1. Introduction
Since the introduction of penicillin for treating bacterial infections, antibiotics have not only prevented and treated a variety of infectious diseases but have also revolutionized modern medicine by advancing clinical practices in surgery, organ transplantation, cancer chemotherapy, and the use of artificial devices. However, a significant threat has emerged: antibiotic resistance [
1]. The incidence of clinically pan-resistant and multi-drug resistant bacteria has continued to rise in recent years, particularly with the appearance of super-resistant strains, pushing antimicrobial therapy towards a critical drug-free situation [
1]. The rapid proliferation of bacterial drug resistance now poses a global health challenge [
2]. According to the Global Tuberculosis Report 2023, the estimated proportion of drug-resistant tuberculosis (TB) patients among new cases was 3.3% in 2022 [
3]. Additionally, the number of TB patients resistant to rifampicin and isoniazid, the most effective anti-tuberculosis drugs, continues to rise [
4,
5], highlighting a critical situation in the resistance of
M. tuberculosis [
6].
Phages provide innovative approaches for treating drug-resistant bacterial infections in humans due to their abundant sources [
7], host specificity, and superior bactericidal properties compared to antibiotics [
8]. Used alone or in combination with antibiotics, phages offer significant antimicrobial benefits and hold wide-ranging application potential [
9,
10,
11]. In recent years, there has been an increase in reports on the use of phages for treating
mycobacterium infections, and the effectiveness of
mycobacterium phages against various
mycobacterium infections has been successfully demonstrated [
12,
13].
In this study, we isolated and characterized a strain of mycobacterium phage named Henu3. We analyzed its biological properties and genome to provide a theoretical foundation for the design of novel M. tuberculosis treatments using phage therapy. The goal of this research is to contribute to the development of future antimicrobial products and offer a new strategy for combating M. tuberculosis.
2. Results
2.1. Isolation and Morphological Characterization of Henu3
Mycobacterium smegmatis mc
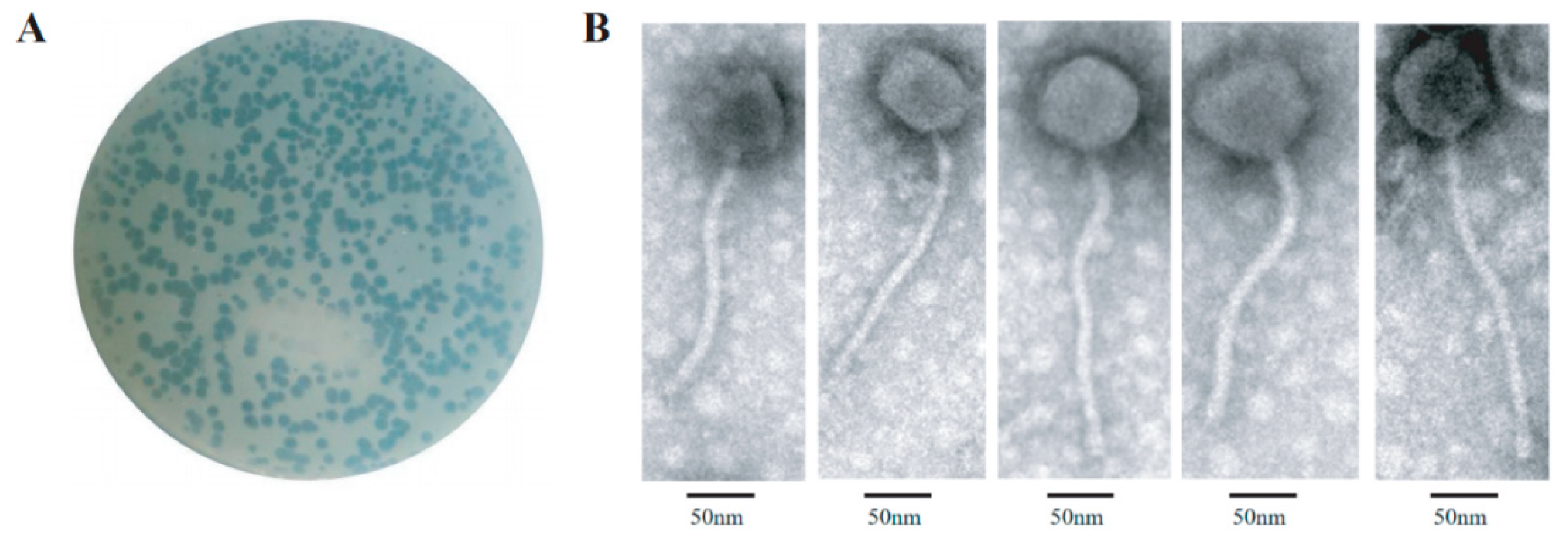
2155 was used as a host for a new mycobacteriophage, Henu3, isolation from soil samples collected around a hospital environment. Phage Henu3 formed clear, circular plaques on the host bacterial lawn, measuring 4 to 6 mm in diameter with well-defined edges, indicating its lytic nature (
Figure 1A). TEM characterization showed that phage Henu3 possesses a non-contractile tail measuring approximately 215 nm in length and an icosahedral head with an average diameter of 54 ± 3 nm (
Figure 1B). Based on these morphological features and according to the latest International Committee on Taxonomy of Viruses (ICTV) guidelines (
https://talk.ictvonline.org/), phage Henu3 has been classified within the
Caudoviricetes class [
14].
2.2. Biological Characterization of Phage Henu3
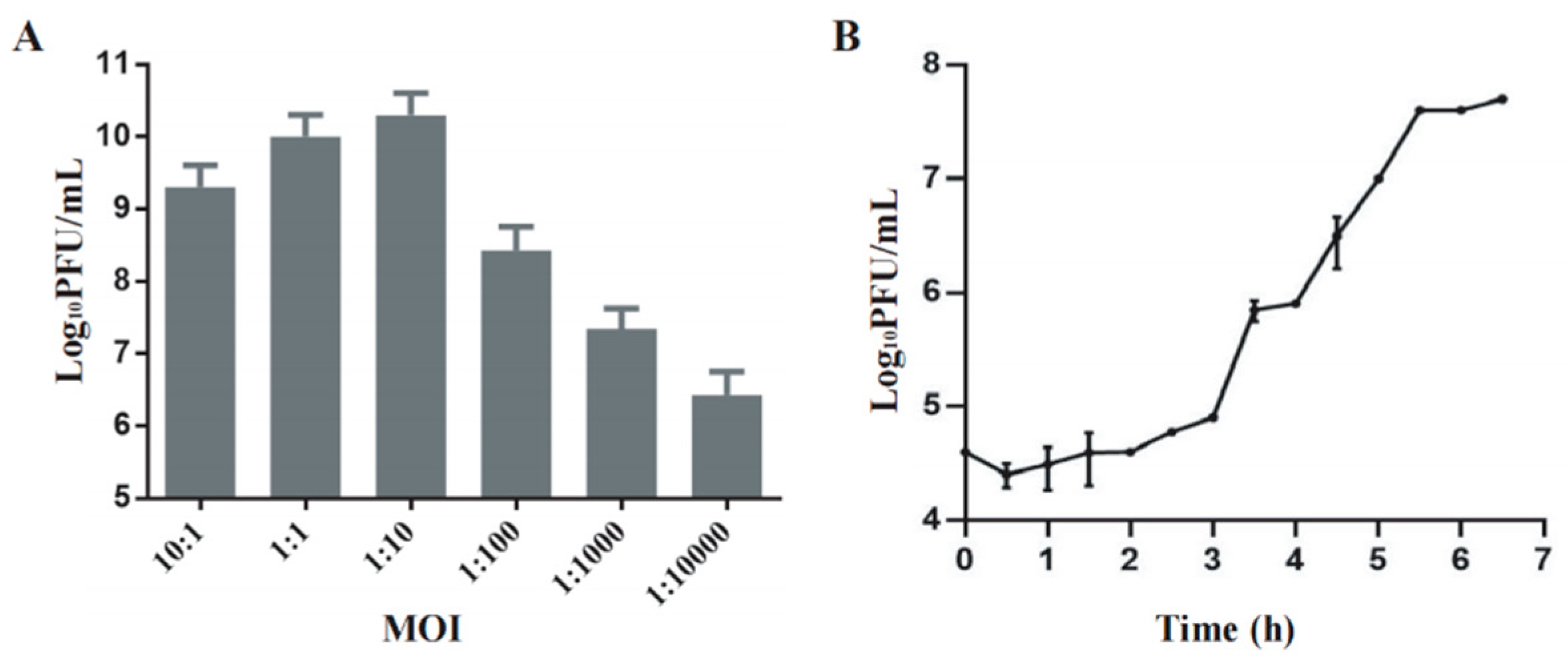
Phage Henu3 produced its highest phage titer of 3×10
10 PFU/mL at a multiplicity of infection (MOI) of 1:10, indicating this to be the optimal MOI for phage Henu3 (
Figure 2A). To further characterize the phage’s lifecycle, a one-step growth curve was performed using
Mycobacterium smegmatis mc
2155 at the optimal MOI. The phage Henu3 showed a latent period of 3 h followed by a lytic period of 2.5 h. The average burst size was 40 PFU/cell (
Figure 2B).
2.3. Influence of Temperature, pH, and Ultraviolet on Phage Henu3 Stability
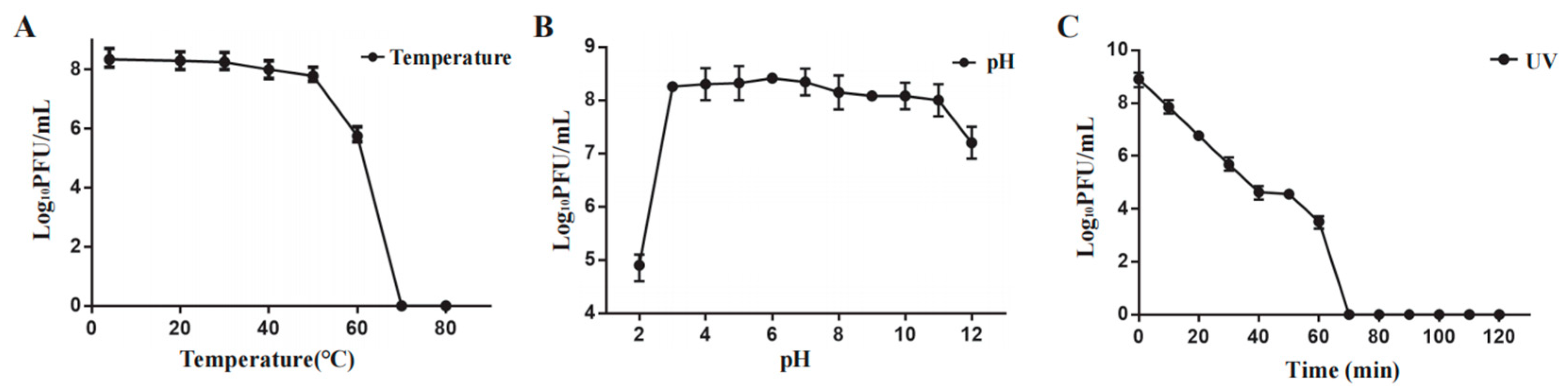
Phage Henu3 stability was carried out under varying temperatures, pH range, and ultraviolet light. Phage Henu3 almost maintained its activity between 4 °C and 50 °C. A notable decline in phage titer was observed at 60 °C, whereas the phage titer was undetected at higher temperatures (
Figure 3A). The acid-base tolerance test revealed that phage Henu3 was stable within a pH range of 3-11, with the highest titer observed at pH 7. The phage titer decreased tenfold in extremely alkaline conditions (pH 12) and thousandfold in highly acidic conditions (pH 2), suggesting that phage Henu3 is more tolerant to alkaline than acidic environments (
Figure 3B). Moreover, under UV light exposure, the phage Henu3 titer progressively decreased over a duration of 60 min, with complete inactivation observed after 70 min, suggesting sensitivity to ultraviolet radiation (
Figure 3C). Altogether, these findings are crucial for considering the practical applications of Henu3 in phage therapy.
2.4. The Whole Genome Sequencing and Bioinformatics Analysis of Phage Henu3
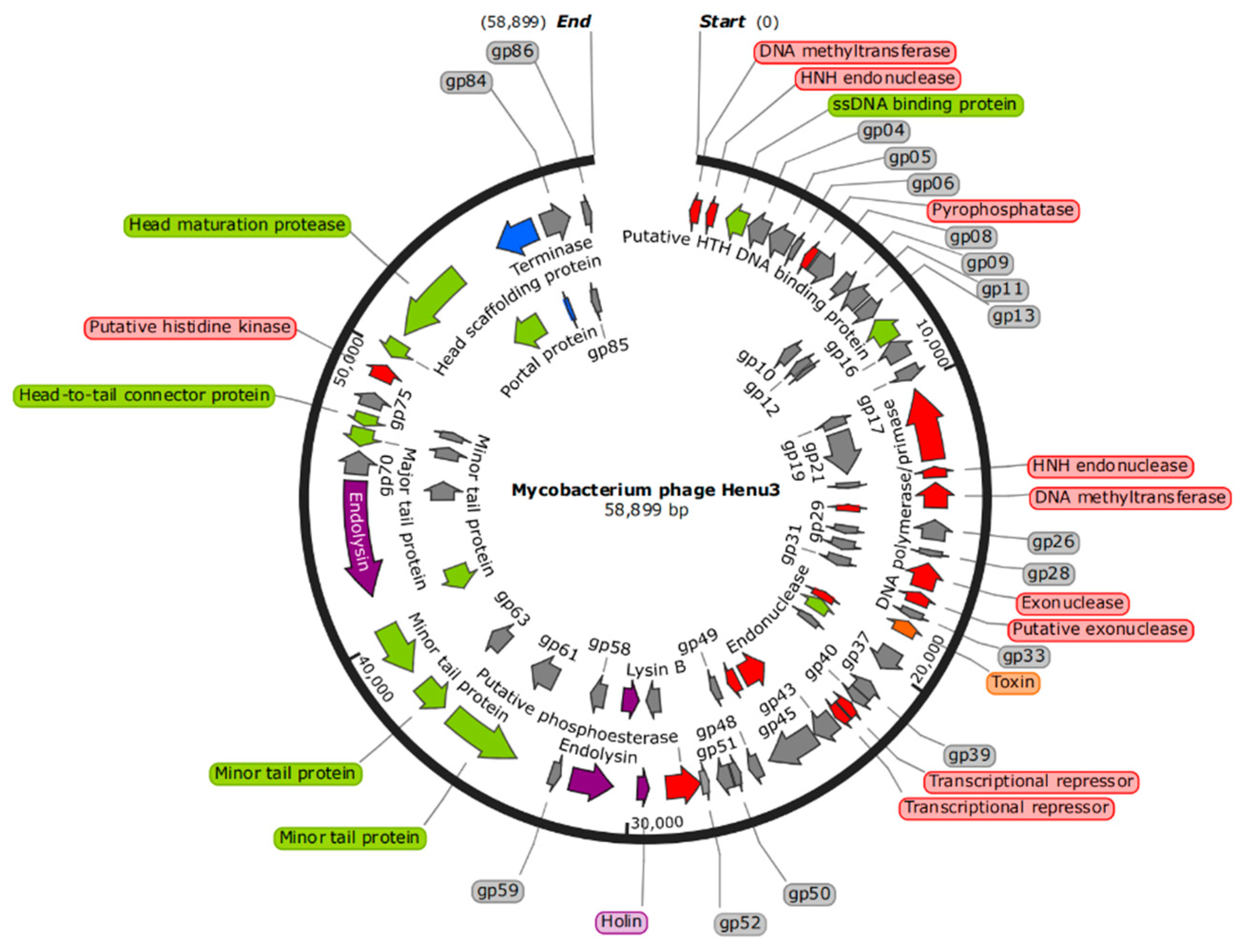
The genome of phage Henu3 had a double-stranded DNA with 58,899 base pairs (bp) in length. The genomic composition is characterized by base contents of A (15.3%), G (32.6%), T (17.1%), and C (35%), resulting in GC content of 67.6% similar to that of its host,
M. smegmatis mc
2155, indicating a possible evolutionary adaptation to its host. The whole-genome sequence of mycobacteriophage Henu3 was deposited in GenBank (Accession Number MK224497). The phage Henu3 genome includes one tRNA gene (tRNA
Lys) and 86 open reading frames (ORFs), with the ORFs encompassing 91.73% of the entire genome. These ORFs have an average length of 711 bp, distributed with 35 encoded in the forward direction and 51 in the reverse direction. Among these, 34 ORFs (39.5% of the total) have been identified with specific functions and are categorized into three functional groups: nucleic acid metabolism, DNA replication, and recombination; phage packaging and morphology; and proteins related to lytic functions (
Figure 4).
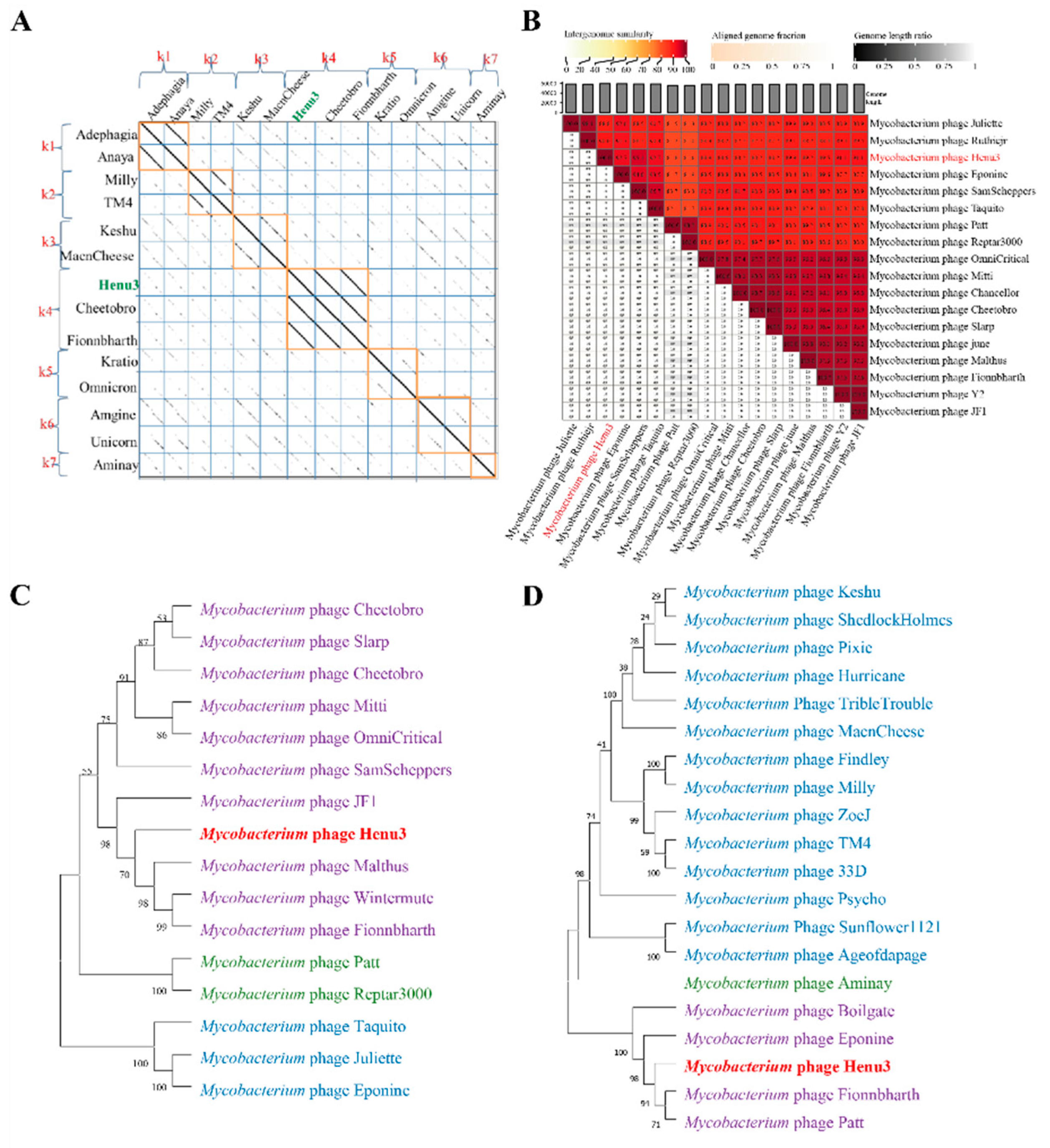
To elucidate the relationships of m
ycobacteriophage Henu3 with other phages, Dot plot analysis, heatmap analysis and phylogenetic trees construction were performed. The dot plot analysis indicated the closest similarity between Henu3 and Fionnbharth within the mycobacteriophage K4 sub-cluster (
Figure 5A). When compared with other phages from the K cluster, the sequence coverage values for Henu3 were all below 40%, underscoring its distinctiveness and supporting its classification as a new member of the K4 sub-cluster. Heatmap analysis of genomic similarities revealed that the entire genome of Henu3 and other phages with greater than 80% blast query coverage on NCBI share significant likeness, including five phages that demonstrate over 90% similarity (
Figure 5B). Phylogenetic trees constructed using the tape measure protein align Henu3 closely with mycobacteriophages Malthus, Wintermute, and Fionnbharth (
Figure 5C), while trees based on the tail protein indicate strong similarities between Henu3, Fionnbharth, and Patt (
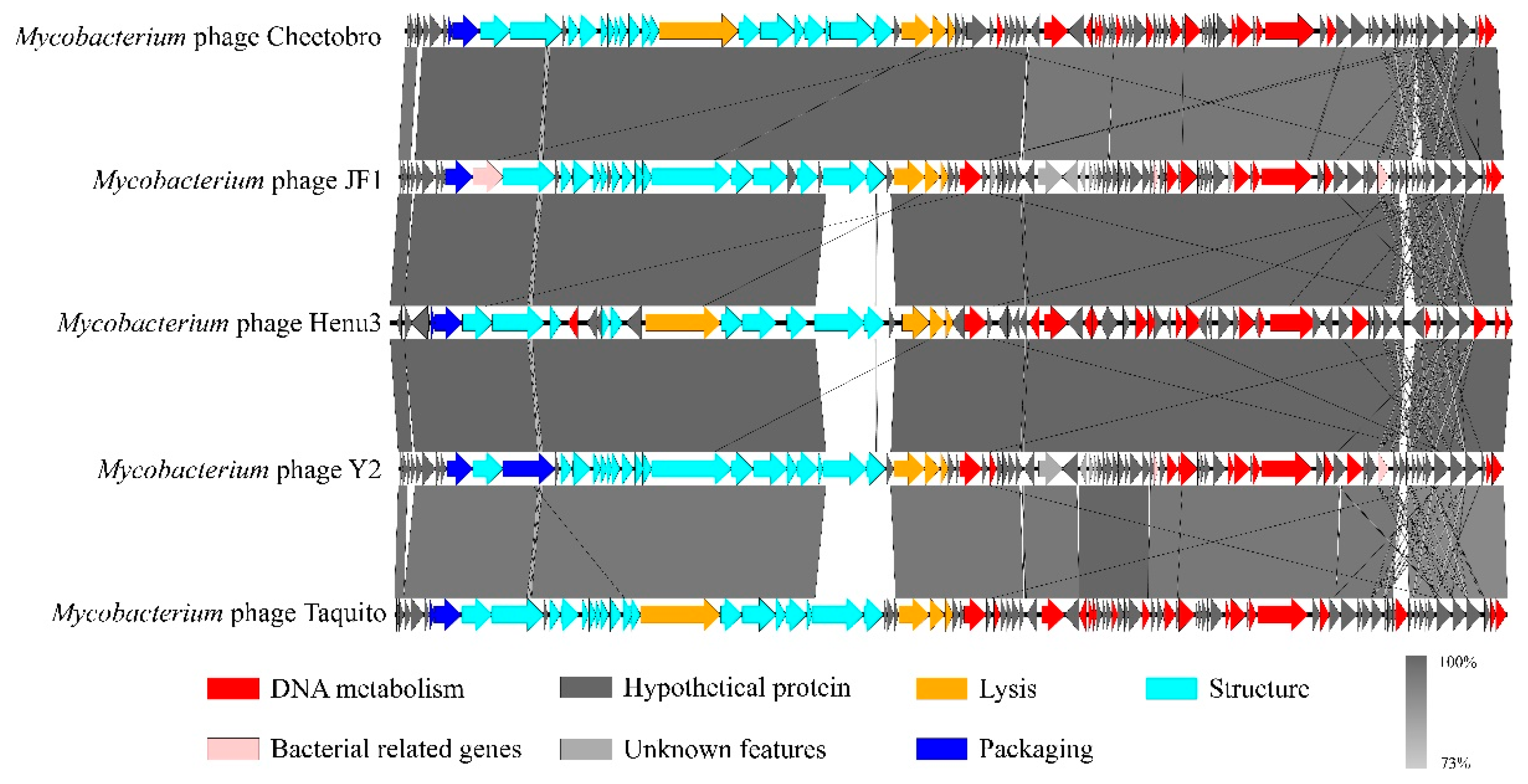
Figure 5D). Further comparative genomic analyses between Henu3 and previously documented mycobacteriophages (Cheetobro, JF1, Taquito, and Y2) revealed that similarities at the amino acid level for DNA metabolism and key functional genes exceed 73% (
Figure 6). Moreover, these phages also share highly similar assembly and lysis proteins, indicating common structural and functional characteristics.
2.5. Isolation of Phage-Resistant Bacteria
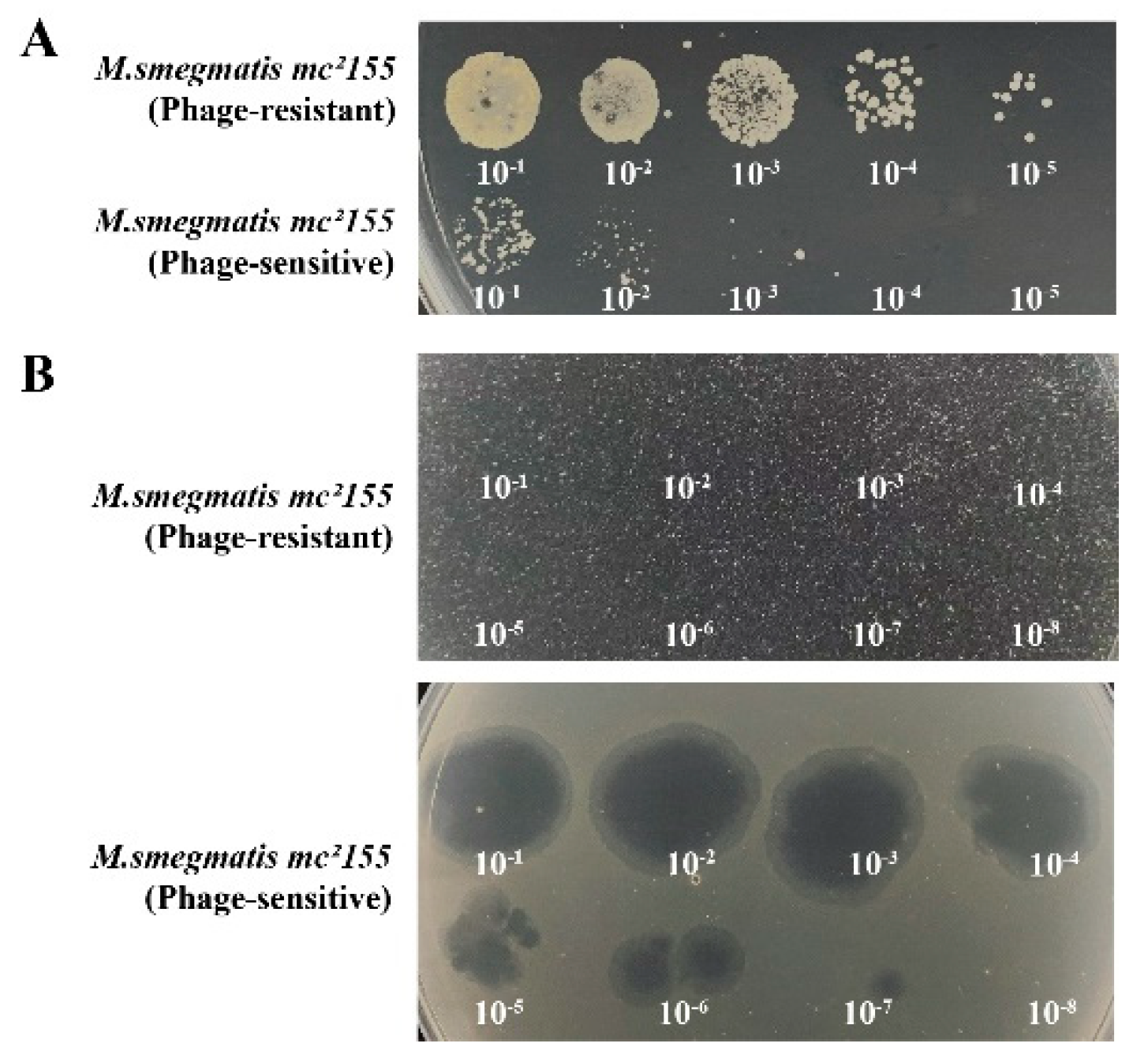
The phage-resistant strains, 0G10 and 2E1, were successfully isolated. During the forward verification procedure, these strains showed confluent growth on plates containing phage Henu3, unlike the phage-sensitive bacteria, which formed only a few colonies (
Figure 7A). Additionally, in the reverse verification procedure, when phage Henu3 was spotted onto 7H10 agar plates containing phage-resistant strains, no lysis was observed. In contrast, lysis zones were evident with the phage-sensitive bacteria (
Figure 7B). This indicates a successful selection of phage-resistant strains with stable resistance traits.
2.6. Phage-Resistant Strain Exhibits Increased Sensitivity to Antibiotics
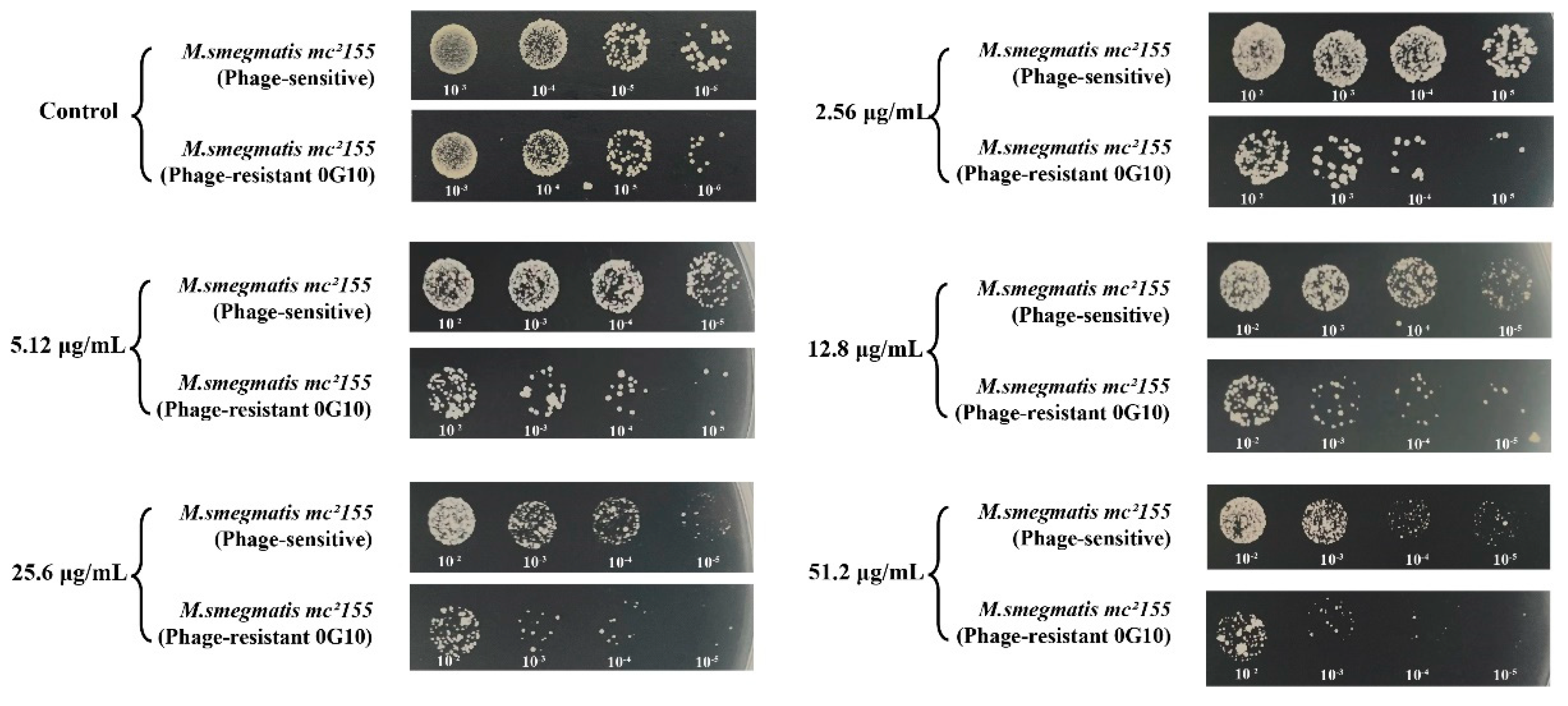
The 0G10 strain displayed increased sensitivity to cefotaxime compared to its sensitive counterparts. Neither the 0G10 strain nor the phage-sensitive strain showed significant differences in CFU counts on plates without cefotaxime. However, at higher cefotaxime concentrations of 2.56 µg/mL, 5.12 µg/mL, 12.8 µg/mL, 25.6 µg/mL and 51.2 µg/mL, the CFU counts for the 0G10 strain were significantly lower than those for the sensitive strain (
Figure 8).
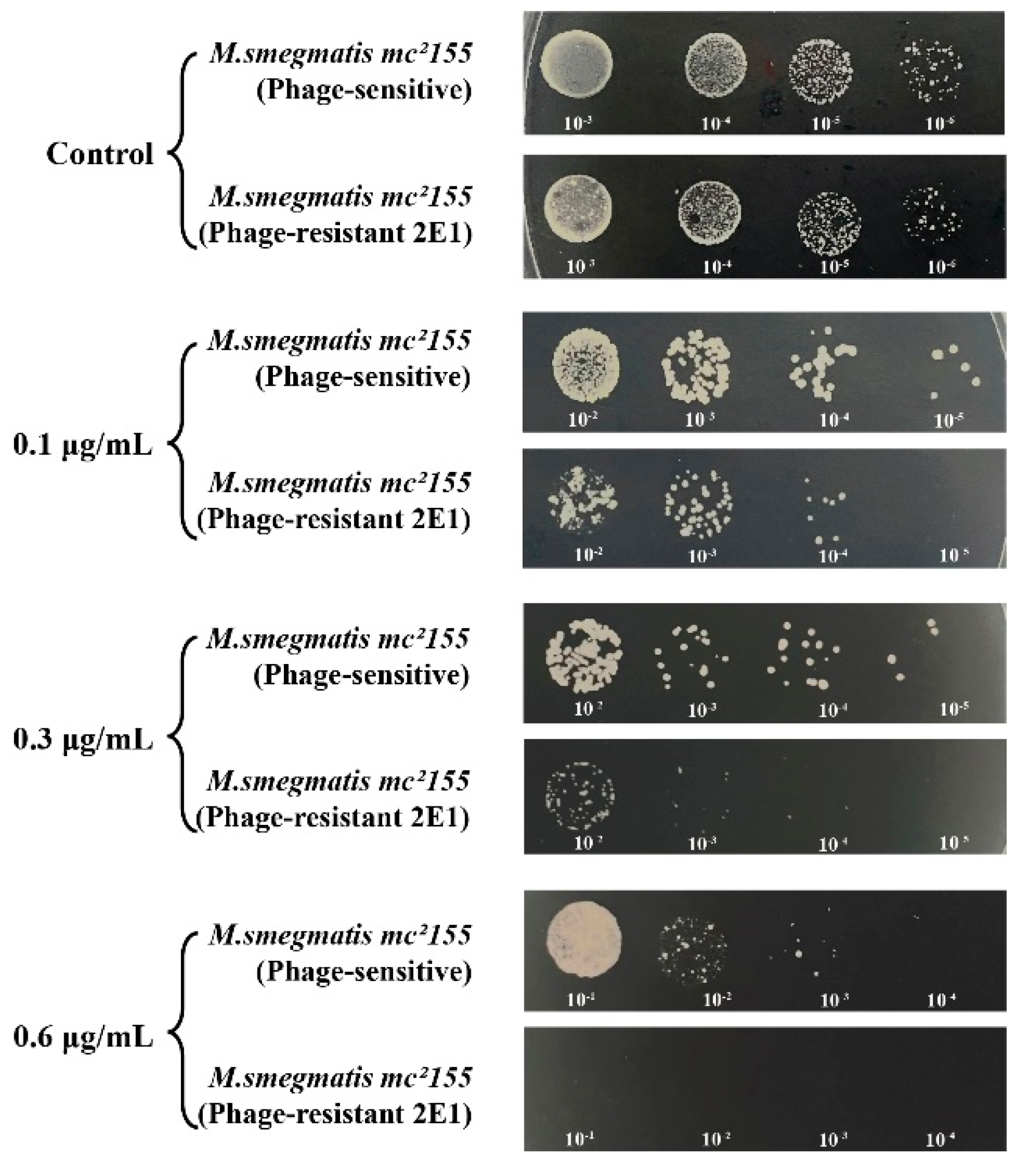
Both phage-resistant 2E1 and sensitive strains showed identical CFU counts on kanamycin-free plates. However, when exposed to kanamycin concentrations of 0.1 µg/mL and 0.2 µg/mL, CFU counts for the 2E1 strain decreased by an order of magnitude compared to the phage-sensitive strain. Interestingly, at a higher concentration of 0.6 µg/mL, the 2E1 strain failed to show any colonies, indicating an extreme sensitivity to kanamycin in contrast to the phage-sensitive strain (
Figure 9).
2.7. Whole-Genome Re-Sequencing Analysis of Phage-Resistant Bacteria 0G10 and 2E1
To investigate the balancing act between phage resistance and antibiotic sensitivity in strains 0G10 and 2E1, whole-genome sequencing analysis was performed. When comparing these genomes with that of the sensitive strain, a total of 12 insertion-deletion (InDel) mutations were identified (
Table 1). Among these mutations, two were exclusive to the 2E1 genome, one was unique to 0G10, and the remaining nine mutations were common to both 0G10 and 2E1. These mutations affected genes encoding proteins associated with cell wall synthesis enzymes, cell membrane transport proteins, transcriptional regulators, and two-component regulatory systems. One notable gene, MSMEG_2348, encodes a glycosyltransferase implicated in the glycosylation of the phospho-wall teichoic acid component of bacterial cell walls. Previous studies in
Streptomyces coelicolor have shown that the absence of this glycosyltransferase enhances bacterial sensitivity to antibiotics, such as vancomycin and β-lactam antibiotics, and increases resistance to phage φC31 invasion [
15]. Additionally, the genes MSMEG_5878, MSMEG_4512, and MSMEG_6720, which encode a keratinase, the mycobactin polyketide synthase MbtD, and a hydrolase, respectively, are involved in the synthesis of cell wall peptidoglycan or phospho-wall teichoic acid. It is hypothesized that mutations in these four genes may alter phage adsorption mechanisms, thereby facilitating the emergence of phage-resistant bacterial strains.
In
M. smegmatis, the genes MSMEG_0916, MSMEG_1060, and MSMEG_6084 encode the transcription factors TetR, Lsr2, and GntR, respectively. These factors regulate various cellular processes, including amino acid metabolism, DNA replication, and repair. Lsr2, acting as a transcriptional repressor, directly controlimped gene expression by binding to promoter regions. Marta Kołodziej and colleagues have shown that Lsr2 regulates the expression of mycobacterial polyketide synthase, which is crucial for synthesizing lipooligosaccharides.
Mycobacteria lacking Lsr2 form smooth colonies and exhibit increased sensitivity to rifampicin and nalidixic acid [
16]. Similarly, Kriti Arora and others have reported that Lsr2 deficiency impeded phage infection [
17]. While it is hypothesized that other transcription factors might influence the antibiotic stress response, further experiments are required for confirmation. Additionally, genes MSMEG_0555, MSMEG_4468, MSMEG_2004, and MSMEG_6758 encode a sugar ABC transporter protease, a substrate-binding protein, an MFS family membrane transporter protein, and an aquaporin, respectively. Research has indicated that the absence of the substrate-binding protein DppA1 in
Pseudomonas aeruginosa inhibits Pf5 phage adsorption and enhances biofilm formation [
18]. It is speculated that similar membrane transport proteins in
M. smegmatis may affect phage DNA entry and contribute to phage resistance.
The gene MSMEG_6236 encodes a two-component regulatory system (MnoR) commonly found in bacteria. This system, comprising a sensor histidine kinase and a response regulator, is crucial for bacteria to detect environmental changes, regulate the expression of growth and virulence factors, and adapt to stress. It functions through the phosphorylation and dephosphorylation of proteins. Lucas-Elío
et al. demonstrated that such systems can regulate the expression of the CRISPR-Cas system, thereby playing a significant role in phage infection prevention. It is hypothesized that the MnoR system encoded by MSMEG_6236 might serve a similar function in mycobacteria [
19], although further experimental validation is necessary.
3. Discussion
Due to the overuse of clinical antibiotics, there has been a surge in drug-resistant bacteria, pushing humanity towards a “post-antibiotic era” where effective antibiotics may no longer be available [
20]. In response to this antibiotic crisis, relevant authorities have implemented a series of policy measures to restrict antibiotic use. Phage therapy has emerged as a novel approach for treating antibiotic-resistant bacterial infections, offering a promising alternative to traditional antibiotics [
21]. Phages offer significant advantages over antibiotics, including high specificity, safety, and a lower propensity for resistance development, thereby garnering considerable global scholarly interest. To provide a theoretical foundation for the clinical choice between phages and antibiotics, the authors have explored the biological characteristics of
mycobacterium phages and their mechanisms for coordinating phage resistance and stress response.
This study involved sampling soil from around hospitals to isolate and purify phage Henu3, using M. smegmatis as the host bacterium. The morphological observation and whole-genome sequencing revealed that phage Henu3 is a lytic phage of the Siphoviridae family, featuring a genome of 58,899 bp with 86 ORFs and one tRNA gene. Phylogenetic analysis placed phage Henu3 within the mycobacterium phage K4 sub-cluster. Thermal stability assays showed that the phage’s titer remains stable from 4°C to 50°C but significantly decreases above this temperature range. Acid resistance tests indicated a thousandfold decrease in titer at a pH of 2, demonstrating poor acid resistance. Additionally, phage Henu3 is sensitive to ultraviolet light; its titer gradually diminishes over 60 minutes under 40W UV light and completely diminishes after 70 minutes. Experiments determining the optimal multiplicity of infection (MOI) and conducting one-step growth curves confirmed that phage Henu3 can rapidly lyse its host and release progeny, highlighting its potent lytic capabilities and providing a foundation for studying phage-resistant strains.
Genomic analysis serves as a fundamental basis for phage classification. In this study, the whole genome of phage Henu3 was sequenced using the Illumina 2100 sequencing platform and analyzed via BlastP. The analysis revealed that out of the 86 ORFs identified, 33 (38.5%) encode proteins with specific functions. The functions of the proteins encoded by the remaining ORFs remain unidentified. The 33 functionally characterized ORFs can be categorized into three groups: nucleic acid metabolism, DNA replication, and recombination; phage packaging and morphology; and proteins associated with lytic activity.
To explore the genetic mechanisms behind phage resistance and stress response coordination in bacteria, this study employed continuous passaging and both positive and negative verification methods to isolate phage-resistant strains. Comparative stress response analyses between the selected phage-resistant strains 0G10 and 2E1 and their sensitive counterparts revealed that strain 0G10 demonstrated increased sensitivity to cefotaxime, while strain 2E1 was more sensitive to kanamycin. Subsequently, genomic DNA from strains 0G10 and 2E1 was extracted and resequenced, uncovering frameshift mutations in 12 genes associated with cell wall synthesis, membrane transport proteins, two-component regulatory systems, and transcriptional regulators. These mutations suggest a potential balance between phage resistance and stress response in these bacteria.
This study presents the isolation and characterization of a new mycobacteriophage Henu3, which exhibits a short latency period, a large burst size, resistance to alkaline conditions, but not to acidic environments, and elevated temperature tolerance. Comparative genomic and phylogenetic analyses classify Henu3 as a new member of the mycobacteriophage K4 subcluster, enriching the research on drug-resistant mycobacteria. Additionally, the study identified phage-resistant strains 0G10 and 2E1, which demonstrated increased sensitivity to cefotaxime and kanamycin, respectively. Whole-genome re-sequencing of these strains revealed mutations in twelve genes involved in cell wall synthesis, membrane transport, two-component regulatory systems, and transcriptional regulation. These findings provide a theoretical foundation for the use of phage therapy in tuberculosis and the study of resistance mechanisms in M. tuberculosis.
4. Materials and Methods
4.1. Isolation and Purification of Phage Henu3
Phage was isolated from the soil sample collected from the hospital of the first affiliated hospital of Henan University. Approximately 15 g of soil from the hospital surroundings were dissolved in 30 mL of SM buffer and subsequently filtered through a 0.22 µm filter to remove bacteria. Two mL of log-phase
M. smegmatis mc
2155 was added to a conical flask containing 30 mL of the filtrate and incubated in a thermostatic shaker at 37 °C for three days. The mixed culture was then centrifuged at 12,000 rpm for six min to precipitate the bacteria, and the supernatant was further filtered through a 0.22 µm filter. The presence of phage in the filtrate was screened using the spot assay. The isolated phage was purified using the double-layer agar (DLA) technique [
22,
23], which was repeated ten times to ensure consistency in plaque morphology.
4.2. Transmission Electron Microscopy (TEM)
Phage Henu3 morphology was examined using TEM. Briefly, 20 μL of purified phage lysate was placed onto a copper grid and allowed to stand for 15 min. Excess liquid was removed using filter paper, and the grid was then stained with 2% phosphotungstic acid for 10 min. before visualizing the phage particles using a Tecnai 10 (Philips) TEM [
24] .
4.3. Genomic Sequencing of Phage Henu3
Phage Henu3 genomic DNA was extracted using a combination of Proteinase K- Sodium dodecyl sulfate (SDS), and phenol extraction methods [
25]. The extracted DNA was then sequenced using next-generation sequencing technology on the Illumina 2100 platform. The genome was fragmented using ultrasonication (Fisher Scientific), generating randomly sized genomic fragments. For DNA library construction, fragments approximately 300 bp in length were isolated using gel electrophoresis and purified with the TruSeq DNA Sample Prep Kit-Set A. Libraries were then amplified using the TruSeq PE Cluster Kit (Illumina, USA), and sequencing reactions were conducted on an Illumina sequencer. For sequence assembly, data were compiled using Newbler software to reconstruct the complete phage genome sequence [
26].
4.4. Bioinformatics Analysis of the Whole Genome Sequence of Henu3
The phage genome sequences were characterized using EditSeq from the DNASTAR package, with analyses conducted on the base composition, G+C content, and the average gene length and density across the genome. The Open Reading Frames (ORFs) were predicted using Softberry (
http://linux1.softberry.com/berry.Phtmltopi) and GeneMark [
27] (
http://exon. edu/GeneMark/), selecting ATG, TTG, and GTG as start codons. For phylogenetic analysis, a neighbor-joining tree was constructed using MEGA7 software [
28] with 1,000 bootstrap replications, and phage family classification was performed using DNAplotter. Multiple genome comparisons were conducted with EasyFig [
29]. Comparisons with antibiotic resistance and virulence genes were performed using ResFinder (
https://cge.cbs.dtu.dk/services/ResFinder/) and VirulenceFinder (
https://cge.cbs.dtu.dk/services/VirulenceFinder/). The Henu3 genome was mapped using Proksee (
https://proksee.ca) based on GenBank data and visualized using GenomeVx [
30] (
http://wolfe.ucd.ie/GenomeVx/).
4.5. Optimal Multiplicity of Infection (MOI) for Phage Henu3
The optimal MOI of phage Henu3 was assessed as previously described [
31]. In brief,
M. smegmatis mc
2155 in log-phase culture (2×10
7 CFU/mL) was mixed with phage Henu3 at different MOI ratios of 10:1, 1:1, 1:10, 1:100, and 1:1000, respectively [
32]. The phage-bacteria co-culture was incubated at 37 °C for 12 h in a shaking incubator, and the phage titer was determined using the DLA agar technique. The MOI that resulted in the highest titer was identified as the optimal MOI for phage Henu3. Each MOI was tested in triplicate to ensure accuracy.
4.6. One-Step Growth Curve of Phage Henu3
A one-step growth curve was conducted to determine the latent period and burst size of phage Henu3 [
23,
31]. Briefly,
M. smegmatis mc
2155 log-phase culture and phage Henu3 was mixed at the optimal MOI ratio (1:10) and incubated at 37 °C for 30 min. Following incubation, the mixture was centrifuged, and the supernatant was discarded. The pellet was re-suspended in 10 mL of 7H9 medium and re-incubated. Samples were taken every 30 min for phage Henu3 titer determination by the DLA technique [
23].
4.7. Thermal, pH, and Ultraviolet Stability of Phage Henu3
Phage stability was evaluated against various conditions, as previously described [
33]. In brief, a highly concentrated phage solution (1 × 10
9 PFU/mL) was incubated for 1 h at different temperatures (4 °C, 20 °C, 30 °C, 40 °C, 50 °C, 60 °C, 70 °C, and 80 °C) or under varying pH values (2, 4, 6, 8, 10, and 12) for 1 h at 27 °C. After incubation, the phage titer was determined using the DLA technique. To evaluate ultraviolet (UV) stability, EP tubes containing phage solution were positioned under a 40W UV lamp at a wavelength of 253.7 nm, with the distance between the tubes and the lamp set to 10 cm. Aliquots of 100 µL were withdrawn every 10 min for titer determination.
4.8. Testing for Phage-Resistant Strains
The phage-resistant strains were screened using the traditional spot assay [
34,
35,
36]. The bacterial colonies that developed within the lysis zones were picked-up and underwent two rounds of streaking onto 7H10 agar plates to obtain pure individual colonies. The phage-resistant colonies were further confirmed using bidirectional validation techniques. For the forward validation procedure, either phage-resistant or phage-sensitive
M. smegmatis mc
2155 strains were tenfold serially diluted, and aliquots of 5 µL of each diluted culture were inoculated onto the surface of 7H10 agar plates containing Phage Henu3 in the top soft agar layer. Following incubation at 37 °C for 72 h, the plates were inspected for the phage-resistant colonies. For reverse validation procedure, phage Henu3 solution (1 × 10
9 PFU/mL) was serially diluted, and 5 µL aliquots from each dilution were spotted onto 7H10 agar plates pre-seeded with either phage-resistant or phage-sensitive
M. smegmatis mc
2155 strains, respectively. No lysis zones following incubation at 37 °C for 48 h confirms the phage-resistance phenotypes [
34]. The entire procedure was repeated three times to ensure consistent results.
4.9. Forward and Reverse Validation of Phage-Resistant Bacteria
Positive Validation Procedure: 300 µL of high-concentration phage solution (1×109 PFU/mL) was evenly spread on a 7H10 agar plate. Both resistant (phage-resistant M. smegmatis mc2155) and sensitive (M. smegmatis mc2155) bacterial strains were serially diluted in tenfold increment using the same high-concentration phage solution and MP buffer, respectively. Subsequently, 5 µL of each diluted culture—both phage-resistant and sensitive—was dropwise added to a 7H10 plate containing phage Henu3. The plates were then incubated in a biochemical incubator at 37°C for 72 h to assess the number of CFUs. This test was performed three times to ensure consistency in the results.
Reverse Validation Procedure: double-layer agar plates were prepared with both phage-resistant and sensitive M. smegmatis mc2155 strains. Concurrently, phage Henu3 at a concentration of 1×109 PFU/mL was serially diluted from 10-1 to 10-8. Subsequently, 5 µL of Phage Henu3 from each dilution gradient was added dropwise onto the respective double-layer agar plates. The plates were allowed to dry and then incubated in a biochemical incubator at 37°C for 48 hours. For plates with resistant bacteria, no plaques (empty spots) are expected, whereas plates with sensitive bacteria should show clear plaque formation. The number of plaques at different dilution gradients was observed and recorded. This procedure was repeated three times to ensure consistent results.
4.10. Detection of Susceptibility of Phage-Resistant Bacteria to Various Antibiotics
The minimum inhibitory concentration (MIC) of cefotaxime and kanamycin was determined against the phage-sensitive and phage-resistant
M. smegmatis mc
2155 strains using the broth microdilution method [
37]. Subsequently, the phage-resistant
M. smegmatis mc
2155 strains were grown in 7H9 medium in the presence of 1/2 MIC, 1/4 MIC, 1/8 MIC, and so forth. In brief, aliquots of 195 µL of 7H9 medium with varying antibiotic concentrations was dispensed into each well of the microtiter plate. Then, aliquots of 5 µL of phage-resistant bacteria in the log-phase were added to each well. The plates were then incubated at 37 °C for 72 h. The phage-sensitive
M. smegmatis mc
2155 strain served as a control. Samples were taken every 12 h for colony forming unit (CFU) counts. This procedure was performed in triplicate to ensure consistency in the results.
4.11. Statistical Analysis
The experiments were replicated three times, and the experimental results are expressed as mean ± standard deviation (SD). All results were analyzed by one-way analysis of variance (Dunnett’s test) for a significance level of p < 0.05. GraphPad Prism 9.0 (GraphPad Software, San Diego, CA, USA) was used for data analysis.
Author Contributions
Conceptualization, T.T. and S.E.G.; methodology, X.L. and Y.W.; validation, T.T., S.E.G. and Y.W.; writing—original draft preparation, X.L.; writing—review and editing, J.X.; supervision, T.T. All authors have read and agreed to the published version of the manuscript.
Funding
This research was funded by the Key R&D and Promotion Projects of Henan Province (232102311139); China Postdoctoral Science Foundation (2021m690095); National Innovation and Entrepreneurship Training Program for College Students (20231022009 and 20231021006).
Data Availability Statement
Data is contained within the article or supplementary material. Complete genome of
mycobacterium phage Henu3 is available in
https://www.ncbi.nlm.nih. gov/nuccore/ under the GenBank accession number MK224497.
Conflicts of Interest
The authors declare no conflicts of interest.
References
- Davies, J.; Davies, D. Origins and Evolution of Antibiotic Resistance. Microbiol. Mol. Biol. Rev. 2010, 74, 417–433. [Google Scholar] [CrossRef] [PubMed]
- Gagneux, S. Fitness cost of drug resistance inMycobacterium tuberculosis. Clin. Microbiol. Infect. 2009, 15, 66–68. [Google Scholar] [CrossRef] [PubMed]
- World Health, O., Global tuberculosis report 2023. World Health Organization: Geneva, 2023.
- Allué-Guardia, A.; Saranathan, R.; Chan, J.; Torrelles, J.B. Mycobacteriophages as Potential Therapeutic Agents against Drug-Resistant Tuberculosis. Int. J. Mol. Sci. 2021, 22, 735. [Google Scholar] [CrossRef] [PubMed]
- Espinosa-Pereiro, J.; Sánchez-Montalvá, A.; Aznar, M.L.; Espiau, M. MDR Tuberculosis Treatment. Medicina 2022, 58, 188. [Google Scholar] [CrossRef] [PubMed]
- Kelishomi, F.Z.; Khanjani, S.; Fardsanei, F.; Sarabi, H.S.; Nikkhahi, F.; Dehghani, B. Bacteriophages of Mycobacterium tuberculosis, their diversity, and potential therapeutic uses: a review. BMC Infect. Dis. 2022, 22, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Sulakvelidze, A.; Alavidze, Z.; Morris, J.G., Jr. Bacteriophage therapy. Antimicrob Agents Chemother 2001, 45, (3), 649–59. [Google Scholar] [CrossRef]
- Harada, L.K.; Silva, E.C.; Campos, W.F.; Del Fiol, F.S.; Vila, M.; Dąbrowska, K.; Krylov, V.N.; Balcão, V.M. Biotechnological applications of bacteriophages: State of the art. Microbiol. Res. 2018, 212–213, 38–58. [Google Scholar] [CrossRef] [PubMed]
- Anand, T.; Virmani, N.; Kumar, S.; Mohanty, A.K.; Pavulraj, S.; Bera, B.C.; Vaid, R.K.; Ahlawat, U.; Tripathi, B. Phage therapy for treatment of virulent Klebsiella pneumoniae infection in a mouse model. J. Glob. Antimicrob. Resist. 2019, 21, 34–41. [Google Scholar] [CrossRef] [PubMed]
- Carvalho, C.M.; Gannon, B.W.; E Halfhide, D.; Santos, S.B.; Hayes, C.M.; Roe, J.M.; Azeredo, J. The in vivo efficacy of two administration routes of a phage cocktail to reduce numbers of Campylobacter coli and Campylobacter jejuni in chickens. BMC Microbiol. 2010, 10, 232–232. [Google Scholar] [CrossRef] [PubMed]
- Seo, B.-J.; Song, E.-T.; Lee, K.; Kim, J.-W.; Jeong, C.-G.; Moon, S.-H.; Son, J.S.; Kang, S.H.; Cho, H.-S.; Jung, B.Y.; et al. Evaluation of the broad-spectrum lytic capability of bacteriophage cocktails against various Salmonella serovars and their effects on weaned pigs infected with Salmonella Typhimurium. J. Veter- Med Sci. 2018, 80, 851–860. [Google Scholar] [CrossRef] [PubMed]
- Dedrick, R.M.; Guerrero-Bustamante, C.A.; Garlena, R.A.; Russell, D.A.; Ford, K.; Harris, K.; Gilmour, K.C.; Soothill, J.; Jacobs-Sera, D.; Schooley, R.T.; et al. Engineered bacteriophages for treatment of a patient with a disseminated drug-resistant Mycobacterium abscessus. Nat. Med. 2019, 25, 730–733. [Google Scholar] [CrossRef]
- Mtimka, S.; Pillay, P.; Kwezi, L.; Pooe, O.J.; Tsekoa, T.L. An Exploratory Review of the Potential of Lytic Proteins and Bacteriophages for the Treatment of Tuberculosis. Microorganisms 2024, 12, 570. [Google Scholar] [CrossRef] [PubMed]
- Turner, D.; Shkoporov, A.N.; Lood, C.; Millard, A.D.; Dutilh, B.E.; Alfenas-Zerbini, P.; van Zyl, L.J.; Aziz, R.K.; Oksanen, H.M.; Poranen, M.M.; et al. Abolishment of morphology-based taxa and change to binomial species names: 2022 taxonomy update of the ICTV bacterial viruses subcommittee. Arch. Virol. 2023, 168, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Holman, N.D.M.; Wilkinson, A.J.; Smith, M.C.M. Alanine-scanning mutagenesis of protein mannosyl-transferase from Streptomyces coelicolor reveals strong activity-stability correlation. Microbiology 2021, 167, 001103. [Google Scholar] [CrossRef] [PubMed]
- Kołodziej, M.; Łebkowski, T.; Płociński, P.; Hołówka, J.; Paściak, M.; Wojtaś, B.; Bury, K.; Konieczny, I.; Dziadek, J.; Zakrzewska-Czerwińska, J. Lsr2 and Its Novel Paralogue Mediate the Adjustment of Mycobacterium smegmatis to Unfavorable Environmental Conditions. mSphere 2021, 6. [Google Scholar] [CrossRef] [PubMed]
- Arora, K.; Whiteford, D.C.; Lau-Bonilla, D.; Davitt, C.M.; Dahl, J.L. Inactivation of lsr2 Results in a Hypermotile Phenotype in Mycobacterium smegmatis. J. Bacteriol. 2008, 190, 4291–4300. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.; Song, S.; Sheng, L.; Zhu, L.; Kim, J.-S.; Wood, T.K. Substrate Binding Protein DppA1 of ABC Transporter DppBCDF Increases Biofilm Formation in Pseudomonas aeruginosa by Inhibiting Pf5 Prophage Lysis. Front. Microbiol. 2018, 9, 30. [Google Scholar] [CrossRef] [PubMed]
- Lucas-Elío, P.; Molina-Quintero, L.R.; Xu, H.; Sánchez-Amat, A. A histidine kinase and a response regulator provide phage resistance to Marinomonas mediterranea via CRISPR-Cas regulation. Sci. Rep. 2021, 11, 1–13. [Google Scholar] [CrossRef]
- Jin, Y.; Li, W.; Zhang, H.; Ba, X.; Li, Z.; Zhou, J. The Post-Antibiotic Era: A New Dawn for Bacteriophages. Biology 2023, 12, 681. [Google Scholar] [CrossRef] [PubMed]
- Tagliaferri, T.L.; Jansen, M.; Horz, H.-P. Fighting Pathogenic Bacteria on Two Fronts: Phages and Antibiotics as Combined Strategy. Front. Cell. Infect. Microbiol. 2019, 9, 22. [Google Scholar] [CrossRef] [PubMed]
- Tan, Y.; Su, J.; Luo, D.; Liang, B.; Liu, S.; Zeng, H. Isolation and genome-wide analysis of the novel Acinetobacter baumannii bacteriophage vB_AbaM_AB3P2. Arch. Virol. 2024, 169, 1–6. [Google Scholar] [CrossRef] [PubMed]
- Shende, R.K.; Hirpurkar, S.D.; Sannat, C.; Rawat, N.; Pandey, V. Isolation and characterization of bacteriophages with lytic activity against common bacterial pathogens. Veter- World 2017, 10, 973–978. [Google Scholar] [CrossRef] [PubMed]
- Tizro, P.; Choi, C.; Khanlou, N. , Sample Preparation for Transmission Electron Microscopy. Methods Mol Biol 2019, 1897, 417–424. [Google Scholar] [PubMed]
- Summer, E.J. , Preparation of a phage DNA fragment library for whole genome shotgun sequencing. Methods Mol Biol 2009, 502, 27–46. [Google Scholar] [PubMed]
- Liu, Y.; Liu, M.; Hu, R.; Bai, J.; He, X.; Jin, Y. Isolation of the Novel Phage PHB09 and Its Potential Use against the Plant Pathogen Pseudomonas syringae pv. actinidiae. Viruses 2021, 13. [Google Scholar] [CrossRef]
- Aziz, R.K.; Bartels, D.; Best, A.A.; DeJongh, M.; Disz, T.; Edwards, R.A.; Formsma, K.; Gerdes, S.; Glass, E.M.; Kubal, M.; et al. The RAST server: Rapid annotations using subsystems technology. BMC Genom. 2008, 9, 75. [Google Scholar] [CrossRef] [PubMed]
- Saitou, N.; Nei, M. The neighbor-joining method: a new method for reconstructing evolutionary trees. Mol. Biol. Evol. 1987, 4, 406–425. [Google Scholar] [CrossRef] [PubMed]
- Sullivan, M.J.; Petty, N.K.; Beatson, S.A. Easyfig: A genome comparison visualizer. Bioinformatics 2011, 27, 1009–1010. [Google Scholar] [CrossRef] [PubMed]
- Moraru, C.; Varsani, A.; Kropinski, A.M. VIRIDIC—A Novel Tool to Calculate the Intergenomic Similarities of Prokaryote-Infecting Viruses. Viruses 2020, 12, 1268. [Google Scholar] [CrossRef] [PubMed]
- Nayak, T.; Kakkar, A.; Singh, R.K.; Jaiswal, L.K.; Singh, A.K.; Temple, L.; Gupta, A. Isolation and characterization of a novel mycobacteriophage Kashi-VT1 infecting Mycobacterium species. Front. Cell. Infect. Microbiol. 2023, 13, 1173894. [Google Scholar] [CrossRef] [PubMed]
- Hyman, P.; Abedon, S.T. Bacteriophage host range and bacterial resistance. Adv. Appl. Microbiol. 2010, 70, 217–248. [Google Scholar] [PubMed]
- Park, D.-W.; Lim, G.-y.; Lee, Y.-d.; Park, J.-H. Characteristics of lytic phage vB_EcoM-ECP26 and reduction of shiga-toxin producing Escherichia coli on produce romaine. Appl. Biol. Chem. 2020, 63, 19. [Google Scholar] [CrossRef]
- Gillis, A.; Mahillon, J. An improved method for rapid generation and screening of Bacillus thuringiensis phage-resistant mutants. J. Microbiol. Methods 2014, 106, 101–103. [Google Scholar] [CrossRef]
- Shen, Q.; Zhou, X.-T.; Guo, Q.; Xue, Y.-P.; Zheng, Y.-G. Engineering laboratory/factory-specific phage-resistant strains of Escherichia coli by mutagenesis and screening. World J. Microbiol. Biotechnol. 2022, 38, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Li, P.; Lin, H.; Mi, Z.; Xing, S.; Tong, Y.; Wang, J. Screening of Polyvalent Phage-Resistant Escherichia coli Strains Based on Phage Receptor Analysis. Front. Microbiol. 2019, 10, 850. [Google Scholar] [CrossRef] [PubMed]
- Tang, X.; Shen, Y.; Song, X.; Benghezal, M.; Marshall, B.J.; Tang, H.; Li, H. Reassessment of the Broth Microdilution Method for Susceptibility Testing of Helicobacter pylori. J. Infect. Dis. 2022, 226, S486–S492. [Google Scholar] [CrossRef] [PubMed]
|
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).