Submitted:
25 September 2024
Posted:
26 September 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and methods
2.1. Virus Strains, Cell Culture, and Virus Isolation
2.2. Animal Experiments and Collection of Diagnostic Samples
2.3. Optimization of Virus Capture Using NAMPs
2.4. Assessment of Virus Capture by PCR and Virus Titration
2.5. Virus Capture from Suspensions Containing Multiple Viruses
2.6. Virus Capture from Whole Blood
2.7. Virus Capture from Experimental and Clinical Samples and Diagnostic Sensitivity
2.8. Nucleic Acid (DNA/RNA) Extractions and qPCR/RT-qPCR
- ASFV qPCR: one cycle of 95 oC for 20s followed by 40 cycles of amplification with each cycle consisting of 95 oC for 10s and 60 oC for 30s;
- CaPV qPCR: one cycle of 95 oC for 10 min (enzyme activation/template denaturation) followed by 40 cycles of amplification with each cycle consisting of 95 oC for 15 s and 60 oC for 60 s; and
- PPRV RT-qPCR: one cycle of 45 oC for 10 min (reverse transcription), one cycle of 95 oC for 10 min (enzyme activation/template denaturation) and 40 cycles of amplification with each cycle consisting of 95 oC for 15 s and 60 oC for 60 s.
3. Results
3.1. Optimization of Nucleic Acids (Viral DNA/RNA) Extractions
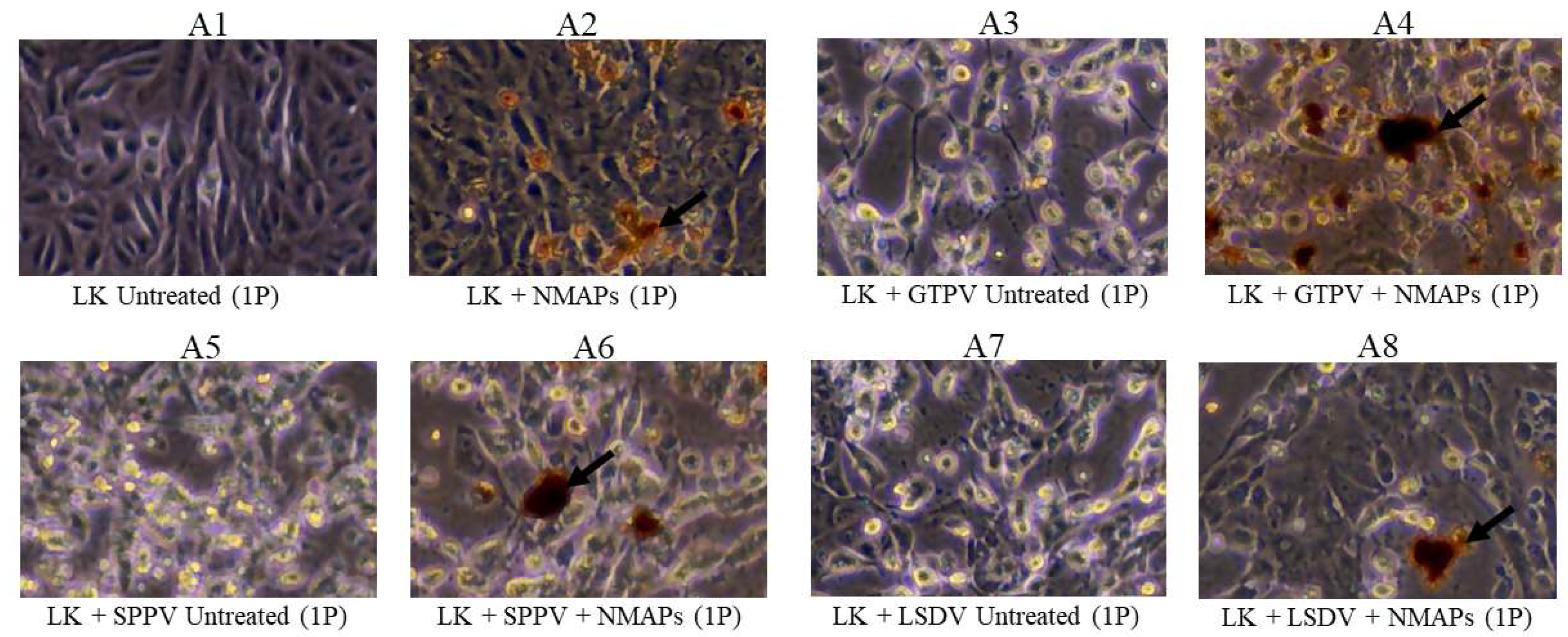
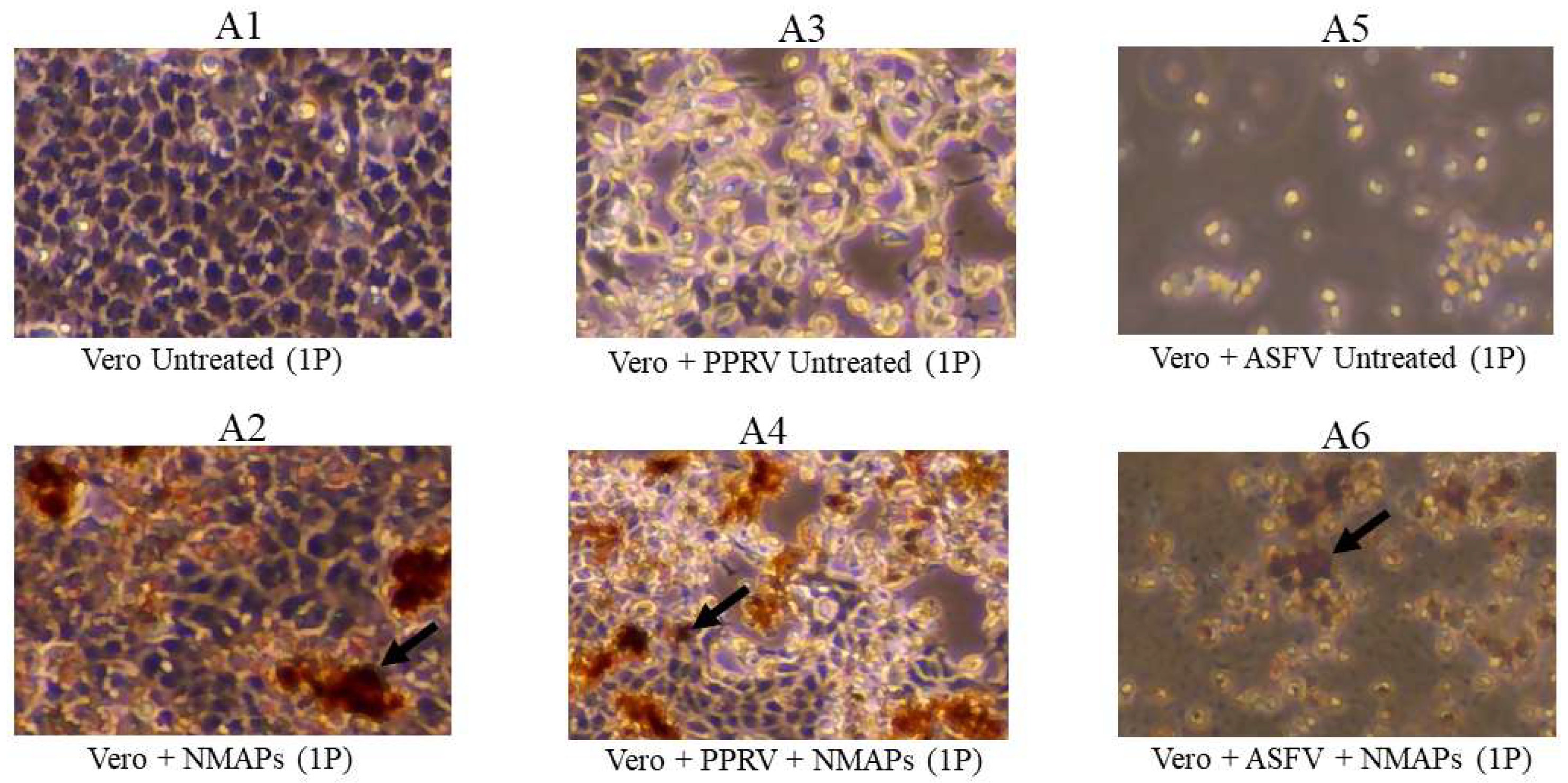
3.2. Assessment of Virus Capture Analyzed by VI, Virus Titration and Virus-Specific qPCR/RT-qPCR
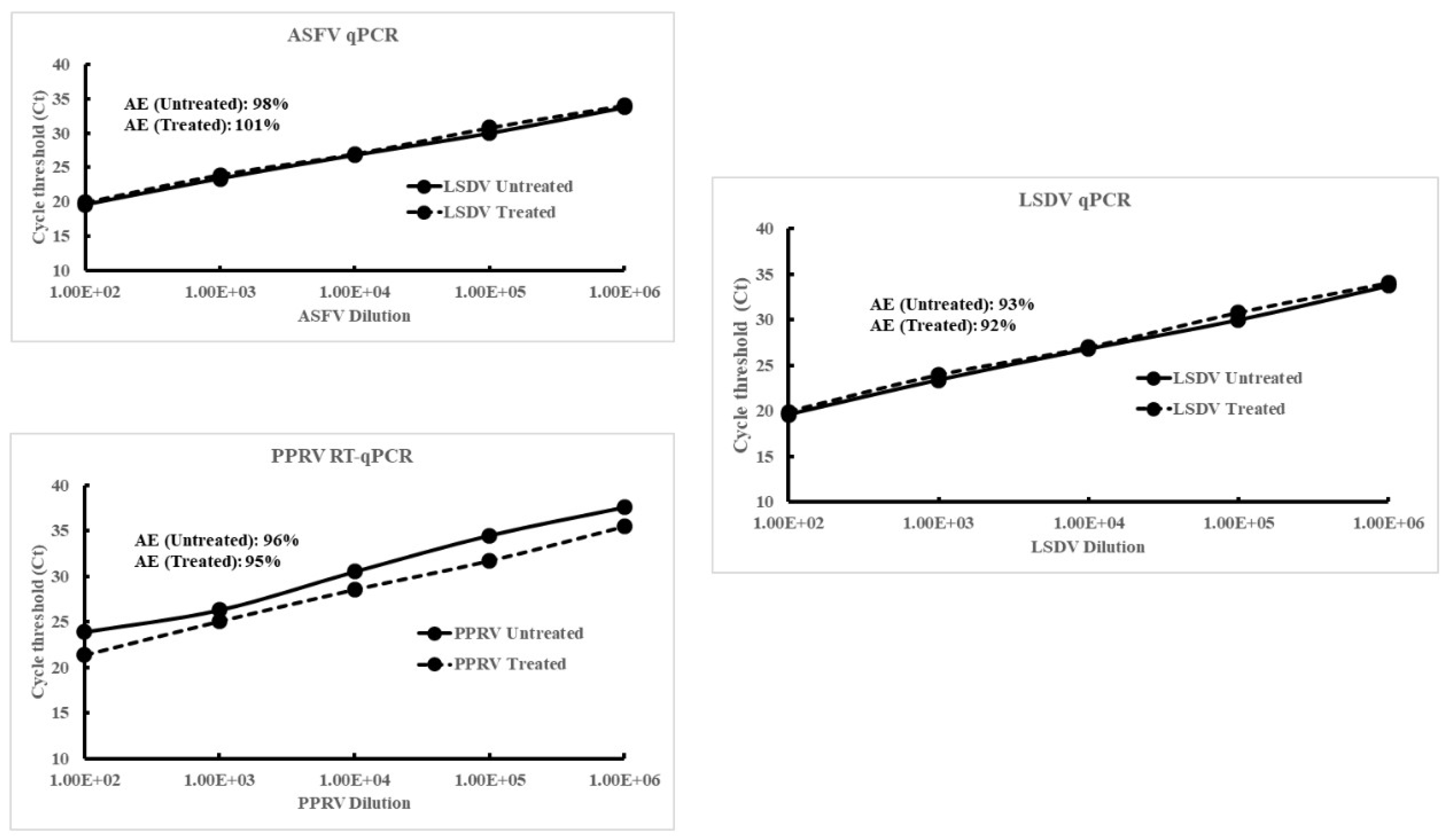
3.3. Enhanced Recovery and Sensitivity of Detection of the Viruses Using NMAPs
3.4. Virus Capture and Recovery from EWB
3.5. Virus Capture from Experimental and Clinical Samples and Diagnostic Sensitivity
3.6. Virus Capture from Suspensions Containing Multiple Viruses
4. Discussion
Author Contributions
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
| Virus | Host | # Animals | Dpi | Specimen | Animal ID | Cycle threshold (qPCR) | |
| Untreated* (-NMAPs) |
Treated# (+NMAPs) |
||||||
| ASFV | Swine | 20 | 7 | EWB | 52468 | 17.939 | 17.111 |
| 52469 | 17.481 | 17.084 | |||||
| 52470 | 16.115 | 16.124 | |||||
| 52471 | 16.855 | 16.243 | |||||
| 52.792 | 18.479 | 18.120 | |||||
| 52793 | 20.388 | 20.072 | |||||
| 52795 | 19.375 | 19.165 | |||||
| 52796 | 17.905 | 17.649 | |||||
| 53465 | 28.951 | 28.541 | |||||
| 53466 | 20.162 | 19.326 | |||||
| 52467 | 22.593 | 21.554 | |||||
| 53468 | 20.900 | 20.495 | |||||
| 53469 | 19.637 | 19.100 | |||||
| 53470 | 16.111 | 15.224 | |||||
| 53471 | 16.855 | 16.243 | |||||
| 53838 | 30.569 | 30.211 | |||||
| 53839 | 21.758 | 21.123 | |||||
| 53841 | 17.283 | 17.032 | |||||
| 53842 | 18.500 | 17.632 | |||||
| 53843 | 16.752 | 16.068 | |||||
| Virus | Host | # Animals | Dpi | Specimen | Animal ID | Cycle threshold (qPCR) | |
| Untreated* (-NMAPs) |
Treated# (+NMAPs) |
||||||
| PPRV | Goat | 8 | 8 | EWB | 23-1 | 29.814 | 29.156 |
| 23-2 | 28.097 | 28.898 | |||||
| 23-3 | 30.107 | 27.365 | |||||
| 23-4 | 28.025 | 27.430 | |||||
| 23-13 | 32.076 | 30.060 | |||||
| 23-14 | 28.198 | 27.535 | |||||
| 23-15 | 33.507 | 33.197 | |||||
| 23-16 | 31.424 | 31.160 | |||||
| Nasal swab | 23-1 | 20.993 | 19.201 | ||||
| 23-2 | 28.477 | 26.065 | |||||
| 23-3 | 29.109 | 27.197 | |||||
| 23-4 | 20.364 | 18.708 | |||||
| 23-13 | 24.098 | 25.993 | |||||
| 23-14 | 26.279 | 27.479 | |||||
| 23-15 | 28.097 | 30.306 | |||||
| 23-16 | 26.814 | 24.128 | |||||
| Oral | 23-13 | 25.407 | 27.432 | ||||
| 23-14 | 27.801 | 29.069 | |||||
| 23-15 | 31.847 | 30.905 | |||||
| 23-16 | 28.884 | 27.814 | |||||
| Conjunctival swab | 23-1 | 21.767 | 19.994 | ||||
| 23-2 | 25.942 | 25.302 | |||||
| 23-3 | 29.991 | 26.679 | |||||
| 23-4 | 26.068 | 25.326 | |||||
| 23-13 | 23.608 | 27.526 | |||||
| 23-14 | 26.845 | 26.492 | |||||
| 23-15 | 22.841 | 23.954 | |||||
| 23-16 | 22.502 | 23.016 | |||||
| Virus | Host | # Animals | Dpi | Specimen | Animal ID | Cycle threshold (qPCR) | |
| Untreated* (-NMAPs) |
Treated# (+NMAPs) | ||||||
| SPPV | Sheep | 6 | 10 | Oral swab | 23-20 | 26.344 | 31.046 |
| 23-21 | 28.602 | 29.857 | |||||
| 23-22 | 33.764 | 32.054 | |||||
| 23-28 | 30.836 | 32.176 | |||||
| 23-29 | 33.162 | 32.513 | |||||
| 23-30 | 32.458 | 32.871 | |||||
| Nasal swab | 23-20 | 29.088 | 22.960 | ||||
| 23.21 | 25.898 | 26.889 | |||||
| 23-22 | 37.022 | 37.407 | |||||
| 23-28 | 24.412 | 25.454 | |||||
| 23-29 | 29.240 | 31.502 | |||||
| 23-30 | 29.284 | 31.072 | |||||
| Conjunctival; swab | 23-20 | 27.779 | 29.219 | ||||
| 23-21 | 27.600 | 29.739 | |||||
| 23-22 | 35.546 | 35.869 | |||||
| 23-28 | 18.085 | 19.427 | |||||
| 23-29 | 26.118 | 28.356 | |||||
| 23-30 | 28.167 | 28.871 | |||||
| EWB | 23-20 | 24.550 | 23.989 | ||||
| 23-21 | 27.415 | 27.479 | |||||
| 23-22 | 23.393 | 21.898 | |||||
| 23-28 | 27.8097 | 27.066 | |||||
| 23-29 | 23.406 | 23.050 | |||||
| 23-30 | 21.470 | 20.604 | |||||
| Virus | Host | # Animals | Specimen | Animal ID | Cycle threshold (qPCR) | |
| Untreated* (-NMAPs) |
Treated# (+NMAPs) |
|||||
| ASFV | Swine | 20 | EWB | 14 | 18.06 | 17.470 |
| 27 | 19.526 | 19.056 | ||||
| 29 | 19.423 | 19.154 | ||||
| 30 | 18.722 | 18.372 | ||||
| 32 | 22.583 | 20,540 | ||||
| 36 | 22.432 | 21.468 | ||||
| 37 | 20.704 | 19.779 | ||||
| 39 | 18.155 | 17.823 | ||||
| 41 | 18.618 | 18.214 | ||||
| 42 | 22.822 | 22.890 | ||||
| 43 | 36.399 | 35.919 | ||||
| 44 | 22.325 | 22.221 | ||||
| 45 | 18.859 | 17.599 | ||||
| 46 | 20.383 | 20.061 | ||||
| 52 | 19.443 | 19.137 | ||||
| 53 | 19.309 | 18.443 | ||||
| 54 | 18.489 | 18.786 | ||||
| 58 | 17.624 | 17.362 | ||||
| 59 | 22.920 | 22.330 | ||||
| 60 | 20.967 | 20.260 | ||||
References
- Banyard, A.C.; Parida, S.; Batten, C.; Oura, C.; Kwiatek, O.; Libeau, G. Global distribution of Peste des petits ruminants virus and prospects for improved diagnosis and control. J. Gen. Virol. 2010, 91, 2885-2897. [CrossRef]
- Nanda, Y.P.; Chatterjee, A.; Purohit, A.K.; Diallo, A.; Innui, K.; Sharma, R.N.; Libeau, G.; Thevasagayam, J.A.; Bruning, A.; Kitching, R.; Anderson, J.; Barrett, Taylor W.P. The isolation of Peste des petits ruminants virus from northern India. Vet. Microbiol. 1996, 51, 207–216. [CrossRef]
- Ozkul, A.; Akca, Y.; Alkan, F.; Barrett, T.; Karaoglu, T.; Dagalp, S.B.; Anderson, J.; Yesilbag, K.; Cokcaliskan, C.; Gencay, A.; Burgu, I. Prevalence, distribution, and host range of peste des petits ruminants virus, Turkey. Emerg. Infect. Dis. 2002, 8, 708–712. [CrossRef]
- Khalafalla, A.I.; Saeed, I.K.; Ali, Y.H.; Abdurrahman, M.B.; Kwiatek, O.; Libeau, G.; Obeida, A.A.; Abbas, Z. An outbreak of peste des petits ruminants (PPR) in camels in the Sudan. Acta Trop. 2010, 116, 161–165. [CrossRef]
- Suresh, K.P.; Bhavya, A.P.; Shivamallu, C.; Achar, R.; Silina, E.; Stupin, V.; Kollur, S.P.; Shome, B.R.; S. Patil, S.S. Seroprevalence of sheeppox and goatpox virus in Asia and African continent: A systematic review and meta-analysis (Scientometrics). Veterinary world, 2022, 2231-0916. http://www.veterinaryworld.org/Vol.15/February-2022/29.pdf.
- Moudgil, G.; Chadha, J.; Khullar, L.; Chhibber, S.; Harjai, K. Lumpy skin disease: Insights into current status and geographical expansion of a transboundary viral disease. Microbial Pathogens, 2024, 86, 106485. [CrossRef]
- Tulman, E.R.; Afonso, C.L.; Lu, Z.; Zsak, Z.; Sur, J.H.; Sandybaev, N.T.; Kerembekova, U.Z.; Zaitsev, V.L.; Kutish, G.F.; Rock, D.L. The Genomes of Sheep pox and Goat pox Viruses. J. Virol. 2002. 76, 6054-6061. [CrossRef]
- WOAH (World Organization of Animal Health). https://www.woah.org/app/uploads/2024/04/asf-report49.pdf.
- Gonzales, W.; Moreno, C.; Duran, U.; Henao, N.; Bencosme, M.; Lora, P.; Reyes, R.; Núñez, R.; De Gracia, A.; Perez, A.M. African swine fever in Dominican Republic. Transbound. Emerg. Dis. 2021, 68, 3018-3019. [CrossRef]
- Dixon, L.K.; Chapman, D.A.; Netherton, C.L.; Upton, C. African swine fever virus replication and genomics. Virus Res. 2013. 173, 3-14. [CrossRef]
- Mahapatra, M.; Howson, E.; Fowler, V.; Batten, C.; Flannery, J.; Selvaraj, M.; Parida, S. Rapid Detection of Peste des Petits Ruminants Virus (PPRV) Nucleic Acid Using a Novel Low-Cost Reverse Transcription Loop-Mediated Isothermal Amplification (RT-LAMP) Assay for Future Use in Nascent PPR Eradication Program. Viruses, 2019, 11, 699. [CrossRef]
- Batten, C.A.; Banyard, A.C.; King, D.P.; Henstock, M.R.; Edwards, L.; Sanders, A.; Buczkowski, H.; Oura, C.C.L.; Barrett, T. A real time RT-PCR assay for the specific detection of Peste des petits ruminants virus. J. Virol. Methods 2011, 171, 401-404. [CrossRef]
- Balinsky, C.A.; Delhon, G.; Smoliga, G.; Parat, M.; French, R.A.; Geary, S.J.; Rock, D.L.; Rodriguez, L.L. Rapid Preclinical Detection of Sheep pox Virus by a Real-Time PCR Assay. J. Clin. Microbiol. 2008. 46, 438-442. [CrossRef]
- Das, A.; Deng, M.Y.; Babiuk, S.; McIntosh, M.T. Modification of two capripox virus quantitative real-time PCR assays to improve diagnostic sensitivity and include beta-actin as an internal positive control. J. Vet. Diag. Invest. 2017. 29, 351-356.
- Das, A.; Babiuk, S.; McIntosh, M.T. Development of a Loop-Mediated Isothermal Amplification Assay for Rapid Detection of Capripoxviruses. J. Clin. Microbiol. 2012. 50, 1613-1620. http://jcm.asm.org/content/50/5/1613.
- Zsak, L.; Borca, M.V.; Risatti, G.R.; Zsak, A.; French, R.A.; Lu, Z.; Kutish, G.F.; Neilan, J.G.; Callahan, J.D.; Nelson, W.M.; Rock, D.L. Preclinical diagnosis of African swine fever in contact-exposed swine by a real-time PCR assay. J. Clin. Microbiol, 2005. 43, 112-119. [CrossRef]
- Wang, Y.; Xu, L.; Noll, L.; Stoy, C.; Porter, E.; Fu, J.; Feng, Y.; Peddireddi, L.; Liu, X.; Dodd, K.A.; Jia, W.; Bai, J. Development of a real-time PCR assay for detection of African swine fever virus with an endogenous internal control. Transbound. Emerg. Dis. 2020, 67, 2446-2454. [CrossRef]
- Wang, D.; Yu, J.; Wang, Y.; Zhang, M.; Li, P.; Liu, M.; Liu, Y. (2019). Development of a real-time loop-mediated isothermal amplification (LAMP) assay and visual LAMP assay for detection of African swine fever virus (ASFV). J. Virol. Methods, 2019, 276, 113775. [CrossRef]
- Xu, W.; Xu, N.; Zhang, M.; Wang, Y.; Ling, G.; Yuan, Y.; Zhang, P. Nanotraps based on multifunctional materials for trapping and enrichment. Acta Biomaterialia 2022, 138, 57-72. [CrossRef]
- Shafagati, N.; Fite K, Patanarut, A.; Baer A, Pinkham, C.; An S.; Foote, B.; Lepene, B.; Kehn-Hall, K. Enhanced detection of respiratory pathogens with nanotrap particles. Virulence 2016, 7, 756-769. [CrossRef]
- Das, A.; Ahmed, Z.; Xu, L.; Jia, W. Assessment and Verification of Chemical Inactivation of Peste des Petits Ruminants Virus by Virus Isolation Following Virus Capture using Nanotrap® Magnetic Virus Particles. Microbiol. Spectrum 2023, 11. [CrossRef]
- Cottral, G.E. 1978. Manual of Standardized Methods for Veterinary Microbiology. 1st Edition, Cornell University Press, Ithaca, NY 1978, pp. 81-82.
- Shafagati N, Narayanan A, Baer A, Fite K, Pinkham C, Charles Bailey C, Kashanchi F, Lepene B, Kehn-Hall K. 2013. The use of Nanotrap particles as a sample enrichment method to enhance the detection of Rift Valley Fever Virus. PLOS One, 2023, v7: e2296. [CrossRef]
- Barclay, R.A.; Akhrymuk, I.; Patnaik1, A.; Callahan, V.; Lehman, C.; Andersen, P.; Barbero, R.; Barksdale, S.; Dunlap, R.; Goldfarb, D.; Jones-Roe, T.; Kelly, R.; Kim, B.; Miao, S.; Munns, A.; Munns, D.; Patel, S.; Porter, E.; Ramsey, R.; Sahoo, S.; Swahn, O.; Warsh, J.; Kehn-Hall, K.; Lepene, B. Hydrogel particles improve detection of SARS-CoV-2 RNA from multiple sample types. Scientific Reports 2020, 10, 22425. [CrossRef]
- Akhrymuk, I.; Lin, S.C.; Sun, M.; Patnaik, A.; Lehman, C.; Altamura, L.; Minoguw, T.; Lepene, B.; van Hoek, M.L.; Kehn-Hall, K. Magnetic Nanotrap particles preserve the stability of Venezuelan Equine Encephalitis virus in blood for laboratory detection. Frontiers in Veterinary Science 2020, 6, article 509. [CrossRef]
- Rådström, P.; Knutsson, R.; Wolfs, P.; Lövenklev, M.; Löfström, C. Pre-PCR Processing. Molecular Biotechnology 2004. 26, 133-146. [CrossRef]
- Schrader, C.; Schielke, A.; Ellerbroek, L.; Johne, R. PCR inhibitors – occurrence, properties, and removal. J. Applied Microbiol. 2012, 113, 1014-1026. https://doi:10.1111/j.1365-2672.2012.05384.x.
- Kermekchiev, M.B.; Kirilova, L.I.; Erika E. Vail, E.E.; Wayne M. Barnes, W.M. Mutants of Taq DNA polymerase resistant to PCR inhibitors allow DNA amplification from whole blood and crude soil samples. Nucleic Acids Research 2009, 37, e40. [CrossRef]
- Śpibida, M.; Krawczyk, B.; Olszewski, M.; Kur, J. Modified DNA polymerases for PCR troubleshooting. J. Appl. Genetics 2017, 58, 133-142. [CrossRef]
- Abu Al-Saud, W.; Rådström, P. Capacity of nine thermostable DNA polymerases to mediate DNA amplification in the presence of PCR-inhibiting samples. Appl. Env. Microbiol. 1998. 64, 3748-3753. [CrossRef]
- Das, A.; Spackman, E.; Pantin-Jackwood, M.J.; Suarez, D.L. Removal of real-time reverse transcription polymerase chain reaction (RT-PCR) inhibitors associated with cloacal swab samples and tissues for improved diagnosis of Avian influenza virus by RT-PCR. J. Vet. Diag. Invest. 2009, 21, 771-778. [CrossRef]
- Das, A.; Spackman, E.; Senne, D.; Pedersen, J.; Suarez, D.L. Development of an Internal Positive Control for Rapid Diagnosis of Avian Influenza Virus Infections by Real-Time Reverse transcription-PCR with Lyophilized Reagents. J. Clin. Microbiol. 2006, 44, 3065-3073. https://doi:10.1128/JCM.00639-06.
- Liu, P.; Guo, L.; Cavallo, M.; Cantrell, C.; Hilton, S-P.; Nguyen, A.; Long, A.; Dunbar, J.; Barbero, R.; Barclay, R.; Sablon III, O.; Wolfe, M.; Lepene, B.; Moe, C. Comparison of Nanotrap® Microbiome A Particles, membrane filtration, and skim milk workflows for SARS-CoV-2 concentration in wastewater. Frontiers in Microbiol. 2023. [CrossRef]
- Ahmed, W.; Bivins, A.; Korajkic, A.; Metcalfe, S.; Smith, W.J.M.; Simpson, S.L. Comparative analysis of Adsorption-Extraction (AE) and Nanotrap® Magnetic Virus Particles (NMAP) workflows for the recovery of endogenous enveloped and non-enveloped viruses in wastewater. Sc. Total Env. 2023, 160072. http://dx.doi.org/10.1016/j.scitotenv.2022.160072.
- Thakali, O.; Shalina Shahin, S.; Sherchan, S.P. Wastewater Surveillance of SARS-CoV-2 RNA in a Prison Facility. Water 2024, 16, 570. [CrossRef]
- Sharma, V.; Takamura, H.; Biyani, M.; Honda, R. Real-Time On-Site Monitoring of Viruses in Wastewater Using Nanotrap Particles and RICCA Technologies. Biosensors 2024, 14, 115. [CrossRef]
- Ahmed, W.; Smith, W.J.M.; Ananda Tiwari, A.; Bivins, A.; Simpson, S.L. Unveiling indicator, enteric, and respiratory viruses in aircraft lavatory wastewater using adsorption-extraction and Nanotrap® Microbiome A Particles workflow. Sc. Total Env. 2023, 896, 165007. http://dx.doi.org/10.1016/j.scitotenv.2023.165007.
- Brighton, K.; Fish, S.; Wu, H.; Vigil, K.; Aw, T.G. Targeted community wastewater surveillance for SARS-CoV-2 and Mpox virus during a festival mass-gathering event. Sc. Total Env. 2024, 906, 167443. [CrossRef]
- Toribio-Avedillo, D.; Gómez-Gómez, C.; Sala-Comorera, L.; Rodríguez-Rubio, L.; Carcereny, A.; García-Pedemonte, D.; Pintó, R.M.; Guix, S.; Galofré, B.; Bosch, A.; Susana Merino, S.; Muniesa, M. Monitoring influenza and respiratory syncytial virus in wastewater. Beyond COVID-19. Sc. Total Env. 2023, 892, 164495. http://dx.doi.org/10.1016/j.scitotenv.2023.164495.
- Ahmed, W.; Liua, Y.; Smith, W.; Ingall, W.; Belby, M.; Bivins, A.; Bertsch, P.; Williams, D.; Richards, K.; Simpson, S. Harnessing Wastewater Surveillance to Detect Viral Diseases in Livestock Settings. Sc. Total Env. 2024, in press. [CrossRef]
- Shafagati, N.; Patanarut, A.; Luchini, A.; Lundberg, L.; Bailey, C.; Petricoin III, E’; Liotta, L.; Narayanan, A.; Lepene, B.; Kehn-Hall, K. The use of Nanotrap particles for biodefense and emerging infectious disease diagnostics. Pathogens and Diseases 2014, 71, 164-176. [CrossRef]
| PPRV-Egypt | PPRV RT-qPCR (Cycle threshold) |
Virus titer (TCID50/ml) |
|
|
Serial dilution Untreated* |
Undiluted | 17.333 | 5.5 |
| 10-1 Dilution | 20.374 | 4.3 | |
| 10-2 Dilution | 22.914 | 3.2 | |
| 10-3 Dilution | 26.582 | 2.4 | |
| 10-4 Dilution | 30.440 | Below LOD | |
| 10-5 Dilution | 32.318 | Below LOD | |
| 10-6 - 10-10 Dilution | Undetectable | Below LOD | |
| Serial dilution Treated# |
Undiluted | 18.547 | 5.1 |
| 10-1 Dilution | 20.991 | 4.5 | |
| 10-2 Dilution | 24.846 | 3.5 | |
| 10-3 Dilution | 27.147 | 2.4 | |
| 10-4 Dilution | 29.810 | 1.9 | |
| 10-5 Dilution | 32.772 | Below LOD | |
| 10-6 - 10-10 Dilution | Undetectable | Below LOD | |
| Working stock/ Dilution |
Cycle Threshold (qPCR/RT-qPCR) | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| PPRV | GTPV | SPPV | LSDV | ASFV | ||||||
| UT# | Treated¶ | UT | Treated | UT | Treated | UT | Treated | UT | Treated | |
| WS* (0:0) | 32.279 | ND§ | 31.754 | ND | 31.141 | ND | 32.700 | ND | 32.289 | ND |
| WS 1:10 | 35.285 | 35.543 | 34.281 | 34.455 | 34.295 | 34.298 | 35.180 | 35.756 | 35.958 | 35.289 |
| WS 1:25 | UD† | 35.869 | UD | 34.722 | UD | 34.208 | UD | 35.980 | UD | 35.448 |
| WS 1:50 | UD | 35.577 | UD | 34.929 | UD | 34.627 | UD | 35.237 | UD | 35.238 |
| WS 1:100 | UD | 35.587 | UD | 35.993 | UD | 34.124 | UD | 35.970 | UD | 35.316 |
| WS1:250 | UD | 35.554 | UD | 34.889 | UD | 34.688 | UD | 35.605 | UD | 35.636 |
| WS 1:500 | UD | 35.931 | UD | 34.559 | UD | 35.943 | UD | 35.642 | UD | 35.920 |
| WS 1:1000 | UD | 35.339 | UD | 34.986 | UD | 34.995 | UD | 35.192 | UD | 35.509 |
| Virus | Cycle threshold (spiked virus) of qPCR/RT-qPCR with or without virus capture or HGB treatment |
|||||||
|---|---|---|---|---|---|---|---|---|
| PBS | EWB (0:0) |
EWB (1:10) | EWB (0:0) + HGB¶ (Supernatant) |
|||||
| Untreated* | Treated# | Untreated | Treated | Untreated | Treated | Untreated | Treated | |
| GTPV | 16.907 | 17.928 | 21.033 | 21.375 | 19.235 | 18.174 | 21.439 | 22.177 |
| SPPV | 17.956 | 18.350 | 20.231 | 20.669 | 20.056 | 19.523 | 21.617 | 22.221 |
| LSDV | 17.776 | 18.711 | 20.623 | 19.864 | 19.742 | 18.995 | 21.731 | 22.108 |
| PPRV | 18.561 | 19.552 | 20.615 | 20.718 | 20.145 | 19.630 | 21.846 | 22.604 |
| ASFV | 19.915 | 20.155 | 22.033 | 22.298 | 19.444 | 20.603 | 22.525 | 23.138 |
|
Virus (spiked) |
EWB (host) |
EWB (dilution/treatment) |
NMVPs Added (Y/N) |
CPE (VI) | Cycle threshold (qPCR/RT-qPCR) |
||
| 1P | 2P | 1P | 2P | ||||
| GTPV | Goat | 0:0 | Y | N | Y | 23.342 | 18.045 |
| 1:10 | Y | Y | Y | 17.631 | 16.495 | ||
| HGB (Sup) | Y | Y | Y | 18.569 | 17.663 | ||
| SPPV | Sheep | 0:0 | Y | N | Y | 23.657 | 18.322 |
| 1:10 | Y | Y | Y | 17.913 | 16.724 | ||
| HGB (Sup) | Y | Y | Y | 17.812 | 16.583 | ||
| LSDV | Cattle | 0:0 | Y | N | Y | 23.644 | 18.509 |
| 1:10 | Y | Y | Y | 19.766 | 18.609 | ||
| HGB (Sup) | Y | Y | Y | 18.578 | 17.455 | ||
| PPRV | Goat | 0:0 | Y | N | Y | 25.257 | 20.311 |
| 1:10 | Y | Y | Y | 19.984 | 18.766 | ||
| HGB (Sup) | Y | Y | Y | 20.572 | 19.427 | ||
| ASFV | Swine | 0:0 | Y | N | Y | 23.712 | 18.116 |
| 1:10 | Y | Y | Y | 17.913 | 16.893 | ||
| HGB (Sup) | Y | Y | Y | 18.579 | 17.398 | ||
| Virus | Host animal | # of Animals | # of Specimens |
Infection route |
Ct (qPCR/RT-qPCR) | Diagnostic sensitivity (%) |
|||
| Untreated (-NMAPs) |
Treated (+NMAPs) |
||||||||
| # Pos | # Neg | # Pos | # Neg | ||||||
| PPRV | Goat | 8 | 28 | EI# | 28 | 0 | 28 | 0 | 100 |
| SPPV | Sheep | 6 | 24 | 24 | 0 | 24 | 0 | 100 | |
| ASFV | Swine | 20 | 20 | 20 | 0 | 20 | 0 | 100 | |
| ASFV | Swine | 20 | 20 | NI¶ | 20 | 0 | 20 | 0 | 100 |
| Virus | Virus suspension (PBS) Single (S)/Mixed (M) |
Cycle threshold (Ct) (qPCR/RT-qPCR) |
|---|---|---|
| CaPV | GTPV (S) | 27.420 (GTPV) |
| GTPV + PPRV + ASFV (M) | 27.078 (GTPV) | |
| SPPV (S) | 27.182 (SPPV) | |
| SPPV + PPRV + ASFV (M) | 27.470 (SPPV) | |
| LSDV (S) | 29.754 (LSDV) | |
| LSDV + PPRV + ASFV (M) | 29.116 (LSDV) | |
| PPRV | PPRV (S) | 29.001 (PPRV) |
| PPRV + GTPV + ASFV (M) | 29.157 (PPRV) | |
| PPRV + SPPV + ASFV (M) | 29.893 (PPRV) | |
| PPRV + LSDV + ASFV (M) | 29.846 (PPRV) | |
| ASFV | ASFV (S) | 31.659 (ASFV) |
| ASFV + GTPV + PPRV (M) | 31.397 (ASFV) | |
| ASFV + SPPV + PPRV (M) | 31.785 (ASFV) | |
| ASFV + LSDV + PPRV (M) | 31.307 (ASFV) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




