Submitted:
18 July 2024
Posted:
19 July 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results
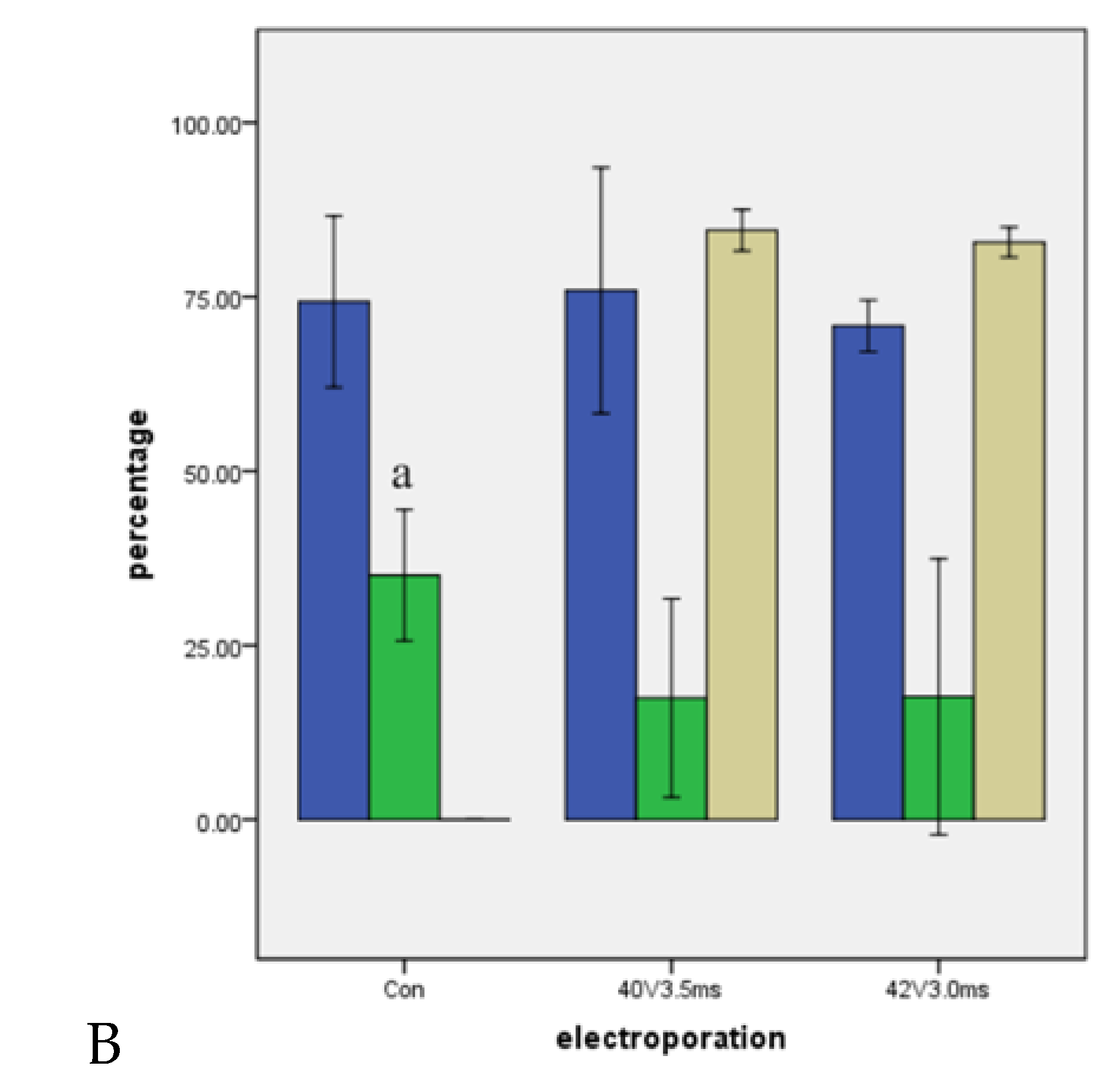
2.1. Effect of Different Electroporation Settings on Cleavage and Blastocyst Rates
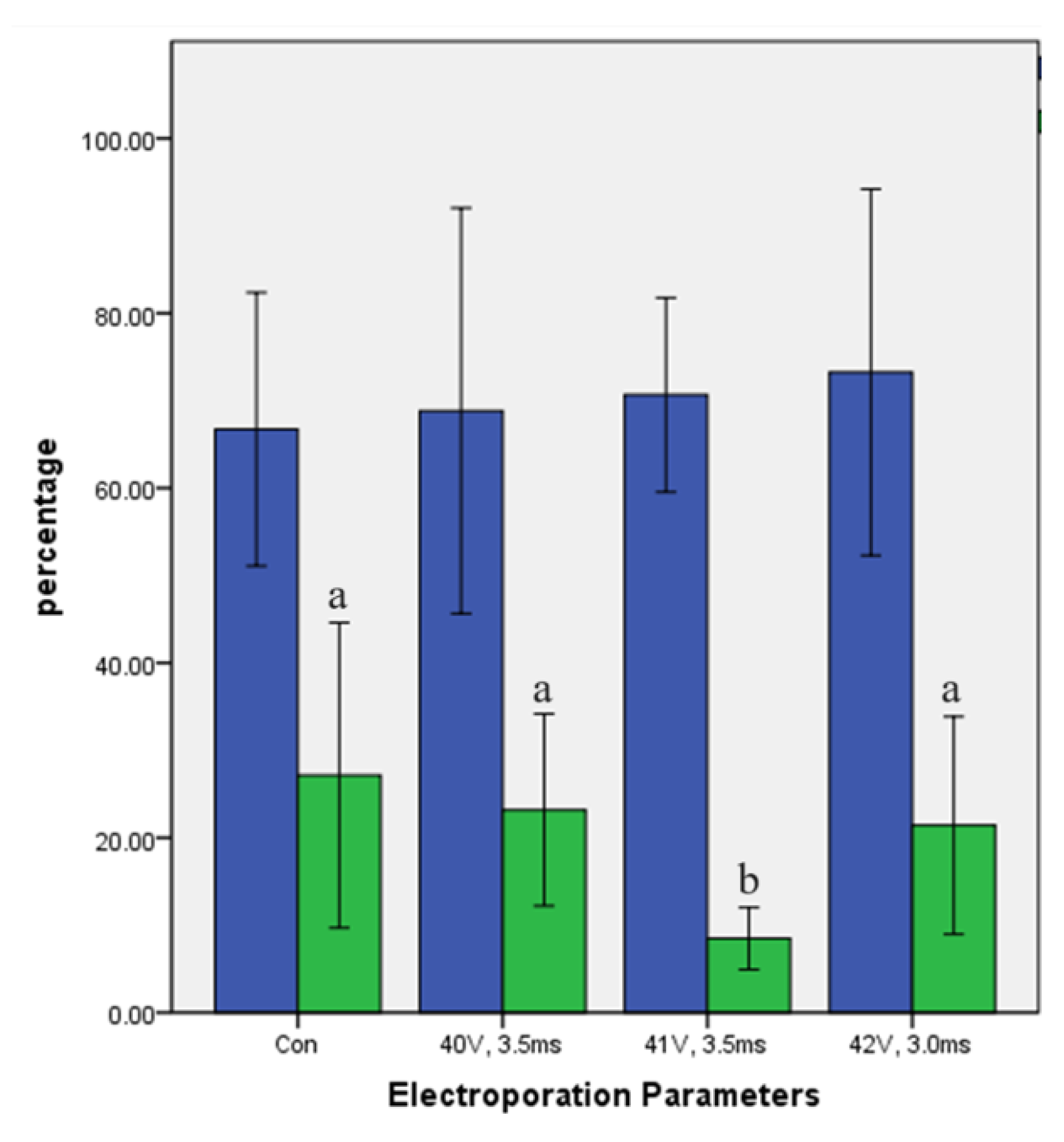
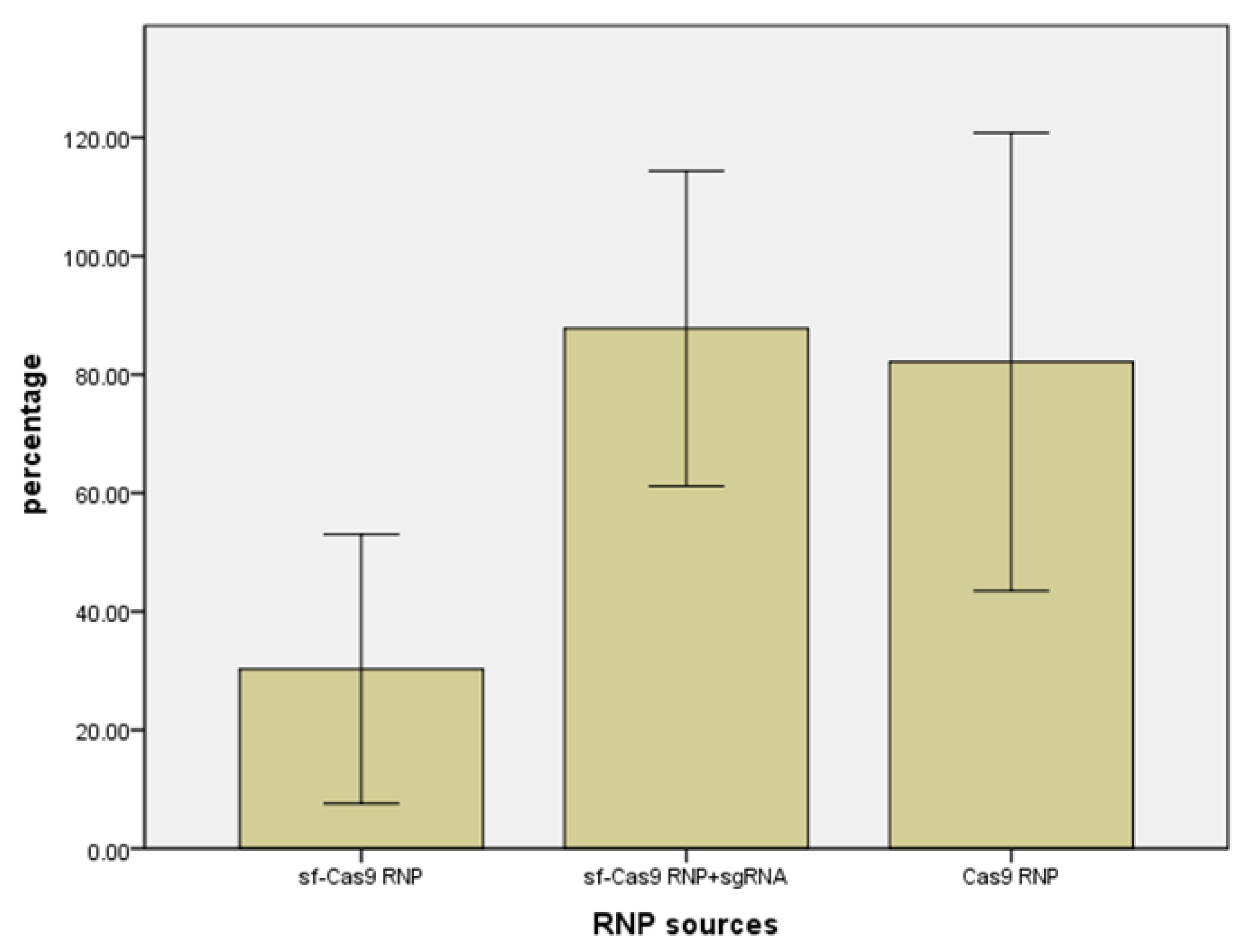
2.2. Effect of three formulations of Cas9 RNPs on the mutation of electroporation on ACTG1 gene in sheep.
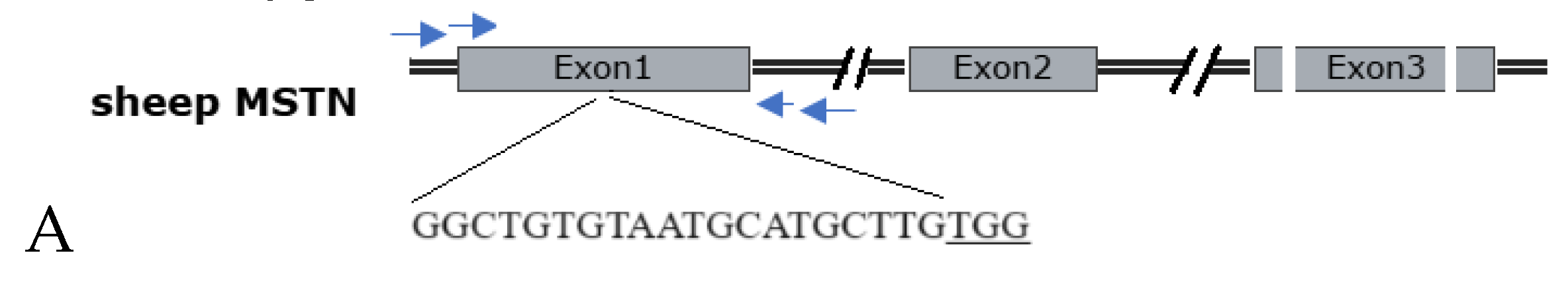
2.3. Effect of Electroporation on MSTN Mutation Rates
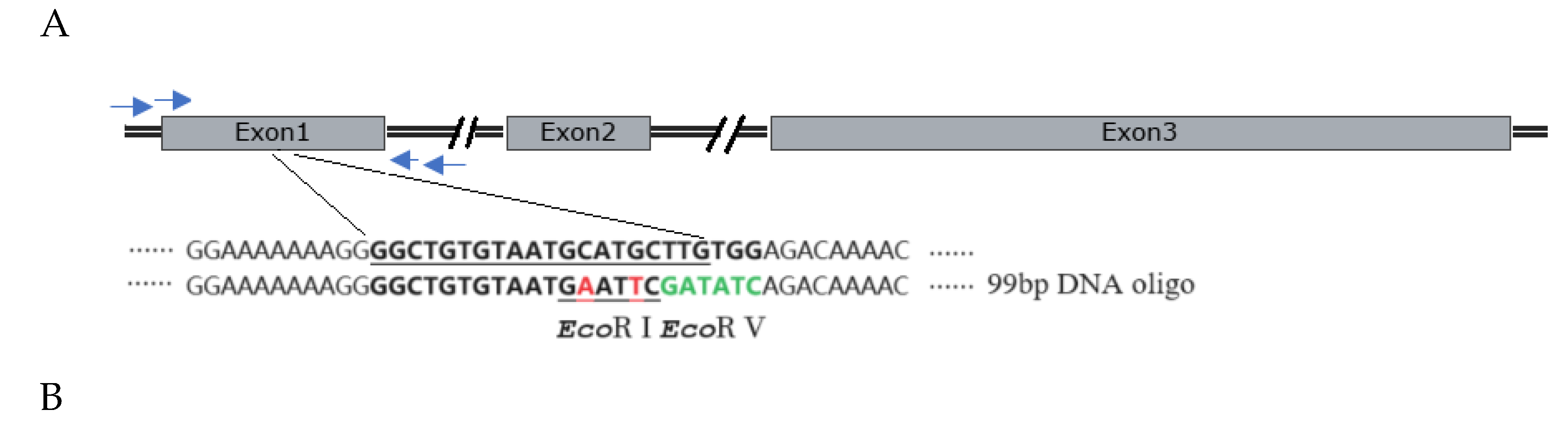
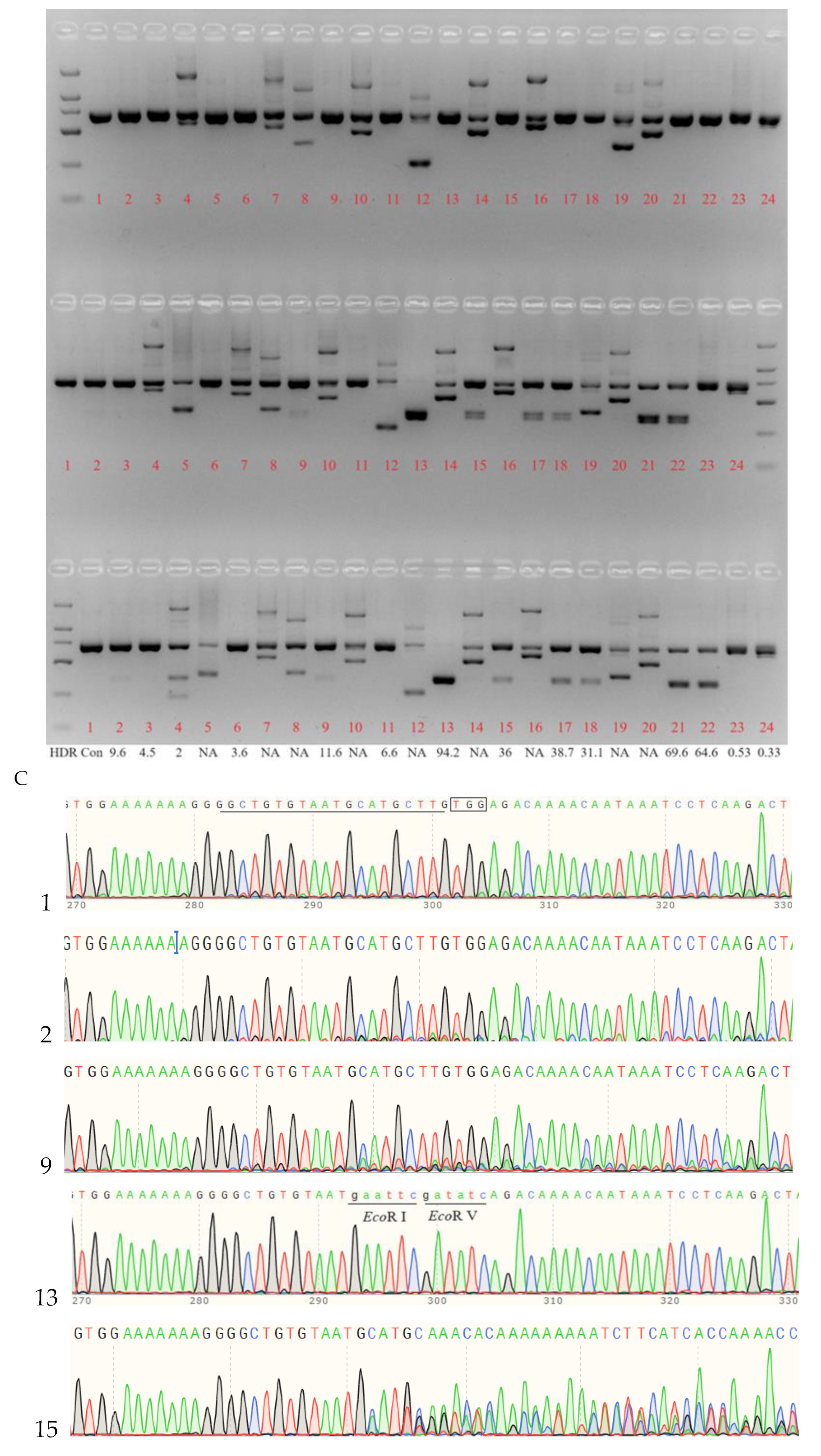
2.4. Cas9 RNP-Mediated ssODN Knock-in Efficiency in Sheep Zygotes
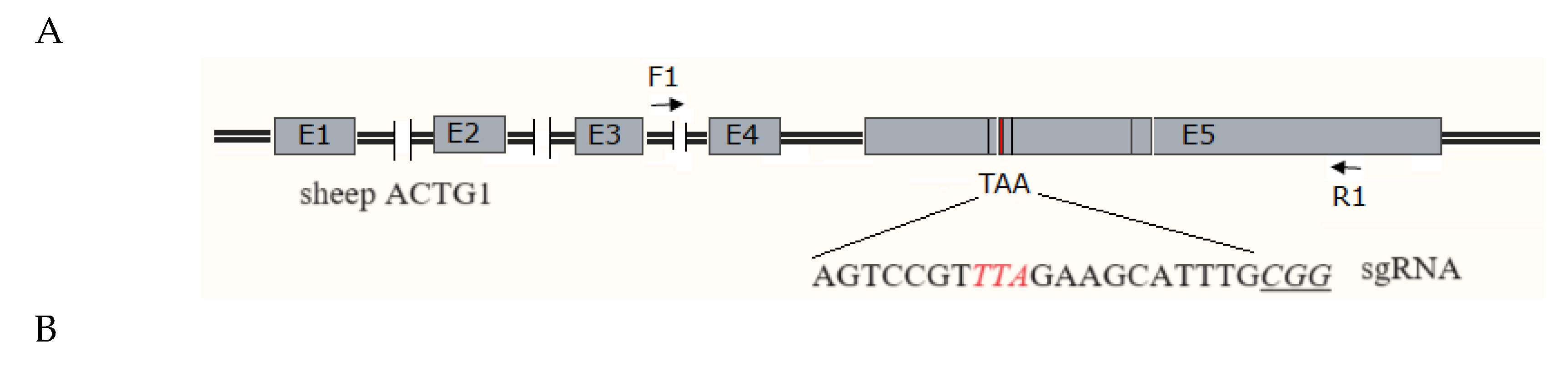
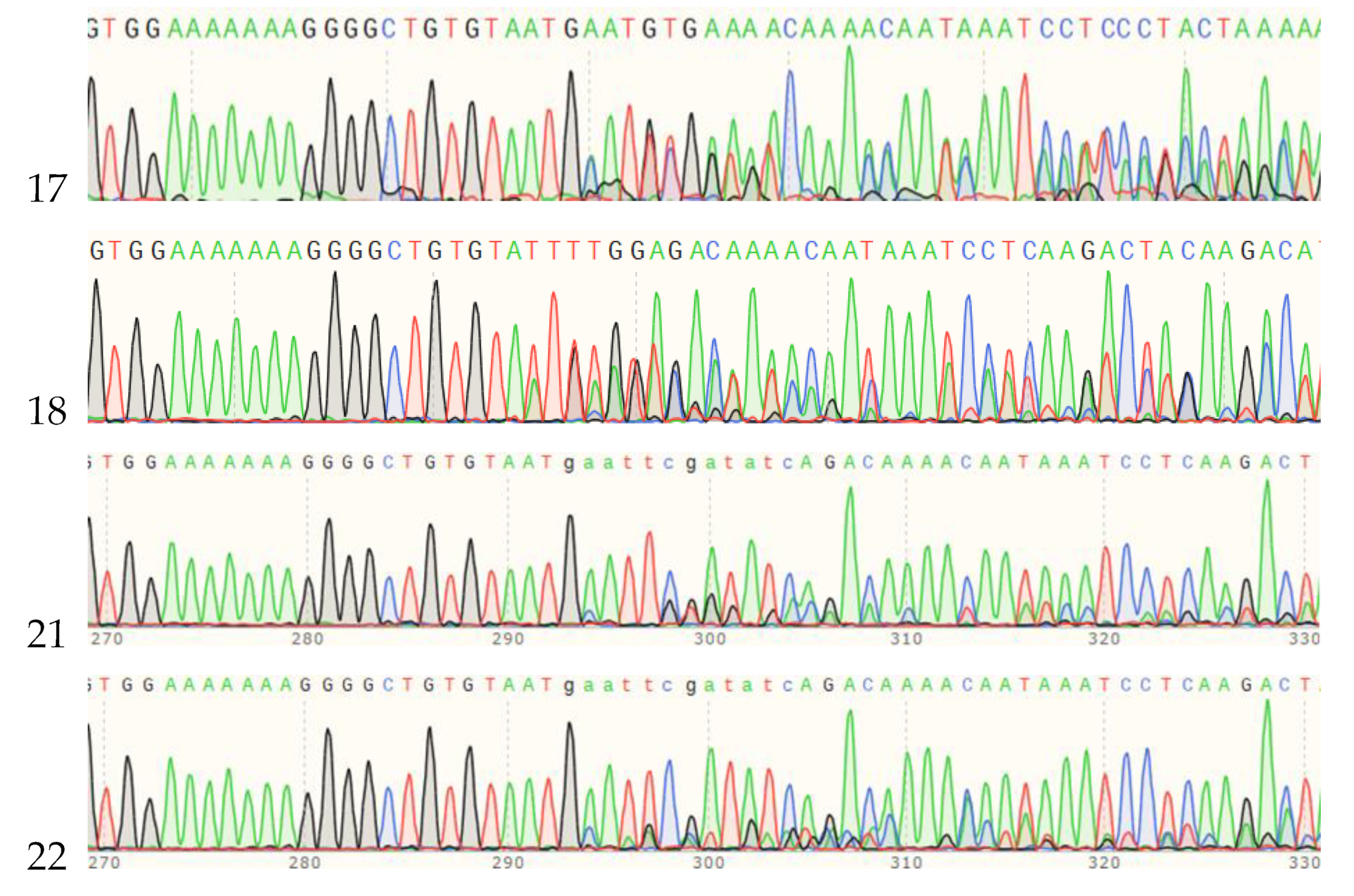
2.5. Identification of MSTN Mutation Animals
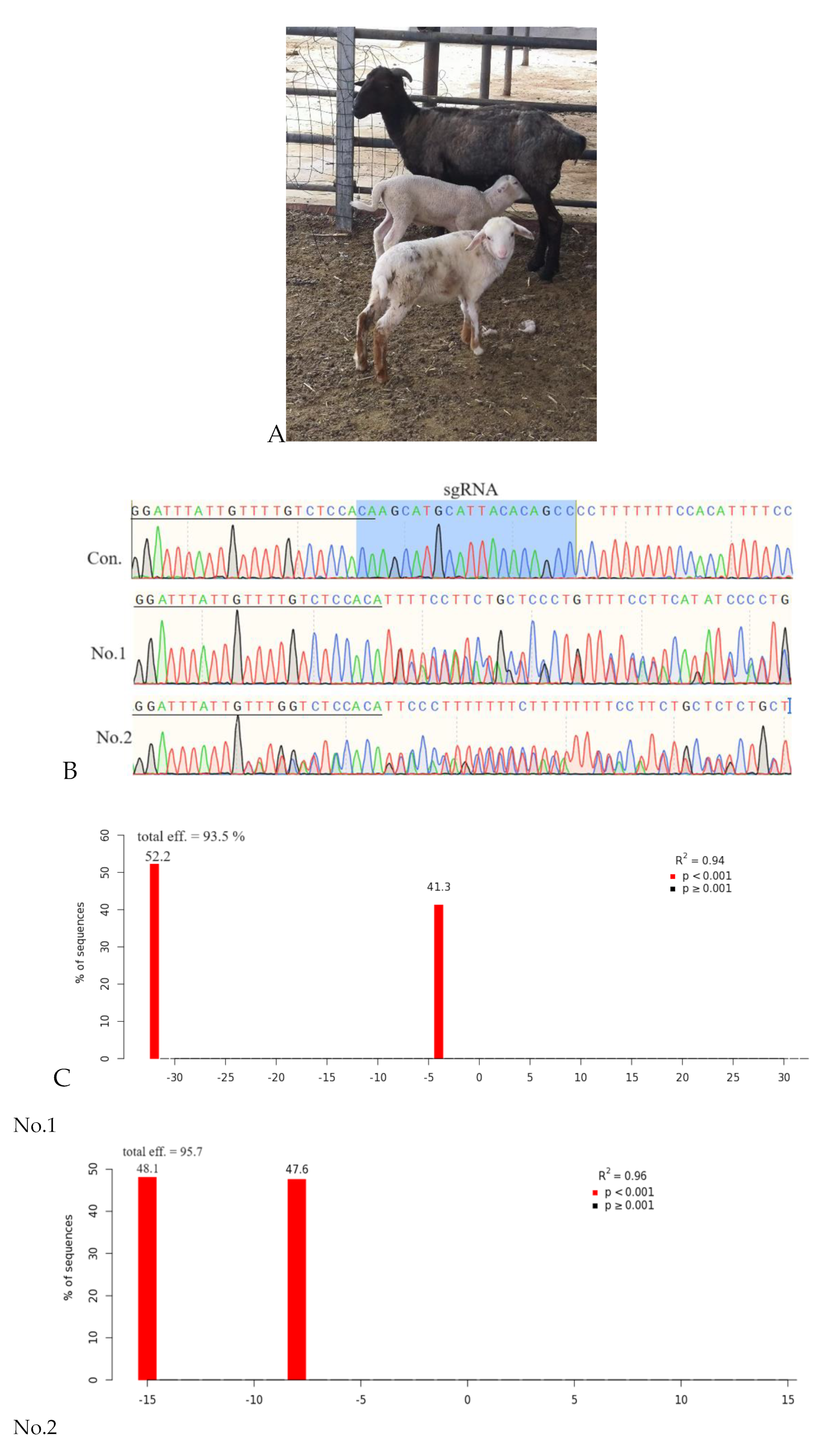
3. Discussion
4. Materials and Methods
4.1. Experimental Design
4.2. Reagents and Solutions
4.3. Oocyte Recovery and In Vitro Maturation (IVM)
4.4. In vitro fertilization (IVF)
4.5. In Vitro Culture (IVC)
4.7. Cas9 RNP Preparation
4.9. Electroporation Programs Optimization
4.10. Electroporation
4.11. Mutation Detection of Sheep Blastocysts
4.12. Embryo Transfer and Pregnancy Diagnosis
4.13. Identification of MSTN Mutation Lambs
4.14. Statistical Analysis
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Acknowledgments
Conflicts of Interest
References
- Woolley, S. A., Salavati, M., & Clark, E. L. (2023). Recent advances in the genomic resources for sheep. Mammalian genome : official journal of the International Mammalian Genome Society, 34(4), 545–558. [CrossRef]
- Kalds, P., Zhou, S., Cai, B., Liu, J., Wang, Y., Petersen, B., Sonstegard, T., Wang, X., & Chen, Y. (2019). Sheep and Goat Genome Engineering: From Random Transgenesis to the CRISPR Era. Frontiers in genetics, 10, 750. [CrossRef]
- Wang, J. Y., & Doudna, J. A. (2023). CRISPR technology: A decade of genome editing is only the beginning. Science (New York, N.Y.), 379(6629), eadd8643.
- Wang, X., Niu, Y., Zhou, J., Yu, H., Kou, Q., Lei, A., Zhao, X., Yan, H., Cai, B., Shen, Q., Zhou, S., Zhu, H., Zhou, G., Niu, W., Hua, J., Jiang, Y., Huang, X., Ma, B., & Chen, Y. (2016). Multiplex gene editing via CRISPR/Cas9 exhibits desirable muscle hypertrophy without detectable off-target effects in sheep. Scientific reports, 6, 32271. [CrossRef]
- Crispo, M., Mulet, A. P., Tesson, L., Barrera, N., Cuadro, F., dos Santos-Neto, P. C., Nguyen, T. H., Crénéguy, A., Brusselle, L., Anegón, I., & Menchaca, A. (2015). Efficient Generation of Myostatin Knock-Out Sheep Using CRISPR/Cas9 Technology and Microinjection into Zygotes. PloS one, 10(8), e0136690. [CrossRef]
- Zhang, X., Li, W., Liu, C., Peng, X., Lin, J., He, S., Li, X., Han, B., Zhang, N., Wu, Y., Chen, L., Wang, L., MaYila, Huang, J., & Liu, M. (2017). Alteration of sheep coat color pattern by disruption of ASIP gene via CRISPR Cas9. Scientific reports, 7(1), 8149. [CrossRef]
- Menchaca, A., Dos Santos-Neto, P. C., Souza-Neves, M., Cuadro, F., Mulet, A. P., Tesson, L., Chenouard, V., Guiffès, A., Heslan, J. M., Gantier, M., Anegón, I., & Crispo, M. (2020). Otoferlin gene editing in sheep via CRISPR-assisted ssODN-mediated Homology Directed Repair. Scientific reports, 10(1), 5995.
- Lin, Y., Li, J., Li, C., Tu, Z., Li, S., Li, X. J., & Yan, S. (2022). Application of CRISPR/Cas9 System in Establishing Large Animal Models. Frontiers in cell and developmental biology, 10, 919155.
- Modzelewski, A. J., Chen, S., Willis, B. J., Lloyd, K. C. K., Wood, J. A., & He, L. (2018). Efficient mouse genome engineering by CRISPR-EZ technology. Nature protocols, 13(6), 1253–1274.
- Lin, J. C., & Van Eenennaam, A. L. (2021). Electroporation-Mediated Genome Editing of Livestock Zygotes. Frontiers in genetics, 12, 648482.
- Kaneko, T., Sakuma, T., Yamamoto, T., & Mashimo, T. (2014). Simple knockout by electroporation of engineered endonucleases into intact rat embryos. Scientific reports, 4, 6382.
- Qin, W., Dion, S. L., Kutny, P. M., Zhang, Y., Cheng, A. W., Jillette, N. L., Malhotra, A., Geurts, A. M., Chen, Y. G., & Wang, H. (2015). Efficient CRISPR/Cas9-Mediated Genome Editing in Mice by Zygote Electroporation of Nuclease. Genetics, 200(2), 423–430.
- Wang, W., Kutny, P. M., Byers, S. L., Longstaff, C. J., DaCosta, M. J., Pang, C., Zhang, Y., Taft, R. A., Buaas, F. W., & Wang, H. (2016). Delivery of Cas9 Protein into Mouse Zygotes through a Series of Electroporation Dramatically Increases the Efficiency of Model Creation. Journal of genetics and genomics = Yi chuan xue bao, 43(5), 319–327.
- Ciccarelli, M., Giassetti, M. I., Miao, D., Oatley, M. J., Robbins, C., Lopez-Biladeau, B., Waqas, M. S., Tibary, A., Whitelaw, B., Lillico, S., Park, C. H., Park, K. E., Telugu, B., Fan, Z., Liu, Y., Regouski, M., Polejaeva, I. A., & Oatley, J. M. (2020). Donor-derived spermatogenesis following stem cell transplantation in sterile NANOS2 knockout males. Proceedings of the National Academy of Sciences of the United States of America, 117(39), 24195–24204.
- Tanihara, F., Hirata, M., Nguyen, N. T., Le, Q. A., Hirano, T., Takemoto, T., Nakai, M., Fuchimoto, D. I., & Otoi, T. (2018). Generation of a TP53-modified porcine cancer model by CRISPR/Cas9-mediated gene modification in porcine zygotes via electroporation. PloS one, 13(10), e0206360.
- Sundby, L. J., Southern, W. M., Hawbaker, K. M., Trujillo, J. M., Perrin, B. J., & Ervasti, J. M. (2022). Nucleotide- and Protein-Dependent Functions of Actg1. Molecular biology of the cell, 33(9), ar77.
- McPherron, A. C., Lawler, A. M., & Lee, S. J. (1997). Regulation of skeletal muscle mass in mice by a new TGF-beta superfamily member. Nature, 387(6628), 83–90.
- Zhao, J., Lai, L., Ji, W., & Zhou, Q. (2019). Genome editing in large animals: current status and future prospects. National science review, 6(3), 402–420.
- Kalds, P., Zhou, S., Cai, B., Liu, J., Wang, Y., Petersen, B., Sonstegard, T., Wang, X., & Chen, Y. (2019). Sheep and Goat Genome Engineering: From Random Transgenesis to the CRISPR Era. Frontiers in genetics, 10, 750.
- Guo, R., Wang, H., Meng, C., Gui, H., Li, Y., Chen, F., Zhang, C., Zhang, H., Ding, Q., Zhang, J., Zhang, J., Qian, Y., Zhong, J., & Cao, S. (2023). Efficient and Specific Generation of MSTN-Edited Hu Sheep Using C-CRISPR. Genes, 14(6), 1216.
- Mahdi, A. K., Medrano, J. F., & Ross, P. J. (2022). Single-Step Genome Editing of Small Ruminant Embryos by Electroporation. International journal of molecular sciences, 23(18), 10218.
- Jordan, E. T., Collins, M., Terefe, J., Ugozzoli, L., & Rubio, T. (2008). Optimizing electroporation conditions in primary and other difficult-to-transfect cells. Journal of biomolecular techniques : JBT, 19(5), 328–334.
- Nishio, K., Tanihara, F., Nguyen, T. V., Kunihara, T., Nii, M., Hirata, M., Takemoto, T., & Otoi, T. (2018). Effects of voltage strength during electroporation on the development and quality of in vitro-produced porcine embryos. Reproduction in domestic animals, 53(2), 313–318.
- Anzalone, D. A., Palazzese, L., Czernik, M., Sabatucci, A., Valbonetti, L., Capra, E., & Loi, P. (2021). Controlled spermatozoa-oocyte interaction improves embryo quality in sheep. Scientific reports, 11(1), 22629.
- Hennig, S. L., Owen, J. R., Lin, J. C., Young, A. E., Ross, P. J., Van Eenennaam, A. L., & Murray, J. D. (2020). Evaluation of mutation rates, mosaicism and off target mutations when injecting Cas9 mRNA or protein for genome editing of bovine embryos. Scientific reports, 10(1), 22309.
- Miskel, D., Poirier, M., Beunink, L., Rings, F., Held, E., Tholen, E., Tesfaye, D., Schellander, K., Salilew-Wondim, D., Blaschka, C., Große-Brinkhaus, C., Brenig, B., & Hoelker, M. (2022). The cell cycle stage of bovine zygotes electroporated with CRISPR/Cas9-RNP affects frequency of Loss-of-heterozygosity editing events. Scientific reports, 12(1), 10793.
- Vilarino, M., Rashid, S. T., Suchy, F. P., McNabb, B. R., van der Meulen, T., Fine, E. J., Ahsan, S. D., Mursaliyev, N., Sebastiano, V., Diab, S. S., Huising, M. O., Nakauchi, H., & Ross, P. J. (2017). CRISPR/Cas9 microinjection in oocytes disables pancreas development in sheep. Scientific reports, 7(1), 17472.
- Namula, Z., Wittayarat, M., Hirata, M., Hirano, T., Nguyen, N. T., Le, Q. A., Fahrudin, M., Tanihara, F., & Otoi, T. (2019). Genome mutation after the introduction of the gene editing by electroporation of Cas9 protein (GEEP) system into bovine putative zygotes. In vitro cellular & developmental biology. Animal, 55(8), 598–603.
- Qiao, J., Li, W., Lin, S., Sun, W., Ma, L., & Liu, Y. (2019). Co-expression of Cas9 and single-guided RNAs in Escherichia coli streamlines production of Cas9 ribonucleoproteins. Communications biology, 2, 161.
- Kagita, A., Lung, M. S. Y., Xu, H., Kita, Y., Sasakawa, N., Iguchi, T., Ono, M., Wang, X. H., Gee, P., & Hotta, A. (2021). Efficient ssODN-Mediated Targeting by Avoiding Cellular Inhibitory RNAs through Precomplexed CRISPR-Cas9/sgRNA Ribonucleoprotein. Stem cell reports, 16(4), 985–996.
- Hsu, P. D., Scott, D. A., Weinstein, J. A., Ran, F. A., Konermann, S., Agarwala, V., Li, Y., Fine, E. J., Wu, X., Shalem, O., Cradick, T. J., Marraffini, L. A., Bao, G., & Zhang, F. (2013). DNA targeting specificity of RNA-guided Cas9 nucleases. Nature biotechnology, 31(9), 827–832.
- Jinek, M., East, A., Cheng, A., Lin, S., Ma, E., & Doudna, J. (2013). RNA-programmed genome editing in human cells. eLife, 2, e00471.
- Mekler, V., Minakhin, L., Semenova, E., Kuznedelov, K., & Severinov, K. (2016). Kinetics of the CRISPR-Cas9 effector complex assembly and the role of 3′-terminal segment of guide RNA. Nucleic acids research, 44(6), 2837–2845.
- Tanihara, F., Hirata, M., Namula, Z., Do, L. T. K., Yoshimura, N., Lin, Q., Takebayashi, K., Sakuma, T., Yamamoto, T., & Otoi, T. (2023). Pigs with an INS point mutation derived from zygotes electroporated with CRISPR/Cas9 and ssODN. Frontiers in cell and developmental biology, 11, 884340.
- Remy, S., Chenouard, V., Tesson, L., Usal, C., Ménoret, S., Brusselle, L., Heslan, J. M., Nguyen, T. H., Bellien, J., Merot, J., De Cian, A., Giovannangeli, C., Concordet, J. P., & Anegon, I. (2017). Generation of gene-edited rats by delivery of CRISPR/Cas9 protein and donor DNA into intact zygotes using electroporation. Scientific reports, 7(1), 16554.
- Speckhart, S. L., Wooldridge, L. K., & Ealy, A. D. (2023). An updated protocol for in vitro bovine embryo production. STAR protocols, 4(1), 101924.
- Guo Yanhua, Pi Wenhui. (2021). Use of the CRISPR-Cas9 system in sheep fibroblasts to lead fluorescent tags into ACTG1 gene. Xinjiang Nongye Kexue (China), 58(9), 1747-1755.
| Scheme 1. | Electroporation Parameters | ||||||
|---|---|---|---|---|---|---|---|
| Pulse Type | Voltages (V) | Pulse Length (ms) | Pulse Interval (ms) | Pulses Number | Decay Rate (%) | Polarity | |
| Con. | —— | —— | —— | —— | —— | —— | —— |
| 1 | Poring | 40 | 3.5 | 50 | 4 | 10 | + |
| Transfer | 5 | 50 | 50 | 5 | 40 | +/- | |
| 2 | Poring | 41 | 3.5 | 50 | 4 | 10 | + |
| Transfer | 5 | 50 | 50 | 5 | 40 | +/- | |
| 3 | Poring | 42 | 3.0 | 50 | 4 | 10 | + |
| Transfer | 5 | 50 | 50 | 5 | 40 | +/- | |
| Primer. | Scheme 5. |
| shACTG1F1 | AGCATGACTGACCTCCCTTTG |
| shACTG1R1 | CCCAACCCCATGTAAGACCG |
| shACTG1F2 | CACCATGTACCCTGGCAT |
| shACTG1R2 | ACATTCTCACCTCAGCTAC |
| shMSTNF1 | GTGACTTGTGACAGACAGGGTT |
| shMSTNR1 | AATGTAGCAGCTTTCAGTCTCAT |
| shMSTNF2 | AATCACAGATCCCGACGACAC |
| shMSTNR2 | TCCTTACGTACAAGCCAGCAG |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




