Submitted:
17 July 2024
Posted:
18 July 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
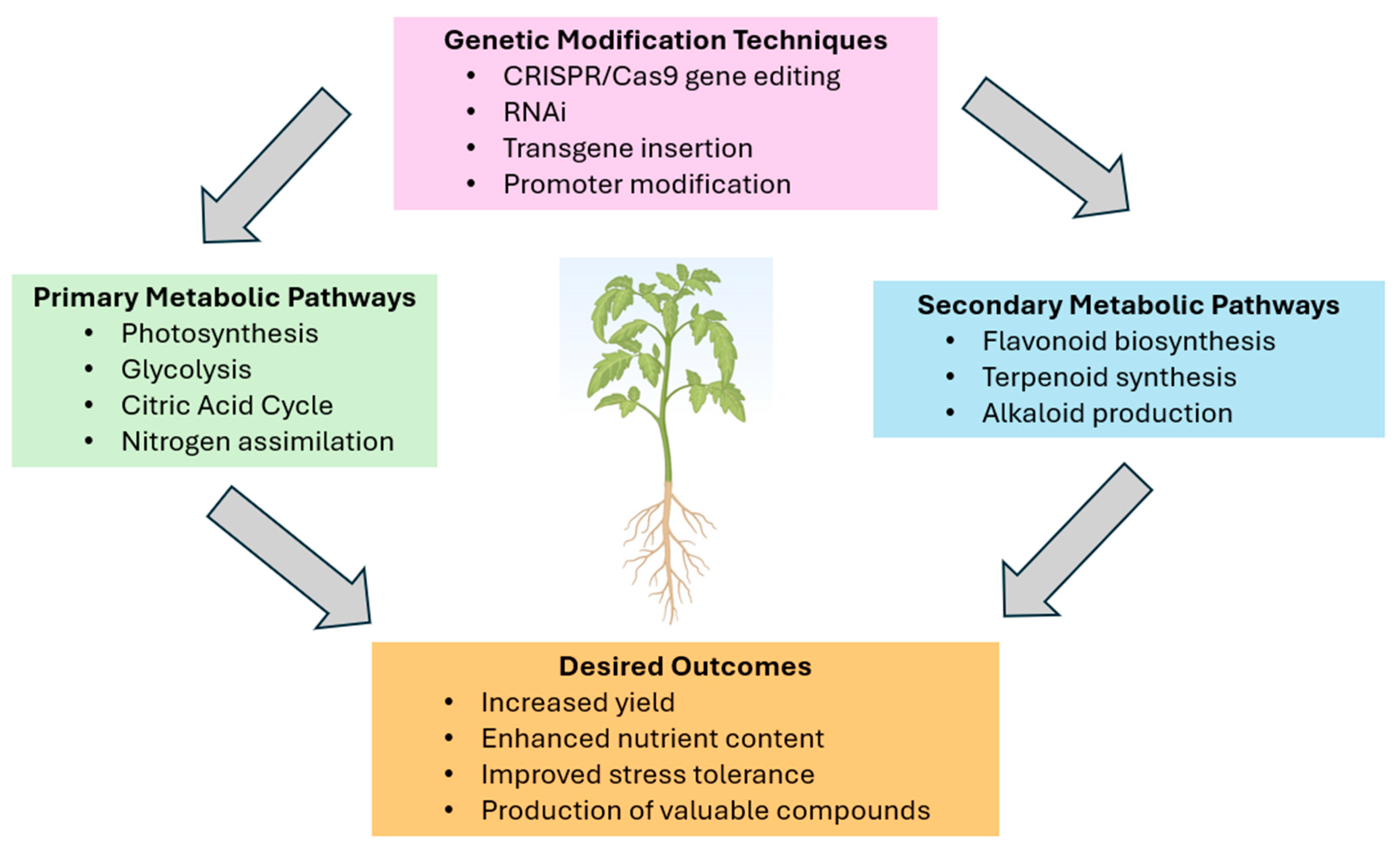
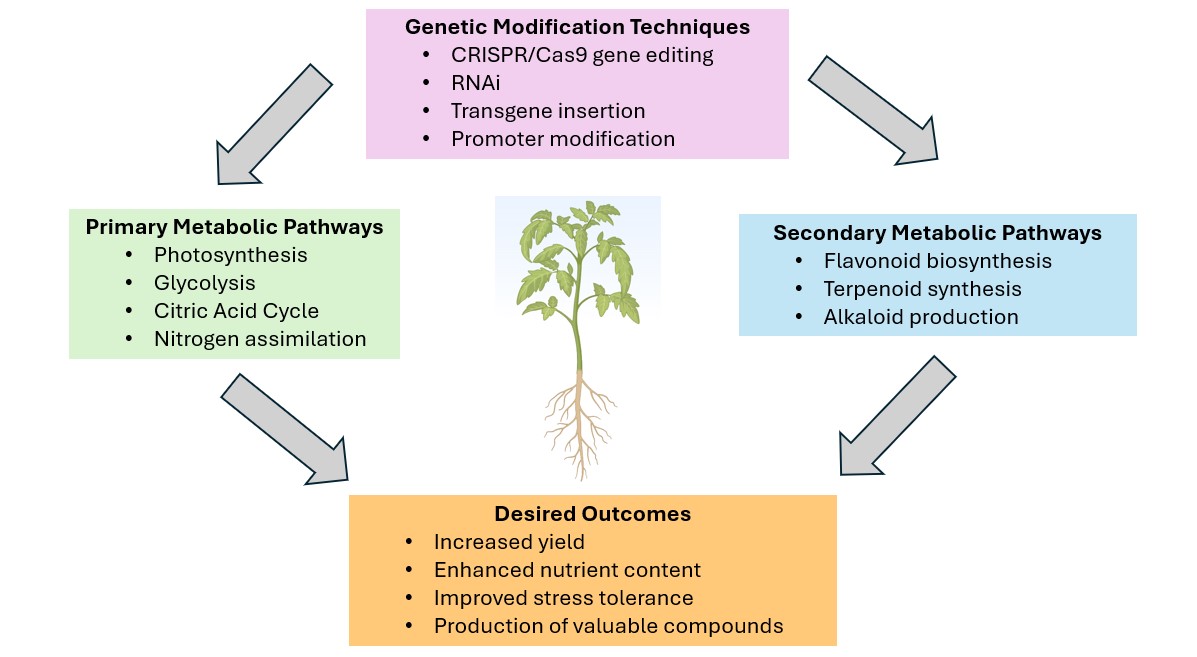
2. Principles of Metabolic Engineering in Plants
2.1. Overview of Plant Metabolic Pathways
2.2. Strategies for Manipulating Metabolic Pathways
2.3. Tools and Techniques for Genetic Modification
3. Applications in Crop Improvement
3.1. Enhancing Photosynthetic Efficiency
3.2. Improving Nutrient Use Efficiency
3.3. Enhancing Stress Tolerance
3.4. Production of Valuable Metabolites
4. Emerging Trends and Future Perspectives
4.1. Synthetic Biology Approaches
4.2. Multi-Gene Trait Stacking
4.3. Integration of Omics and Computational Modeling
4.4. Regulatory and Biosafety Considerations
5. Conclusions
References
- Lee, S. Y., Nielsen, J. & Stephanopoulos, G. Metabolic Engineering: Concepts and Applications. (2021).
- van Dijk, M.; Morley, T.; Rau, M.L.; Saghai, Y. A meta-analysis of projected global food demand and population at risk of hunger for the period 2010–2050. Nat. Food 2021, 2, 494–501, . [CrossRef]
- Antoniewicz, M.R. A guide to metabolic flux analysis in metabolic engineering: Methods, tools and applications. Metab. Eng. 2020, 63, 2–12, . [CrossRef]
- Ran, F.A.; Hsu, P.D.; Wright, J.; Agarwala, V.; Scott, D.A.; Zhang, F. Genome engineering using the CRISPR-Cas9 system. Nat. Protoc. 2013, 8, 2281–2308, doi:10.1038/nprot.2013.143.
- Sweetlove, L.J.; Nielsen, J.; Fernie, A.R. Engineering central metabolism – a grand challenge for plant biologists. Plant J. 2016, 90, 749–763, . [CrossRef]
- Hefferon, K.L. Nutritionally Enhanced Food Crops; Progress and Perspectives. Int. J. Mol. Sci. 2015, 16, 3895–3914, . [CrossRef]
- Petrie, J.R.; Shrestha, P.; Belide, S.; Kennedy, Y.; Lester, G.; Liu, Q.; Divi, U.K.; Mulder, R.J.; Mansour, M.P.; Nichols, P.D.; et al. Metabolic Engineering Camelina sativa with Fish Oil-Like Levels of DHA. PLOS ONE 2014, 9, e85061, . [CrossRef]
- Boghigian, B.A.; Seth, G.; Kiss, R.; Pfeifer, B.A. Metabolic flux analysis and pharmaceutical production. Metab. Eng. 2010, 12, 81–95, . [CrossRef]
- Henry, C.S.; Broadbelt, L.J.; Hatzimanikatis, V. Thermodynamics-Based Metabolic Flux Analysis. Biophys. J. 2007, 92, 1792–1805, . [CrossRef]
- Dai, Z.; Locasale, J.W. Understanding metabolism with flux analysis: From theory to application. Metab. Eng. 2016, 43, 94–102, . [CrossRef]
- Xu, Y. Metabolomics study on Arabidopsis thaliana abiotic stress responses for priming, recovery, and stress combinations. (2018).
- Holms, H. Flux analysis and control of the central metabolic pathways in Escherichia coli. FEMS Microbiol. Rev. 19, 85–116 (1996).
- Xu, Y.; Fu, X.; Sharkey, T.D.; Shachar-Hill, Y.; Walker, B.J. The metabolic origins of non-photorespiratory CO2 release during photosynthesis: a metabolic flux analysis. Plant Physiol. 2021, 186, 297–314, . [CrossRef]
- Lee, D.-Y.; Yun, H.; Park, S.; Lee, S.Y. MetaFluxNet: the management of metabolic reaction information and quantitative metabolic flux analysis. Bioinformatics 2003, 19, 2144–2146, . [CrossRef]
- Fan, K.-T.; Xu, Y.; Hegeman, A.D. Elevated Temperature Effects on Protein Turnover Dynamics in Arabidopsis thaliana Seedlings Revealed by 15N-Stable Isotope Labeling and ProteinTurnover Algorithm. Int. J. Mol. Sci. 2024, 25, 5882, . [CrossRef]
- Long, C.P.; Antoniewicz, M.R. High-resolution 13C metabolic flux analysis. Nat. Protoc. 2019, 14, 2856–2877, . [CrossRef]
- Quek, L.-E.; Wittmann, C.; Nielsen, L.K.; O Krömer, J. OpenFLUX: efficient modelling software for 13C-based metabolic flux analysis. Microb. Cell Factories 2009, 8, 25–25, . [CrossRef]
- Xu, Y.; Wieloch, T.; Kaste, J.A.M.; Shachar-Hill, Y.; Sharkey, T.D. Reimport of carbon from cytosolic and vacuolar sugar pools into the Calvin–Benson cycle explains photosynthesis labeling anomalies. Proc. Natl. Acad. Sci. 2022, 119, . [CrossRef]
- Quek, L.-E.; Dietmair, S.; Krömer, J.O.; Nielsen, L.K. Metabolic flux analysis in mammalian cell culture. Metab. Eng. 2010, 12, 161–171, . [CrossRef]
- Fan KaiTing, F. K. et al. Application of data-independent acquisition approach to study the proteome change from early to later phases of tomato pathogenesis responses. (2019).
- Wittmann, C. & Heinzle, E. Mass spectrometry for metabolic flux analysis. Biotechnol. Bioeng. 62, 739–750 (1999).
- Bar-Even, A.; Noor, E.; Savir, Y.; Liebermeister, W.; Davidi, D.; Tawfik, D.S.; Milo, R. The Moderately Efficient Enzyme: Evolutionary and Physicochemical Trends Shaping Enzyme Parameters. Biochemistry 2011, 50, 4402–4410, . [CrossRef]
- E Dueber, J.; Wu, G.C.; Malmirchegini, G.R.; Moon, T.S.; Petzold, C.J.; Ullal, A.V.; Prather, K.L.J.; Keasling, J.D. Synthetic protein scaffolds provide modular control over metabolic flux. Nat. Biotechnol. 2009, 27, 753–759, . [CrossRef]
- López-Calcagno, P. E. et al. Stimulating photosynthetic processes increases productivity and water-use efficiency in the field. Nat. Plants 6, 1054–1063 (2020).
- Gelvin, S. B. Agrobacterium-mediated plant transformation: the biology behind the “gene-jockeying” tool. Microbiol. Mol. Biol. Rev. 67, 16–37 (2003).
- Christou, P. Particle gun mediated transformation. Curr. Opin. Biotechnol. 1993, 4, 135–141, . [CrossRef]
- Rojo, F.P.; Nyman, R.K.M.; Johnson, A.A.T.; Navarro, M.P.; Ryan, M.H.; Erskine, W.; Kaur, P. CRISPR-Cas systems: ushering in the new genome editing era. Bioengineered 2018, 9, 214–221, . [CrossRef]
- Anzalone, A.V.; Randolph, P.B.; Davis, J.R.; Sousa, A.A.; Koblan, L.W.; Levy, J.M.; Chen, P.J.; Wilson, C.; Newby, G.A.; Raguram, A.; et al. Search-and-replace genome editing without double-strand breaks or donor DNA. Nature 2019, 576, 149–157, . [CrossRef]
- Kaplan, F.; Guy, C.L. RNA interference of Arabidopsis beta-amylase8 prevents maltose accumulation upon cold shock and increases sensitivity of PSII photochemical efficiency to freezing stress. Plant J. 2005, 44, 730–743, . [CrossRef]
- Liu, Y., Schiff, M. & Dinesh-Kumar, S. P. Virus-induced gene silencing in tomato. Plant J. 31, 777–786 (2002).
- Khakhar, A., Leydon, A. R., Lemmex, A. C., Klavins, E. & Nemhauser, J. L. Synthetic hormone-responsive transcription factors can monitor and re-program plant development. Elife 7, e34702 (2018).
- Weber, W.; Fussenegger, M. Emerging biomedical applications of synthetic biology. Nat. Rev. Genet. 2011, 13, 21–35, . [CrossRef]
- Urano, K., Kurihara, Y., Seki, M. & Shinozaki, K. ‘Omics’ analyses of regulatory networks in plant abiotic stress responses. Curr. Opin. Plant Biol. 13, 132–138 (2010).
- Parry, M.A.J.; Andralojc, P.J.; Scales, J.C.; Salvucci, M.E.; Carmo-Silva, A.E.; Alonso, H.; Whitney, S.M. Rubisco activity and regulation as targets for crop improvement. J. Exp. Bot. 2012, 64, 717–730, . [CrossRef]
- Adebiyi, A.O.; Jazmin, L.J.; Young, J.D. 13C flux analysis of cyanobacterial metabolism. Photosynth. Res. 2014, 126, 19–32, . [CrossRef]
- Long, S.P.; Marshall-Colon, A.; Zhu, X.-G. Meeting the Global Food Demand of the Future by Engineering Crop Photosynthesis and Yield Potential. Cell 2015, 161, 56–66, . [CrossRef]
- South, P.F.; Cavanagh, A.P.; Liu, H.W.; Ort, D.R. Synthetic glycolate metabolism pathways stimulate crop growth and productivity in the field. Science 2019, 363, 45–+, . [CrossRef]
- Forde, B.G.; Lea, P.J. Glutamate in plants: metabolism, regulation, and signalling. J. Exp. Bot. 2007, 58, 2339–2358, . [CrossRef]
- Veneklaas, E.J.; Lambers, H.; Bragg, J.; Finnegan, P.M.; Lovelock, C.E.; Plaxton, W.C.; Price, C.A.; Scheible, W.; Shane, M.W.; White, P.J.; et al. Opportunities for improving phosphorus-use efficiency in crop plants. New Phytol. 2012, 195, 306–320, . [CrossRef]
- Garcia-Oliveira, A.L.; Tan, L.; Fu, Y.; Sun, C. Genetic Identification of Quantitative Trait Loci for Contents of Mineral Nutrients in Rice Grain. J. Integr. Plant Biol. 2008, 51, 84–92, . [CrossRef]
- Igarashi, D.; Bethke, G.; Xu, Y.; Tsuda, K.; Glazebrook, J.; Katagiri, F. Pattern-Triggered Immunity Suppresses Programmed Cell Death Triggered by Fumonisin B1. PLOS ONE 2013, 8, e60769, . [CrossRef]
- Li, T.; Pang, N.; He, L.; Xu, Y.; Fu, X.; Tang, Y.; Shachar-Hill, Y.; Chen, S. Re-Programing Glucose Catabolism in the Microalga Chlorella sorokiniana under Light Condition. Biomolecules 2022, 12, 939, . [CrossRef]
- Chen, Y., Fan, K., Hung, S. & Chen, Y. The role of peptides cleaved from protein precursors in eliciting plant stress reactions. New Phytol. 225, 2267–2282 (2020).
- Umezawa, T.; Fujita, M.; Fujita, Y.; Yamaguchi-Shinozaki, K.; Shinozaki, K. Engineering drought tolerance in plants: discovering and tailoring genes unlock the future. Curr. Opin. Biotechnol. 2006, 17, 113–122, doi:10.1016/j.copbio.2006.02.002.
- Asada, K. Production and Scavenging of Reactive Oxygen Species in Chloroplasts and Their Functions. Plant Physiol. 2006, 141, 391–396, . [CrossRef]
- Wani, S.H.; Kumar, V.; Shriram, V.; Sah, S.K. Phytohormones and their metabolic engineering for abiotic stress tolerance in crop plants. Crop. J. 2016, 4, 162–176, . [CrossRef]
- Xu, Y.; Fu, X. Reprogramming of Plant Central Metabolism in Response to Abiotic Stresses: A Metabolomics View. Int. J. Mol. Sci. 2022, 23, 5716, . [CrossRef]
- Malik, M.R.; Tang, J.; Sharma, N.; Burkitt, C.; Ji, Y.; Mykytyshyn, M.; Bohmert-Tatarev, K.; Peoples, O.; Snell, K.D. Camelina sativa, an oilseed at the nexus between model system and commercial crop. Plant Cell Rep. 2018, 37, 1367–1381, . [CrossRef]
- Poirier, Y.; Nawrath, C.; Somerville, C. Production of Polyhydroxyalkanoates, a Family of Biodegradable Plastics and Elastomers, in Bacteria and Plants. Nat. Biotechnol. 1995, 13, 142–150, . [CrossRef]
- Xu, Y. Metabolic Flux Redistribution in Engineered Microorganisms for Biofuel Production. Innov. Life Sci. J. 7, 1–12 (2021).
- Kromdijk, J.; Long, S.P. One crop breeding cycle from starvation? How engineering crop photosynthesis for rising CO2and temperature could be one important route to alleviation. Proc. R. Soc. B: Biol. Sci. 2016, 283, 20152578, . [CrossRef]
- Bar-Even, A.; Noor, E.; Lewis, N.E.; Milo, R. Design and analysis of synthetic carbon fixation pathways. Proc. Natl. Acad. Sci. 2010, 107, 8889–8894, . [CrossRef]
- Medford, J.I.; Prasad, A. Plant synthetic biology takes root. Science 2014, 346, 162–163, . [CrossRef]
- Xu, Y.; A Koroma, A.; E Weise, S.; Fu, X.; Sharkey, T.D.; Shachar-Hill, Y. Daylength variation affects growth, photosynthesis, leaf metabolism, partitioning, and metabolic fluxes. Plant Physiol. 2023, 194, 475–490, . [CrossRef]
- Wang, H.H.; Isaacs, F.J.; Carr, P.A.; Sun, Z.Z.; Xu, G.; Forest, C.R.; Church, G.M. Programming cells by multiplex genome engineering and accelerated evolution. Nature 2009, 460, 894–898, . [CrossRef]
- Fan, K.-T.; Rendahl, A.K.; Chen, W.-P.; Freund, D.M.; Gray, W.M.; Cohen, J.D.; Hegeman, A.D. Proteome Scale-Protein Turnover Analysis Using High Resolution Mass Spectrometric Data from Stable-Isotope Labeled Plants. J. Proteome Res. 2016, 15, 851–867, . [CrossRef]
- Simkin, A.J.; McAusland, L.; Lawson, T.; Raines, C.A. Overexpression of the RieskeFeS Protein Increases Electron Transport Rates and Biomass Yield. Plant Physiol. 2017, 175, 134–145, . [CrossRef]
- Naqvi, S.; Zhu, C.; Farre, G.; Ramessar, K.; Bassie, L.; Breitenbach, J.; Conesa, D.P.; Ros, G.; Sandmann, G.; Capell, T.; et al. Transgenic multivitamin corn through biofortification of endosperm with three vitamins representing three distinct metabolic pathways. Proc. Natl. Acad. Sci. 2009, 106, 7762–7767, . [CrossRef]
- Xu, Z.; Li, J.; Guo, X.; Jin, S.; Zhang, X. Metabolic engineering of cottonseed oil biosynthesis pathway via RNA interference. Sci. Rep. 2016, 6, 33342, . [CrossRef]
- Mittler, R.; Blumwald, E. Genetic Engineering for Modern Agriculture: Challenges and Perspectives. Annu. Rev. Plant Biol. 2010, 61, 443–462, . [CrossRef]
- Fu, X.; Xu, Y. Dynamic Metabolic Changes in Arabidopsis Seedlings under Hypoxia Stress and Subsequent Reoxygenation Recovery. Stresses 2023, 3, 86–101, . [CrossRef]
- Sweetlove, L.J.; Fernie, A.R. The role of dynamic enzyme assemblies and substrate channelling in metabolic regulation. Nat. Commun. 2018, 9, 1–12, . [CrossRef]
- Iba, K. et al. A gene encoding a chloroplast ω-3 fatty acid desaturase complements alterations in fatty acid desaturation and chloroplast copy number of the fad7 mutant of Arabidopsis thaliana. J. Biol. Chem. 268, 24099–24105 (1993).
- Cheung, C.M.; Poolman, M.G.; Fell, D.A.; Ratcliffe, R.G.; Sweetlove, L.J. A Diel Flux Balance Model Captures Interactions between Light and Dark Metabolism during Day-Night Cycles in C3 and Crassulacean Acid Metabolism Leaves. Plant Physiol. 2014, 165, 917–929, . [CrossRef]
- Zamboni, N.; Fendt, S.-M.; Rühl, M.; Sauer, U. 13C-based metabolic flux analysis. Nat. Protoc. 2009, 4, 878–892, . [CrossRef]
- Fan, K.-T.; Hsu, Y.; Yeh, C.-F.; Chang, C.-H.; Chang, W.-H.; Chen, Y.-R. Quantitative Proteomics Reveals the Dynamic Regulation of the Tomato Proteome in Response to Phytophthora infestans. Int. J. Mol. Sci. 2021, 22, 4174, . [CrossRef]
- Fan, K.; Hsu, C.; Chen, Y. Mass spectrometry in the discovery of peptides involved in intercellular communication: From targeted to untargeted peptidomics approaches. Mass Spectrom. Rev. 2023, 42, 2404–2425, . [CrossRef]
- Schwartz, J.-M.; Gaugain, C.; Nacher, J.C.; de Daruvar, A.; Kanehisa, M. Observing metabolic functions at the genome scale. Genome Biol. 2007, 8, R123–R123, . [CrossRef]
- Saito, K.; Yonekura-Sakakibara, K.; Nakabayashi, R.; Higashi, Y.; Yamazaki, M.; Tohge, T.; Fernie, A.R. The flavonoid biosynthetic pathway in Arabidopsis: Structural and genetic diversity. Plant Physiol. Biochem. 2013, 72, 21–34, . [CrossRef]
- Xu, Y.; Freund, D.M.; Hegeman, A.D.; Cohen, J.D. Metabolic signatures of Arabidopsis thaliana abiotic stress responses elucidate patterns in stress priming, acclimation, and recovery. Stress Biol. 2022, 2, 1–16, . [CrossRef]
- Stanford, N.J.; Lubitz, T.; Smallbone, K.; Klipp, E.; Mendes, P.; Liebermeister, W. Systematic Construction of Kinetic Models from Genome-Scale Metabolic Networks. PLOS ONE 2013, 8, e79195, . [CrossRef]
- Lasanthi-Kudahettige, R.; Magneschi, L.; Loreti, E.; Gonzali, S.; Licausi, F.; Novi, G.; Beretta, O.; Vitulli, F.; Alpi, A.; Perata, P. Transcript Profiling of the Anoxic Rice Coleoptile. Plant Physiol. 2007, 144, 218–231, . [CrossRef]
- Harwood, W.; Ricroch, A.; Clairand, P. Use of CRISPR systems in plant genome editing: toward new opportunities in agriculture. Emerg. Top. Life Sci. 2017, 1, 169–182, . [CrossRef]
- Li, Y.; Hallerman, E.M.; Liu, Q.; Wu, K.; Peng, Y. The development and status of Bt rice in China. Plant Biotechnol. J. 2015, 14, 839–848, . [CrossRef]
- Zilberman, D., Kaplan, S., Kim, E. & Waterfield, G. Lessons from the California GM labelling proposition on the state of crop biotechnology. in Handbook on Agriculture, Biotechnology and Development 538–549 (Edward Elgar Publishing, 2014).
- Fraiture, M.-A.; Herman, P.; Taverniers, I.; De Loose, M.; Deforce, D.; Roosens, N.H. Current and New Approaches in GMO Detection: Challenges and Solutions. BioMed Res. Int. 2015, 2015, 1–22, . [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



