Submitted:
20 July 2024
Posted:
24 July 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Input Files
2.2. Performing Mixture Deconvolution
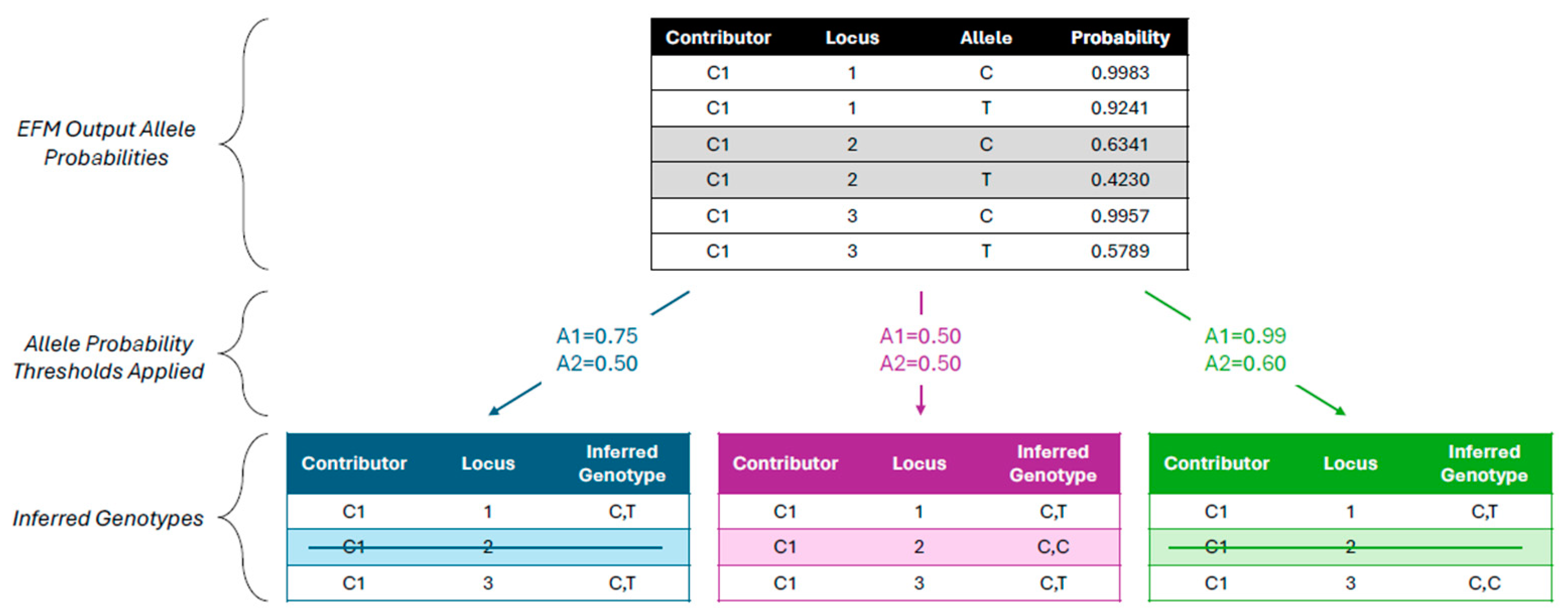
2.3. Inferring SNP Genotypes from EFM Results
2.4. Creating GEDmatch® PRO-Formatted Reports
2.5. Calculating Validation Metrics
2.6. Verification of Software Functions and Generation of Example Results
3. Results and Discussion
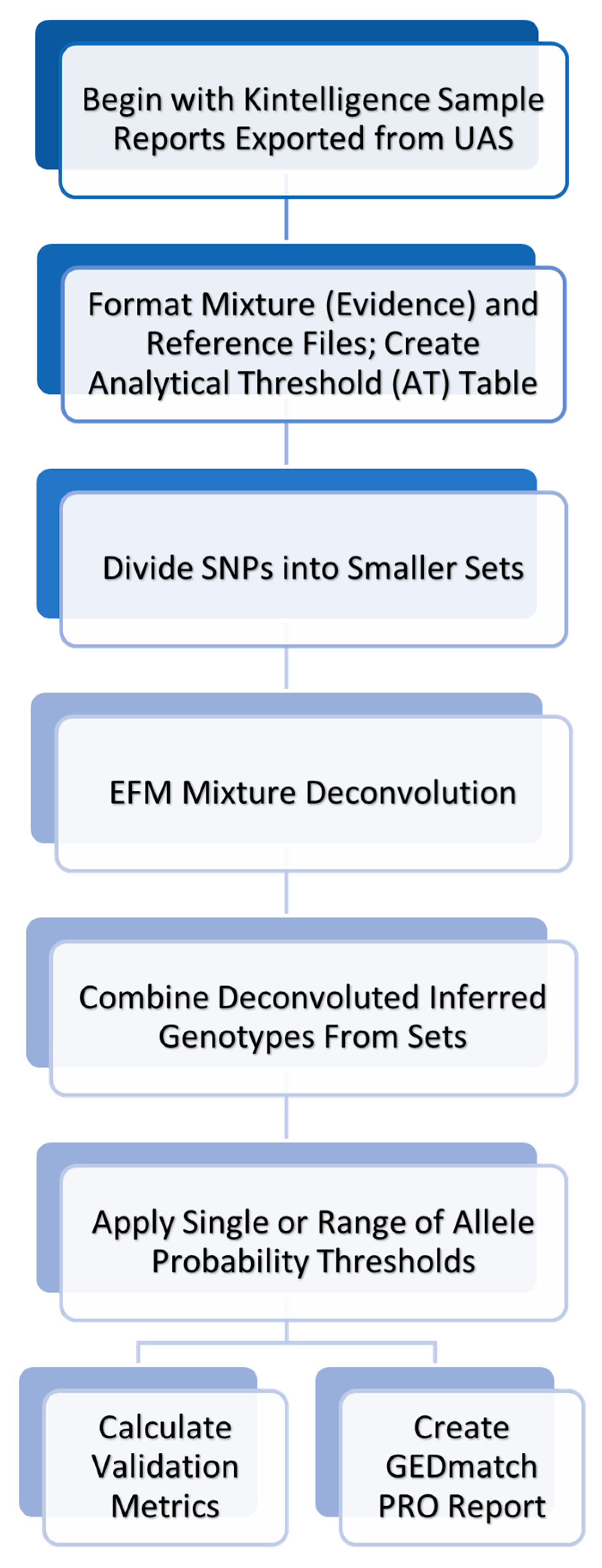
3.1. MixDeR Workflow
3.1.1. Use of EFM Allele Probabilities Rather Than Genotype Probabilities
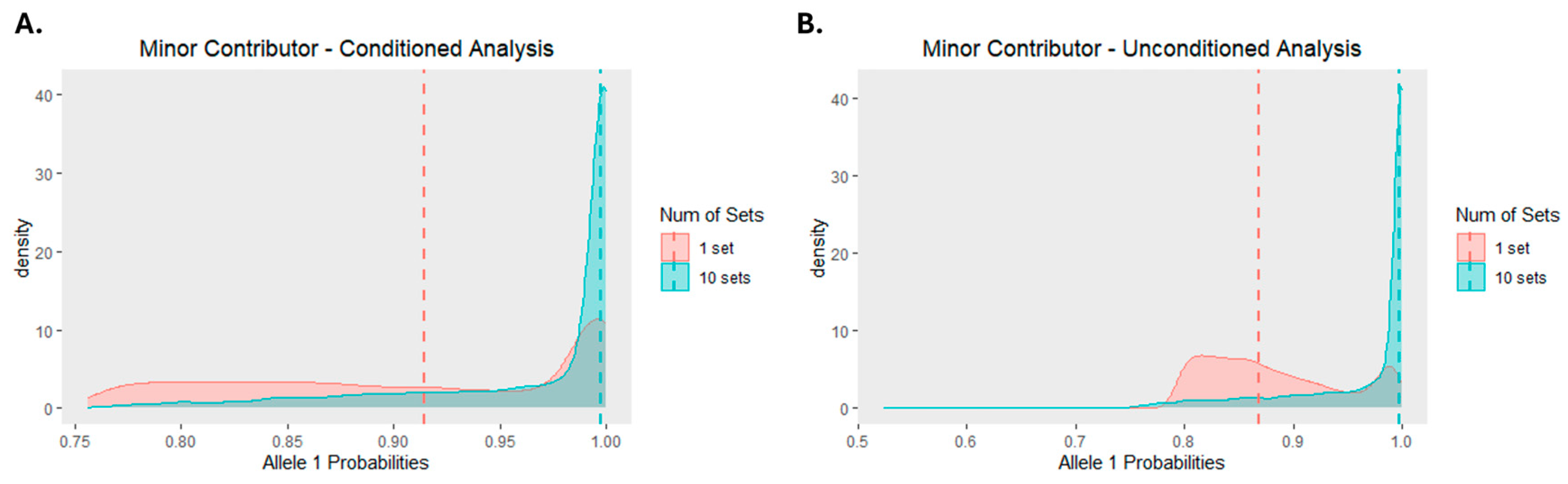
3.1.2. Separation of SNPs into Sets
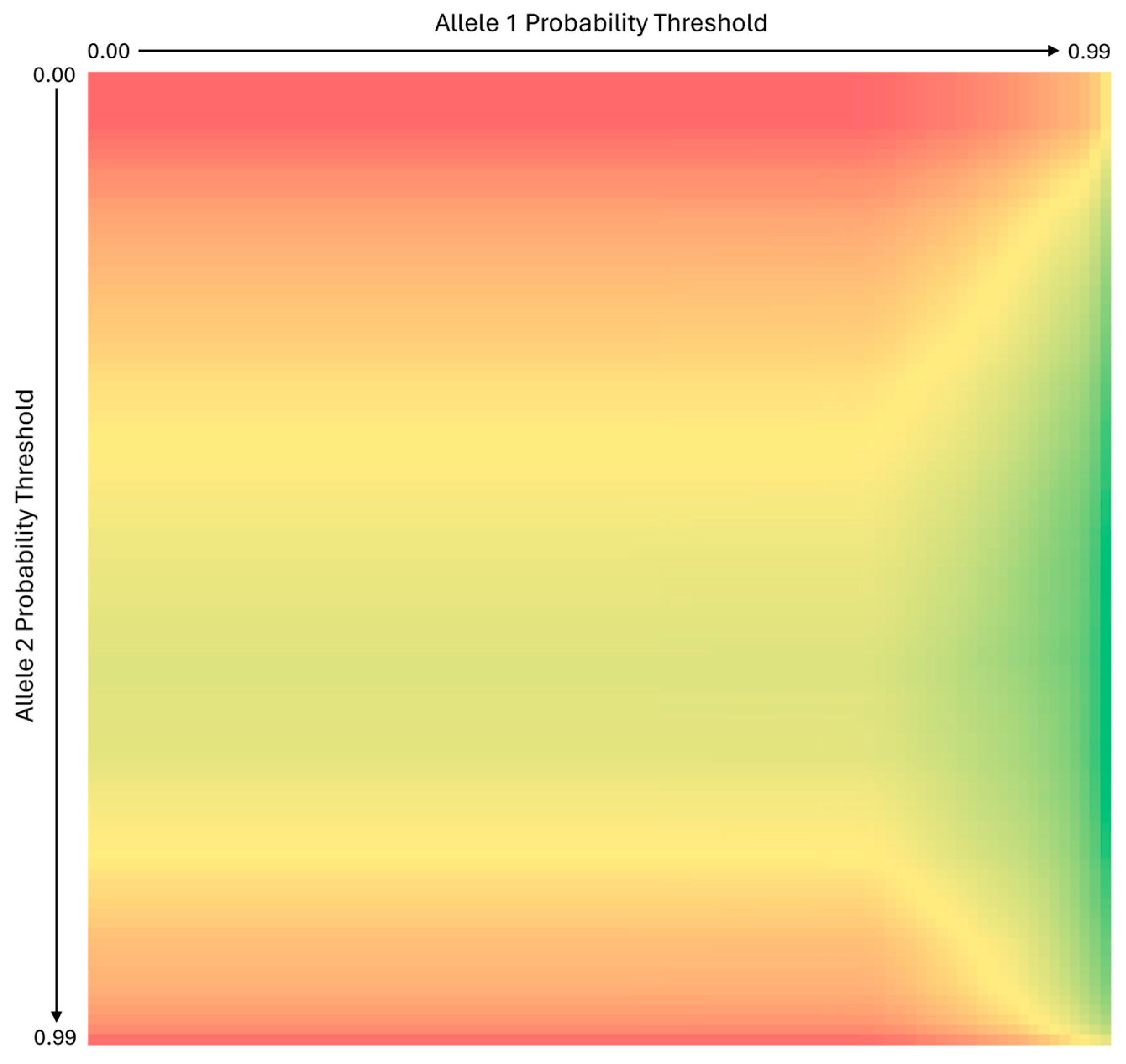
3.1.3. Use of Independent Allele 1 and Allele 2 Probability Thresholds
3.1.4. Use of a Minimum Number of SNPs
3.2. MixDeR GUI
- using different allele probability threshold combinations;
- using different minimum SNP numbers;
- from mixture profiles developed using differing DNA inputs;
- from mixtures with differing contributor ratios;
- from conditioned versus unconditioned deconvolutions; and
- from single versus replicate profiles.
3.3. Software Verification and Example Results
3.4. Challenges
- If a mixture ratio of exactly 1:1 is predicted by EFM, the alleles and allele probabilities for both contributors in the EFM output will be the same. However, even when the EFM-predicted mixture ratio was not exactly 1:1, we encountered instances in which the alleles and allele probabilities for both contributors in the EFM output were identical.
- In the EFM “All Marginal (A)” output, the contributor 1 allele probabilities should always be higher than contributor 2 allele probabilities. However, we encountered instances in which the opposite occurred (the contributor 2 allele probabilities were higher than the contributor 1 allele probabilities).
3.5. Limitations
3.6. Current and Future Work
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Gill, P. An assessment of the utility of single nucleotide polymorphisms (SNPs) for forensic purposes. Int. J. Leg. Med. 2001, 114, 204–210. [Google Scholar] [CrossRef] [PubMed]
- Budowle, B. SNP typing strategies. Forensic Science International 2004, 2, 139–142. [Google Scholar]
- Sobrino, B.; Carracedo, A. SNP Typing in Forensic Genetics: A Review. in Methods in Molecular Biology, 2005, 297, 107–126. [Google Scholar] [CrossRef]
- Kayser, M.; de Knijff, P. Improving human forensics through advances in genetics, genomics and molecular biology. Nature Reviews 2011, 12, 179–192. [Google Scholar] [CrossRef] [PubMed]
- Sanchez, J.J.; Phillips, C.; Børsting, C.; Balogh, K.; Bogus, M.; Fondevila, M.; Harrison, C.D.; Musgrave-Brown, E.; Salas, A.; Syndercombe-Court, D.; et al. A multiplex assay with 52 single nucleotide polymorphisms for human identification. Electrophoresis 2006, 27, 1713–1724. [Google Scholar] [CrossRef] [PubMed]
- Kidd, K.K.; Pakstis, A.J.; Speed, W.C.; Grigorenko, E.L.; Kajuna, S.L.; Karoma, N.J.; Kungulilo, S.; Kim, J.-J.; Lu, R.-B.; Odunsi, A.; et al. Developing a SNP panel for forensic identification of individuals. Forensic Sci. Int. 2006, 164, 20–32. [Google Scholar] [CrossRef] [PubMed]
- Phillips, C.; Fang, R.; Ballard, D.; Fondevila, M.; Harrison, C.; Hyland, F.; Musgrave-Brown, E.; Proff, C.; Ramos-Luis, E.; Sobrino, B.; et al. Evaluation of the Genplex SNP typing system and a 49plex forensic marker panel. Forensic Sci. Int. Genet. 2007, 1, 180–185. [Google Scholar] [CrossRef] [PubMed]
- Phillips, C.; Salas, A.; Sánchez, J.; Fondevila, M.; Gómez-Tato, A.; Álvarez-Dios, J.; Calaza, M.; de Cal, M.C.; Ballard, D.; Lareu, M.; et al. Inferring ancestral origin using a single multiplex assay of ancestry-informative marker SNPs. Forensic Sci. Int. Genet. 2007, 1, 273–280. [Google Scholar] [CrossRef]
- Tomas, C.; Stangegaard, M.; Børsting, C.; Hansen, A.J.; Morling, N.; The SNPforID Consortium. Typing of 48 autosomal SNPs and amelogenin with GenPlex SNP genotyping system in forensic genetics. Forensic Sci. Int. Genet. 2008, 3, 1–6. [Google Scholar] [CrossRef]
- Sanchez, J.; Børsting, C.; Balogh, K.; Berger, B.; Bogus, M.; Butler, J.; Carracedo, A.; Court, D.S.; Dixon, L.; Filipović, B.; et al. Forensic typing of autosomal SNPs with a 29 SNP-multiplex—Results of a collaborative EDNAP exercise. Forensic Sci. Int. Genet. 2008, 2, 176–183. [Google Scholar] [CrossRef]
- Fang, R.; Pakstis, A.J.; Hyland, F.; Wang, D.; Shewale, J.; Kidd, J.R.; Kidd, K.K.; Furtado, M.R. Multiplexed SNP detection panels for human identification. Forensic Science International: Genetics Supplement Series 2009, 2, 538–539. [Google Scholar] [CrossRef]
- Kosoy, R.; Nassir, R.; Tian, C.; White, P.A.; Butler, L.M.; Silva, G.; Kittles, R.; Alarcon-Riquelme, M.E.; Gregersen, P.K.; Belmont, J.W.; et al. Ancestry informative marker sets for determining continental origin and admixture proportions in common populations in America. Hum. Mutat. 2009, 30, 69–78. [Google Scholar] [CrossRef] [PubMed]
- Liu, F.; van Duijn, K.; Vingerling, J.R.; Hofman, A.; Uitterlinden, A.G.; Janssens, A.C.J.; Kayser, M. Eye color and the prediction of complex phenotypes from genotypes. Curr. Biol. 2009, 19, R192–R193. [Google Scholar] [CrossRef] [PubMed]
- Pakstis, A.J.; Speed, W.C.; Fang, R.; Hyland, F.C.L.; Furtado, M.R.; Kidd, J.R.; Kidd, K.K. SNPs for a universal individual identification panel. Hum. Genet. 2009, 127, 315–324. [Google Scholar] [CrossRef] [PubMed]
- Walsh, S.; Liu, F.; Ballantyne, K.N.; van Oven, M.; Lao, O.; Kayser, M. IrisPlex: A sensitive DNA tool for accurate prediction of blue and brown eye colour in the absence of ancestry information. Forensic Sci. Int. Genet. 2010, 5, 170–180. [Google Scholar] [CrossRef] [PubMed]
- Branicki, W.; Liu, F.; van Duijn, K.; Draus-Barini, J.; Pośpiech, E.; Walsh, S.; Kupiec, T.; Wojas-Pelc, A.; Kayser, M. Model-based prediction of human hair color using DNA variants. Hum. Genet. 2011, 129, 443–454. [Google Scholar] [CrossRef]
- Kayser, M. Forensic DNA Phenotyping: Predicting human appearance from crime scene material for investigative purposes. Forensic Sci. Int. Genet. 2015, 18, 33–48. [Google Scholar] [CrossRef] [PubMed]
- Phillips, C. Forensic genetic analysis of bio-geographical ancestry. Forensic Sci. Int. Genet. 2015, 18, 49–65. [Google Scholar] [CrossRef]
- Daniel, R.; Santos, C.; Phillips, C.; Fondevila, M.; van Oorschot, R.; Carracedo, A.; Lareu, M.; McNevin, D. A SNaPshot of next generation sequencing for forensic SNP analysis. Forensic Sci. Int. Genet. 2014, 14, 50–60. [Google Scholar] [CrossRef]
- Köcher, S.; Müller, P.; Berger, B.; Bodner, M.; Parson, W.; Roewer, L.; Willuweit, S. Inter-laboratory validation study of the ForenSeq™ DNA Signature Prep Kit. Forensic Sci. Int. Genet. 2018, 36, 77–85. [Google Scholar] [CrossRef]
- Hollard, C.; Ausset, L.; Chantrel, Y.; Jullien, S.; Clot, M.; Faivre, M.; Suzanne, E.; Pène, L.; Laurent, F.-X. Automation and developmental validation of the ForenSeq™ DNA Signature Preparation kit for high-throughput analysis in forensic laboratories. Forensic Sci. Int. Genet. 2019, 40, 37–45. [Google Scholar] [CrossRef]
- Brandhagen, M.D.; Just, R.S.; Irwin, J.A. Validation of NGS for mitochondrial DNA casework at the FBI Laboratory. Forensic Sci. Int. Genet. 2020, 44, 102151. [Google Scholar] [CrossRef]
- Silvery, J.; Ganschow, S.; Wiegand, P.; Tiemann, C. Developmental validation of the monSTR identity panel, a forensic STR multiplex assay for massively parallel sequencing. Forensic Sci. Int. Genet. 2020, 46, 102236. [Google Scholar] [CrossRef] [PubMed]
- Cihlar, J.C.; Amory, C.; Lagacé, R.; Roth, C.; Parson, W.; Budowle, B. Developmental Validation of a MPS Workflow with a PCR-Based Short Amplicon Whole Mitochondrial Genome Panel. Genes 2020, 11, 1345. [Google Scholar] [CrossRef]
- Cuenca, D.; Battaglia, J.; Halsing, M.; Sheehan, S. Mitochondrial Sequencing of Missing Persons DNA Casework by Implementing Thermo Fisher’s Precision ID mtDNA Whole Genome Assay. Genes 2020, 11, 1303. [Google Scholar] [CrossRef] [PubMed]
- Gandotra, N.; Speed, W.C.; Qin, W.; Tang, Y.; Pakstis, A.J.; Kidd, K.K.; Scharfe, C. Validation of novel forensic DNA markers using multiplex microhaplotype sequencing. Forensic Sci. Int. Genet. 2020, 47, 102275–102275. [Google Scholar] [CrossRef]
- Senst, A.; Caliebe, A.; Scheurer, E.; Schulz, I. Validation and beyond: Next generation sequencing of forensic casework samples including challenging tissue samples from altered human corpses using the MiSeq FGx system. J. Forensic Sci. 2022. [Google Scholar] [CrossRef] [PubMed]
- Kocsis, B.; Matrai, N.; Barany, G.; Tomory, G.; Heinrich, A.; Egyed, B. Internal Validation of the Precision Id Globalfiler Ngs Str Panel V2 Kit with Locus-Specific Analytical Threshold, and with Special Regard to Mixtures and Low Template DNA Detection. [CrossRef]
- de Vries, J.H.; Kling, D.; Vidaki, A.; Arp, P.; Kalamara, V.; Verbiest, M.M.; Piniewska-Róg, D.; Parsons, T.J.; Uitterlinden, A.G.; Kayser, M. Impact of SNP microarray analysis of compromised DNA on kinship classification success in the context of investigative genetic genealogy. Forensic Sci. Int. Genet. 2022, 56, 102625. [Google Scholar] [CrossRef] [PubMed]
- Davawala, A.; Stock, A.; Spiden, M.; Daniel, R.; McBain, J.; Hartman, D. Forensic genetic genealogy using microarrays for the identification of human remains: The need for good quality samples—A pilot study. Forensic Sci. Int. 2022, 334, 111242. [Google Scholar] [CrossRef]
- Russell, D.A.; Gorden, E.M.; Peck, M.A.; Neal, C.M.; Heaton, M.C.; Bouchet, J.L.; Koeppel, A.F.; Ciuzio, E.; Turner, S.D.; Reedy, C.R. Developmental validation of the illumina infinium assay using the global screening array (GSA) on the iScan system for use in forensic laboratories. Forensic Genomics 2023, 3, 15–24. [Google Scholar] [CrossRef]
- Phillis, C. The Golden State Killer investigation and the nascent field of forensic genealogy. Forensic Science International: Genetics 2018, 36, 186–188. [Google Scholar] [CrossRef]
- Wickenheiser, R.A. Forensic genealogy, bioethics and the Golden State Killer case. Forensic Sci. Int. Synerg. 2019, 1, 114–125. [Google Scholar] [CrossRef] [PubMed]
- Kling, D.; Phillips, C.; Kennett, D.; Tillmar, A. Investigative genetic genealogy: Current methods, knowledge and practice. Forensic Sci. Int. Genet. 2021, 52, 102474. [Google Scholar] [CrossRef]
- Dowdeswell, T.L. Forensic genetic genealogy: A profile of cases solved. Forensic Sci. Int. Genet. 2022, 58, 102679. [Google Scholar] [CrossRef]
- Dowdeswell, T.L. Forensic genetic genealogy project v. 2022. Mendeley Data, vol. 1, 2023.
- Snedecor, J.; Fennell, T.; Stadick, S.; Homer, N.; Antunes, J.; Stephens, K.; Holt, C. Fast and accurate kinship estimation using sparse SNPs in relatively large database searches. Forensic Sci. Int. Genet. 2022, 61, 102769. [Google Scholar] [CrossRef]
- Antunes, J.; Walichiewicz, P.; Forouzmand, E.; Barta, R.; Didier, M.; Han, Y.; Perez, J.; Snedecor, J.; Zlatkov, C.; Padmabandu, G.; et al. Developmental validation of the ForenSeq® Kintelligence kit, MiSeq FGx® sequencing system and ForenSeq Universal Analysis Software. Forensic Science International: Genetics 2024, 71, 103055. [Google Scholar] [CrossRef]
- Mitchell, R.; Enke, S.; Eskey, K.; Ferguson, T.; Just, R. A method to enable forensic genetic genealogy investigations from DNA mixtures. Forensic Sci. Int. Genet. Suppl. Ser. 2022, 8, 159–161. [Google Scholar] [CrossRef]
- Bleka. ; Storvik, G.; Gill, P. EuroForMix: An open source software based on a continuous model to evaluate STR DNA profiles from a mixture of contributors with artefacts. Forensic Sci. Int. Genet. 2015, 21, 35–44. [Google Scholar] [CrossRef]
- Verogen. Universal Analysis Software v2.0 and above - Reference Guide. 2021.
- Genomes Project Consortium. A global reference for human genetic variation. Nature 2015, 526, 68–74. [Google Scholar] [CrossRef]
- Karczewski, K.J.; Francioli, L.C.; Tiao, G.; Cummings, B.B.; Alfoldi, J.; Wang, Q.; Collins, R.L.; Laricchia, K.M.; Ganna, A.; Birnbaum, D.P.; et al. The mutational constraint spectrum quantified from variation in 141,456 humans. Nature 2020, 581, 434–443. [Google Scholar] [CrossRef]
- Peck, M.A.; Koeppel, A.F.; Gorden, E.M.; Bouchet, J.L.; Heaton, M.C.; Russell, D.A.; Reedy, C.R.; Neal, C.M.; Turner, S.D. Internal Validation of the ForenSeq Kintelligence Kit for Application to Forensic Genetic Genealogy. Forensic Genom. 2022, 2, 103–114. [Google Scholar] [CrossRef]
- Bleka, O.; Eduardoff, M.; Santos, C.; Phillips, C.; Parson, W.; Gill, P. Open source software EuroForMix can be used to analyse complex SNP mixtures. Forensic Sci. Int. Genet. 2017, 31, 105–110. [Google Scholar] [CrossRef] [PubMed]
- Gill, P.; Just, R.; Parson, W.; Phillips, C.; Bleka, O. Interpretation of complex DNA profiles generated by massively parallel sequencing. In Forensic Practitioner's Guide to the Interpretation of Complex DNA Mixtures; Elsevier, 2020; pp. 419–451.
- Hwa, H.-L.; Wu, M.-Y.; Chung, W.-C.; Ko, T.-M.; Lin, C.-P.; Yin, H.-I.; Lee, T.-T.; Lee, J.C.-I. Massively parallel sequencing analysis of nondegraded and degraded DNA mixtures using the ForenSeq™ system in combination with EuroForMix software. Int. J. Leg. Med. 2018, 133, 25–37. [Google Scholar] [CrossRef] [PubMed]
- Hwa, H.-L.; Wu, M.-Y.; Hsu, P.-M.; Yin, H.-I.; Lee, T.-T.; Su, C.-W. DNA mixture analyses of autosomal single nucleotide polymorphisms for individual identification using droplet digital polymerase-chain reaction and massively parallel sequencing in combination with EuroFormix. Aust. J. Forensic Sci. 2023, 1–14. [Google Scholar] [CrossRef]
- Woerner, A.E.; Crysup, B.; King, J.L.; Novroski, N.M.; Coble, M.D. Mixture detection with Demixtify. Forensic Science International: Genetics 2024, 69, 102980. [Google Scholar] [CrossRef]
| Deconvolution Type | Unknown Contributor | Number of SNP Sets |
Number of SNPs in Inferred Genotype |
Inferred Genotype Accuracy |
|---|---|---|---|---|
| Conditioned | Major | 1 | 6,000 | 92.32% |
| Major | 10 | 9,898 | 99.11% | |
| Conditioned | Minor | 1 | 6,000 | 73.46% |
| Minor | 10 | 6,000 | 84.19% | |
| Unconditioned | Major | 1 | 6,000 | 62.12% |
| Major | 10 | 9,632 | 98.91% | |
| Unconditioned | Minor | 1 | 6,000 | 62.25% |
| Minor | 10 | 6,000 | 83.32% |
| Deconvolution Type | Unknown Contributor | Kintelligence Profile(s) Used |
Number of SNPS in Inferred Genotypes |
Inferred Genotype Accuracy | Heterozygosity |
|---|---|---|---|---|---|
| Conditioned | Major | Single | 9,898 | 99.11% | 45.77% |
| Major | Replicates | 10,035 | 99.27% | 46.44% | |
| Conditioned | Minor | Single | 6,000 | 84.19% | 33.30% |
| Minor | Replicates | 6,000 | 90.17% | 32.90% | |
| Unconditioned | Major | Single | 9,632 | 98.91% | 44.38% |
| Major | Replicates | 10,036 | 99.05% | 46.52% | |
| Unconditioned | Minor | Single | 6,000 | 83.32% | 32.50% |
| Minor | Replicates | 6,000 | 90.45% | 32.12% |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




