Submitted:
08 August 2024
Posted:
12 August 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Advantages of Mass Spectrometry in Clinical Applications
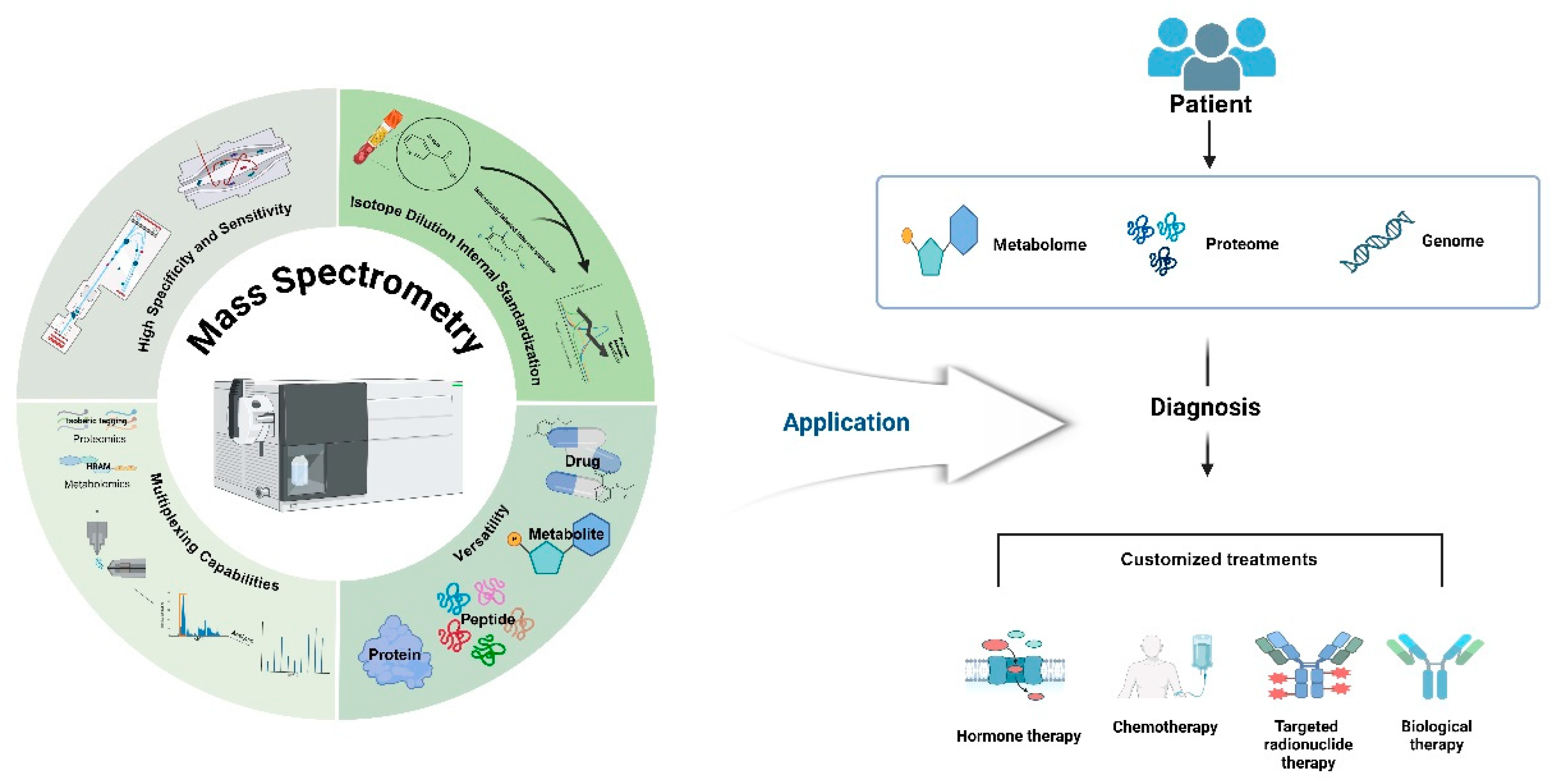
2.1. High Specificity and Sensitivity
2.2. Multiplexing Capabilities
2.3. Versatility
2.4. Isotope Dilution Internal Standardization
3. Applications in Biomarker Discovery and Personalized Medicine
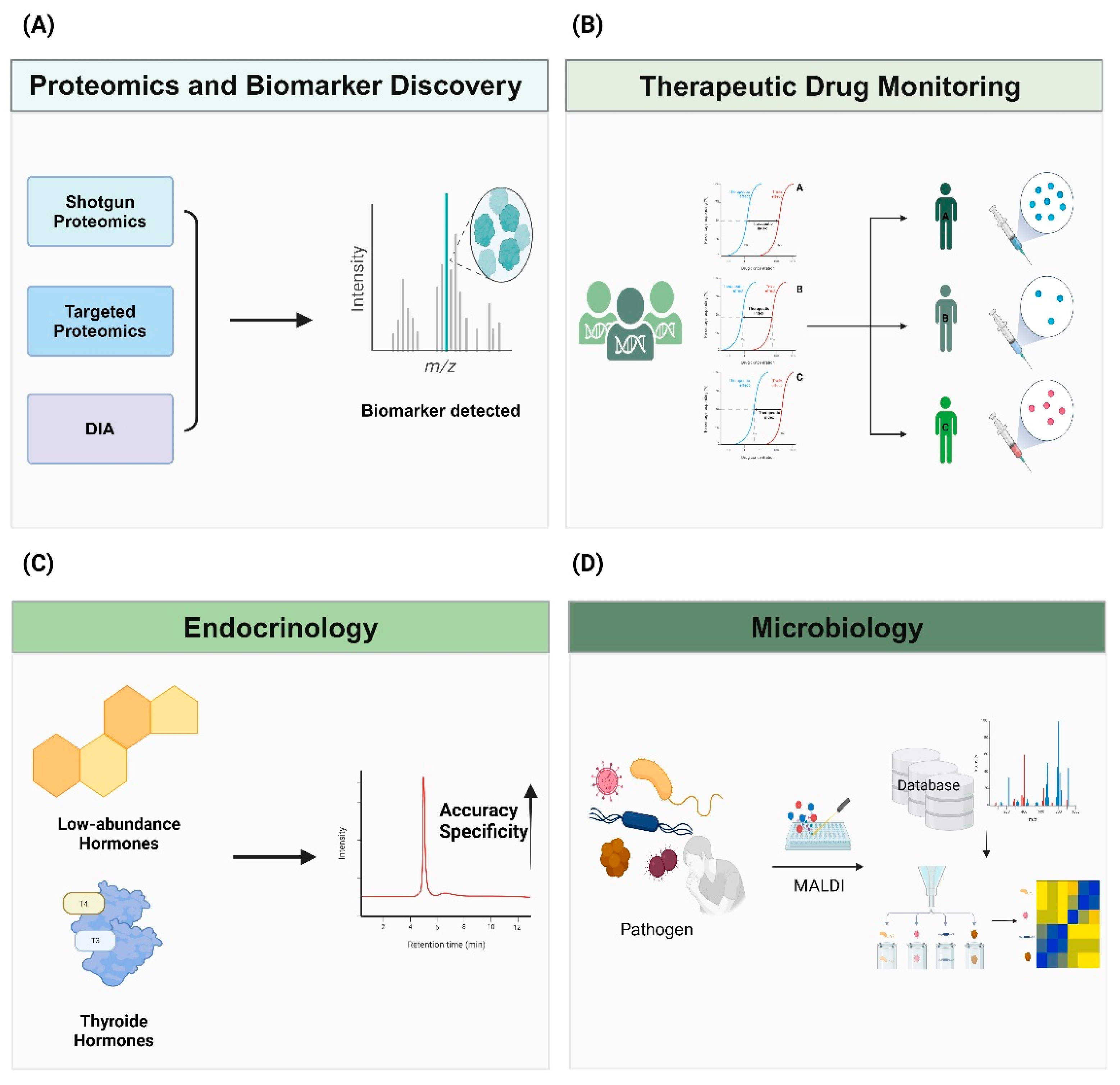
3.1. Proteomics and Biomarker Discovery
3.2. Therapeutic Drug Monitoring
3.3. Endocrinology
3.4. Microbiology
4. Enhancing Accessibility and Integration of Mass Spectrometry in Clinical Laboratories
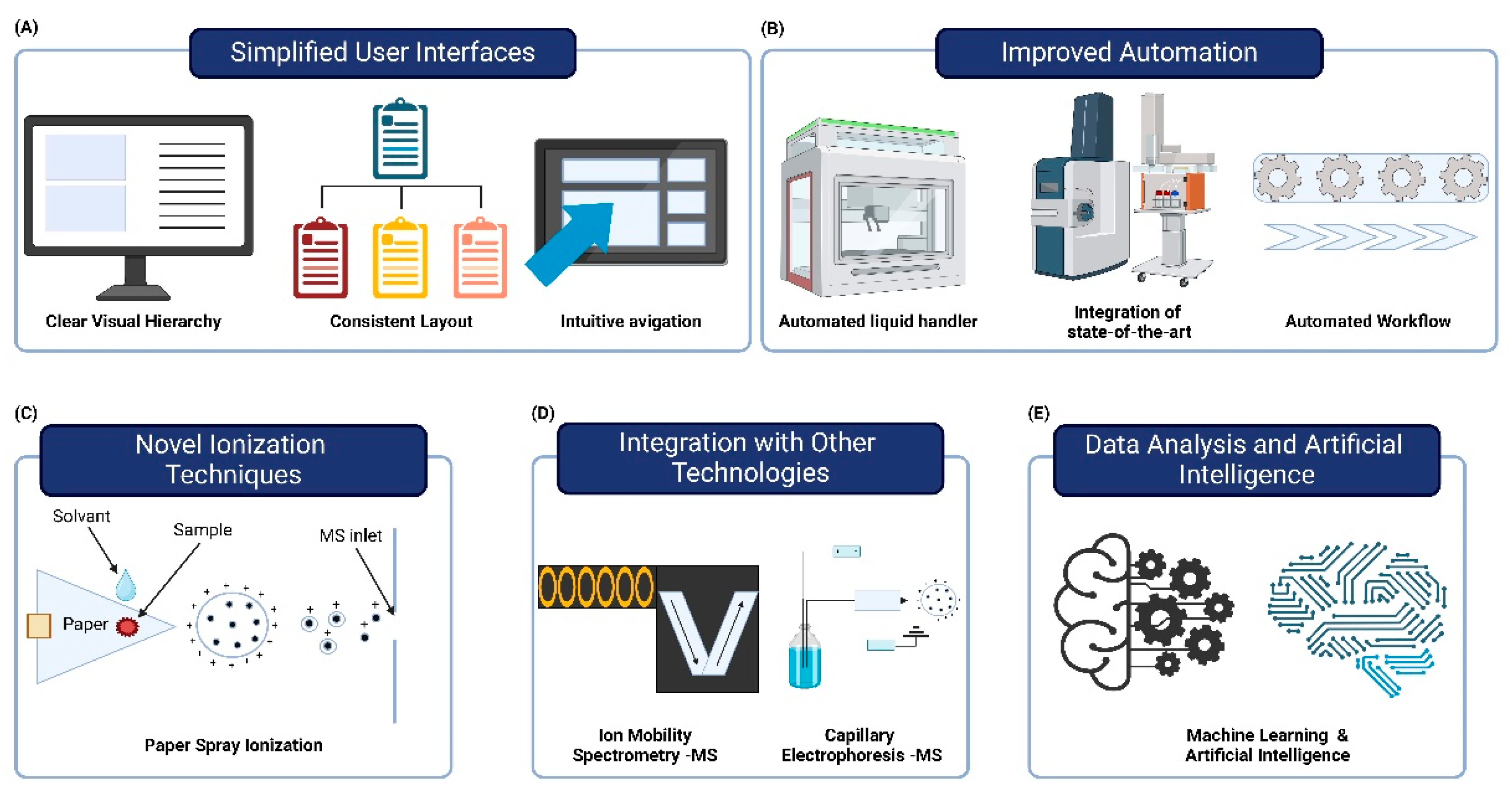
4.1. Simplified User Interfaces
4.2. Improved Automation
4.3. Novel Ionization Techniques
4.4. Integration with Other Technologies
4.5. Data Analysis and Artificial Intelligence
5. Challenges and Opportunities in Implementing Mass Spectrometry in Clinical Laboratories
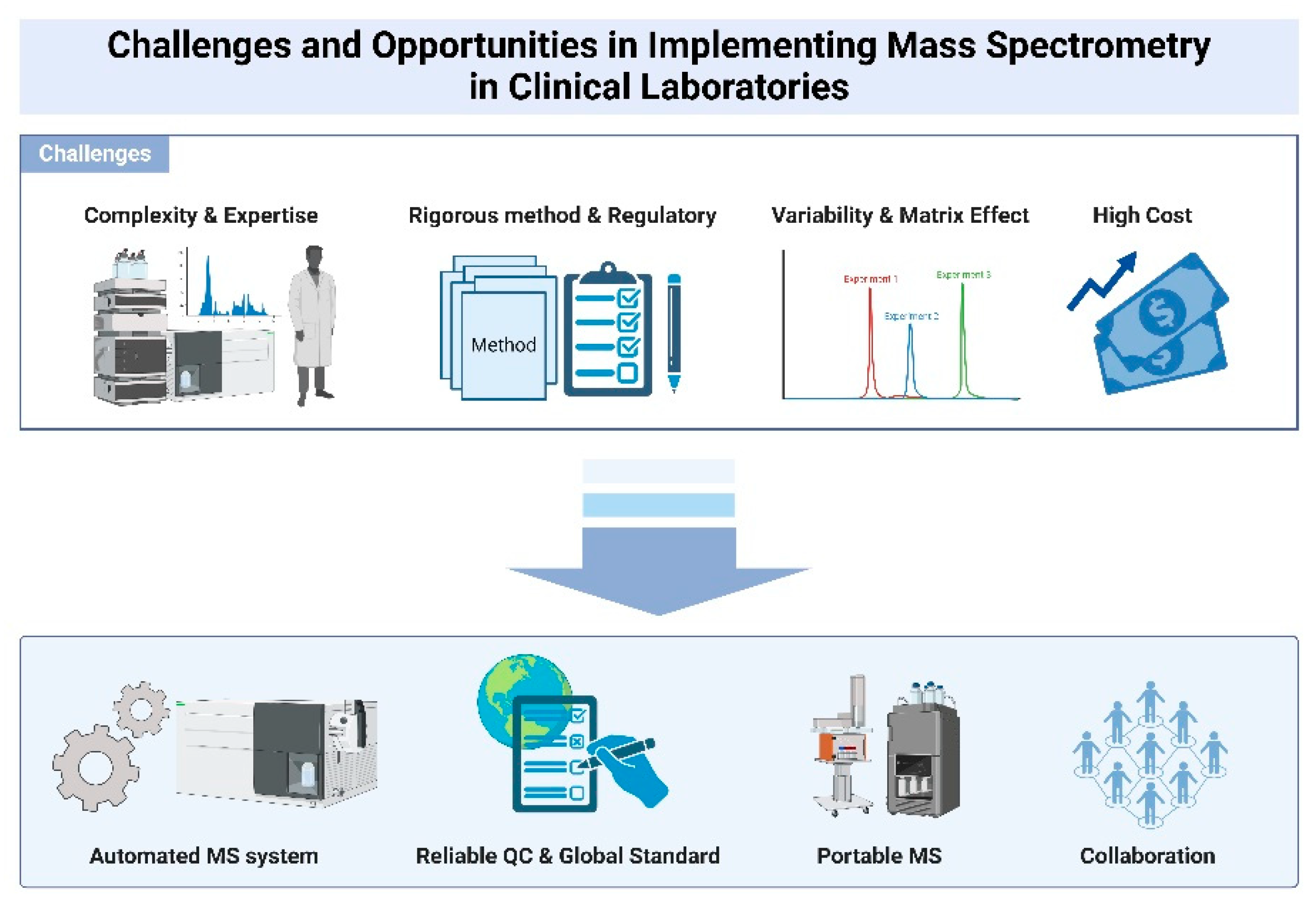
5.1. Complexity and Expertise
5.2. Standardization
5.3. Automation
5.4. Cost
6. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Zhou, X.; Zhang, W.; Ouyang, Z. Recent advances in on-site mass spectrometry analysis for clinical applications. Trends Analyt Chem 2022, 149, 116548. [Google Scholar] [CrossRef]
- Wenk, D.; Zuo, C.; Kislinger, T.; Sepiashvili, L. Recent developments in mass-spectrometry-based targeted proteomics of clinical cancer biomarkers. Clin Proteomics 2024, 21, 6. [Google Scholar] [CrossRef] [PubMed]
- Thomas, S.N.; French, D.; Jannetto, P.J.; Rappold, B.A.; Clarke, W.A. Liquid chromatography-tandem mass spectrometry for clinical diagnostics. Nat Rev Methods Primers 2022, 2, 96. [Google Scholar] [CrossRef]
- Cui, J.J.; Wang, L.Y.; Tan, Z.R.; Zhou, H.H.; Zhan, X.; Yin, J.Y. Mass Spectrometry-Based Personalized Drug Therapy. Mass Spectrom Rev 2020, 39, 523–552. [Google Scholar] [CrossRef]
- Birhanu, A.G. Mass spectrometry-based proteomics as an emerging tool in clinical laboratories. Clin Proteomics 2023, 20, 32. [Google Scholar] [CrossRef] [PubMed]
- Rischke, S.; Hahnefeld, L.; Burla, B.; Behrens, F.; Gurke, R.; Garrett, T.J. Small molecule biomarker discovery: Proposed workflow for LC-MS-based clinical research projects. J Mass Spectrom Adv Clin Lab 2023, 28, 47–55. [Google Scholar] [CrossRef]
- Grebe, S.K.; Singh, R.J. LC-MS/MS in the Clinical Laboratory - Where to From Here? Clin Biochem Rev 2011, 32, 5–31. [Google Scholar] [PubMed]
- Beck, O.; Olin, A.C.; Mirgorodskaya, E. Potential of Mass Spectrometry in Developing Clinical Laboratory Biomarkers of Nonvolatiles in Exhaled Breath. Clin Chem 2016, 62, 84–91. [Google Scholar] [CrossRef]
- Koomen, D.C.; May, J.C.; McLean, J.A. Insights and prospects for ion mobility-mass spectrometry in clinical chemistry. Expert Rev Proteomics 2022, 19, 17–31. [Google Scholar] [CrossRef]
- Chouinard, C.D.; Wei, M.S.; Beekman, C.R.; Kemperman, R.H.; Yost, R.A. Ion Mobility in Clinical Analysis: Current Progress and Future Perspectives. Clin Chem 2016, 62, 124–133. [Google Scholar] [CrossRef]
- Huffman, R.G.; Leduc, A.; Wichmann, C.; Di Gioia, M.; Borriello, F.; Specht, H.; Derks, J.; Khan, S.; Khoury, L.; Emmott, E.; et al. Prioritized mass spectrometry increases the depth, sensitivity and data completeness of single-cell proteomics. Nat Methods 2023, 20, 714–722. [Google Scholar] [CrossRef]
- Arul, A.B.; Robinson, R.A.S. Sample Multiplexing Strategies in Quantitative Proteomics. Anal Chem 2019, 91, 178–189. [Google Scholar] [CrossRef]
- Recchia, M.J.J.; Baumeister, T.U.H.; Liu, D.Y.; Linington, R.G. MultiplexMS: A Mass Spectrometry-Based Multiplexing Strategy for Ultra-High-Throughput Analysis of Complex Mixtures. Anal Chem 2023, 95, 11908–11917. [Google Scholar] [CrossRef]
- Tsumagari, K.; Isobe, Y.; Imami, K.; Arita, M. Exploring protein lipidation by mass spectrometry-based proteomics. J Biochem 2024, 175, 225–233. [Google Scholar] [CrossRef]
- Egertson, J.D.; Kuehn, A.; Merrihew, G.E.; Bateman, N.W.; MacLean, B.X.; Ting, Y.S.; Canterbury, J.D.; Marsh, D.M.; Kellmann, M.; Zabrouskov, V.; et al. Multiplexed MS/MS for improved data-independent acquisition. Nat Methods 2013, 10, 744–746. [Google Scholar] [CrossRef]
- Swiner, D.J.; Jackson, S.; Burris, B.J.; Badu-Tawiah, A.K. Applications of Mass Spectrometry for Clinical Diagnostics: The Influence of Turnaround Time. Anal Chem 2020, 92, 183–202. [Google Scholar] [CrossRef]
- Holbrook, J.H.; Kemper, G.E.; Hummon, A.B. Quantitative mass spectrometry imaging: therapeutics & biomolecules. Chem Commun (Camb) 2024, 60, 2137–2151. [Google Scholar] [CrossRef]
- Banerjee, S. Empowering Clinical Diagnostics with Mass Spectrometry. ACS Omega 2020, 5, 2041–2048. [Google Scholar] [CrossRef]
- Zhang, G.; Annan, R.S.; Carr, S.A.; Neubert, T.A. Overview of peptide and protein analysis by mass spectrometry. Curr Protoc Mol Biol 2014, 108, 10 21 11–10 21 30. [Google Scholar] [CrossRef]
- Lanekoff, I.; Stevens, S.L.; Stenzel-Poore, M.P.; Laskin, J. Matrix effects in biological mass spectrometry imaging: identification and compensation. Analyst 2014, 139, 3528–3532. [Google Scholar] [CrossRef]
- Zheng, X.; Chen, X.; Liu, T.; Jiang, J.; Cui, X.; Zhao, Q.; Hu, P. Liquid chromatography-tandem mass spectrometry methods for quantification of roxadustat (FG-4592) in human plasma and urine and the applications in two clinical pharmacokinetic studies. J Chromatogr B Analyt Technol Biomed Life Sci 2022, 1203, 123274. [Google Scholar] [CrossRef] [PubMed]
- Cheng, W.L.; Markus, C.; Lim, C.Y.; Tan, R.Z.; Sethi, S.K.; Loh, T.P.; Protocols, I.W.G.o.M.E. Calibration Practices in Clinical Mass Spectrometry: Review and Recommendations. Ann Lab Med 2023, 43, 5–18. [Google Scholar] [CrossRef] [PubMed]
- Villanueva, J.; Carrascal, M.; Abian, J. Isotope dilution mass spectrometry for absolute quantification in proteomics: concepts and strategies. J Proteomics 2014, 96, 184–199. [Google Scholar] [CrossRef] [PubMed]
- Faria, S.S.; Morris, C.F.; Silva, A.R.; Fonseca, M.P.; Forget, P.; Castro, M.S.; Fontes, W. A Timely Shift from Shotgun to Targeted Proteomics and How It Can Be Groundbreaking for Cancer Research. Front Oncol 2017, 7, 13. [Google Scholar] [CrossRef] [PubMed]
- Nakayasu, E.S.; Gritsenko, M.; Piehowski, P.D.; Gao, Y.; Orton, D.J.; Schepmoes, A.A.; Fillmore, T.L.; Frohnert, B.I.; Rewers, M.; Krischer, J.P.; et al. Tutorial: best practices and considerations for mass-spectrometry-based protein biomarker discovery and validation. Nat Protoc 2021, 16, 3737–3760. [Google Scholar] [CrossRef] [PubMed]
- Amiri-Dashatan, N.; Koushki, M.; Abbaszadeh, H.A.; Rostami-Nejad, M.; Rezaei-Tavirani, M. Proteomics Applications in Health: Biomarker and Drug Discovery and Food Industry. Iran J Pharm Res 2018, 17, 1523–1536. [Google Scholar] [PubMed]
- Azevedo, R.; Jacquemin, C.; Villain, N.; Fenaille, F.; Lamari, F.; Becher, F. Mass Spectrometry for Neurobiomarker Discovery: The Relevance of Post-Translational Modifications. Cells 2022, 11. [Google Scholar] [CrossRef]
- Kang, J.S.; Lee, M.H. Overview of therapeutic drug monitoring. Korean J Intern Med 2009, 24, 1–10. [Google Scholar] [CrossRef]
- Liu, L.; Zhang, L.; Zheng, X.; Liu, X.; Liu, W.; Wu, J. LC-MS/MS-based multiplex antibacterial platform for therapeutic drug monitoring in intensive care unit patients. Front Pharmacol 2023, 14, 1116071. [Google Scholar] [CrossRef]
- Barco, S.; Mesini, A.; Barbagallo, L.; Maffia, A.; Tripodi, G.; Pea, F.; Saffioti, C.; Castagnola, E.; Cangemi, G. A liquid chromatography-tandem mass spectrometry platform for the routine therapeutic drug monitoring of 14 antibiotics: Application to critically ill pediatric patients. J Pharm Biomed Anal 2020, 186, 113273. [Google Scholar] [CrossRef]
- Gaspar, V.P.; Ibrahim, S.; Zahedi, R.P.; Borchers, C.H. Utility, promise, and limitations of liquid chromatography-mass spectrometry-based therapeutic drug monitoring in precision medicine. J Mass Spectrom 2021, 56, e4788. [Google Scholar] [CrossRef]
- van der Kleij, M.B.A.; Guchelaar, N.A.D.; Mathijssen, R.H.J.; Versluis, J.; Huitema, A.D.R.; Koolen, S.L.W.; Steeghs, N. Therapeutic Drug Monitoring of Kinase Inhibitors in Oncology. Clin Pharmacokinet 2023, 62, 1333–1364. [Google Scholar] [CrossRef]
- Vogeser, M.; Parhofer, K.G. Liquid chromatography tandem-mass spectrometry (LC-MS/MS)--technique and applications in endocrinology. Exp Clin Endocrinol Diabetes 2007, 115, 559–570. [Google Scholar] [CrossRef]
- Holmes, D.T. A brief update on mass spectrometry applications to routine clinical endocrinology. Clin Mass Spectrom 2019, 13, 18–20. [Google Scholar] [CrossRef]
- French, D. Clinical utility of laboratory developed mass spectrometry assays for steroid hormone testing. J Mass Spectrom Adv Clin Lab 2023, 28, 13–19. [Google Scholar] [CrossRef]
- Taylor, A.E.; Keevil, B.; Huhtaniemi, I.T. Mass spectrometry and immunoassay: how to measure steroid hormones today and tomorrow. Eur J Endocrinol 2015, 173, D1–D12. [Google Scholar] [CrossRef]
- Singhal, N.; Kumar, M.; Kanaujia, P.K.; Virdi, J.S. MALDI-TOF mass spectrometry: an emerging technology for microbial identification and diagnosis. Front Microbiol 2015, 6, 791. [Google Scholar] [CrossRef]
- Grenga, L.; Pible, O.; Armengaud, J. Pathogen proteotyping: A rapidly developing application of mass spectrometry to address clinical concerns. Clin Mass Spectrom 2019, 14 Pt A, 9–17. [Google Scholar] [CrossRef]
- Calderaro, A.; Chezzi, C. MALDI-TOF MS: A Reliable Tool in the Real Life of the Clinical Microbiology Laboratory. Microorganisms 2024, 12. [Google Scholar] [CrossRef]
- Al-Manei, K.; Ghorbani, M.; Naud, S.; Al-Manei, K.K.; Sobkowiak, M.J.; Lund, B.; Hazirolan, G.; Sallberg Chen, M.; Ozenci, V. Clinical Microbial Identification of Severe Oral Infections by MALDI-TOF Mass Spectrometry in Stockholm County: an 11-Year (2010 to 2020) Epidemiological Investigation. Microbiol Spectr 2022, 10, e0248722. [Google Scholar] [CrossRef] [PubMed]
- Diehl, C.; Martins, A.; Almeida, A.; Silva, T.; Ribeiro, O.; Santinha, G.; Rocha, N.; Silva, A.G. Defining Recommendations to Guide User Interface Design: Multimethod Approach. JMIR Hum Factors 2022, 9, e37894. [Google Scholar] [CrossRef]
- Silva, A.G.; Caravau, H.; Martins, A.; Almeida, A.M.P.; Silva, T.; Ribeiro, O.; Santinha, G.; Rocha, N.P. Procedures of User-Centered Usability Assessment for Digital Solutions: Scoping Review of Reviews Reporting on Digital Solutions Relevant for Older Adults. JMIR Hum Factors 2021, 8, e22774. [Google Scholar] [CrossRef] [PubMed]
- Fu, Q.; Murray, C.I.; Karpov, O.A.; Van Eyk, J.E. Automated proteomic sample preparation: The key component for high throughput and quantitative mass spectrometry analysis. Mass Spectrom Rev 2023, 42, 873–886. [Google Scholar] [CrossRef]
- Waldenmaier, H.E.; Gorre, E.; Poltash, M.L.; Gunawardena, H.P.; Zhai, X.A.; Li, J.; Zhai, B.; Beil, E.J.; Terzo, J.C.; Lawler, R.; et al. "Lab of the Future" horizontal line Today: Fully Automated System for High-Throughput Mass Spectrometry Analysis of Biotherapeutics. J Am Soc Mass Spectrom 2023, 34, 1073–1085. [Google Scholar] [CrossRef] [PubMed]
- Reilly, L.; Lara, E.; Ramos, D.; Li, Z.; Pantazis, C.B.; Stadler, J.; Santiana, M.; Roberts, J.; Faghri, F.; Hao, Y.; et al. A fully automated FAIMS-DIA mass spectrometry-based proteomic pipeline. Cell Rep Methods 2023, 3, 100593. [Google Scholar] [CrossRef]
- Yang, Q.; Wang, H.; Maas, J.D.; Chappell, W.J.; Manicke, N.E.; Cooks, R.G.; Ouyang, Z. Paper spray ionization devices for direct, biomedical analysis using mass spectrometry. Int J Mass Spectrom 2012, 312, 201–207. [Google Scholar] [CrossRef]
- Nguyen, T.M.H.; Song, W.Y.; Kim, T.Y. Characterization of Spray Modes and Factors Affecting the Ionization Efficiency of Paper Spray Ionization. Front Chem 2022, 10, 864184. [Google Scholar] [CrossRef] [PubMed]
- Chiang, S.; Zhang, W.; Ouyang, Z. Paper spray ionization mass spectrometry: recent advances and clinical applications. Expert Rev Proteomics 2018, 15, 781–789. [Google Scholar] [CrossRef] [PubMed]
- Feider, C.L.; Krieger, A.; DeHoog, R.J.; Eberlin, L.S. Ambient Ionization Mass Spectrometry: Recent Developments and Applications. Anal Chem 2019, 91, 4266–4290. [Google Scholar] [CrossRef]
- Helena, H.; Ivona, V.; Roman, R.; Frantisek, F. Current applications of capillary electrophoresis-mass spectrometry for the analysis of biologically important analytes in urine (2017 to mid-2021): A review. J Sep Sci 2022, 45, 305–324. [Google Scholar] [CrossRef]
- Mischak, H.; Coon, J.J.; Novak, J.; Weissinger, E.M.; Schanstra, J.P.; Dominiczak, A.F. Capillary electrophoresis-mass spectrometry as a powerful tool in biomarker discovery and clinical diagnosis: an update of recent developments. Mass Spectrom Rev 2009, 28, 703–724. [Google Scholar] [CrossRef]
- Ebbels, T.M.D.; van der Hooft, J.J.J.; Chatelaine, H.; Broeckling, C.; Zamboni, N.; Hassoun, S.; Mathe, E.A. Recent advances in mass spectrometry-based computational metabolomics. Curr Opin Chem Biol 2023, 74, 102288. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Jonassen, I.; Goksoyr, A. Machine Learning Approaches for Biomarker Discovery Using Gene Expression Data. In Bioinformatics, Helder, I.N., Ed.; Brisbane (AU), 2021.
- Beck, A.G.; Muhoberac, M.; Randolph, C.E.; Beveridge, C.H.; Wijewardhane, P.R.; Kenttamaa, H.I.; Chopra, G. Recent Developments in Machine Learning for Mass Spectrometry. ACS Meas Sci Au 2024, 4, 233–246. [Google Scholar] [CrossRef] [PubMed]
- Swan, A.L.; Mobasheri, A.; Allaway, D.; Liddell, S.; Bacardit, J. Application of machine learning to proteomics data: classification and biomarker identification in postgenomics biology. OMICS 2013, 17, 595–610. [Google Scholar] [CrossRef]
- Naji, Y.; Mahdaoui, M.; Klevor, R.; Kissani, N. Artificial Intelligence and Multiple Sclerosis: Up-to-Date Review. Cureus 2023, 15, e45412. [Google Scholar] [CrossRef]
- Bonacchi, R.; Filippi, M.; Rocca, M.A. Role of artificial intelligence in MS clinical practice. Neuroimage Clin 2022, 35, 103065. [Google Scholar] [CrossRef]
- Adam, G.; Rampasek, L.; Safikhani, Z.; Smirnov, P.; Haibe-Kains, B.; Goldenberg, A. Machine learning approaches to drug response prediction: challenges and recent progress. NPJ Precis Oncol 2020, 4, 19. [Google Scholar] [CrossRef]
- Boiko, D.A.; Kozlov, K.S.; Burykina, J.V.; Ilyushenkova, V.V.; Ananikov, V.P. Fully Automated Unconstrained Analysis of High-Resolution Mass Spectrometry Data with Machine Learning. J Am Chem Soc 2022, 144, 14590–14606. [Google Scholar] [CrossRef] [PubMed]
- Torun, F.M.; Virreira Winter, S.; Doll, S.; Riese, F.M.; Vorobyev, A.; Mueller-Reif, J.B.; Geyer, P.E.; Strauss, M.T. Transparent Exploration of Machine Learning for Biomarker Discovery from Proteomics and Omics Data. J Proteome Res 2023, 22, 359–367. [Google Scholar] [CrossRef]
- Abdelmoula, W.M.; Lopez, B.G.; Randall, E.C.; Kapur, T.; Sarkaria, J.N.; White, F.M.; Agar, J.N.; Wells, W.M.; Agar, N.Y.R. Peak learning of mass spectrometry imaging data using artificial neural networks. Nat Commun 2021, 12, 5544. [Google Scholar] [CrossRef]
- Chace, D.H. Mass spectrometry in the clinical laboratory. Chem Rev 2001, 101, 445–477. [Google Scholar] [CrossRef]
- Annesley, T.; Majzoub, J.; Hsing, A.; Wu, A.; Rockwood, A.; Mason, D. Mass spectrometry in the clinical laboratory: how have we done, and where do we need to be? Clin Chem 2009, 55, 1236–1239. [Google Scholar] [CrossRef] [PubMed]
- Clarke, W.; Rhea, J.M.; Molinaro, R. Challenges in implementing clinical liquid chromatography-tandem mass spectrometry methods--the light at the end of the tunnel. J Mass Spectrom 2013, 48, 755–767. [Google Scholar] [CrossRef]
- Mugueta, C.; Gonzalez, A.; Deza, S.; Roca, C.A.; Contreras, T.; Puig, N.; Nerea, V. Mass spectrometry in clinical protein laboratories. Adv Lab Med 2024, 5, 97–99. [Google Scholar] [CrossRef] [PubMed]
- Vesper, H.W.; Myers, G.L.; Miller, W.G. Current practices and challenges in the standardization and harmonization of clinical laboratory tests. Am J Clin Nutr 2016, 104 Suppl 3, 907S–912S. [Google Scholar] [CrossRef]
- Annesley, T.M.; Cooks, R.G.; Herold, D.A.; Hoofnagle, A.N. Clinical Mass Spectrometry-Achieving Prominence in Laboratory Medicine. Clin Chem 2016, 62, 1–3. [Google Scholar] [CrossRef] [PubMed]
- Bittremieux, W.; Tabb, D.L.; Impens, F.; Staes, A.; Timmerman, E.; Martens, L.; Laukens, K. Quality control in mass spectrometry-based proteomics. Mass Spectrom Rev 2018, 37, 697–711. [Google Scholar] [CrossRef]
- Rankin-Turner, S.; Heaney, L.M. Mass spectrometry in the clinical laboratory. A short journey through the contribution to the scientific literature by CCLM. Clin Chem Lab Med 2023, 61, 873–879. [Google Scholar] [CrossRef]
- Vicente, F.B.; Lin, D.C.; Haymond, S. Automation of chromatographic peak review and order to result data transfer in a clinical mass spectrometry laboratory. Clin Chim Acta 2019, 498, 84–89. [Google Scholar] [CrossRef]
- Lunt, A.M.; Fakhruldeen, H.; Pizzuto, G.; Longley, L.; White, A.; Rankin, N.; Clowes, R.; Alston, B.; Gigli, L.; Day, G.M.; et al. Modular, multi-robot integration of laboratories: an autonomous workflow for solid-state chemistry. Chem Sci 2024, 15, 2456–2463. [Google Scholar] [CrossRef]
- Junger, S.; Hoene, M.; Shipkova, M.; Danzl, G.; Schoberl, C.; Peter, A.; Lehmann, R.; Wieland, E.; Braitmaier, H. Automated LC-MS/MS: Ready for the clinical routine Laboratory? J Mass Spectrom Adv Clin Lab 2023, 30, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Pursell, M.E.; DeVor, A.; Awoyemi, O.; Valentine, S.J.; Li, P. Portable mass spectrometry system: instrumentation, applications, and path to 'omics analysis. Proteomics 2022, 22, e2200112. [Google Scholar] [CrossRef] [PubMed]
- Houfani, A.A.; Foster, L.J. Review of the Real and Sometimes Hidden Costs in Proteomics Experimental Workflows. Methods Mol Biol 2022, 2456, 1–14. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



