Submitted:
16 May 2024
Posted:
17 May 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. RNA-Seq
2.2. Normalization
2.3. Up-Regulated Genes
2.4. Shannon Entropy
2.5. Statistics
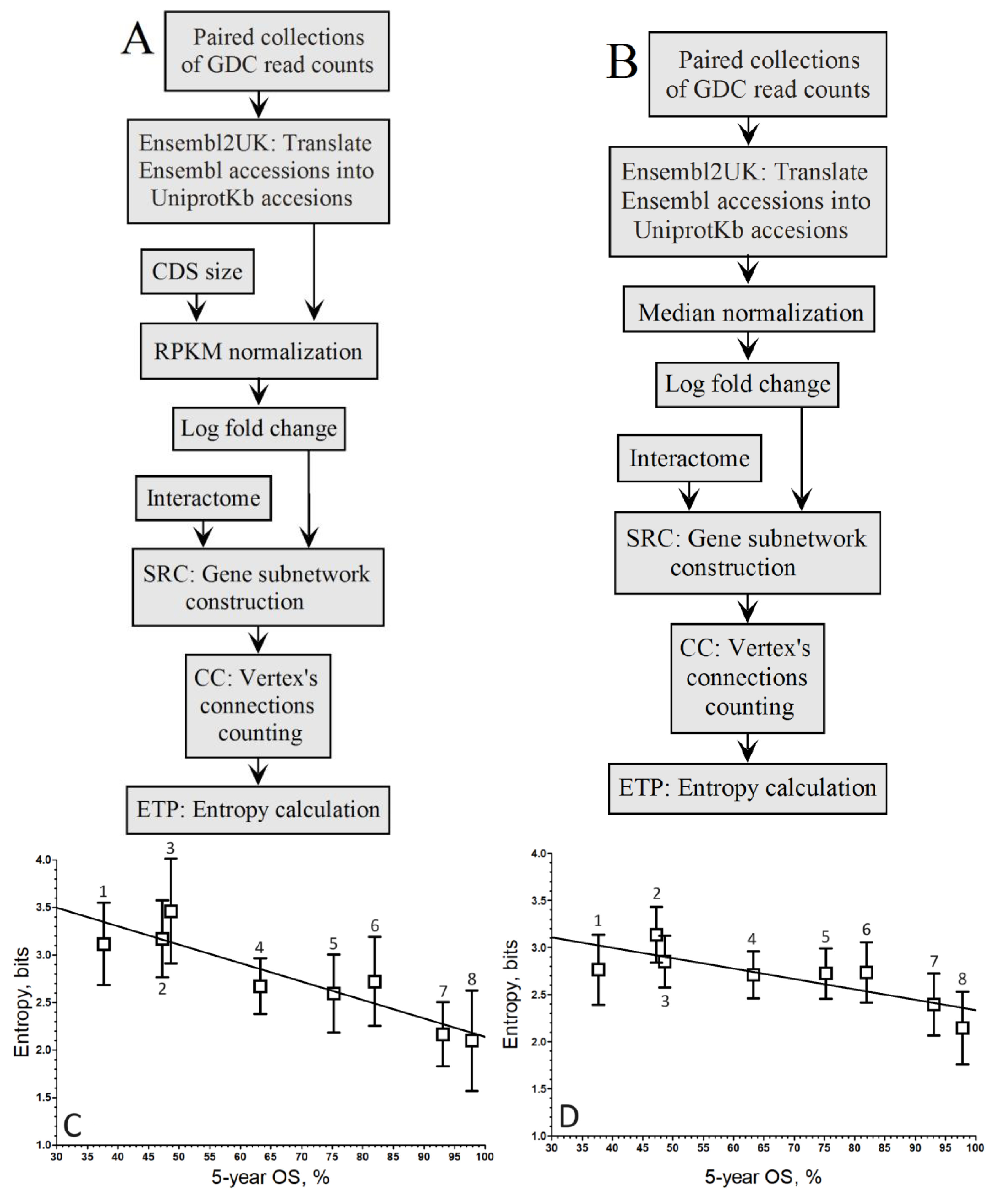
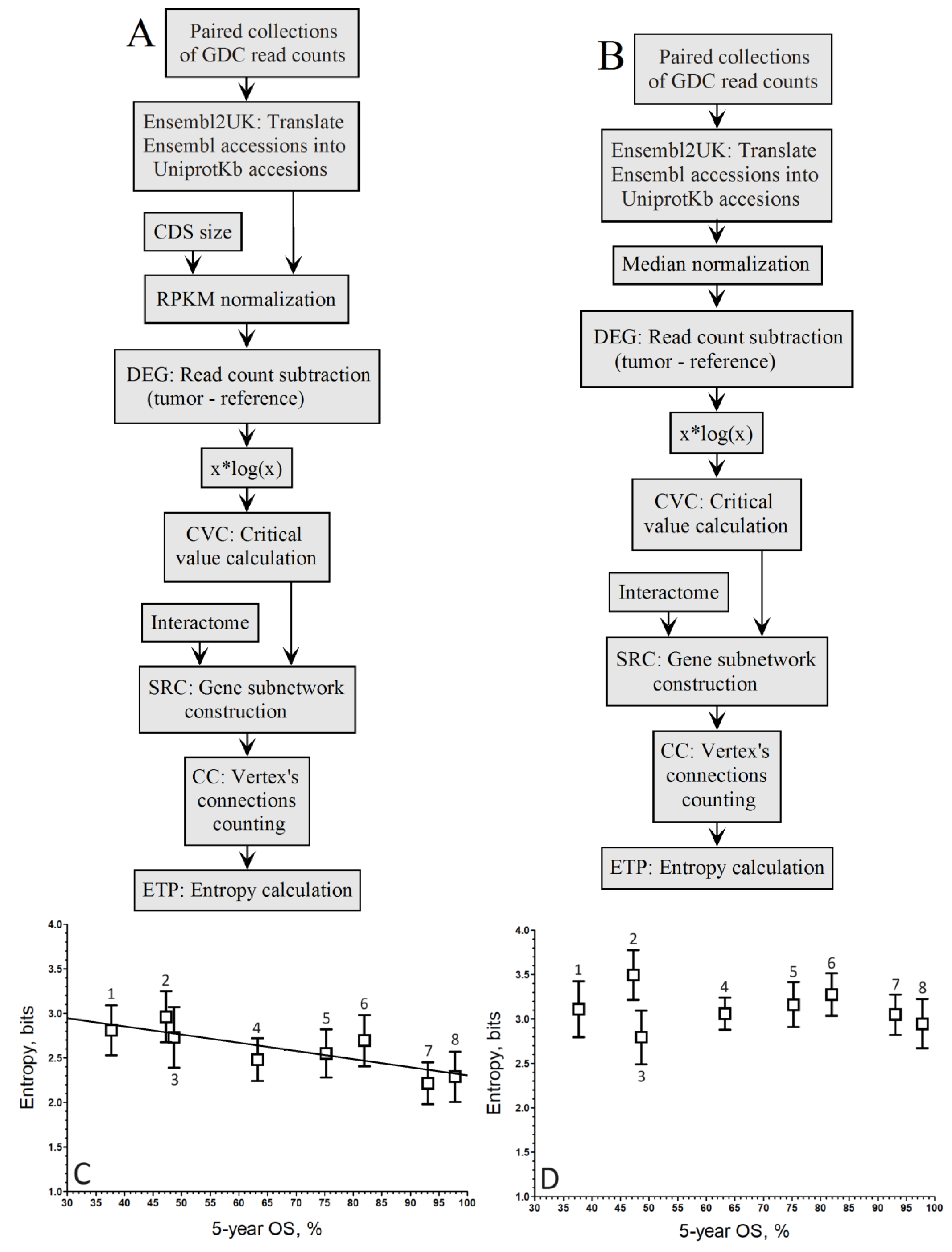
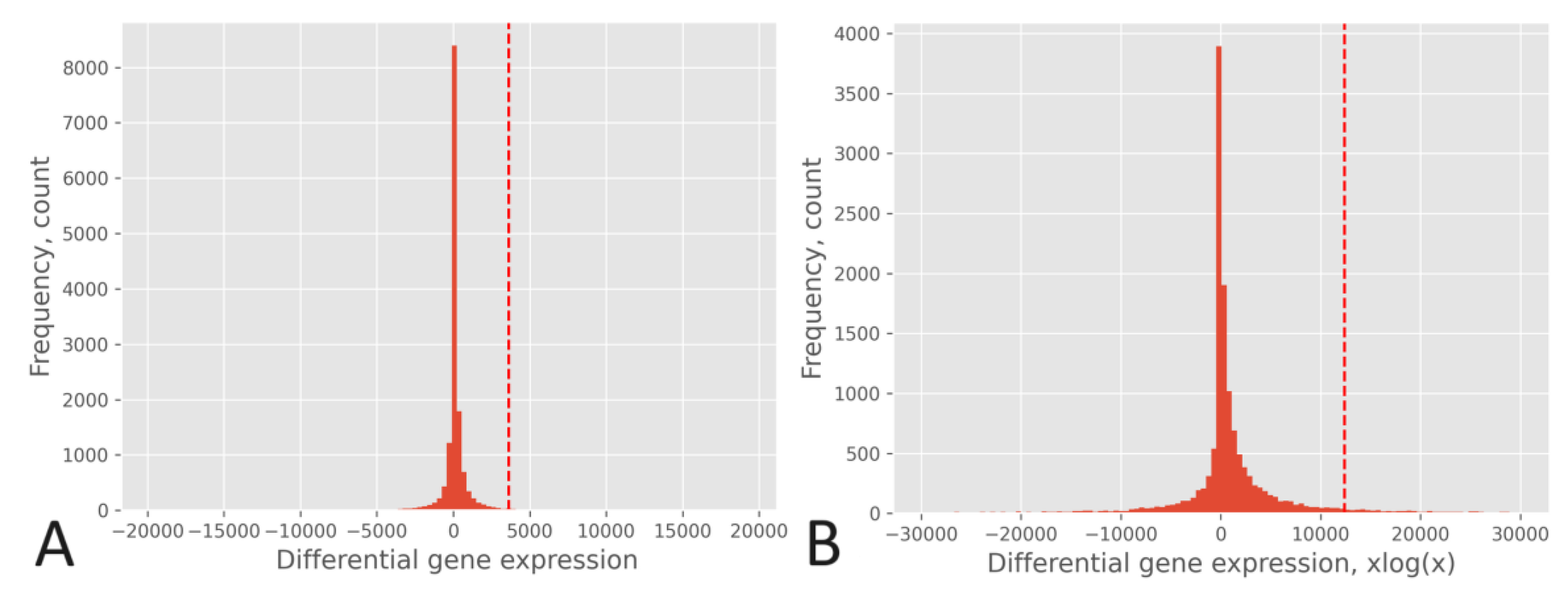
3. Results
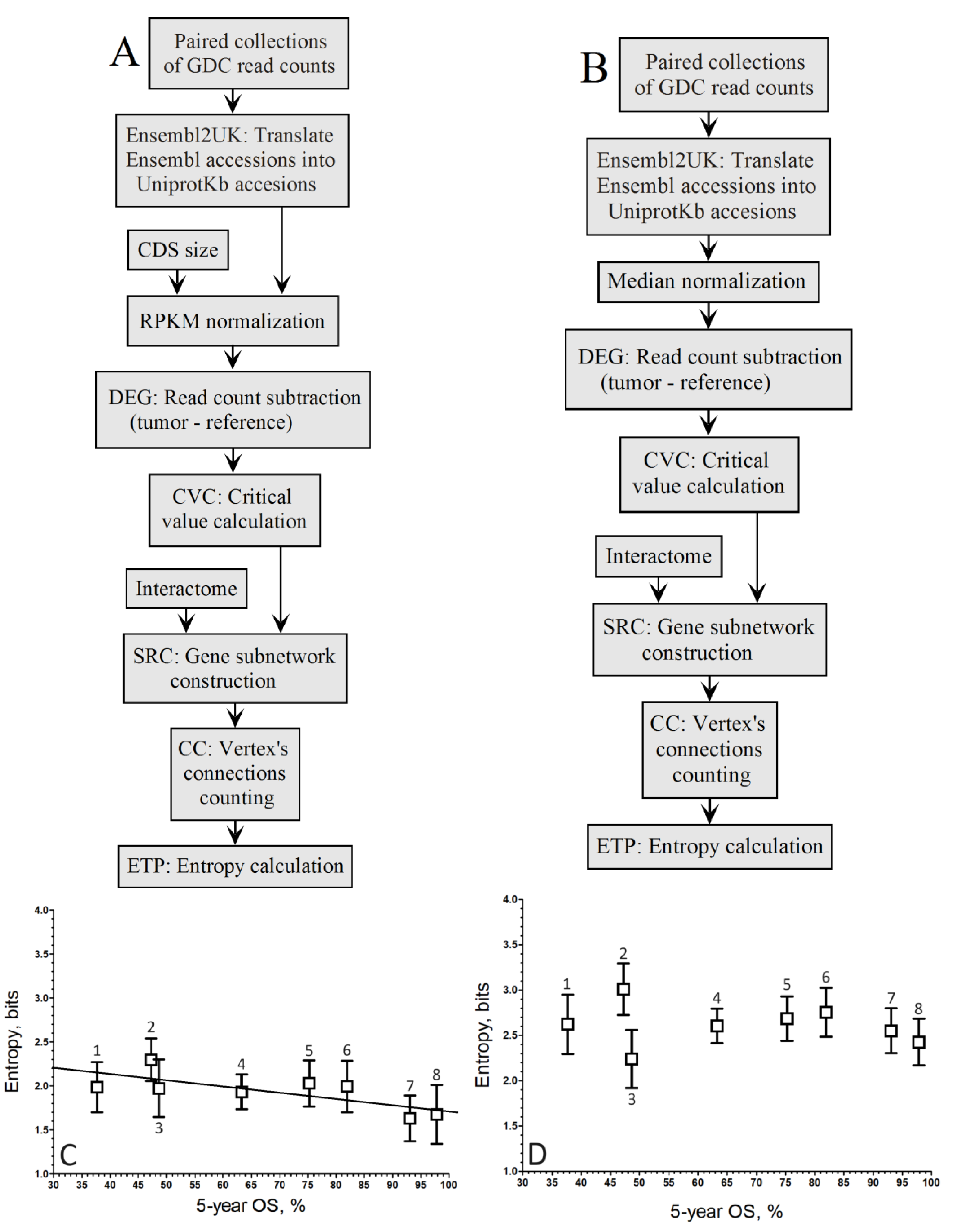
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Hong, M.; Tao, S.; Zhang, L.; Diao, L.T.; Huang, X.; Huang, S.; Xie, S.J.; Xiao, Z.D.; Zhang, H. RNA sequencing: new technologies and applications in cancer research. J. Hematol. Oncol. 2020, 13, 166. [Google Scholar] [CrossRef]
- Kasi, P.M.; Smirnova, S.; Nikulin, V.; Brown, J.H.; Almog, N.; Ogloblina, A.; Yam, C.; Peguero, J.A.; Pandya, D.M.; Kerr, D.; Fowler, N. RNA sequencing as a confirmatory assay and its impact on patient care in multiple cancer types. J. Clin. Oncol. 2023, 41, e15058. [Google Scholar] [CrossRef]
- Nagalakshmi, U.; Wang, Z.; Waern, K.; Shou, C.; Raha, D.; Gerstein, M.; Snyder, M. The transcriptional landscape of the yeast genome defined by RNA sequencing. Science 2008, 320, 1344–1349. [Google Scholar] [CrossRef] [PubMed]
- Holt, R.A.; Jones, S.J. The new paradigm of flow cell sequencing. Genome Res. 2008, 18, 839–846. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Wang, C.Y. From bulk, single-cell to spatial RNA sequencing. Int. J. Oral. Sci. 2021, 13, 36. [Google Scholar] [CrossRef]
- Everaert, C.; Luypaert, M.; Maag, J.L.V.; Cheng, Q.X.; Dinger, M.E.; Hellemans, J.; Mestdagh, P. Benchmarking of RNA-sequencing analysis workflows using whole-transcriptome RT-qPCR expression data. Sci. Rep. 2017, 7, 1559. [Google Scholar] [CrossRef] [PubMed]
- Koch, C.M.; Chiu, S.F.; Akbarpour, M.; Bharat, A.; Ridge, K.M.; Bartom, E.T.; Winter, D.R. A Beginner's guide to analysis of RNA sequencing data. Am. J. Respir. Cell Mol. Biol. 2018, 59, 145–157. [Google Scholar] [CrossRef] [PubMed]
- Evans, C.; Hardin, J.; Stoebel, D.M. Selecting between-sample RNA-seq normalization methods from the perspective of their assumptions. Brief. Bioinform. 2018, 19, 776–792. [Google Scholar] [CrossRef]
- Gerard, D. Data-based RNA-seq simulations by binomial thinning. BMC Bioinformatics 2020, 21, 206. [Google Scholar] [CrossRef]
- Conforte, A.J.; Tuszynski, J.A.; da Silva, F.A.B.; Carels, N. Signaling complexity measured by Shannon entropy and its application in personalized medicine. Front. Genet. 2019, 10, 930. [Google Scholar] [CrossRef]
- Pires, J.G.; da Silva, G.F.; Weyssow, T.; Conforte, A.J.; Pagnoncelli, D.; da Silva, F.A.B.; Carels, N. Galaxy and MEAN Stack to create a user-friendly workflow for the rational optimization of cancer chemotherapy. Front. Genet. 2021, 12, 624259. [Google Scholar] [CrossRef] [PubMed]
- Abrams, Z.B.; Johnson, T.S.; Huang, K.; Payne, P.R.O.; Coombes, K. A protocol to evaluate RNA sequencing normalization methods. BMC Bioinformatics 2019, 20 (Suppl 24). [Google Scholar] [CrossRef] [PubMed]
- Roberts, A.; Trapnell, C.; Donaghey, J.; Rinn, J.L.; Pachter, L. Improving RNA-seq expression estimates by correcting for fragment bias. Genome Biol. 2011, 12, R22. [Google Scholar] [CrossRef] [PubMed]
- Dillies, M.A.; Rau, A.; Aubert, J.; Hennequet-Antier, C.; Jeanmougin, M.; Servant, N.; Keime, C.; Marot, G.; Castel, D.; Estelle, J.; et al. A comprehensive evaluation of normalization methods for Illumina high-throughput RNA sequencing data analysis. Brief. Bioinform. 2013, 14, 671–83. [Google Scholar] [CrossRef] [PubMed]
- Conesa, A.; Madrigal, P.; Tarazona, S.; Gomez-Cabrero, D.; Cervera, A.; McPherson, A.; Szcześniak, M.W.; Gaffney, D.J.; Elo, L.L.; Zhang, X.; Mortazavi, A. A survey of best practices for RNA-seq data analysis. Genome Biol. 2016, 17, 13. [Google Scholar] [CrossRef]
- Everett, L.J.; Mav, D.; Phadke, D.P.; Balik-Meisner, M.R.; Shah, R.R. Impact of aligner, normalization method, and sequencing depth on TempO-seq accuracy. Bioinform. Biol Insights. 2022, 16, 11779322221095216. [Google Scholar] [CrossRef]
- Goll, J.B.; Bosinger, S.E.; Jensen, T.L.; Walum, H.; Grimes, T.; Tharp, G.K.; Natrajan, M.S.; Blazevic, A.; Head, R.D.; Gelber, C.E.; et al. The Vacc-SeqQC project: Benchmarking RNA-seq for clinical vaccine studies. Front. Immunol. 2023, 13, 1093242. [Google Scholar] [CrossRef] [PubMed]
- Ge, S.X.; Son, E.W.; Yao, R. iDEP: an integrated web application for differential expression and pathway analysis of RNA-Seq data. BMC Bioinformatics 2018, 19, 534. [Google Scholar] [CrossRef]
- Le, G.B.H.H.; Steenwyk, J.L.; Manske, N.; Smolin, M.; Abdulali, A.; Kamat, A.; Kanchana, R.; Giffin, K.; Andere, A.; Workman, K. Latch Verified Bulk-RNA Seq toolkit: a cloud-based suite of workflows for bulk RNA-seq quality control, analysis, and functional enrichment. bioRxiv 2022, 2022.11.10.516016. [Google Scholar] [CrossRef]
- Etoh, K.; Nakao, M. A web-based integrative transcriptome analysis, RNAseqChef, uncovers the cell/tissue type-dependent action of sulforaphane. J. Biol. Chem. 2023, 299, 104810. [Google Scholar] [CrossRef]
- Scheepbouwer, C.; Hackenberg, M.; van Eijndhoven, M.A.J.; Gerber, A.; Pegtel, M.; Gómez-Martín, C. NORMSEQ: a tool for evaluation, selection and visualization of RNA-Seq normalization methods. Nucleic Acids Res. 2023, 51, W372–W378. [Google Scholar] [CrossRef] [PubMed]
- Costa-Silva, J.; Domingues, D.; Lopes, F.M. RNA-Seq differential expression analysis: An extended review and a software tool. PLoS One 2017, 12, e0190152. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.; Wong, L.; Goh, W.W.B. How to do quantile normalization correctly for gene expression data analyses. Sci. Rep. 2020, 10, 15534. [Google Scholar] [CrossRef] [PubMed]
- Wang, D.; Cheng, L.; Wang, M.; Wu, R.; Li, P.; Li, B.; Zhang, Y.; Gu, Y.; Zhao, W.; Wang, C.; Guo, Z. Extensive increase of microarray signals in cancers calls for novel normalization assumptions. Comput. Biol. Chem. 2011, 35, 126–130. [Google Scholar] [CrossRef]
- Corchete, L.A.; Rojas, E.A.; Alonso-López, D.; De Las Rivas, J.; Gutiérrez, N.C.; Burguillo, F.J. Systematic comparison and assessment of RNA-seq procedures for gene expression quantitative analysis. Sci. Rep. 2020, 10, 19737. [Google Scholar] [CrossRef] [PubMed]
- Teng, M.; Love, M.I.; Davis, C.A.; Djebali, S.; Dobin, A.; Graveley, B.R.; Li, S.; Mason, C.E.; Olson, S.; Pervouchine, D.; et al. A benchmark for RNA-seq quantification pipelines. Genome Biol. 2016, 17, 74. [Google Scholar] [CrossRef]
- Sampathkumar, N.K.; Sundaram, V.K.; Danthi, P.S.; Barakat, R.; Solomon, S.; Mondal, M.; Carre, I.; El Jalkh, T.; Padilla-Ferrer, A.; Grenier, J.; et al. RNA-seq is not required to determine stable reference genes for qPCR normalization. PLoS Comput. Biol. 2022, 18, e1009868. [Google Scholar] [CrossRef] [PubMed]
- Baik, B.; Yoon, S.; Nam, D. Benchmarking RNA-seq differential expression analysis methods using spike-in and simulation data. PLoS One 2020, 15, e0232271. [Google Scholar] [CrossRef]
- Risso, D.; Ngai, J.; Speed, T.P.; Dudoit, S. Normalization of RNA-seq data using factor analysis of control genes or samples. Nat. Biotechnol. 2014, 32, 896–902. [Google Scholar] [CrossRef]
- Mortazavi, A.; Williams, B.A.; McCue, K.; Schaeffer, L.; Wold, B. Mapping and quantifying mammalian transcriptomes by RNA-seq. Nat. Methods 2008, 5, 621–628. [Google Scholar] [CrossRef]
- Zhao, S.; Ye, Z.; Stanton, R. Misuse of RPKM or TPM normalization when comparing across samples and sequencing protocols. RNA 2020, 26, 903–909. [Google Scholar] [CrossRef] [PubMed]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef] [PubMed]
- Anders, S.; Huber, W. Differential expression analysis for sequence count data. Genome biology 2010, 11, R106. [Google Scholar] [CrossRef]
- Carels, N.; Tilli, T.; Tuszynski, J.A. A computational strategy to select optimized protein targets for drug development toward the control of cancer diseases. PLoS One 2015, 10, e0115054. [Google Scholar] [CrossRef] [PubMed]
- Tilli, T.M.; Castro, C.S.; Tuszynski, J.A.; Carels, N. A strategy to identify housekeeping genes suitable for analysis in breast cancer diseases. BMC Genomics 2016, 17, 639. [Google Scholar] [CrossRef] [PubMed]
- Barbosa-Silva, A.; Magalhães, M.; da Silva, G.F.; da Silva, F.A.B.; Carneiro, F.R.G.; Carels, N. A data science approach for the identification of molecular signatures of aggressive cancers. Cancers 2022, 14, 2325. [Google Scholar] [CrossRef] [PubMed]
- Breitkreutz, D.; Hlatky, L.; Rietman, E.; Tuszynski, J.A. Molecular signaling network complexity is correlated with cancer patient survivability. Proc. Natl. Acad. Sci. USA 2012, 109, 9209–9212. [Google Scholar] [CrossRef]
- Liu, J.; Lichtenberg, T.; Hoadley, K.; Poisson, L.; Lazar, A.; Cherniack, A.D.; Kovatich, A.J.; Benz, C.C.; Levine, D.A.; Lee, A.V.; et al. An integrated TCGA pan-cancer clinical data resource to drive high-quality survival outcome analytics. Cell 2018, 173, 400–416. [Google Scholar] [CrossRef]
- Zenil, H.; Kiani, N.A.; Tegnér, J. A review of graph and network complexity from an algorithmic information perspective. Entropy 2018, 20, 551. [Google Scholar] [CrossRef]
- Jolicoeur, P. Bivariate allometry: interval estimation of the slopes of the ordinary and standardized normal major axes and structural relationship. J. Theor. Biol. 1990, 144, 275–85. [Google Scholar] [CrossRef]
- van Wieringen, W.N.; van der Vaart, A.W. Statistical analysis of the cancer cell’s molecular entropy using high-throughput data. Bioinformatics 2011, 27, 556–563. [Google Scholar] [CrossRef] [PubMed]
- Winterbach, W.; Mieghem, P.; Reinders, M.; Wang, H.; de Ridder, D. Topology of molecular interaction networks. BMC Syst. Biol. 2013, 7, 90. [Google Scholar] [CrossRef] [PubMed]
- Banerji, C.R.S.; Severini, S.; Caldas, C.; Teschendorff, A.E. Intra-tumour signalling entropy determines clinical outcome in breast and lung cancer. PLoS Comput. Biol. 2015, 11, e1004115. [Google Scholar] [CrossRef] [PubMed]
| Cancer type | Abbreviation | OS1 | GDC, n2 |
|---|---|---|---|
| Stomach adenocarcinoma | STAD | 38 | 27 |
| Lung squamous cell carcinoma | LUSC | 47 | 48 |
| Liver hepatocellular carcinoma | LIHC | 49 | 50 |
| Kidney renal clear cell carcinoma | KIRC | 63 | 71 |
| Kidney renal papillary cell carcinoma | KIRP | 75 | 31 |
| Breast cancer | BRCA | 82 | 46 |
| Thyroid cancer | THCA | 93 | 56 |
| Prostate cancer | PRAD | 98 | 50 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




