Submitted:
19 July 2024
Posted:
22 July 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Impact of Salt and Drought Stress on Rice
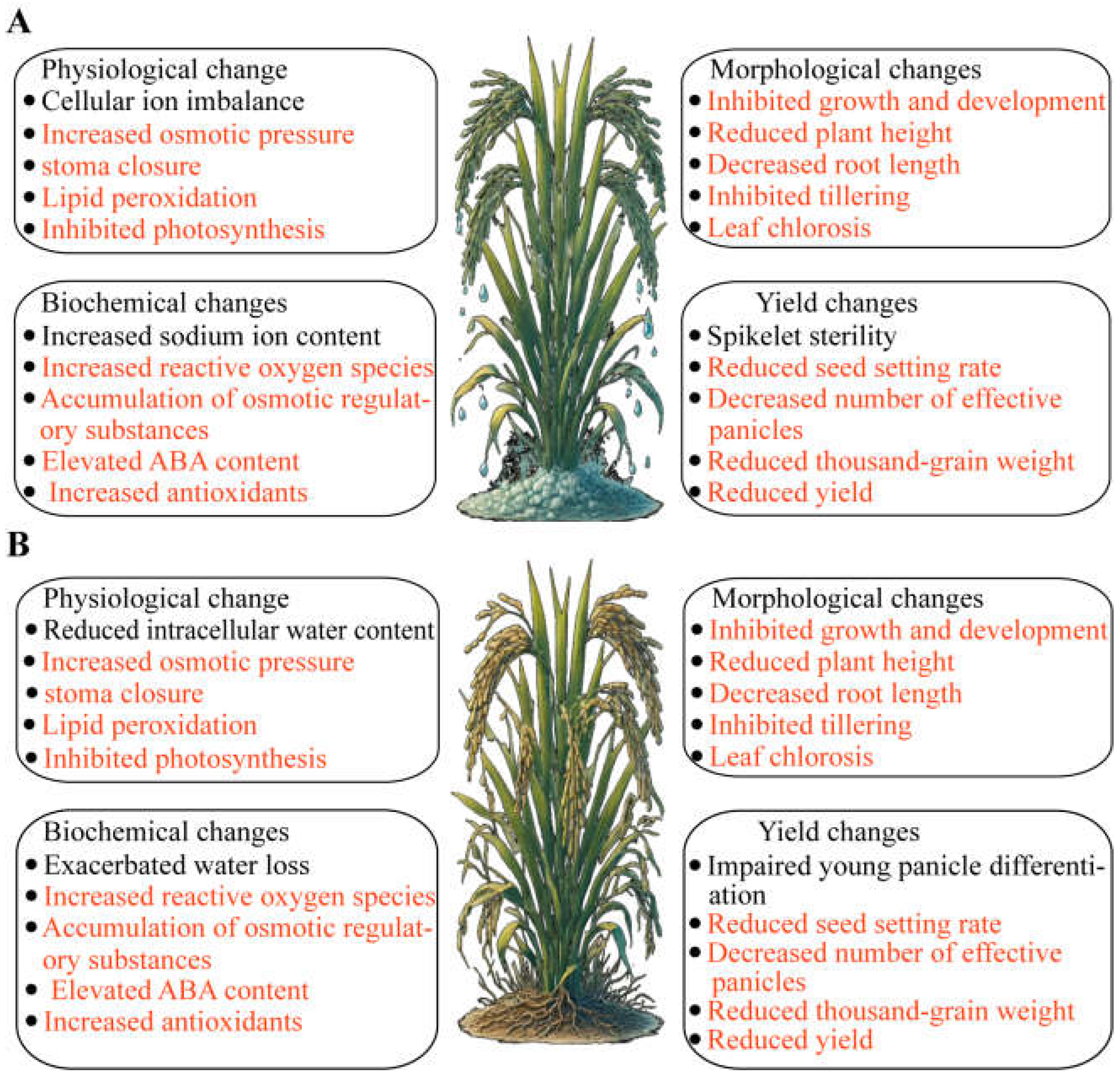
2.1. Effects on Morphology, Physiology, Biochemistry, and Yield
2.2. Impact on Rice Quality
3. Mechanisms of Rice Responses to Salt and Drought Stress
3.1. Synthesis and Accumulation of Osmolytes
3.2. Regulation of Ion Balance
3.3. Regulation of the Antioxidant System
4. Transcriptional Regulatory Networks
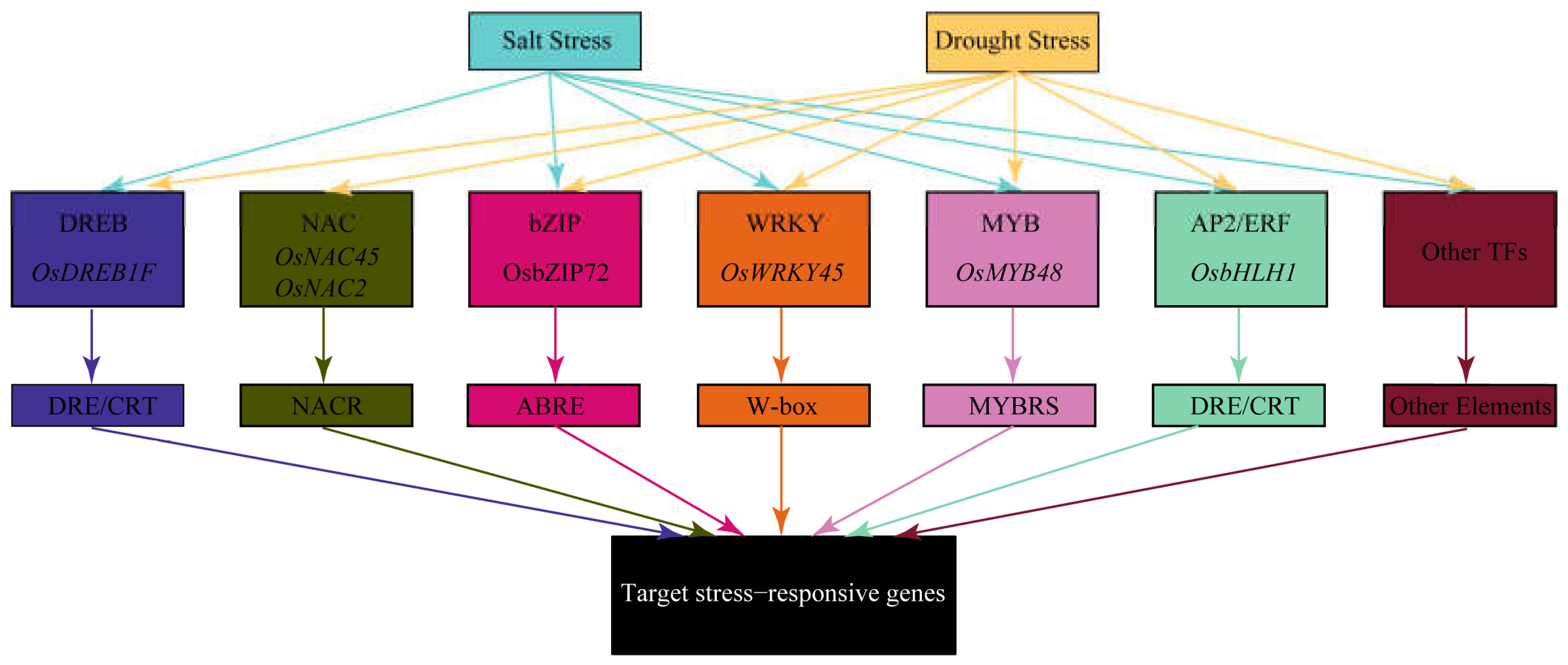
4.1. DREB Transcription Factors
4.2. NAC Transcription Factors
4.3. bZIP Transcription Factors
4.4. WRKY Transcription Factors
4.5. MYB Transcription Factors
4.6. AP2/ERF Transcription Factors
5. Regulation by Plant Hormones
5.1. Abscisic Acid (ABA)
5.2. Gibberellins (GA)
5.3. Other Hormones
6. Breeding for Salt and Drought Tolerance in Rice
7. Future Research Directions and Application Prospects
7.1. Epigenetic Regulation
7.2. Microbial Symbiosis
7.3. Smart Agriculture Technologies
7.4. Research on Combined Stresses and Multi-Stage Stress Tolerance
7.5. Improvement of Water-Saving and Drought-Resistant Rice for Salt-Alkaline Tolerance
8. Conclusions
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
Ethical Approval
References
- Shaar-Moshe, L., Blumwald, E., & Peleg, Z. (2017). Unique Physiological and Transcriptional Shifts under Combinations of Salinity, Drought, and Heat. Plant physiology, 174(1), 421–434. [CrossRef]
- Munns, R., & Tester, M. (2008). Mechanisms of salinity tolerance. Annual review of plant biology, 59, 651–681. [CrossRef]
- Zhu J. K. (2002). Salt and drought stress signal transduction in plants. Annual review of plant biology, 53, 247–273. [CrossRef]
- Alfatih, A., Zhang, J., Song, Y., Jan, S. U., Zhang, Z. S., Xia, J. Q., Zhang, Z. Y., Nazish, T., Wu, J., Zhao, P. X., & Xiang, C. B. (2023). Nitrate-responsive OsMADS27 promotes salt tolerance in rice. Plant communications, 4(2), 100458. [CrossRef]
- Nongpiur, R. C., Singla-Pareek, S. L., & Pareek, A. (2016). Genomics Approaches For Improving Salinity Stress Tolerance in Crop Plants. Current genomics, 17(4), 343–357. [CrossRef]
- Zhu J. K. (2001). Plant salt tolerance. Trends in plant science, 6(2), 66–71. [CrossRef]
- Razzaq, A., Ali, A., Safdar, L. B., Zafar, M. M., Rui, Y., Shakeel, A., Shaukat, A., Ashraf, M., Gong, W., & Yuan, Y. (2020). Salt stress induces physiochemical alterations in rice grain composition and quality. Journal of food science, 85(1), 14–20. [CrossRef]
- Khan, M. I. R., Palakolanu, S. R., Chopra, P., Rajurkar, A. B., Gupta, R., Iqbal, N., & Maheshwari, C. (2021). Improving drought tolerance in rice: Ensuring food security through multi-dimensional approaches. Physiologia plantarum, 172(2), 645–668. [CrossRef]
- Farooq, M., Wahid, A., Kobayashi, N., Fujita, D., & Basra, S. M. A. (2009). Plant drought stress: Effects, mechanisms and management. Agronomy for Sustainable Development, 29(1), 185-212. [CrossRef]
- Zheng, C., Niu, S., Yan, Y., Zhou, G., Peng, Y., He, Y., Zhou, J., Li, Y., & Xie, X. (2024). Moderate Salinity Stress Affects Rice Quality by Influencing Expression of Amylose- and Protein-Content-Associated Genes. International journal of molecular sciences, 25(7), 4042. [CrossRef]
- Thitisaksakul, M., Tananuwong, K., Shoemaker, C. F., Chun, A., Tanadul, O. U., Labavitch, J. M., & Beckles, D. M. (2015). Effects of timing and severity of salinity stress on rice (Oryza sativa L.) yield, grain composition, and starch functionality. Journal of agricultural and food chemistry, 63(8), 2296–2304. [CrossRef]
- Shabala, S., & Cuin, T.A. (2008). Potassium transport and plant salt tolerance. Physiologia plantarum, 133 4, 651-69. [CrossRef]
- Zhan, C., Lei, L., Guo, H., Zhou, S., Xu, C., Liu, Z., Wu, Z., Deng, Y., Miao, Y., Han, Y., Zhang, M., Li, H., Huang, S., Yang, C., Zhang, F., Li, Y., Liu, L., Liu, X., Abbas, H. M. K., Fernie, A. R., … Luo, J. (2023). Disease resistance conferred by components of essential chrysanthemum oil and the epigenetic regulation of OsTPS1. Science China. Life sciences, 66(5), 1108–1118. [CrossRef]
- Wang, Q., Ni, L., Cui, Z., Jiang, J., Chen, C., & Jiang, M. (2022). The NADPH oxidase OsRbohA increases salt tolerance by modulating K+ homeostasis in rice. The Crop Journal. [CrossRef]
- Pandey, V., & Shukla, A.K. (2015). Acclimation and Tolerance Strategies of Rice under Drought Stress. Rice Science, 22, 147-161. [CrossRef]
- Sripinyowanich, S., Klomsakul, P., Boonburapong, B., Bangyeekhun, T., Asami, T., Gu, H., Buaboocha, T., & Chadchawan, S. (2013). Exogenous ABA induces salt tolerance in indica rice (Oryza sativa L.): The role of OsP5CS1 and OsP5CR gene expression during salt stress. Environmental and Experimental Botany, 86, 94-105. [CrossRef]
- Zang, B., Li, H., Li, W., Deng, X. W., & Wang, X. (2011). Analysis of trehalose-6-phosphate synthase (TPS) gene family suggests the formation of TPS complexes in rice. Plant molecular biology, 76(6), 507–522. [CrossRef]
- Wang, J., Nan, N., Li, N., Liu, Y., Wang, T. J., Hwang, I., Liu, B., & Xu, Z. Y. (2020). A DNA Methylation Reader-Chaperone Regulator-Transcription Factor Complex Activates OsHKT1;5 Expression during Salinity Stress. The Plant cell, 32(11), 3535–3558. [CrossRef]
- Wei, H., Wang, X., He, Y., Xu, H., & Wang, L. (2021). Clock component OsPRR73 positively regulates rice salt tolerance by modulating OsHKT2;1-mediated sodium homeostasis. The EMBO journal, 40(3), e105086. [CrossRef]
- Prodjinoto, H., Irakoze, W., Gandonou, C., Quinet, M., & Lutts, S. (2023). Comparison between the impact of osmotic and NaCl treatments on the expression of genes coding for ion transporters in Oryza glaberrima Steud. PloS one, 18(11), e0290752. [CrossRef]
- Wei, H., Wang, X., He, Y., Xu, H., & Wang, L. (2021). Clock component OsPRR73 positively regulates rice salt tolerance by modulating OsHKT2;1-mediated sodium homeostasis. The EMBO journal, 40(3), e105086. [CrossRef]
- Xiao, L., Shi, Y., Wang, R., Feng, Y., Wang, L., Zhang, H., Shi, X., Jing, G., Deng, P., Song, T., Jing, W., & Zhang, W. (2022). The transcription factor OsMYBc and an E3 ligase regulate expression of a K+ transporter during salt stress. Plant physiology, 190(1), 843–859. [CrossRef]
- Almeida, P., Katschnig, D., & de Boer, A. H. (2013). HKT transporters--state of the art. International journal of molecular sciences, 14(10), 20359–20385. [CrossRef]
- Zhang, M., Cao, Y., Wang, Z., Wang, Z. Q., Shi, J., Liang, X., Song, W., Chen, Q., Lai, J., & Jiang, C. (2018). A retrotransposon in an HKT1 family sodium transporter causes variation of leaf Na+ exclusion and salt tolerance in maize. The New phytologist, 217(3), 1161–1176. [CrossRef]
- Gill, S. S., & Tuteja, N. (2010). Reactive oxygen species and antioxidant machinery in abiotic stress tolerance in crop plants. Plant physiology and biochemistry: PPB, 48(12), 909–930. [CrossRef]
- Ray, P. D., Huang, B. W., & Tsuji, Y. (2012). Reactive oxygen species (ROS) homeostasis and redox regulation in cellular signaling. Cellular signalling, 24(5), 981–990. [CrossRef]
- Huang, X., Huang, L., Zhao, X., Jia, J., Zhang, G., Zhang, M., & Jiang, M. (2022). A J-Protein OsDjC46 Interacts with ZFP36 to Participate in ABA-Mediated Antioxidant Defense in Rice. Antioxidants (Basel, Switzerland), 11(2), 207. [CrossRef]
- Deng, P., Cao, C., Shi, X., Jiang, Q., Ge, J., Shen, L., Guo, C., Jiang, L., Jing, W., & Zhang, W. (2023). OsCYBDOMG1, a cytochrome b561 domain-containing protein, regulates salt tolerance and grain yield in rice. TAG. Theoretical and applied genetics. Theoretische und angewandte Genetik, 136(4), 76. [CrossRef]
- Wang, A., Shu, X., Jing, X., Jiao, C., Chen, L., Zhang, J., Ma, L., Jiang, Y., Yamamoto, N., Li, S., Deng, Q., Wang, S., Zhu, J., Liang, Y., Zou, T., Liu, H., Wang, L., Huang, Y., Li, P., & Zheng, A. (2021). Identification of rice (Oryza sativa L.) genes involved in sheath blight resistance via a genome-wide association study. Plant biotechnology journal, 19(8), 1553–1566. [CrossRef]
- Smita, S., Katiyar, A., Chinnusamy, V., Pandey, D. M., & Bansal, K. C. (2015). Transcriptional Regulatory Network Analysis of MYB Transcription Factor Family Genes in Rice. Frontiers in plant science, 6, 1157. [CrossRef]
- Jwa, N. S., Agrawal, G. K., Tamogami, S., Yonekura, M., Han, O., Iwahashi, H., & Rakwal, R. (2006). Role of defense/stress-related marker genes, proteins and secondary metabolites in defining rice self-defense mechanisms. Plant physiology and biochemistry: PPB, 44(5-6), 261–273. [CrossRef]
- Nuruzzaman, M., Sharoni, A. M., & Kikuchi, S. (2013). Roles of NAC transcription factors in the regulation of biotic and abiotic stress responses in plants. Frontiers in microbiology, 4, 248. [CrossRef]
- Liu, Q., Kasuga, M., Sakuma, Y., Abe, H., Miura, S., Yamaguchi-Shinozaki, K., & Shinozaki, K. (1998). Two transcription factors, DREB1 and DREB2, with an EREBP/AP2 DNA binding domain separate two cellular signal transduction pathways in drought- and low-temperature-responsive gene expression, respectively, in Arabidopsis. The Plant cell, 10(8), 1391–1406. [CrossRef]
- Sakuma, Y., Maruyama, K., Qin, F., Osakabe, Y., Shinozaki, K., & Yamaguchi-Shinozaki, K. (2006). Dual function of an Arabidopsis transcription factor DREB2A in water-stress-responsive and heat-stress-responsive gene expression. Proceedings of the National Academy of Sciences of the United States of America, 103(49), 18822–18827. [CrossRef]
- Mizoi, J., Shinozaki, K., & Yamaguchi-Shinozaki, K. (2012). AP2/ERF family transcription factors in plant abiotic stress responses. Biochimica et biophysica acta, 1819(2), 86–96. [CrossRef]
- Ke, Y., Yang, Z., Yu, S., Li, T., Wu, J., Gao, H., Fu, Y., & Luo, L. (2014). Characterization of OsDREB6 responsive to osmotic and cold stresses in rice. Journal of Plant Biology, 57, 150 - 161. [CrossRef]
- Wang, H., Lu, S., Guan, X., Jiang, Y., Wang, B., Hua, J., & Zou, B. (2022). Dehydration-Responsive Element Binding Protein 1C, 1E, and 1G Promote Stress Tolerance to Chilling, Heat, Drought, and Salt in Rice. Frontiers in plant science, 13, 851731. [CrossRef]
- Olsen, A. N., Ernst, H. A., Leggio, L. L., & Skriver, K. (2005). NAC transcription factors: structurally distinct, functionally diverse. Trends in plant science, 10(2), 79–87. [CrossRef]
- Yu, S., Huang, A., Li, J., Gao, L., Feng, Y., Pemberton, E., & Chen, C. (2018). OsNAC45 plays complex roles by mediating POD activity and the expression of development-related genes under various abiotic stresses in rice root. Plant Growth Regulation, 84, 519-531. [CrossRef]
- Jiang, D., Zhou, L., Chen, W., Ye, N., Xia, J., & Zhuang, C. (2019). Overexpression of a microRNA-targeted NAC transcription factor improves drought and salt tolerance in Rice via ABA-mediated pathways. Rice (New York, N.Y.), 12(1), 76. [CrossRef]
- Nijhawan, A., Jain, M., Tyagi, A. K., & Khurana, J. P. (2008). Genomic survey and gene expression analysis of the basic leucine zipper transcription factor family in rice. Plant physiology, 146(2), 333–350. [CrossRef]
- Fu, X., Liu, C., Li, Y., Liao, S., Cheng, H., Tu, Y., Zhu, X., Chen, K., He, Y., & Wang, G. (2021). The coordination of OsbZIP72 and OsMYBS2 with reverse roles regulates the transcription of OsPsbS1 in rice. The New phytologist, 229(1), 370–387. [CrossRef]
- Eulgem, T., Rushton, P. J., Robatzek, S., & Somssich, I. E. (2000). The WRKY superfamily of plant transcription factors. Trends in plant science, 5(5), 199–206. [CrossRef]
- Tao, Z., Kou, Y., Liu, H., Li, X., Xiao, J., & Wang, S. (2011). OsWRKY45 alleles play different roles in abscisic acid signalling and salt stress tolerance but similar roles in drought and cold tolerance in rice. Journal of experimental botany, 62(14), 4863–4874. [CrossRef]
- Katiyar, A., Smita, S., Lenka, S. K., Rajwanshi, R., Chinnusamy, V., & Bansal, K. C. (2012). Genome-wide classification and expression analysis of MYB transcription factor families in rice and Arabidopsis. BMC genomics, 13, 544. [CrossRef]
- Zhao, W., Wang, X., Zhang, Q., Zheng, Q., Yao, H., Gu, X., Liu, D., Tian, X., Wang, X., Li, Y., & Zhu, Z. (2022). H3K36 demethylase JMJ710 negatively regulates drought tolerance by suppressing MYB48-1 expression in rice. Plant physiology, 189(2), 1050–1064. [CrossRef]
- Licausi, F., Ohme-Takagi, M., & Perata, P. (2013). APETALA2/Ethylene Responsive Factor (AP2/ERF) transcription factors: mediators of stress responses and developmental programs. The New phytologist, 199(3), 639–649. [CrossRef]
- Wang, Y., Wang, J., Zhao, X., Yang, S., Huang, L., Du, F., Li, Z., Zhao, X., Fu, B., & Wang, W. (2020). Overexpression of the Transcription Factor Gene OsSTAP1 Increases Salt Tolerance in Rice. Rice (New York, N.Y.), 13(1), 50. [CrossRef]
- Verma, V., Ravindran, P., & Kumar, P. P. (2016). Plant hormone-mediated regulation of stress responses. BMC plant biology, 16, 86. [CrossRef]
- Cutler, S. R., Rodriguez, P. L., Finkelstein, R. R., & Abrams, S. R. (2010). Abscisic acid: emergence of a core signaling network. Annual review of plant biology, 61, 651–679. [CrossRef]
- Zhou, Z., Fan, J., Zhang, J., Yang, Y., Zhang, Y., Zan, X., Li, X., Wan, J., Gao, X., Chen, R., Huang, Z., Xu, Z., & Li, L. (2022). OsMLP423 Is a Positive Regulator of Tolerance to Drought and Salt Stresses in Rice. Plants (Basel, Switzerland), 11(13), 1653. [CrossRef]
- Choi, J., Lee, W., An, G., & Kim, S. R. (2021). OsCBE1, a Substrate Receptor of Cullin4-Based E3 Ubiquitin Ligase, Functions as a Regulator of Abiotic Stress Response and Productivity in Rice. International journal of molecular sciences, 22(5), 2487. [CrossRef]
- Colebrook, E. H., Thomas, S. G., Phillips, A. L., & Hedden, P. (2014). The role of gibberellin signalling in plant responses to abiotic stress. The Journal of experimental biology, 217(Pt 1), 67–75. [CrossRef]
- Wang, J., Qin, H., Zhou, S., Wei, P., Zhang, H., Zhou, Y., Miao, Y., & Huang, R. (2020). The Ubiquitin-Binding Protein OsDSK2a Mediates Seedling Growth and Salt Responses by Regulating Gibberellin Metabolism in Rice. The Plant cell, 32(2), 414–428. [CrossRef]
- Shan, C., Mei, Z., Duan, J., Chen, H., Feng, H., & Cai, W. (2014). OsGA2ox5, a gibberellin metabolism enzyme, is involved in plant growth, the root gravity response and salt stress. PloS one, 9(1), e87110. [CrossRef]
- Yang, C., Ma, B., He, S. J., Xiong, Q., Duan, K. X., Yin, C. C., Chen, H., Lu, X., Chen, S. Y., & Zhang, J. S. (2015). MAOHUZI6/ETHYLENE INSENSITIVE3-LIKE1 and ETHYLENE INSENSITIVE3-LIKE2 Regulate Ethylene Response of Roots and Coleoptiles and Negatively Affect Salt Tolerance in Rice. Plant physiology, 169(1), 148–165. [CrossRef]
- Yin, W., Xiao, Y., Niu, M., Meng, W., Li, L., Zhang, X., Liu, D., Zhang, G., Qian, Y., Sun, Z., Huang, R., Wang, S., Liu, C. M., Chu, C., & Tong, H. (2020). ARGONAUTE2 Enhances Grain Length and Salt Tolerance by Activating BIG GRAIN3 to Modulate Cytokinin Distribution in Rice. The Plant cell, 32(7), 2292–2306. [CrossRef]
- Singh, A. P., Mani, B., & Giri, J. (2021). OsJAZ9 is involved in water-deficit stress tolerance by regulating leaf width and stomatal density in rice. Plant physiology and biochemistry: PPB, 162, 161–170. [CrossRef]
- Singh, V. K., Singh, B. D., Kumar, A., Maurya, S., Krishnan, S. G., Vinod, K. K., Singh, M. P., Ellur, R. K., Bhowmick, P. K., & Singh, A. K. (2018). Marker-Assisted Introgression of Saltol QTL Enhances Seedling Stage Salt Tolerance in the Rice Variety "Pusa Basmati 1". International journal of genomics, 2018, 8319879. [CrossRef]
- Saminadane, T., Geddam, S., Krishnaswamy, P., Jothiganapathy, K., Tamilselvan, A., Ramadoss, B. R., Sri Hari Reddy, P., Singh, U. S., Singh, R. K., Platten, J. D., Gregorio, G. B., Singh, N. K., Bisht, D. S., Kota, S., Ponnuvel, S., & Guntupalli, P. (2024). Development of early maturing salt-tolerant rice variety KKL(R) 3 using a combination of conventional and molecular breeding approaches. Frontiers in genetics, 14, 1332691. [CrossRef]
- Dwivedi, P., Ramawat, N., Raju, D., Dhawan, G., Gopala Krishnan, S., Chinnusamy, V., Bhowmick, P. K., Vinod, K. K., Pal, M., Nagarajan, M., Ellur, R. K., Bollinedi, H., & Singh, A. K. (2022). Drought Tolerant Near Isogenic Lines of Pusa 44 Pyramided With qDTY2.1 and qDTY3.1, Show Accelerated Recovery Response in a High Throughput Phenomics Based Phenotyping. Frontiers in plant science, 12, 752730. [CrossRef]
- Shamsudin, N. A., Swamy, B. P., Ratnam, W., Sta Cruz, M. T., Raman, A., & Kumar, A. (2016). Marker assisted pyramiding of drought yield QTLs into a popular Malaysian rice cultivar, MR219. BMC genetics, 17, 30. [CrossRef]
- Dixit, S., Singh, A., Sandhu, N., Bhandari, A., Vikram, P., & Kumar, A. (2017). Combining drought and submergence tolerance in rice: marker-assisted breeding and QTL combination effects. Molecular breeding: new strategies in plant improvement, 37(12), 143. [CrossRef]
- Rekha, G., Abhilash Kumar, V., Gokulan, C. G., Koushik, M. B. V. N., Laxmi Prasanna, B., Kulkarni, S., Aleena, D., Harika, G., Hajira, S. K., Pranathi, K., Punniakoti, E., Kale, R. R., Dilip Kumar, T., Ayyappa, D., Anila, M., Sinha, P., Manohara, K. K., Padmavathi, G., Subba Rao, L. V., Laha, G. S., … Sundaram, R. M. (2022). DRR Dhan 58, a Seedling Stage Salinity Tolerant NIL of Improved Samba Mahsuri Shows Superior Performance in Multi-location Trials. Rice (New York, N.Y.), 15(1), 45. [CrossRef]
- Bizimana, J. B., Luzi-Kihupi, A., W Murori, R., & Singh, R. K. (2017). Identification of quantitative trait loci for salinity tolerance in rice (Oryza sativa L.) using IR29/Hasawi mapping population. Journal of genetics, 96(4), 571–582. [CrossRef]
- Mondal, S., Septiningsih, E. M., Singh, R. K., & Thomson, M. J. (2022). Mapping QTLs for Reproductive Stage Salinity Tolerance in Rice Using a Cross between Hasawi and BRRI dhan28. International journal of molecular sciences, 23(19), 11376. [CrossRef]
- Tiwari, S., Jain, M., Singla-Pareek, S. L., Bhalla, P. L., Singh, M. B., & Pareek, A. (2023). Pokkali: A Naturally Evolved Salt-Tolerant Rice Shows a Distinguished Set of lncRNAs Possibly Contributing to the Tolerant Phenotype. International journal of molecular sciences, 24(14), 11677. [CrossRef]
- Qin, H., Li, Y., & Huang, R. (2020). Advances and Challenges in the Breeding of Salt-Tolerant Rice. International journal of molecular sciences, 21(21), 8385. [CrossRef]
- Luo L. J. (2010). Breeding for water-saving and drought-resistance rice (WDR) in China. Journal of experimental botany, 61(13), 3509–3517. [CrossRef]
- Bizimana, J. B., Luzi-Kihupi, A., W Murori, R., & Singh, R. K. (2017). Identification of quantitative trait loci for salinity tolerance in rice (Oryza sativa L.) using IR29/Hasawi mapping population. Journal of genetics, 96(4), 571–582. [CrossRef]
- Mondal, S., Septiningsih, E. M., Singh, R. K., & Thomson, M. J. (2022). Mapping QTLs for Reproductive Stage Salinity Tolerance in Rice Using a Cross between Hasawi and BRRI dhan28. International journal of molecular sciences, 23(19), 11376. [CrossRef]
- de Ocampo, M. P., Ho, V. T., Thomson, M. J., Mitsuya, S., Yamauchi, A., & Ismail, A. M. (2022). QTL mapping under salt stress in rice using a Kalarata-Azucena population. Euphytica: Netherlands journal of plant breeding, 218(6), 74. [CrossRef]
- Miao, C., Wang, Z., Zhang, L., Yao, J., Hua, K., Liu, X., Shi, H., & Zhu, J. K. (2019). The grain yield modulator miR156 regulates seed dormancy through the gibberellin pathway in rice. Nature communications, 10(1), 3822. [CrossRef]
- Zhou, Y., Zhou, S., Wang, L., Wu, D., Cheng, H., Du, X., Mao, D., Zhang, C., & Jiang, X. (2020). miR164c and miR168a regulate seed vigor in rice. Journal of integrative plant biology, 62(4), 470–486. [CrossRef]
- Gao, Y., Zou, H., Wang, B., & Yuan, F. (2022). Progress and Applications of Plant Growth-Promoting Bacteria in Salt Tolerance of Crops. International journal of molecular sciences, 23(13), 7036. [CrossRef]
- Navarro, E., Costa, N., & Pereira, A. (2020). A Systematic Review of IoT Solutions for Smart Farming. Sensors (Basel, Switzerland), 20(15), 4231. [CrossRef]
- Zandalinas, S. I., & Mittler, R. (2022). Plant responses to multifactorial stress combination. The New phytologist, 234(4), 1161–1167. [CrossRef]
- Rivero, R. M., Mittler, R., Blumwald, E., & Zandalinas, S. I. (2022). Developing climate-resilient crops: improving plant tolerance to stress combination. The Plant journal: for cell and molecular biology, 109(2), 373–389. [CrossRef]
- Xia, H., Zhang, X., Liu, Y., Bi, J., Ma, X., Zhang, A., Liu, H., Chen, L., Zhou, S., Gao, H., Xu, K., Wei, H., Liu, G., Wang, F., Zhao, H., Luo, X., Hou, D., Lou, Q., Feng, F., Zhou, L., … Luo, L. (2022). Blue revolution for food security under carbon neutrality: A case from the water-saving and drought-resistance rice. Molecular plant, 15(9), 1401–1404. [CrossRef]


| QTL | Recipient Parent | Improvement Effect | Reference |
|---|---|---|---|
| Saltol | PusaBasmati1 (PB1) | 24 NILs with enhanced seedling-stage salt tolerance and similar traits to the parent | [59] |
| qDTY2.1; qDTY3.1 | Pusa 44 | 14 NILs with significantly better yield and grain quality under drought compared to the parent | [60] |
| qDTY2.2; qDTY3.1; qDTY12.1 | MR219 | NILs with higher yield and improved drought tolerance; different QTL combinations show varying performances | [61] |
| qDTY3.1; qDTY6.1; qDTY6.2 | TDK1 | High-yielding and drought-tolerant NILs | [62] |
| qDTY1.1; qDTY2.1; qDTY3.1; qDTY11.1 | Samba Mahsuri | Higher yield under drought conditions | [63] |
| qRL6.1; qRL12.1 | Hasawi × IR29 | Increased root length under salt stress | [64] |
| qPT3.1 | Hasawi × BRRI dhan28 | Improved tillering under salt stress | [65] |
| rkc3.1; rnc3.1 | Kalarata × Azucena | Enhanced root K+ and Na+ concentrations | [66] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




