Submitted:
09 May 2024
Posted:
10 May 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
- 2.1.1. Animals
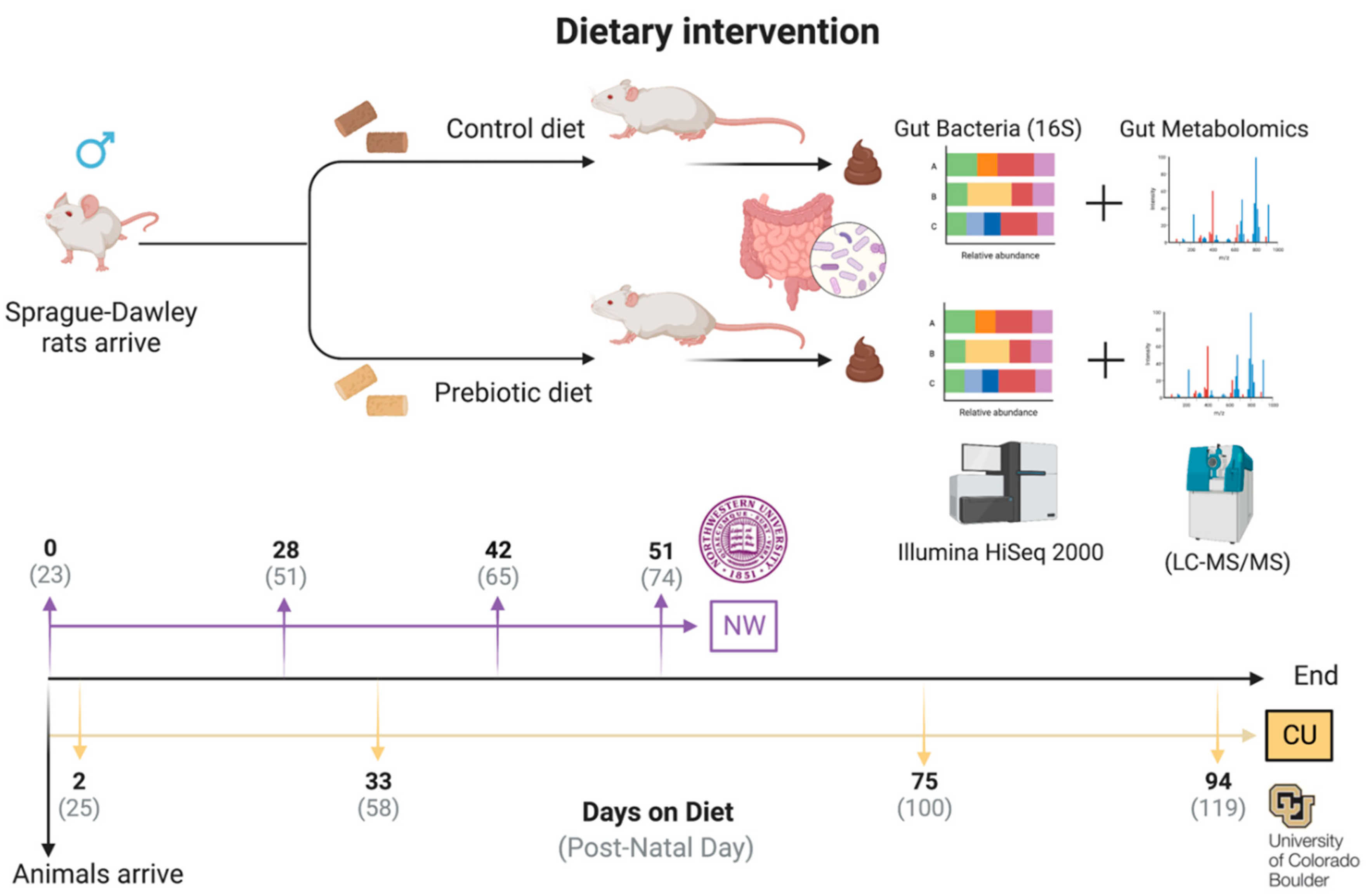
- 2.1.2. Experimental Design
- 2.1.3. Diets
- 2.1.4. Fecal Sample Collection Procedures
- 2.1.5. 16S rRNA Gene Sequencing
- 2.1.6. LC-MS/MS Metabolomics
- 2.1.7. Statistical Analysis
3. Results
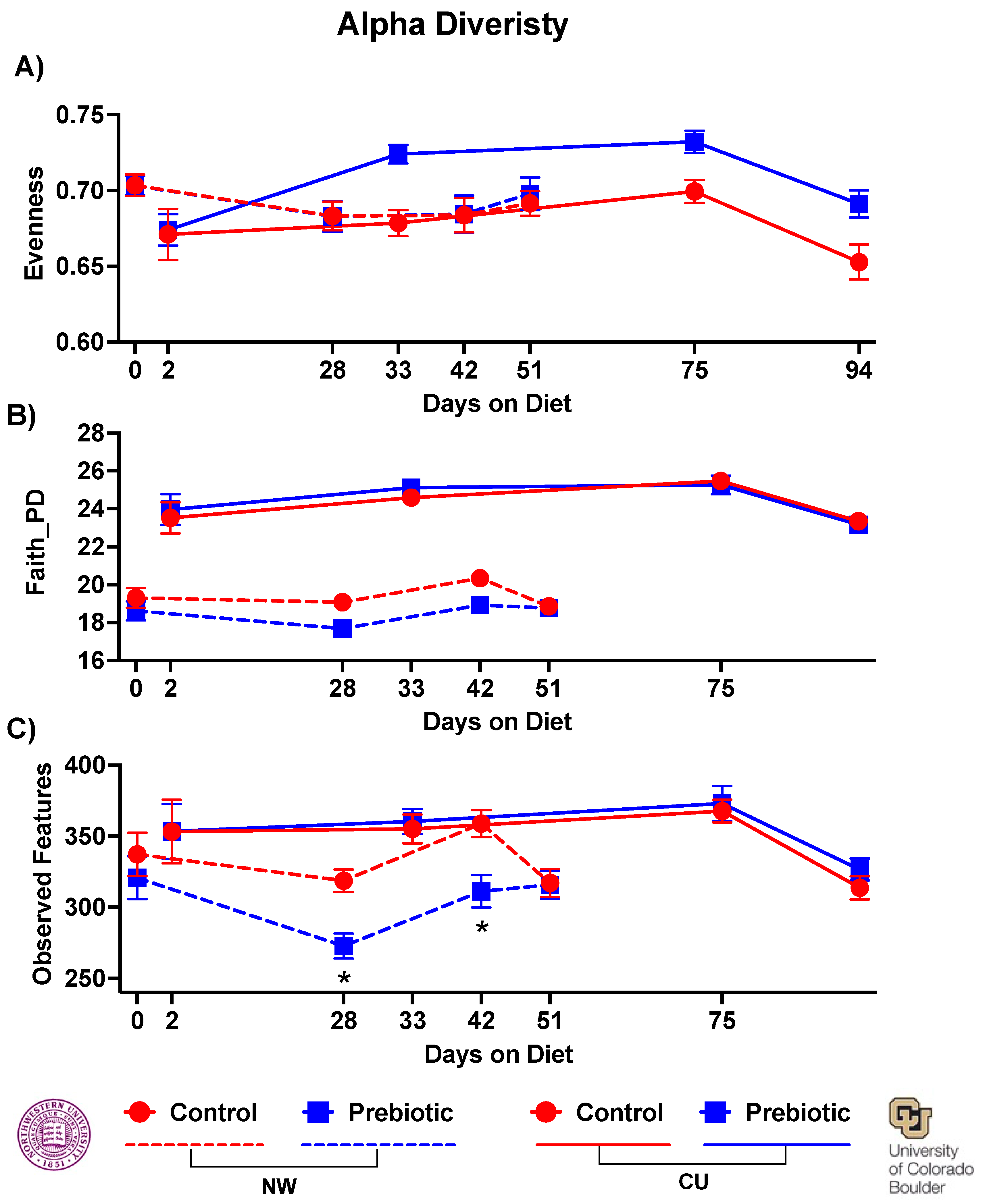
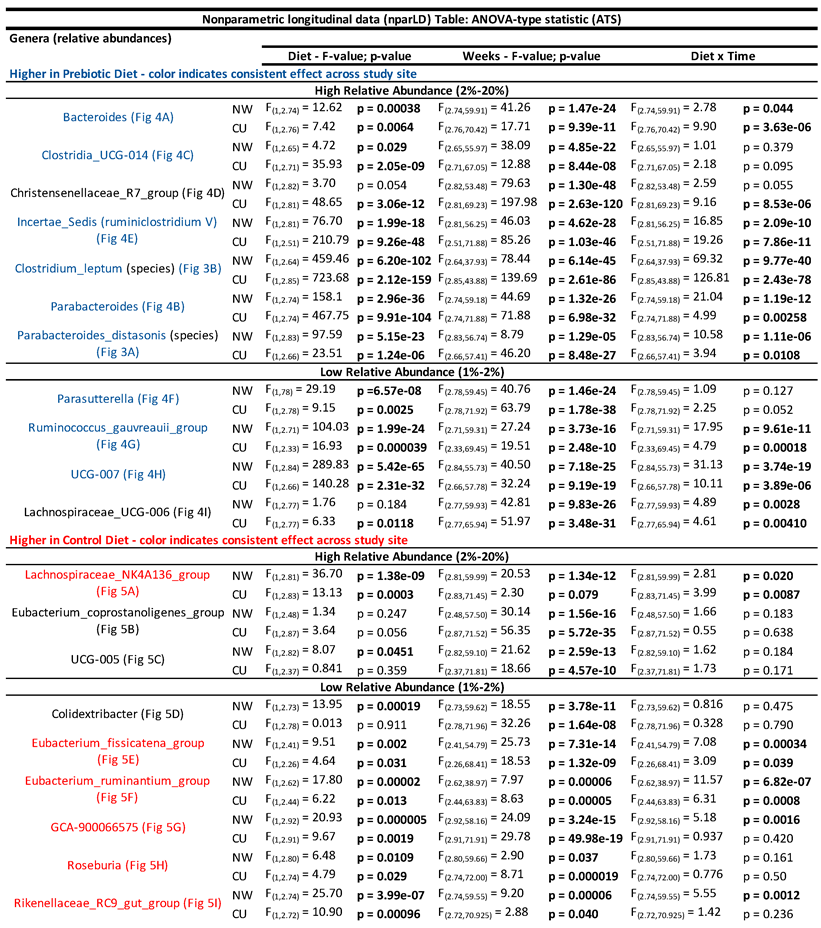
- 3.1.1. Microbiome
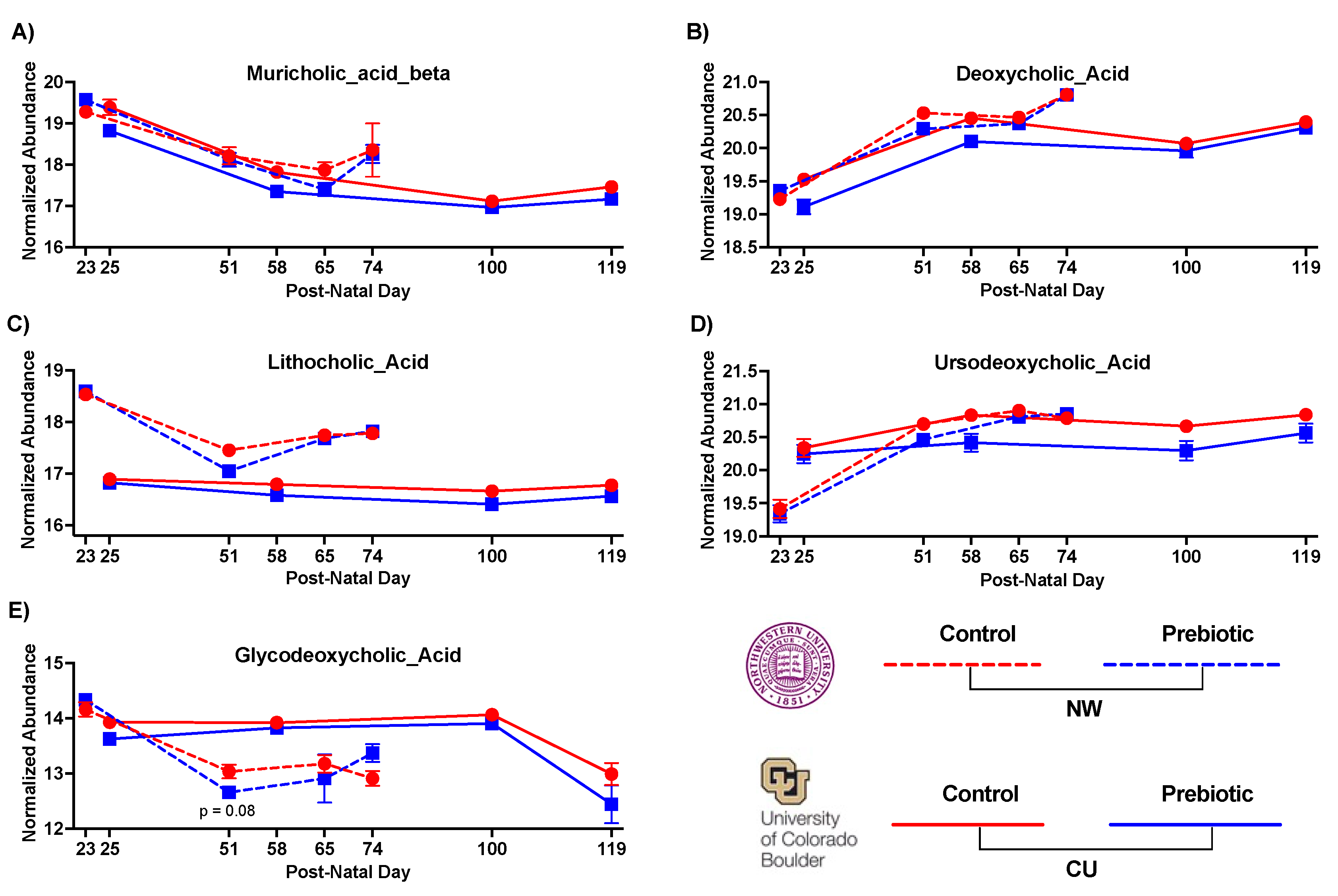
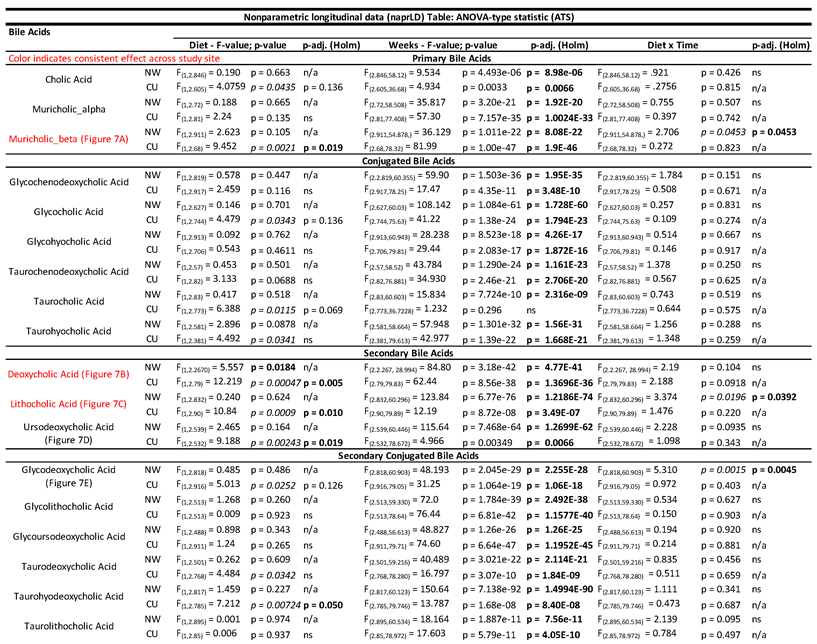
- 3.1.2. Metabolome - Bile Acids
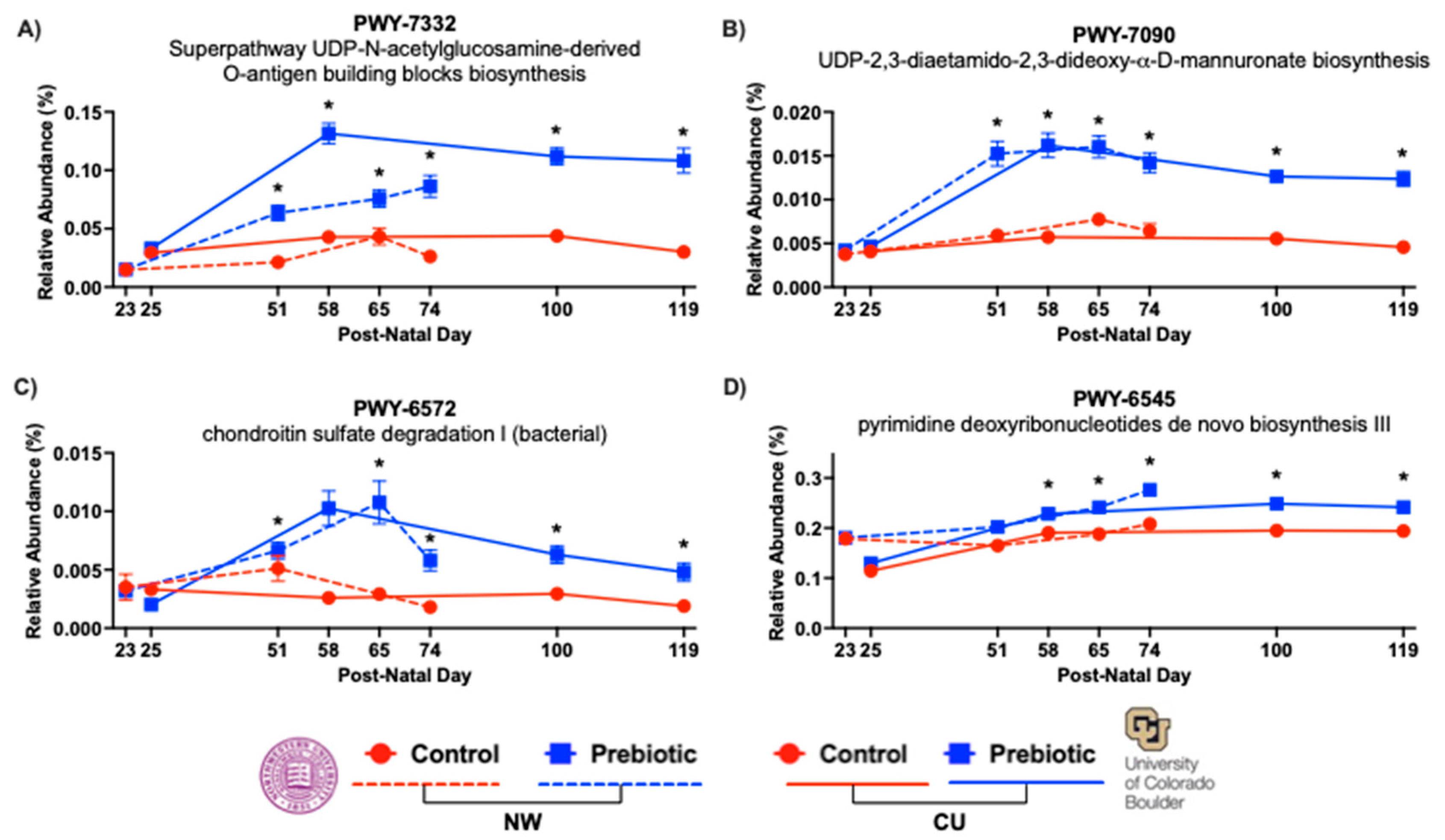
- 3.1.3. PICRUSt2 – pathways
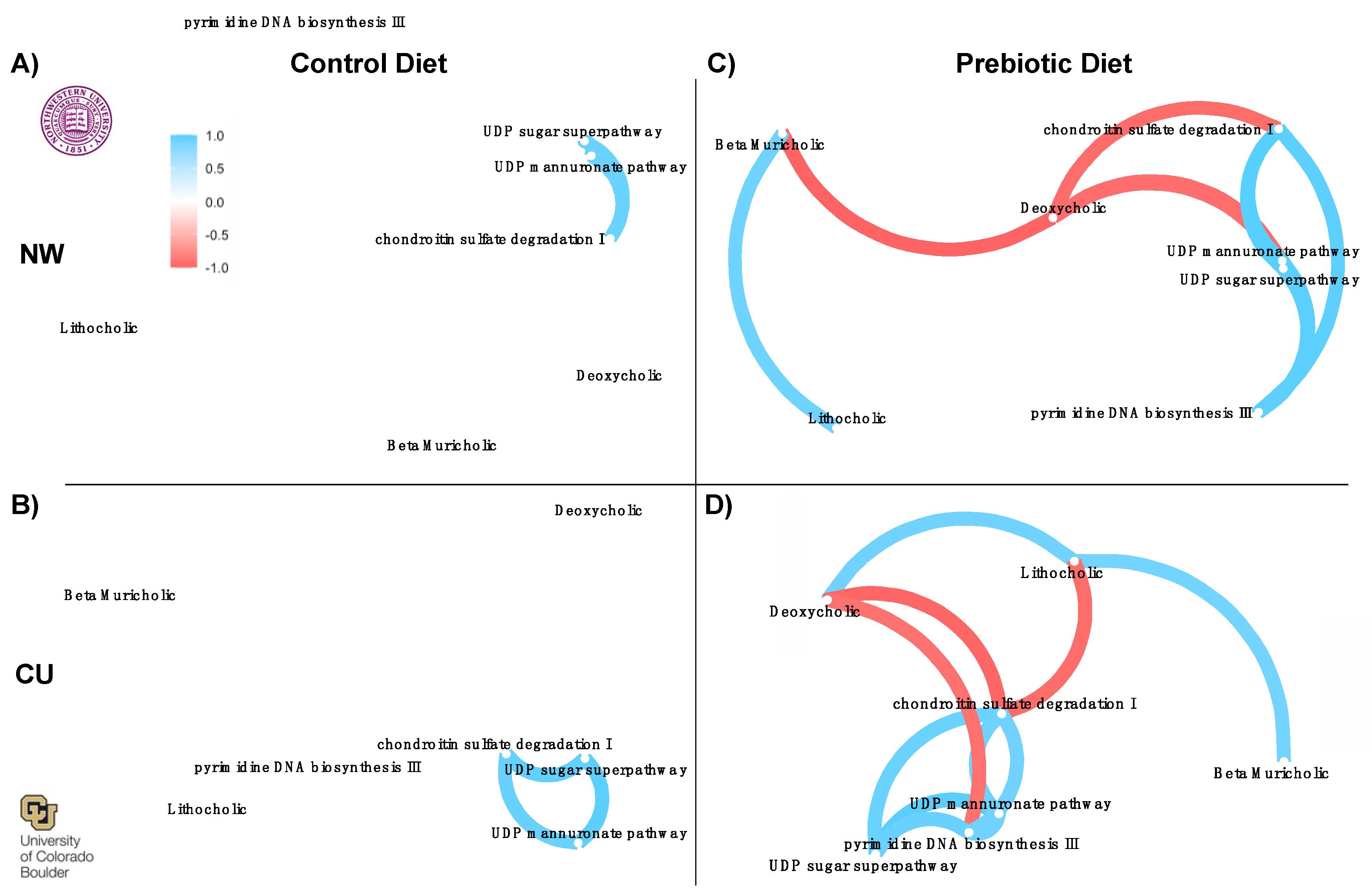
- 3.1.4. Correlation Network Analysis
4. Discussion
Supplementary Materials
Funding
Conflicts of Interest
References
- Zhang, Y.; Wang, R. The human gut phageome: composition, development, and alterations in disease. Front Microbiol 2023, 14, 1213625. [Google Scholar] [CrossRef]
- Zhang, X.; Meng, H.; Hu, X.; Yuan, Z. Diversity and functional profile of gut symbiotic bacteria between Lysinibacillus sphaericus C(3)-41 susceptible and resistant Culex quinquefasciatus Say as revealed by 16S rRNA gene high-throughput sequencing. Front Microbiol 2022, 13, 991105. [Google Scholar] [CrossRef] [PubMed]
- Naya-Catala, F.; Piazzon, M.C.; Calduch-Giner, J.A.; Sitja-Bobadilla, A.; Perez-Sanchez, J. Diet and Host Genetics Drive the Bacterial and Fungal Intestinal Metatranscriptome of Gilthead Sea Bream. Front Microbiol 2022, 13, 883738. [Google Scholar] [CrossRef] [PubMed]
- Vincenot, C.E.; Giannino, F.; Rietkerk, M.; Moriya, K.; Mazzoleni, S. Theoretical considerations on the combined use of System Dynamics and individual-based modeling in ecology. Ecol Model 2011, 222, 210–218. [Google Scholar] [CrossRef]
- Molly, K.; Vandewoestyne, M.; Desmet, I.; Verstraete, W. Validation of the Simulator of the Human Intestinal Microbial Ecosystem (Shime) Reactor Using Microorganism-Associated Activities. Microb Ecol Health D 1994, 7, 191–200. [Google Scholar] [CrossRef]
- Wu, G.D.; Chen, J.; Hoffmann, C.; Bittinger, K.; Chen, Y.Y.; Keilbaugh, S.A.; Bewtra, M.; Knights, D.; Walters, W.A.; Knight, R.; et al. Linking long-term dietary patterns with gut microbial enterotypes. Science 2011, 334, 105–108. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; Qin, P.; Wang, J. High-Fat Diet Alters the Intestinal Microbiota in Streptozotocin-Induced Type 2 Diabetic Mice. Microorganisms 2019, 7. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Blouin, J.M.; Santacruz, A.; Lan, A.; Andriamihaja, M.; Wilkanowicz, S.; Benetti, P.H.; Tome, D.; Sanz, Y.; Blachier, F.; et al. High-protein diet modifies colonic microbiota and luminal environment but not colonocyte metabolism in the rat model: the increased luminal bulk connection. Am J Physiol Gastrointest Liver Physiol 2014, 307, G459–G470. [Google Scholar] [CrossRef] [PubMed]
- Barra, N.G.; Anhe, F.F.; Cavallari, J.F.; Singh, A.M.; Chan, D.Y.; Schertzer, J.D. Micronutrients impact the gut microbiota and blood glucose. J Endocrinol 2021, 250, R1–R21. [Google Scholar] [CrossRef]
- Gibson, G.R.; Hutkins, R.; Sanders, M.E.; Prescott, S.L.; Reimer, R.A.; Salminen, S.J.; Scott, K.; Stanton, C.; Swanson, K.S.; Cani, P.D.; et al. Expert consensus document: The International Scientific Association for Probiotics and Prebiotics (ISAPP) consensus statement on the definition and scope of prebiotics. Nat Rev Gastroenterol Hepatol 2017, 14, 491–502. [Google Scholar] [CrossRef]
- Schloss, P.D. Identifying and Overcoming Threats to Reproducibility, Replicability, Robustness, and Generalizability in Microbiome Research. mBio 2018, 9. [Google Scholar] [CrossRef] [PubMed]
- Vitek, J.; Kalibera, T. R-3 - Repeatability, Reproducibility and Rigor. Acm Sigplan Notices 2012, 47, 30–36. [Google Scholar] [CrossRef]
- Eaton, K.; Pirani, A.; Snitkin, E.S.; Reproducibility Project: Cancer, B. Replication Study: Intestinal inflammation targets cancer-inducing activity of the microbiota. Elife 2018, 7. [Google Scholar] [CrossRef] [PubMed]
- Repass, J.; Reproducibility Project: Cancer, B. Replication Study: Fusobacterium nucleatum infection is prevalent in human colorectal carcinoma. Elife 2018, 7. [Google Scholar] [CrossRef] [PubMed]
- Knight, R.; Vrbanac, A.; Taylor, B.C.; Aksenov, A.; Callewaert, C.; Debelius, J.; Gonzalez, A.; Kosciolek, T.; McCall, L.I.; McDonald, D.; et al. Best practices for analysing microbiomes. Nat Rev Microbiol 2018, 16, 410–422. [Google Scholar] [CrossRef] [PubMed]
- Panek, M.; Cipcic Paljetak, H.; Baresic, A.; Peric, M.; Matijasic, M.; Lojkic, I.; Vranesic Bender, D.; Krznaric, Z.; Verbanac, D. Methodology challenges in studying human gut microbiota - effects of collection, storage, DNA extraction and next generation sequencing technologies. Sci Rep 2018, 8, 5143. [Google Scholar] [CrossRef] [PubMed]
- De Filippo, C.; Cavalieri, D.; Di Paola, M.; Ramazzotti, M.; Poullet, J.B.; Massart, S.; Collini, S.; Pieraccini, G.; Lionetti, P. Impact of diet in shaping gut microbiota revealed by a comparative study in children from Europe and rural Africa. Proc Natl Acad Sci U S A 2010, 107, 14691–14696. [Google Scholar] [CrossRef] [PubMed]
- Gupta, V.K.; Paul, S.; Dutta, C. Geography, Ethnicity or Subsistence-Specific Variations in Human Microbiome Composition and Diversity. Front Microbiol 2017, 8, 1162. [Google Scholar] [CrossRef] [PubMed]
- Kumari, M.; Bhushan, B.; Eslavath, M.R.; Srivastava, A.K.; Meena, R.C.; Varshney, R.; Ganju, L. Impact of high altitude on composition and functional profiling of oral microbiome in Indian male population. Sci Rep 2023, 13, 4038. [Google Scholar] [CrossRef]
- Dong, W.; Ma, L.; Huang, Q.; Yang, X.; Mei, Z.; Kong, M.; Sun, Z.; Zhang, Z.; Li, J.; Zou, J.; et al. Gut microbiome alterations in pulmonary hypertension in highlanders and lowlanders. ERJ Open Res 2023, 9. [Google Scholar] [CrossRef]
- Rothschild, D.; Weissbrod, O.; Barkan, E.; Kurilshikov, A.; Korem, T.; Zeevi, D.; Costea, P.I.; Godneva, A.; Kalka, I.N.; Bar, N.; et al. Environment dominates over host genetics in shaping human gut microbiota. Nature 2018, 555, 210–215. [Google Scholar] [CrossRef]
- Ericsson, A.C.; Franklin, C.L. The gut microbiome of laboratory mice: considerations and best practices for translational research. Mamm Genome 2021, 32, 239–250. [Google Scholar] [CrossRef] [PubMed]
- Bowers, S.J.; Summa, K.C.; Thompson, R.S.; Gonzalez, A.; Vargas, F.; Olker, C.; Jiang, P.; Lowry, C.A.; Dorrestein, P.C.; Knight, R.; et al. A Prebiotic Diet Alters the Fecal Microbiome and Improves Sleep in Response to Sleep Disruption in Rats. Front Neurosci 2022, 16, 889211. [Google Scholar] [CrossRef]
- Thompson, R.S.; Gaffney, M.; Hopkins, S.; Kelley, T.; Gonzalez, A.; Bowers, S.J.; Vitaterna, M.H.; Turek, F.W.; Foxx, C.L.; Lowry, C.A.; et al. Ruminiclostridium 5, Parabacteroides distasonis, and bile acid profile are modulated by prebiotic diet and associate with facilitated sleep/clock realignment after chronic disruption of rhythms. Brain Behav Immun 2021, 97, 150–166. [Google Scholar] [CrossRef] [PubMed]
- Yang, Z.D.; Guo, Y.S.; Huang, J.S.; Gao, Y.F.; Peng, F.; Xu, R.Y.; Su, H.H.; Zhang, P.J. Isomaltulose Exhibits Prebiotic Activity, and Modulates Gut Microbiota, the Production of Short Chain Fatty Acids, and Secondary Bile Acids in Rats. Molecules 2021, 26. [Google Scholar] [CrossRef] [PubMed]
- Phungviwatnikul, T.; Lee, A.H.; Belchik, S.E.; Suchodolski, J.S.; Swanson, K.S. Weight loss and high-protein, high-fiber diet consumption impact blood metabolite profiles, body composition, voluntary physical activity, fecal microbiota, and fecal metabolites of adult dogs. J Anim Sci 2022, 100. [Google Scholar] [CrossRef] [PubMed]
- McMillin, M.; DeMorrow, S. Effects of bile acids on neurological function and disease. FASEB J 2016, 30, 3658–3668. [Google Scholar] [CrossRef] [PubMed]
- Perino, A.; Demagny, H.; Velazquez-Villegas, L.A.; Schoonjans, K. Molecular Physiology of Bile Acid Signaling in Health, Disease and Aging. Physiol Rev 2020. [Google Scholar] [CrossRef] [PubMed]
- Thompson, R.S.; Roller, R.; Mika, A.; Greenwood, B.N.; Knight, R.; Chichlowski, M.; Berg, B.M.; Fleshner, M. Dietary Prebiotics and Bioactive Milk Fractions Improve NREM Sleep, Enhance REM Sleep Rebound and Attenuate the Stress-Induced Decrease in Diurnal Temperature and Gut Microbial Alpha Diversity. Front Behav Neurosci 2017, 10, 240. [Google Scholar] [CrossRef]
- Thompson, R.S.; Vargas, F.; Dorrestein, P.C.; Chichlowski, M.; Berg, B.M.; Fleshner, M. Dietary prebiotics alter novel microbial dependent fecal metabolites that improve sleep. Sci Rep 2020, 10, 3848. [Google Scholar] [CrossRef]
- Mika, A.; Day, H.E.; Martinez, A.; Rumian, N.L.; Greenwood, B.N.; Chichlowski, M.; Berg, B.M.; Fleshner, M. Early life diets with prebiotics and bioactive milk fractions attenuate the impact of stress on learned helplessness behaviours and alter gene expression within neural circuits important for stress resistance. Eur J Neurosci 2016. [Google Scholar] [CrossRef] [PubMed]
- Mika, A.; Gaffney, M.; Roller, R.; Hills, A.; Bouchet, C.A.; Hulen, K.A.; Thompson, R.S.; Chichlowski, M.; Berg, B.M.; Fleshner, M. Feeding the developing brain: Juvenile rats fed diet rich in prebiotics and bioactive milk fractions exhibit reduced anxiety-related behavior and modified gene expression in emotion circuits. Neurosci Lett 2018. [Google Scholar] [CrossRef] [PubMed]
- Mika, A.; Rumian, N.; Loughridge, A.B.; Fleshner, M. Exercise and Prebiotics Produce Stress Resistance: Converging Impacts on Stress-Protective and Butyrate-Producing Gut Bacteria. Int Rev Neurobiol 2016, 131, 165–191. [Google Scholar] [CrossRef] [PubMed]
- Bowers, S.J.; Vargas, F.; Gonzalez, A.; He, S.; Jiang, P.; Dorrestein, P.C.; Knight, R.; Wright, K.P., Jr.; Lowry, C.A.; Fleshner, M.; et al. Repeated sleep disruption in mice leads to persistent shifts in the fecal microbiome and metabolome. PLoS One 2020, 15, e0229001. [Google Scholar] [CrossRef] [PubMed]
- Mika, A.; Fleshner, M. Early-life exercise may promote lasting brain and metabolic health through gut bacterial metabolites. Immunol Cell Biol 2016, 94, 151–157. [Google Scholar] [CrossRef] [PubMed]
- Maslanik, T.; Mahaffey, L.; Tannura, K.; Beninson, L.; Greenwood, B.N.; Fleshner, M. The inflammasome and danger associated molecular patterns (DAMPs) are implicated in cytokine and chemokine responses following stressor exposure. Brain Behav Immun 2013, 28, 54–62. [Google Scholar] [CrossRef] [PubMed]
- Krauth, S.J.; Coulibaly, J.T.; Knopp, S.; Traore, M.; N'Goran, E.K.; Utzinger, J. An in-depth analysis of a piece of shit: distribution of Schistosoma mansoni and hookworm eggs in human stool. PLoS Negl Trop Dis 2012, 6, e1969. [Google Scholar] [CrossRef] [PubMed]
- Apprill, A.; McNally, S.; Parsons, R.; Weber, L. Minor revision to V4 region SSU rRNA 806R gene primer greatly increases detection of SAR11 bacterioplankton. Aquat Microb Ecol 2015, 75, 129–137. [Google Scholar] [CrossRef]
- Caporaso, J.G.; Lauber, C.L.; Walters, W.A.; Berg-Lyons, D.; Huntley, J.; Fierer, N.; Owens, S.M.; Betley, J.; Fraser, L.; Bauer, M.; et al. Ultra-high-throughput microbial community analysis on the Illumina HiSeq and MiSeq platforms. ISME J 2012, 6, 1621–1624. [Google Scholar] [CrossRef]
- Bolyen, E.; Rideout, J.R.; Dillon, M.R.; Bokulich, N.A.; Abnet, C.C.; Al-Ghalith, G.A.; Alexander, H.; Alm, E.J.; Arumugam, M.; Asnicar, F.; et al. Author Correction: Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat Biotechnol 2019, 37, 1091. [Google Scholar] [CrossRef]
- Bolyen, E.; Rideout, J.R.; Dillon, M.R.; Bokulich, N.A.; Abnet, C.C.; Al-Ghalith, G.A.; Alexander, H.; Alm, E.J.; Arumugam, M.; Asnicar, F.; et al. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat Biotechnol 2019, 37, 852–857. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez, A.; Navas-Molina, J.A.; Kosciolek, T.; McDonald, D.; Vazquez-Baeza, Y.; Ackermann, G.; DeReus, J.; Janssen, S.; Swafford, A.D.; Orchanian, S.B.; et al. Qiita: rapid, web-enabled microbiome meta-analysis. Nat Methods 2018, 15, 796–798. [Google Scholar] [CrossRef] [PubMed]
- Amir, A.; McDonald, D.; Navas-Molina, J.A.; Kopylova, E.; Morton, J.T.; Zech Xu, Z.; Kightley, E.P.; Thompson, L.R.; Hyde, E.R.; Gonzalez, A.; et al. Deblur Rapidly Resolves Single-Nucleotide Community Sequence Patterns. mSystems 2017, 2. [Google Scholar] [CrossRef] [PubMed]
- Janssen, S.; McDonald, D.; Gonzalez, A.; Navas-Molina, J.A.; Jiang, L.; Xu, Z.Z.; Winker, K.; Kado, D.M.; Orwoll, E.; Manary, M.; et al. Phylogenetic Placement of Exact Amplicon Sequences Improves Associations with Clinical Information. mSystems 2018, 3. [Google Scholar] [CrossRef] [PubMed]
- Bokulich, N.A.; Kaehler, B.D.; Rideout, J.R.; Dillon, M.; Bolyen, E.; Knight, R.; Huttley, G.A.; Gregory Caporaso, J. Optimizing taxonomic classification of marker-gene amplicon sequences with QIIME 2's q2-feature-classifier plugin. Microbiome 2018, 6, 90. [Google Scholar] [CrossRef] [PubMed]
- Yilmaz, P.; Parfrey, L.W.; Yarza, P.; Gerken, J.; Pruesse, E.; Quast, C.; Schweer, T.; Peplies, J.; Ludwig, W.; Glockner, F.O. The SILVA and "All-species Living Tree Project (LTP)" taxonomic frameworks. Nucleic Acids Res 2014, 42, D643–D648. [Google Scholar] [CrossRef] [PubMed]
- Knights, D.; Kuczynski, J.; Koren, O.; Ley, R.E.; Field, D.; Knight, R.; DeSantis, T.Z.; Kelley, S.T. Supervised classification of microbiota mitigates mislabeling errors. Isme J 2011, 5, 570–573. [Google Scholar] [CrossRef] [PubMed]
- Human Microbiome Project, C. Structure, function and diversity of the healthy human microbiome. Nature 2012, 486, 207–214. [Google Scholar] [CrossRef] [PubMed]
- Lozupone, C.; Lladser, M.E.; Knights, D.; Stombaugh, J.; Knight, R. UniFrac: an effective distance metric for microbial community comparison. ISME J 2011, 5, 169–172. [Google Scholar] [CrossRef]
- Faith, D.P. Phylogenetic pattern and the quantification of organismal biodiversity. Philos Trans R Soc Lond B Biol Sci 1994, 345, 45–58. [Google Scholar] [CrossRef]
- Hagerty, S.L.; Hutchison, K.E.; Lowry, C.A.; Bryan, A.D. An empirically derived method for measuring human gut microbiome alpha diversity: Demonstrated utility in predicting health-related outcomes among a human clinical sample. PLoS One 2020, 15, e0229204. [Google Scholar] [CrossRef] [PubMed]
- Mandal, S.; Van Treuren, W.; White, R.A.; Eggesbo, M.; Knight, R.; Peddada, S.D. Analysis of composition of microbiomes: a novel method for studying microbial composition. Microb Ecol Health Dis 2015, 26, 27663. [Google Scholar] [CrossRef] [PubMed]
- Callahan, B.J.; McMurdie, P.J.; Holmes, S.P. Exact sequence variants should replace operational taxonomic units in marker-gene data analysis. Isme J 2017, 11, 2639–2643. [Google Scholar] [CrossRef] [PubMed]
- Quast, C.; Pruesse, E.; Yilmaz, P.; Gerken, J.; Schweer, T.; Yarza, P.; Peplies, J.; Glockner, F.O. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res 2013, 41, D590–D596. [Google Scholar] [CrossRef]
- Douglas, G.M.; Maffei, V.J.; Zaneveld, J.R.; Yurgel, S.N.; Brown, J.R.; Taylor, C.M.; Huttenhower, C.; Langille, M.G.I. PICRUSt2 for prediction of metagenome functions. Nat Biotechnol 2020, 38, 685–688. [Google Scholar] [CrossRef]
- Melnik, A.V.; da Silva, R.R.; Hyde, E.R.; Aksenov, A.A.; Vargas, F.; Bouslimani, A.; Protsyuk, I.; Jarmusch, A.K.; Tripathi, A.; Alexandrov, T.; et al. Coupling Targeted and Untargeted Mass Spectrometry for Metabolome-Microbiome-Wide Association Studies of Human Fecal Samples. Anal Chem 2017, 89, 7549–7559. [Google Scholar] [CrossRef] [PubMed]
- Kelly, B.J.; Gross, R.; Bittinger, K.; Sherrill-Mix, S.; Lewis, J.D.; Collman, R.G.; Bushman, F.D.; Li, H. Power and sample-size estimation for microbiome studies using pairwise distances and PERMANOVA. Bioinformatics 2015, 31, 2461–2468. [Google Scholar] [CrossRef]
- Tang, Z.Z.; Chen, G.; Alekseyenko, A.V. PERMANOVA-S: association test for microbial community composition that accommodates confounders and multiple distances. Bioinformatics 2016, 32, 2618–2625. [Google Scholar] [CrossRef] [PubMed]
- Lozupone, C.; Knight, R. UniFrac: a new phylogenetic method for comparing microbial communities. Appl Environ Microbiol 2005, 71, 8228–8235. [Google Scholar] [CrossRef]
- Lopez-Almela, I.; Romani-Perez, M.; Bullich-Vilarrubias, C.; Benitez-Paez, A.; Gomez Del Pulgar, E.M.; Frances, R.; Liebisch, G.; Sanz, Y. Bacteroides uniformis combined with fiber amplifies metabolic and immune benefits in obese mice. Gut Microbes 2021, 13, 1–20. [Google Scholar] [CrossRef]
- Morita, H.; Kano, C.; Ishii, C.; Kagata, N.; Ishikawa, T.; Hirayama, A.; Uchiyama, Y.; Hara, S.; Nakamura, T.; Fukuda, S. Bacteroides uniformis and its preferred substrate, alpha-cyclodextrin, enhance endurance exercise performance in mice and human males. Sci Adv 2023, 9, eadd2120. [Google Scholar] [CrossRef] [PubMed]
- Carbajo-Pescador, S.; Porras, D.; Garcia-Mediavilla, M.V.; Martinez-Florez, S.; Juarez-Fernandez, M.; Cuevas, M.J.; Mauriz, J.L.; Gonzalez-Gallego, J.; Nistal, E.; Sanchez-Campos, S. Beneficial effects of exercise on gut microbiota functionality and barrier integrity, and gut-liver crosstalk in an in vivo model of early obesity and non-alcoholic fatty liver disease. Dis Model Mech 2019, 12. [Google Scholar] [CrossRef] [PubMed]
- Ezeji, J.C.; Sarikonda, D.K.; Hopperton, A.; Erkkila, H.L.; Cohen, D.E.; Martinez, S.P.; Cominelli, F.; Kuwahara, T.; Dichosa, A.E.K.; Good, C.E.; et al. Parabacteroides distasonis: intriguing aerotolerant gut anaerobe with emerging antimicrobial resistance and pathogenic and probiotic roles in human health. Gut Microbes 2021, 13, 1922241. [Google Scholar] [CrossRef] [PubMed]
- Berding, K.; Long-Smith, C.M.; Carbia, C.; Bastiaanssen, T.F.S.; van de Wouw, M.; Wiley, N.; Strain, C.R.; Fouhy, F.; Stanton, C.; Cryan, J.F.; et al. A specific dietary fibre supplementation improves cognitive performance-an exploratory randomised, placebo-controlled, crossover study. Psychopharmacology (Berl) 2021, 238, 149–163. [Google Scholar] [CrossRef] [PubMed]
- Tang, R.; Jiang, Y.; Tan, A.; Ye, J.; Xian, X.; Xie, Y.; Wang, Q.; Yao, Z.; Mo, Z. 16S rRNA gene sequencing reveals altered composition of gut microbiota in individuals with kidney stones. Urolithiasis 2018, 46, 503–514. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Huang, C.; Wang, J.; Zhou, H.; Lu, Y.; Lou, L.; Zheng, J.; Tian, L.; Wang, X.; Cao, Z.; et al. Dysbiosis of intestinal microbiota and decrease in paneth cell antimicrobial peptide level during acute necrotizing pancreatitis in rats. PLoS One 2017, 12, e0176583. [Google Scholar] [CrossRef]
- Verheggen, R.; Konstanti, P.; Smidt, H.; Hermus, A.; Thijssen, D.H.J.; Hopman, M.T.E. Eight-week exercise training in humans with obesity: Marked improvements in insulin sensitivity and modest changes in gut microbiome. Obesity (Silver Spring) 2021, 29, 1615–1624. [Google Scholar] [CrossRef]
- Toya, T.; Corban, M.T.; Marrietta, E.; Horwath, I.E.; Lerman, L.O.; Murray, J.A.; Lerman, A. Coronary artery disease is associated with an altered gut microbiome composition. PLoS One 2020, 15, e0227147. [Google Scholar] [CrossRef] [PubMed]
- Babacan Yildiz, G.; Kayacan, Z.C.; Karacan, I.; Sumbul, B.; Elibol, B.; Gelisin, O.; Akgul, O. Altered gut microbiota in patients with idiopathic Parkinson's disease: an age-sex matched case-control study. Acta Neurol Belg 2023, 123, 999–1009. [Google Scholar] [CrossRef]
- Yildirim, E.; Ilina, L.; Laptev, G.; Filippova, V.; Brazhnik, E.; Dunyashev, T.; Dubrovin, A.; Novikova, N.; Tiurina, D.; Tarlavin, N.; et al. The structure and functional profile of ruminal microbiota in young and adult reindeers (Rangifer tarandus) consuming natural winter-spring and summer-autumn seasonal diets. PeerJ 2021, 9, e12389. [Google Scholar] [CrossRef]
- Ma, L.; Ni, Y.; Wang, Z.; Tu, W.; Ni, L.; Zhuge, F.; Zheng, A.; Hu, L.; Zhao, Y.; Zheng, L.; et al. Spermidine improves gut barrier integrity and gut microbiota function in diet-induced obese mice. Gut Microbes 2020, 12, 1–19. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.; Wang, T.; Guo, R.; Cui, W.; Yu, W.; Wang, Z.; Jiang, Y.; Jiang, M.; Wang, X.; Liu, C.; et al. Variation of Serum Uric Acid Is Associated With Gut Microbiota in Patients With Diabetes Mellitus. Front Cell Infect Microbiol 2021, 11, 761757. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Zhang, X.; Tang, G.; Deng, P.; Qin, Y.; Han, J.; Wang, S.; Sun, X.; Li, D.; Chen, Z. The causal relationship between gut microbiota and bone mineral density: a Mendelian randomization study. Front Microbiol 2023, 14, 1268935. [Google Scholar] [CrossRef] [PubMed]
- Song, Y.; Shen, H.; Liu, T.; Pan, B.; De Alwis, S.; Zhang, W.; Luo, X.; Li, Z.; Wang, N.; Ma, W.; et al. Effects of three different mannans on obesity and gut microbiota in high-fat diet-fed C57BL/6J mice. Food Funct 2021, 12, 4606–4620. [Google Scholar] [CrossRef] [PubMed]
- Wang, K.; Qi, L.; Zhao, L.; Liu, J.; Guo, Y.; Zhang, C. Degradation of chondroitin sulfate: Mechanism of degradation, influence factors, structure-bioactivity relationship and application. Carbohydr Polym 2023, 301, 120361. [Google Scholar] [CrossRef] [PubMed]
- Liu, F.; Zhang, N.; Li, Z.; Wang, X.; Shi, H.; Xue, C.; Li, R.W.; Tang, Q. Chondroitin sulfate disaccharides modified the structure and function of the murine gut microbiome under healthy and stressed conditions. Sci Rep 2017, 7, 6783. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Gong, Z.; Zhang, X.; Zhu, F.; Liu, Y.; Jin, C.; Du, X.; Xu, C.; Chen, Y.; Cai, W.; et al. Gut microbial bile acid metabolite skews macrophage polarization and contributes to high-fat diet-induced colonic inflammation. Gut Microbes 2020, 12, 1–20. [Google Scholar] [CrossRef] [PubMed]
- Duboc, H.; Tache, Y.; Hofmann, A.F. The bile acid TGR5 membrane receptor: from basic research to clinical application. Dig Liver Dis 2014, 46, 302–312. [Google Scholar] [CrossRef]
- Pols, T.W.; Noriega, L.G.; Nomura, M.; Auwerx, J.; Schoonjans, K. The bile acid membrane receptor TGR5: a valuable metabolic target. Dig Dis 2011, 29, 37–44. [Google Scholar] [CrossRef]
| PERMANOVAs (pseudo-F) | ||||
|---|---|---|---|---|
| Northwestern | ||||
| 0 | 28 | 42 | 51 | |
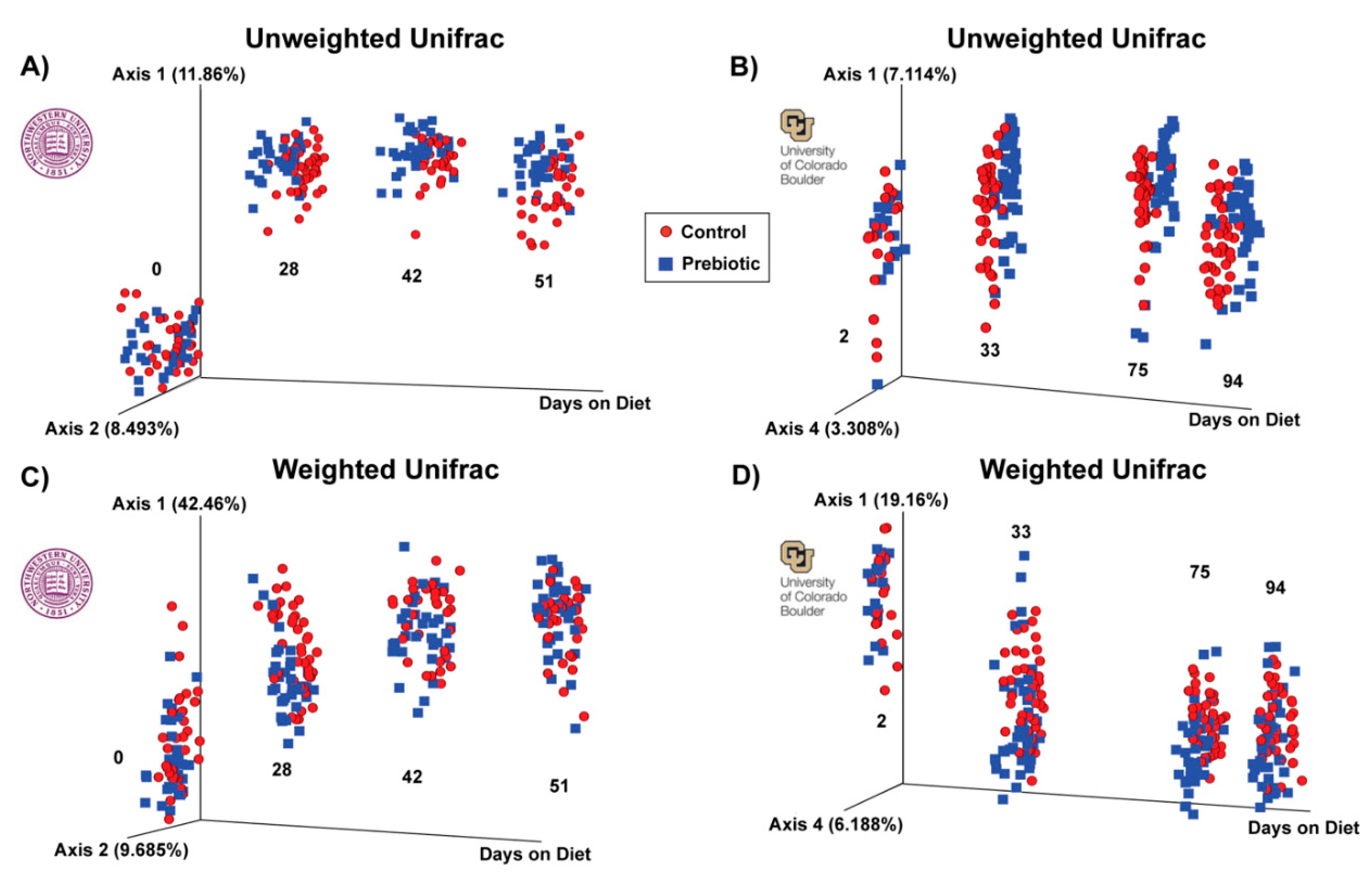
| Unweighted | F(2,68) = 1.24; p = 0.154 | F(2,80) = 7.68; p = 0.001 | F(2,66) = 5.60; p = 0.001 | F(2,69) = 5.87; p = 0.001 |
| Weighted | F(2,68) = 2.19; p = 0.053 | F(2,80) = 9.31; p = 0.001 | F(2,66) = 4.26; p = 0.001 | F(2,66) = 4.34; p = 0.001 |
| University of Colorado Boulder | ||||
| 2 | 33 | 75 | 94 | |
| Unweighted | F(2,48) = 1.31; p = 0.053 | F(2,78) = 4.89; p = 0.001 | F(2,83) = 3.84; p = 0.001 | F(2,84) = 4.16; p = 0.001 |
| Weighted | F(2,48) = 3.97; p = 0.006 | F(2,78) = 10.99; p = 0.001 | F(2,83) = 7.39; p = 0.001 | F(2,84) = 3.93; p = 0.001 |
 |
 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).




