coincident with a Breakpoint Cluster Region Exhibiting Length Variation in Fragile X Syndrome.”. Cell 1991, 65, 905–14. [Google Scholar] [CrossRef] [PubMed]
- Pieretti, M., F. P. Zhang, Y. H. Fu, S. T. Warren, B. A. Oostra, C. T. Caskey, and D. L. Nelson. “Absence of Expression of the Fmr-1 Gene in Fragile X Syndrome.”. Cell 1991, 66, 817–22. [Google Scholar] [CrossRef]
- Hagerman, R. J., E. Berry-Kravis, H. C. Hazlett, D. B. Bailey, Jr., H. Moine, R. F. Kooy, F. Tassone, I. Gantois, N. Sonenberg, J. L. Mandel, and P. J. Hagerman. “Fragile X Syndrome.”. Nat Rev Dis Primers 2017, 3, 17065. [Google Scholar] [CrossRef]
- Tassone, F., A. Beilina, C. Carosi, S. Albertosi, C. Bagni, L. Li, K. Glover, D. Bentley, and P. J. Hagerman. “Elevated Fmr1 Mrna in Premutation Carriers Is Due to Increased Transcription.”. RNA 2007, 13, 555–62. [Google Scholar] [CrossRef]
- Hagerman, R., and P. Hagerman. “Advances in Clinical and Molecular Understanding of the Fmr1 Premutation and Fragile X-Associated Tremor/Ataxia Syndrome.”. Lancet Neurol 2013, 12, 786–98. [Google Scholar] [CrossRef]
- Sutcliffe, J. S., D. L. Nelson, F. Zhang, M. Pieretti, C. T. Caskey, D. Saxe, and S. T. Warren. “DNA Methylation Represses Fmr-1 Transcription in Fragile X Syndrome.”. Hum Mol Genet 1992, 1, 397–400. [Google Scholar] [CrossRef]
- Willemsen, R. , and R. F. Kooy. “Mouse Models of Fragile X-Related Disorders.”. Dis Model Mech 2023, 16. [Google Scholar] [CrossRef]
- Colvin, S. Lea, Q. Zhang, M. Wienisch, T. Kaiser, T. Aida, and G. Feng. “341 Repeats Is Not Enough for Methylation in a New Fragile X Mouse Model.”. eNeuro 2022, 9. [Google Scholar] [CrossRef] [PubMed]
- Kooy, R. F., R. D’Hooge, E. Reyniers, C. E. Bakker, G. Nagels, K. De Boulle, K. Storm, G. Clincke, P. P. De Deyn, B. A. Oostra, and P. J. Willems. “Transgenic Mouse Model for the Fragile X Syndrome.”. Am J Med Genet 1996, 64, 241–5. [Google Scholar] [CrossRef]
- “Fmr1 Knockout Mice: A Model to Study Fragile X Mental Retardation The Dutch-Belgian Fragile X Consortium.”. Cell 1994, 78, 23–33.
- Eiges, R., A. Urbach, M. Malcov, T. Frumkin, T. Schwartz, A. Amit, Y. Yaron, A. Eden, O. Yanuka, N. Benvenisty, and D. Ben-Yosef. “Developmental Study of Fragile X Syndrome Using Human Embryonic Stem Cells Derived from Preimplantation Genetically Diagnosed Embryos.”. Cell Stem Cell 2007, 1, 568–77. [Google Scholar] [CrossRef] [PubMed]
- Urbach, A., O. Bar-Nur, G. Q. Daley, and N. Benvenisty. “Differential Modeling of Fragile X Syndrome by Human Embryonic Stem Cells and Induced Pluripotent Stem Cells.”. Cell Stem Cell 2010, 6, 407–11. [Google Scholar] [CrossRef]
- Bar-Nur, O., I. Caspi, and N. Benvenisty. “Molecular Analysis of Fmr1 Reactivation in Fragile-X Induced Pluripotent Stem Cells and Their Neuronal Derivatives.”. J Mol Cell Biol 2012, 4, 180–3. [Google Scholar] [CrossRef]
- Vershkov, D., N. Fainstein, S. Suissa, T. Golan-Lev, T. Ben-Hur, and N. Benvenisty. “Fmr1 Reactivating Treatments in Fragile X Ipsc-Derived Neural Progenitors in Vitro and in Vivo.”. Cell Rep 2019, 26, 2531–39. [Google Scholar] [CrossRef]
- Vershkov, D., A. Yilmaz, O. Yanuka, A. L. Nielsen, and N. Benvenisty. “Genome-Wide Screening for Genes Involved in the Epigenetic Basis of Fragile X Syndrome.”. Stem Cell Reports 2022, 17, 1048–58. [Google Scholar] [CrossRef]
- Park, C. Y., T. Halevy, D. R. Lee, J. J. Sung, J. S. Lee, O. Yanuka, N. Benvenisty, and D. W. Kim. “Reversion of Fmr1 Methylation and Silencing by Editing the Triplet Repeats in Fragile X Ipsc-Derived Neurons.”. Cell Rep 2015, 13, 234–41. [Google Scholar] [CrossRef]
- Liu, X. S., H. Wu, M. Krzisch, X. Wu, J. Graef, J. Muffat, D. Hnisz, C. H. Li, B. Yuan, C. Xu, Y. Li, D. Vershkov, A. Cacace, R. A. Young, and R. Jaenisch. “Rescue of Fragile X Syndrome Neurons by DNA Methylation Editing of the Fmr1 Gene.”. Cell 2018, 172, 979–92. [Google Scholar] [CrossRef]
- Chong, T. L. L. Ahearn, and L. Cimmino. “Reprogramming the Epigenome with Vitamin C. Frontiers in Cell and Developmental Biology 2019, 7. [Google Scholar]
- Tsukada, Y., J. Fang, H. Erdjument-Bromage, M. E. Warren, C. H. Borchers, P. Tempst, and Y. Zhang. “Histone Demethylation by a Family of Jmjc Domain-Containing Proteins.”. Nature 2006, 439, 811–6. [Google Scholar] [CrossRef] [PubMed]
- Esteban, M. A., T. Wang, B. Qin, J. Yang, D. Qin, J. Cai, W. Li, Z. Weng, J. Chen, S. Ni, K. Chen, Y. Li, X. Liu, J. Xu, S. Zhang, F. Li, W. He, K. Labuda, Y. Song, A. Peterbauer, S. Wolbank, H. Redl, M. Zhong, D. Cai, L. Zeng, and D. Pei. “Vitamin C Enhances the Generation of Mouse and Human Induced Pluripotent Stem Cells.”. Cell Stem Cell 2010, 6, 71–9. [Google Scholar] [CrossRef] [PubMed]
- Wang, T., K. Chen, X. Zeng, J. Yang, Y. Wu, X. Shi, B. Qin, L. Zeng, M. A. Esteban, G. Pan, and D. Pei. “The Histone Demethylases Jhdm1a/1b Enhance Somatic Cell Reprogramming in a Vitamin-C-Dependent Manner.”. Cell Stem Cell 2011, 9, 575–87. [Google Scholar] [CrossRef]
- Stadtfeld, M., E. Apostolou, F. Ferrari, J. Choi, R. M. Walsh, T. Chen, S. S. Ooi, S. Y. Kim, T. H. Bestor, T. Shioda, P. J. Park, and K. Hochedlinger. “Ascorbic Acid Prevents Loss of Dlk1-Dio3 Imprinting and Facilitates Generation of All-Ips Cell Mice from Terminally Differentiated B Cells.”. Nat Genet 2012, 44, 398–405, S1. [Google Scholar] [CrossRef]
- Yue, X., and A. Rao. “Tet Family Dioxygenases and the Tet Activator Vitamin C in Immune Responses and Cancer.”. Blood 2020, 136, 1394–401. [Google Scholar] [CrossRef]
- Kaplanek, R., Z. Kejik, J. Hajduch, K. Vesela, K. Kucnirova, M. Skalickova, A. Venhauerova, B. Hosnedlova, R. Hromadka, P. Dytrych, P. Novotny, N. Abramenko, V. Antonyova, D. Hoskovec, P. Babula, M. Masarik, P. Martasek, and M. Jakubek. “Tet Protein Inhibitors: Potential and Limitations.”. Biomed Pharmacother 2023, 166, 115324. [Google Scholar]
- Lancaster, M. A., and J. A. Knoblich. “Organogenesis in a Dish: Modeling Development and Disease Using Organoid Technologies.”. Science 2014, 345, 1247125. [Google Scholar] [CrossRef]
- Lancaster, M. A., M. Renner, C. A. Martin, D. Wenzel, L. S. Bicknell, M. E. Hurles, T. Homfray, J. M. Penninger, A. P. Jackson, and J. A. Knoblich. “Cerebral Organoids Model Human Brain Development and Microcephaly.”. Nature 2013, 501, 373–9. [Google Scholar] [CrossRef]
- Luo, C., M. A. Lancaster, R. Castanon, J. R. Nery, J. A. Knoblich, and J. R. Ecker. “Cerebral Organoids Recapitulate Epigenomic Signatures of the Human Fetal Brain.”. Cell Rep 2016, 17, 3369–84. [Google Scholar] [CrossRef]
- Chiaradia, I., I. Imaz-Rosshandler, B. S. Nilges, J. Boulanger, L. Pellegrini, R. Das, N. D. Kashikar, and M. A. Lancaster. “Tissue Morphology Influences the Temporal Program of Human Brain Organoid Development.”. Cell Stem Cell 2023, 30, 1351–67. [Google Scholar] [CrossRef] [PubMed]
- Giandomenico, S. L., S. B. Mierau, G. M. Gibbons, L. M. D. Wenger, L. Masullo, T. Sit, M. Sutcliffe, J. Boulanger, M. Tripodi, E. Derivery, O. Paulsen, A. Lakatos, and M. A. Lancaster. “Cerebral Organoids at the Air-Liquid Interface Generate Diverse Nerve Tracts with Functional Output.”. Nat Neurosci 2019, 22, 669–79. [Google Scholar] [CrossRef] [PubMed]
- Lancaster, M. A., and J. A. Knoblich. “Generation of Cerebral Organoids from Human Pluripotent Stem Cells.”. Nat Protoc 2014, 9, 2329–40. [Google Scholar] [CrossRef] [PubMed]
- Pellegrini, L. Bonfio, J. Chadwick, F. Begum, M. Skehel, and M. A. Lancaster. “Human Cns Barrier-Forming Organoids with Cerebrospinal Fluid Production.”. Science 2020, 369. [Google Scholar] [CrossRef] [PubMed]
- Drozd, M. Bardoni, and M. Capovilla. “Modeling Fragile X Syndrome in Drosophila.”. Front Mol Neurosci 2018, 11, 124. [Google Scholar] [CrossRef]
- Dahlhaus, R. “Of Men and Mice: Modeling the Fragile X Syndrome. ” Front Mol Neurosci 2018, 11, 41. [Google Scholar] [CrossRef]
- Berry-Kravis, E. M., L. Lindemann, A. E. Jonch, G. Apostol, M. F. Bear, R. L. Carpenter, J. N. Crawley, A. Curie, V. Des Portes, F. Hossain, F. Gasparini, B. Gomez-Mancilla, D. Hessl, E. Loth, S. H. Scharf, P. P. Wang, F. Von Raison, R. Hagerman, W. Spooren, and S. Jacquemont. “Drug Development for Neurodevelopmental Disorders: Lessons Learned from Fragile X Syndrome.”. Nat Rev Drug Discov 2018, 17, 280–99. [Google Scholar] [CrossRef]
- Lee, H. G., S. Imaichi, E. Kraeutler, R. Aguilar, Y. W. Lee, S. D. Sheridan, and J. T. Lee. “Site-Specific R-Loops Induce Cgg Repeat Contraction and Fragile X Gene Reactivation.”. Cell 2023, 186, 2593–609. [Google Scholar] [CrossRef]
- Brighi, C., F. Salaris, A. Soloperto, F. Cordella, S. Ghirga, V. de Turris, M. Rosito, P. F. Porceddu, C. D’Antoni, A. Reggiani, A. Rosa, and S. Di Angelantonio. “Novel Fragile X Syndrome 2d and 3d Brain Models Based on Human Isogenic Fmrp-Ko Ipscs.”. Cell Death Dis 2021, 12, 498. [Google Scholar] [CrossRef]
- Kang, Y., Zhou, Y. Li, Y. Han, J. Xu, W. Niu, Z. Li, S. Liu, H. Feng, W. Huang, R. Duan, T. Xu, N. Raj, F. Zhang, J. Dou, C. Xu, H. Wu, G. J. Bassell, S. T. Warren, E. G. Allen, P. Jin, and Z. Wen. “A Human Forebrain Organoid Model of Fragile X Syndrome Exhibits Altered Neurogenesis and Highlights New Treatment Strategies.”. Nat Neurosci 2021, 24, 1377–91. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



