Submitted:
19 June 2024
Posted:
20 June 2024
You are already at the latest version
Abstract
Keywords:
Introduction
Methods
Retrieval of Fold Change Reporting in Biomedical Publications
Common Basic Variants of the Fold-Change Calculation
Definition of Group Average from the Untransformed Data
Definition of Group Average from Transformed Data
Pairwise Test/Reference Ratio Calculation
Comparative Evaluation of Common Calculation Methods
Evaluation of the Role of the Data Distribution for the Correct Calculation of FC
Evaluation of the Role of Variance Equality for the Correct Calculation of FC
Evaluation of the Relationship of FC Calculation to Statistical Outcomes
Evaluation of FC Calculation Method Dependency in Biomedical Data
Results
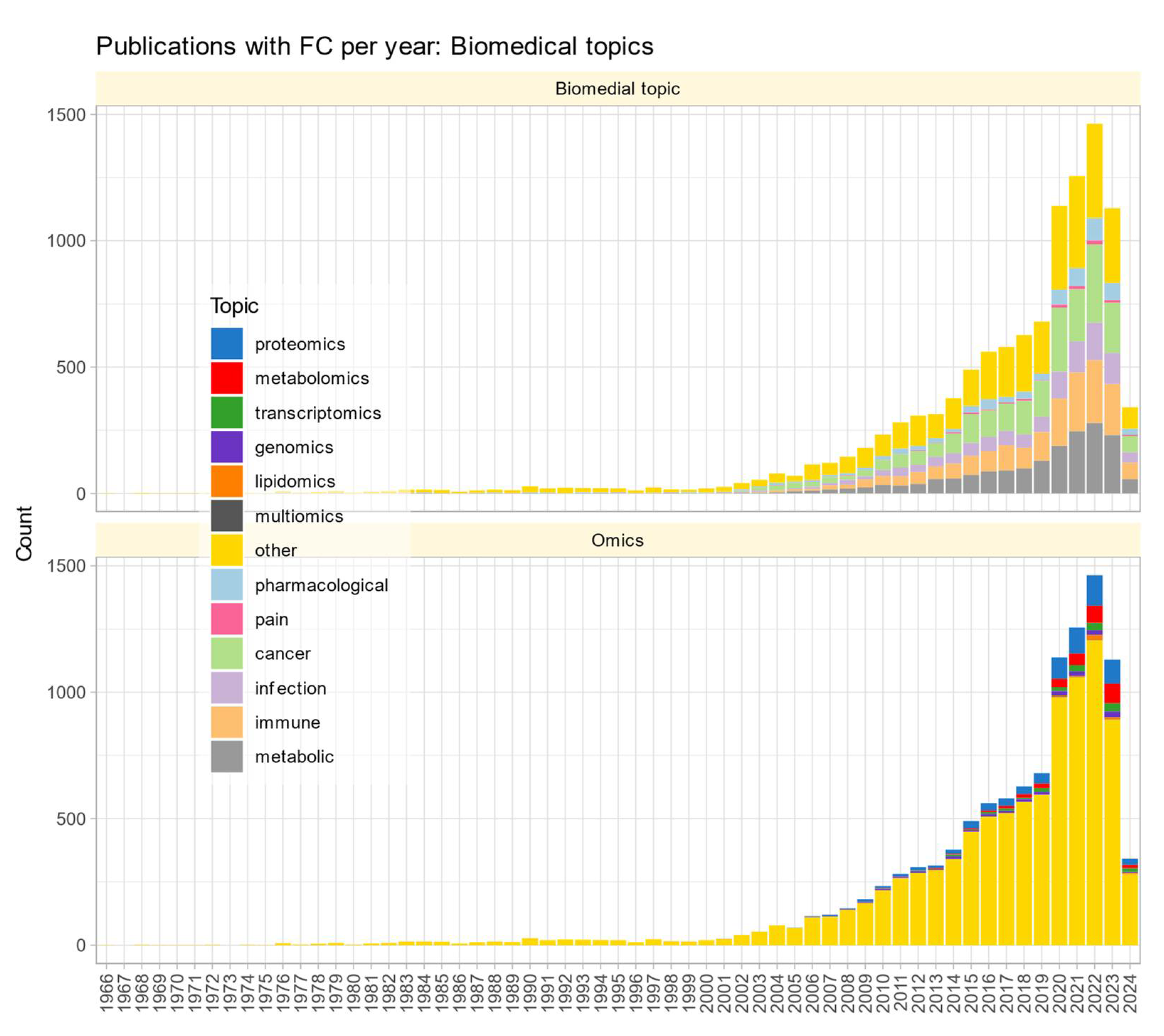
Reporting Styles of Fold-Change Calculation in Biomedical Publications
Role of the Data Distribution for the Correct Calculation of FC
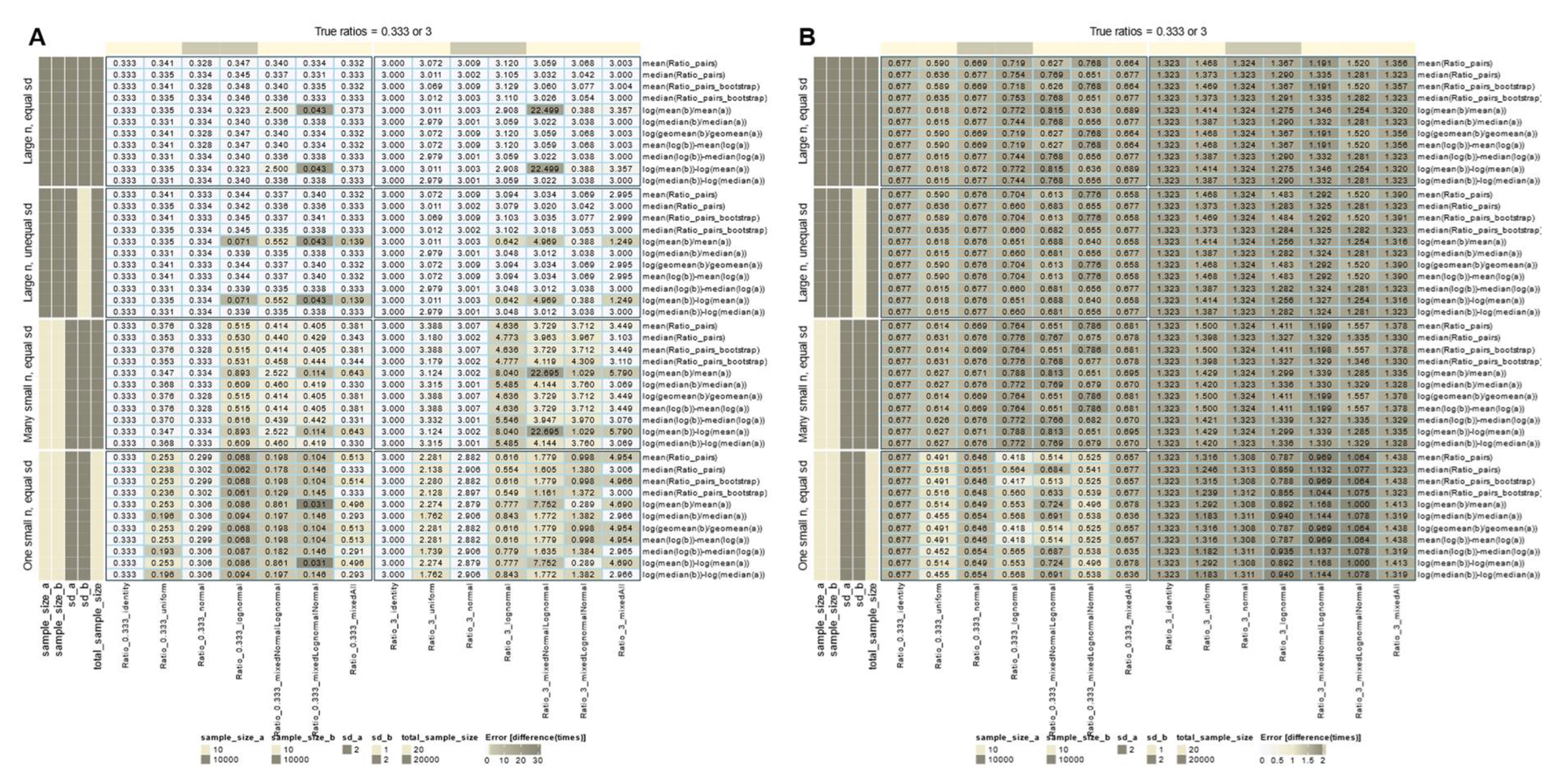
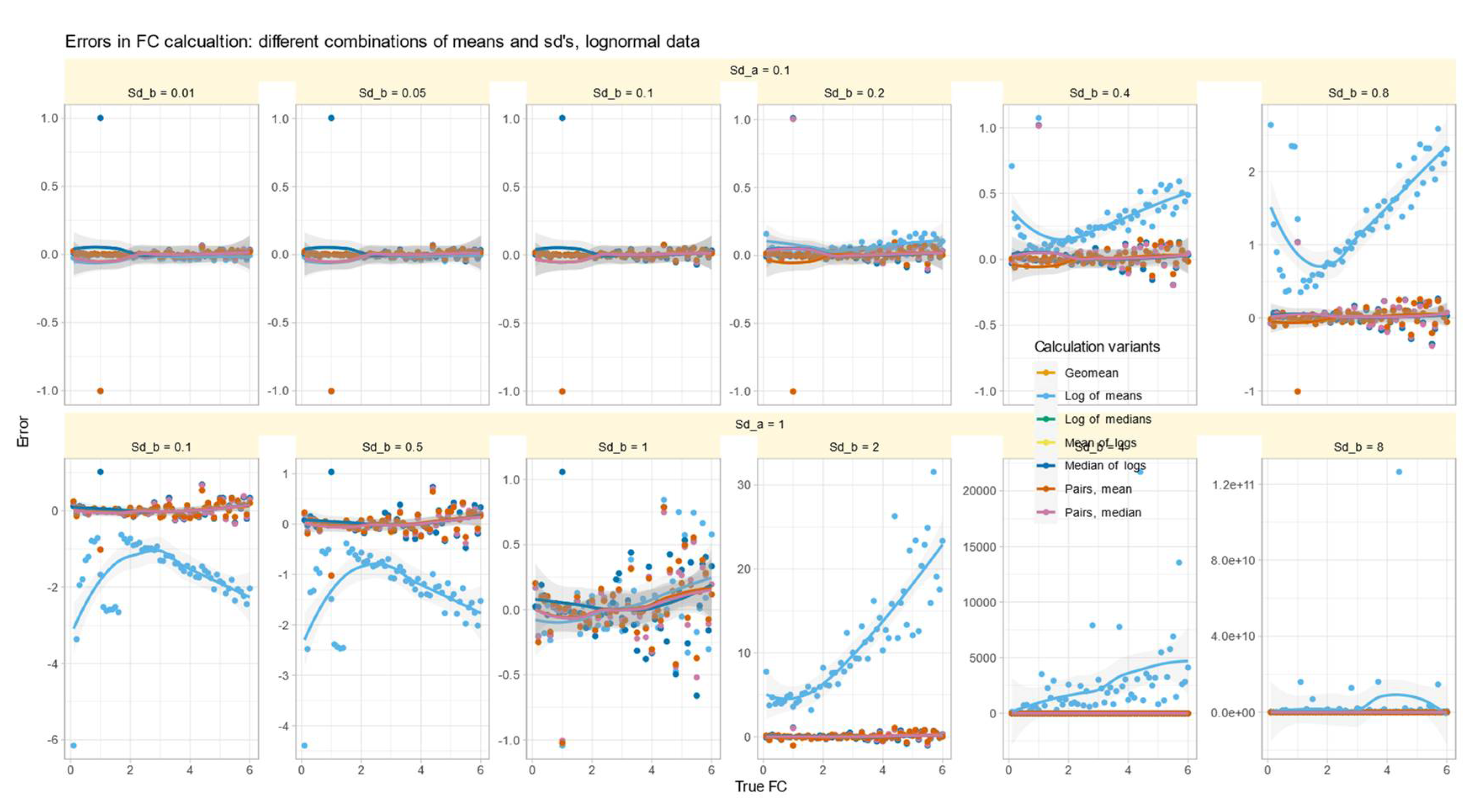
Role of Variance Equality for the Correct Calculation of FC
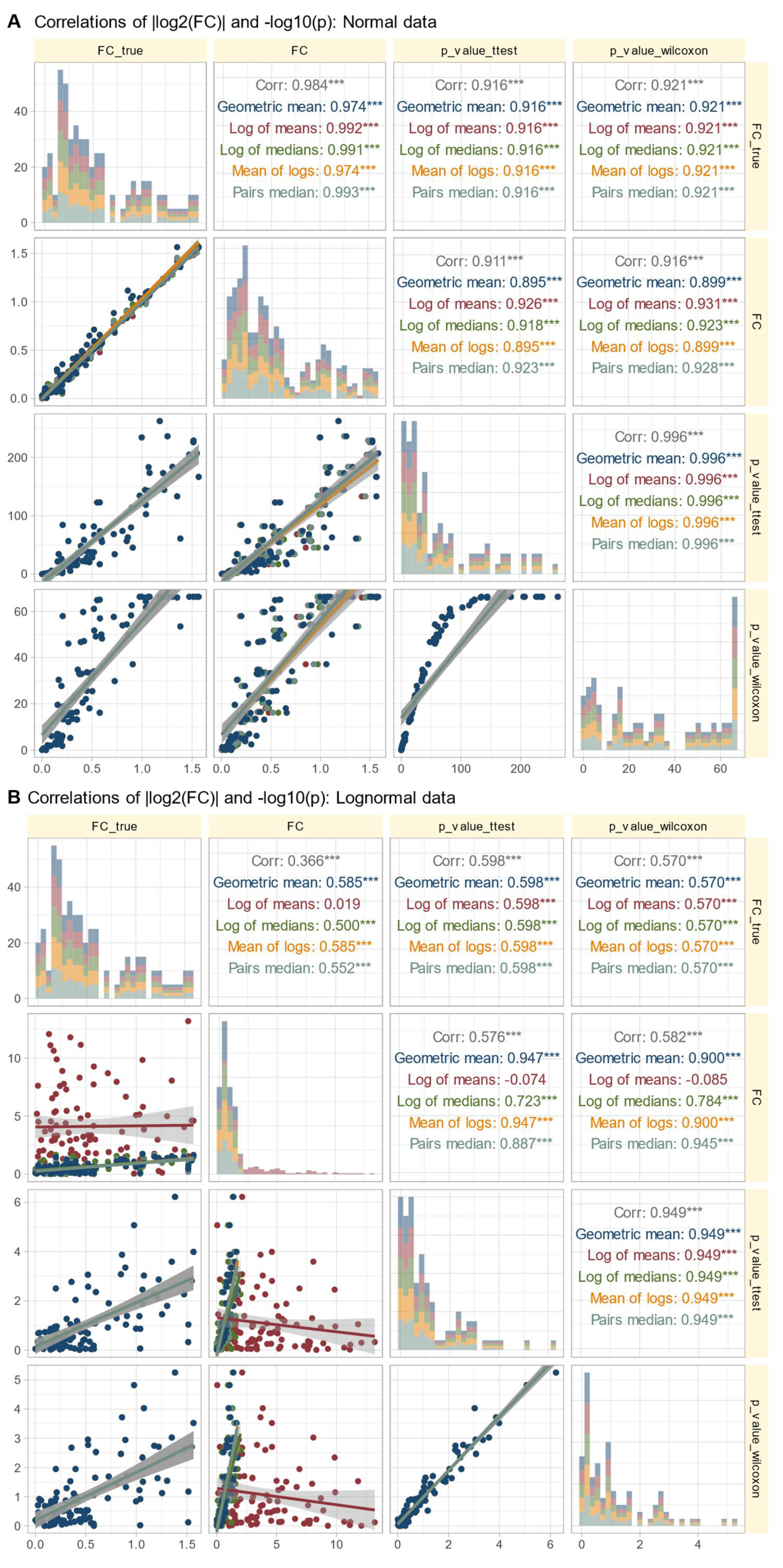
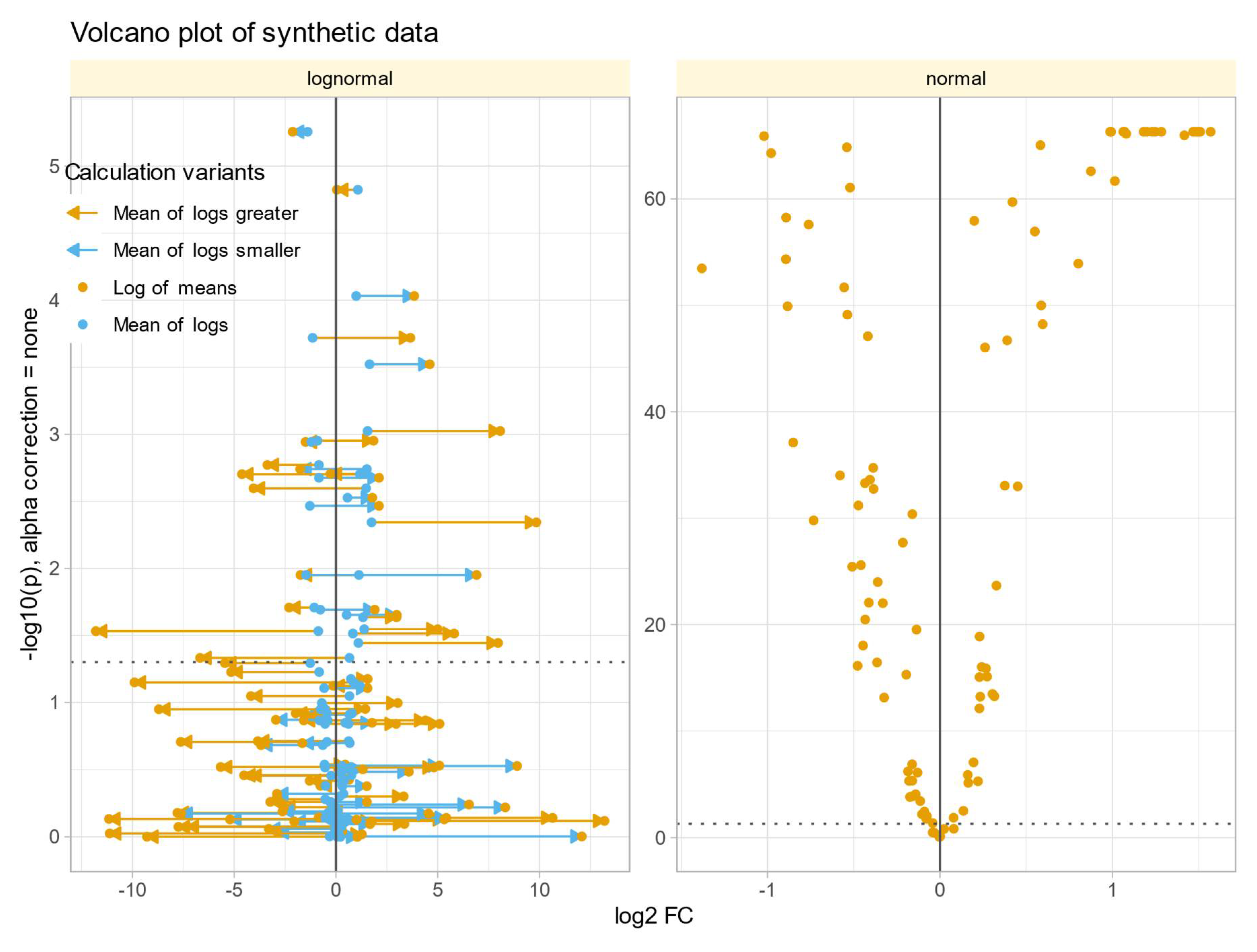
Relationship between Calculated Fold Change and Statistical Significance
FC Calculation Method Dependency in Biomedical Data
Discussion
Conclusions
Declarations
Ethics approval
Consent for Publication
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Guo, L.; Lobenhofer, E.K.; Wang, C.; Shippy, R.; Harris, S.C.; Zhang, L.; Mei, N.; Chen, T.; Herman, D.; Goodsaid, F.M.; et al. Rat toxicogenomic study reveals analytical consistency across microarray platforms. Nat Biotechnol 2006, 24, 1162–1169. [Google Scholar] [CrossRef]
- Dembélé, D.; Kastner, P. Fold change rank ordering statistics: a new method for detecting differentially expressed genes.
- Guyon, I. An introduction to variable and feature selection. J. Mach. Learn. Res. 2003, 3, 1157–1182. [Google Scholar]
- Cheng, J.; Liu, H.-P.; Lin, W.-Y.; Tsai, F.-J. Machine learning compensates fold-change method and highlights oxidative phosphorylation in the brain transcriptome of Alzheimer’s disease. Scientific Reports 2021, 11, 13704. [Google Scholar] [CrossRef] [PubMed]
- Li, W. Volcano plots in analyzing differential expressions with mRNA microarrays. J Bioinform Comput Biol 2012, 10, 1231003. [Google Scholar] [CrossRef]
- Dembélé, D.; Kastner, P. Fold change rank ordering statistics: a new method for detecting differentially expressed genes. BMC Bioinformatics 2014, 15, 14. [Google Scholar] [CrossRef] [PubMed]
- Witten, D.; Tibshirani, R. A comparison of fold-change and the t-statistic for microarray data analysis. 1–17. 2007.
- Fantini, D. easyPubMed: Search and Retrieve Scientific Publication Records from PubMed, 2019.
- Fan, F.Y. PubMedWordcloud: ’Pubmed’ Word Clouds_. R package version 0.3.6, <https://CRAN.R-project.org/package=PubMedWordcloud>, 2019.
- Lötsch, J.; Ultsch, A. Recursive computed ABC (cABC) analysis as a precise method for reducing machine learning based feature sets to their minimum informative size. Sci Rep 2023, 13, 5470. [Google Scholar] [CrossRef]
- Ultsch, A.; Lötsch, J. Computed ABC Analysis for Rational Selection of Most Informative Variables in Multivariate Data. PLoS One 2015, 10, e0129767. [Google Scholar] [CrossRef] [PubMed]
- Tadros, S.F.; D’Souza, M.; Zhu, X.; Frisina, R.D. Gene expression changes for antioxidants pathways in the mouse cochlea: relations to age-related hearing deficits. PLoS One 2014, 9, e90279. [Google Scholar] [CrossRef] [PubMed]
- Olivier, J.; Johnson, W.D.; Marshall, G.D. The logarithmic transformation and the geometric mean in reporting experimental IgE results: what are they and when and why to use them? Ann Allergy Asthma Immunol 2008, 100, 333–337. [Google Scholar] [CrossRef] [PubMed]
- Jain, N.; Thatte, J.; Braciale, T.; Ley, K.; O’Connell, M.; Lee, J.K. Local-pooled-error test for identifying differentially expressed genes with a small number of replicated microarrays. Bioinformatics 2003, 19, 1945–1951. [Google Scholar] [CrossRef] [PubMed]
- Ihaka, R.; Gentleman, R. R: A Language for Data Analysis and Graphics. Journal of Computational and Graphical Statistics 1996, 5, 299–314. [Google Scholar] [CrossRef]
- R Development Core Team. R: A Language and Environment for Statistical Computing. 2008.
- Fechner, G.T. Elemente der Psychophysik; Breitkopf and Härtel: Leipzig, 1860. [Google Scholar]
- Wilcoxon, F. Individual comparisons by ranking methods. Biometrics 1945, 1, 80–83. [Google Scholar] [CrossRef]
- Mann, H.B.; Whitney, D.R. On a test of whether one of two random variables is stochastically larger than the other. Annals of Mathematical Statistics 1947, 18, 50–60. [Google Scholar] [CrossRef]
- Student. The Probable Error of a Mean. Biometrika 1908, 6. [Google Scholar]
- Spearman, C. The proof and measurement of association between two things. Am J Psychol 1904, 15, 72–101. [Google Scholar] [CrossRef]
- Rischke, S.; Poor, S.M.; Gurke, R.; Hahnefeld, L.; Köhm, M.; Ultsch, A.; Geisslinger, G.; Behrens, F.; Lötsch, J. Machine learning identifies right index finger tenderness as key signal of DAS28-CRP based psoriatic arthritis activity. Sci Rep 2023, 13, 22710. [Google Scholar] [CrossRef]
- Kumar, N.; Hoque, M.A.; Sugimoto, M. Robust volcano plot: identification of differential metabolites in the presence of outliers. BMC Bioinformatics 2018, 19, 128. [Google Scholar] [CrossRef] [PubMed]
- Hauber, A.L.; Rosenblatt, M.; Timmer, J. Uncovering specific mechanisms across cell types in dynamical models. PLoS Comput Biol 2023, 19, e1010867. [Google Scholar] [CrossRef] [PubMed]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Fu, W.J.; Hu, J.; Spencer, T.; Carroll, R.; Wu, G. Statistical models in assessing fold change of gene expression in real-time RT-PCR experiments. Comput Biol Chem 2006, 30, 21–26. [Google Scholar] [CrossRef] [PubMed]
- Box, G.E.; Cox, D.R. An analysis of transformations. Journal of the Royal Statistical Society. Series B (Methodological) 1964, 211–252. [Google Scholar] [CrossRef]
- Tukey, J.W. Exploratory Data Analysis; Addison-Wesley: Reading, Mass, 1977. [Google Scholar]
- Tusher, V.G.; Tibshirani, R.; Chu, G. Significance analysis of microarrays applied to the ionizing radiation response. Proc Natl Acad Sci U S A 2001, 98, 5116–5121. [Google Scholar] [CrossRef] [PubMed]
- Li, C.I.; Su, P.F.; Shyr, Y. Sample size calculation based on exact test for assessing differential expression analysis in RNA-seq data. BMC Bioinformatics 2013, 14, 357. [Google Scholar] [CrossRef] [PubMed]
- Bi, R.; Liu, P. Sample size calculation while controlling false discovery rate for differential expression analysis with RNA-sequencing experiments. BMC Bioinformatics 2016, 17, 146. [Google Scholar] [CrossRef] [PubMed]
- Choe, S.E.; Boutros, M.; Michelson, A.M.; Church, G.M.; Halfon, M.S. Preferred analysis methods for Affymetrix GeneChips revealed by a wholly defined control dataset. Genome Biology 2005, 6, R16. [Google Scholar] [CrossRef] [PubMed]
- Newton, M.A.; Kendziorski, C.M.; Richmond, C.S.; Blattner, F.R.; Tsui, K.W. On Differential Variability of Expression Ratios: Improving Statistical Inference about Gene Expression Changes from Microarray Data. Journal of Computational Biology 2001, 8, 37–52. [Google Scholar] [CrossRef] [PubMed]
- Wang, T.; Li, B.; Nelson, C.E.; Nabavi, S. Comparative analysis of differential gene expression analysis tools for single-cell RNA sequencing data. BMC Bioinformatics 2019, 20, 40. [Google Scholar] [CrossRef] [PubMed]
- Dalman, M.R.; Deeter, A.; Nimishakavi, G.; Duan, Z.H. Fold change and p-value cutoffs significantly alter microarray interpretations. BMC Bioinformatics 2012, 13 Suppl 2, S11. [Google Scholar] [CrossRef]
- Kämpke, T. The use of mean values vs. medians in inequality analysis. Journal of economic and social measurement 2010, 35, 43–62. [Google Scholar] [CrossRef]
- Lötsch, J.; Ultsch, A. Comments on the importance of visualizing the distribution of pain-related data. European journal of pain (London, England). [CrossRef]
- National Academies of Sciences, E.; Medicine; Policy; Global, A.; Committee on Science, E.M.; Public, P.; Board on Research, D.; Information; Division on, E.; Physical, S.; et al. In Reproducibility and Replicability in Science; National Academies Press (US): Washington (DC), 2019.
- Wickham, H. ggplot2: Elegant Graphics for Data Analysis; Springer-Verlag New York: 2009.
- Gu, Z.; Eils, R.; Schlesner, M. Complex heatmaps reveal patterns and correlations in multidimensional genomic data. Bioinformatics 2016, 32, 2847–2849. [Google Scholar] [CrossRef] [PubMed]
- Schloerke, B.; Crowley, J.; Cook, D.; Briatte, F.; Marbach, M.; Thoen, E.; Elberg, A.; Larmarange, J. GGally: Extension to ’ggplot2’, 2018.
- Pedersen, T.L. ggforce: Accelerating ’ggplot2’, 2020.


| Definition of expected value | Equation name | Short name | Calculation | Equation # |
|---|---|---|---|---|
| Mean | log(mean(b)/mean(a)) | Log of means | Equation 3 | |
| Mean | log(mean(b))-log(mean(a)) | Log of means | Equation 3 | |
| Median | log(median(b)/median(a)) | Log of medians | Like Equation 3 but median | |
| Median | log(median(b))-log(median(a)) | Log of medians | Like Equation 3 but median | |
| Geometric mean | log(geomean(b)/geomean(a)) | Geometric mean | Equation 4 | |
| Geometric mean | mean(log(b))-mean(log(a)) | Mean of logs | Equation 4 | |
| Mean of logs | median(log(b))-median(log(a)) | Median of logs | Like Equation 4 but median | |
| Paired fold change combinations | mean(Ratio_pairs) | Pairs mean | Equation 5 | |
| Paired fold change combinations | median(Ratio_pairs) | Pairs median | Like Equation 5 but median | |
| Paired fold change combinations | mean(Ratio_pairs_bootstrap) | Pairs mean bootstrap | Like Equation 5 but bootstrapped pairs | |
| Paired fold change combinations | mean(Ratio_pairs_bootstrap) | Pairs median boostrap | Like Equation 5 but bootstrapped pairs |
| Data set | Distribution | Generation |
|---|---|---|
| Data set #1 | Identity |
|
| Uniform |
|
|
| Normal |
|
|
| Log-normal |
|
|
| mixedNormalLognormal |
|
|
| mixedLogormalNormal |
|
|
| Mixed |
|
|
| Data set #2 | Log-normal |
|
| Data set #3 | Normal |
|
| Log-normal |
|
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




