Submitted:
03 May 2024
Posted:
07 May 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
3. Results
4. Discussion
5. Conclusions
Ethics
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Arnaout, Y.; Djelouadji, Z.; Robardet, E.; Cappelle, J.; Cliquet, F.; Touzalin, F.; Jimenez, G.; Hurstel, S.; Borel, C.; Picard-Meyer, E. Genetic Identification of Bat Species for Pathogen Surveillance across France. PLoS One 2022, 17, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Schede mammiferi https://www.mammiferi.org/specie/.
- Ruffo S., S. F. Checklist e Distribuzione Della Fauna Italiana. 10.000 Specie Terrestri e Delle Acque Interne; Agnoli G.L. & Rosa P., Ed.; Verona, 2022.
- Colombi, D.; Serra-Cobo, J.; Métras, R.; Apolloni, A.; Poletto, C.; López-Roig, M.; Bourhy, H.; Colizza, V. Mechanisms for Lyssavirus Persistence in Non-Synanthropic Bats in Europe: Insights from a Modeling Study. Sci. Rep. 2019, 9, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Banyard, A. C.; Evans, J. S.; Luo, T. R.; Fooks, A. R. Lyssaviruses and Bats: Emergence and Zoonotic Threat. Viruses 2014, 6, 2974–2990. [Google Scholar] [CrossRef] [PubMed]
- Černe, D.; Hostnik, P.; Toplak, I.; Presetnik, P.; Maurer-Wernig, J.; Kuhar, U. Discovery of a Novel Bat Lyssavirus in a Long-Fingered Bat (Myotis Capaccinii) from Slovenia. PLoS Negl. Trop. Dis. 2023, 17, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Leopardi, S.; Priori, P.; Zecchin, B.; Poglayen, G.; Trevisiol, K.; Lelli, D.; Zoppi, S.; Scicluna, M. T.; D’Avino, N.; Schiavon, E.; et al. Active and Passive Surveillance for Bat Lyssaviruses in Italy Revealed Serological Evidence for Their Circulation in Three Bat Species. Epidemiol. Infect. 2019, 147. [Google Scholar] [CrossRef] [PubMed]
- Shipley, R.; Wright, E.; Selden, D.; Wu, G.; Aegerter, J.; Fooks, A. R.; Banyard, A. C. Bats and Viruses: Emergence of Novel Lyssaviruses and Association of Bats with Viral Zoonoses in the EU. Trop. Med. Infect. Dis. 2019, 4, 1–22. [Google Scholar] [CrossRef] [PubMed]
- Schatz, J.; Fooks, A. R.; Mcelhinney, L.; Horton, D.; Echevarria, J.; Vázquez-Moron, S.; Kooi, E. A.; Rasmussen, T. B.; Müller, T.; Freuling, C. M. Bat Rabies Surveillance in Europe. Zoonoses Public Health 2013, 60, 22–34. [Google Scholar] [CrossRef] [PubMed]
- Mühldorfer, K.; Speck, S.; Kurth, A.; Lesnik, R.; Freuling, C.; Müller, T.; Kramer-Schadt, S.; Wibbelt, G. Diseases and Causes of Death in European Bats: Dynamics in Disease Susceptibility and Infection Rates. PLoS One 2011, 6(12). [Google Scholar] [CrossRef] [PubMed]
- Letko, M.; Seifert, S. N.; Olival, K. J.; Plowright, R. K.; Munster, V. J. Bat-Borne Virus Diversity, Spillover and Emergence. Nat. Rev. Microbiol. 2020, 18, 461–471. [Google Scholar] [CrossRef] [PubMed]
- Zhou, P.; Yang, X. Lou; Wang, X. G.; Hu, B.; Zhang, L.; Zhang, W.; Si, H. R.; Zhu, Y.; Li, B.; Huang, C. L.; et al. A Pneumonia Outbreak Associated with a New Coronavirus of Probable Bat Origin. Nature 2020, 579, 270–273. [Google Scholar] [CrossRef] [PubMed]
- Banerjee, A.; Baker, M. L.; Kulcsar, K.; Misra, V.; Plowright, R.; Mossman, K. Novel Insights Into Immune Systems of Bats. Front. Immunol. 2020, 11, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Festival, C. B.; Vora, N. M.; Osinubi, M. O. V; Davis, L.; Abdurrahman, M.; Adedire, E. B.; Akpan, H.; Aman-oloniyo, A. F.; Audu, S. W.; Blau, D.; et al. Bat and Lyssavirus Exposure among Humans in Area That. Emerg. Infect. Dis. 2020, 26, 1399–1408. [Google Scholar]
- D.P.R. 8 Settembre 1997, n. 357. 1997.
- Jian Yang. DATABASE OF BAT-ASSOCIATED VIRUSES http://www.mgc.ac.cn/cgi-bin/DBatVir/main.cgi?func=update&update=2023-03.
- Pozo, F.; Juste, J.; Vázquez-Morón, S.; Aznar-López, C.; Ibáñez, C.; Garin, I.; Aihartza, J.; Casas, I.; Tenorio, A.; Echevarría, J. E. Identification of Novel Betaherpesviruses in Iberian Bats Reveals Parallel Evolution. PLoS One 2016, 11, 1–17. [Google Scholar] [CrossRef]
- Baldwin, C. C.; Mounts, J. H.; Smith, D. G.; Weigt, L. A. Genetic Identification and Color Descriptions of Early Life-History Stages of Belizean Phaeoptyx and Astrapogon (Teleostei: Apogonidae) with Comments on Identification of Adult Phaeoptyx. Zootaxa 2009, 22, 1–22. [Google Scholar] [CrossRef]
- Wadhwa, A.; Wilkins, K.; Gao, J.; Condori Condori, R. E.; Gigante, C. M.; Zhao, H.; Ma, X.; Ellison, J. A.; Greenberg, L.; Velasco-Villa, A.; Orciari, L.; Li, Y. A Pan-Lyssavirus Taqman Real-Time RT-PCR Assay for the Detection of Highly Variable Rabies Virus and Other Lyssaviruses. PLoS Negl. Trop. Dis. 2017, 11, 1–17. [Google Scholar] [CrossRef] [PubMed]
- Leopardi, S.; Barneschi, E.; Manna, G.; Zecchin, B.; Priori, P.; Drzewnioková, P.; Festa, F.; Lombardo, A.; Parca, F.; Scaravelli, D.; Ponti, A. M.; De Benedictis, P. Spillover of West Caucasian Bat Lyssavirus (Wcbv) in a Domestic Cat and Westward Expansion in the Palearctic Region. Viruses 2021, 13, 1–17. [Google Scholar] [CrossRef] [PubMed]
- De Benedictis, P.; Leopardi, S.; Markotter, W.; Velasco-Villa, A. The Importance of Accurate Host Species Identification in the Framework of Rabies Surveillance, Control and Elimination. Viruses 2022, 14, 1–11. [Google Scholar] [CrossRef]
- Vandevanter, D. R.; Warrener, P.; Bennett, L.; Schultz, E. R.; Coulter, S.; Garber, R. L.; Rose, T. M. Detection and Analysis of Diverse Herpesviral Species by Consensus Primer PCR. J. Clin. Microbiol. 1996, 34, 1666–1671. [Google Scholar] [CrossRef] [PubMed]
- Ehlers, B.; Borchers, K.; Grund, C.; Frölich, K.; Ludwig, H.; Buhk, H. J. Detection of New DNA Polymerase Genes of Known and Potentially Novel Herpesviruses by PCR with Degenerate and Deoxyinosine-Substituted Primers. Virus Genes 1999, 18, 211–220. [Google Scholar] [CrossRef]
- Wilkinson, D. A.; Joffrin, L.; Lebarbenchon, C.; Mavingui, P. Analysis of Partial Sequences of the RNA-Dependent RNA Polymerase Gene as a Tool for Genus and Subgenus Classification of Coronaviruses. J. Gen. Virol. 2021, 101, 1261–1269. [Google Scholar] [CrossRef] [PubMed]
- Madeira, F.; Pearce, M.; Tivey, A. R. N.; Basutkar, P.; Lee, J.; Edbali, O.; Madhusoodanan, N.; Kolesnikov, A.; Lopez, R. Search and Sequence Analysis Tools Services from EMBL-EBI in 2022. Nucleic Acids Res. 2022, 50, W276–W279. [Google Scholar] [CrossRef] [PubMed]
- Edgar, R. C. MUSCLE: Multiple Sequence Alignment with High Accuracy and High Throughput. Nucleic Acids Res. 2004, 32, 1792–1797. [Google Scholar] [CrossRef] [PubMed]
- Darriba, D.; Taboada, G. L.; Doallo, R.; Posada, D. ProtTest 3: Fast Selection of Best-Fit Models of Protein Evolution. Bioinformatics 2011, 27, 1164–1165. [Google Scholar] [CrossRef]
- Darriba, D.; Taboada, G. L.; Doallo, R.; Posada, D. JModelTest 2: More Models, New Heuristics and Parallel Computing. Nat. Methods 2012, 9(8), 772. [Google Scholar] [CrossRef] [PubMed]
- Guindon, S.; Gascuel, O. A Simple, Fast, and Accurate Algorithm to Estimate Large Phylogenies by Maximum Likelihood. Syst. Biol. 2003, 52(5), 696–704. [Google Scholar] [CrossRef]
- 3.6.3, R. D. C. T. A Language and Environment for Statistical Computing. R Found. Stat. Comput. 2020, 1, https://www.R-project.org.
- Guangchuang, Yu. Data Integration, Manipulation and Visualization of Phylogenetic Trees, 1st ed.; Chapman and Hall/CRC, Ed.; Taylor & Francis: New York, 2022. [Google Scholar] [CrossRef]
- Yu, G. Using Ggtree to Visualize Data on Tree-Like Structures. Curr. Protoc. Bioinforma. 2020, 69, 1–18. [Google Scholar] [CrossRef] [PubMed]
- Paradis, E.; Schliep, K. Ape 5.0: An Environment for Modern Phylogenetics and Evolutionary Analyses in R. Bioinformatics 2019, 35(3), 526–528. [Google Scholar] [CrossRef] [PubMed]
- Leopardi, S.; Desiato, R.; Mazzucato, M.; Orusa, R.; Obber, F.; Averaimo, D.; Berjaoui, S.; Canziani, S.; Capucchio, M. T.; Conti, R.; Di Bella, S.; Festa, F.; Garofalo, L.; Lelli, D.; Madrau, M. P.; Mandola, M. L.; Martin, A. M. M.; Peletto, S.; Pirani, S.; Robetto, S.; Torresi, C.; Varotto, M.; Citterio, C.; Terregino, C. Erratum: One Health Surveillance Strategy for Coronaviruses in Italian Wildlife (Epidemiology & Infection 151 (E96) DOI: 10.1017/S095026882300081X). Epidemiol. Infect. 2023, 151. https://doi.org/10.1017/S0950268823001000.
- Woo, P. C. Y.; Lau, S. K. P.; Li, K. S. M.; Poon, R. W. S.; Wong, B. H. L.; Tsoi, H. wah; Yip, B. C. K.; Huang, Y.; Chan, K. hung; Yuen, K. yung. Molecular Diversity of Coronaviruses in Bats. Virology 2006, 351, 180–187. [Google Scholar] [CrossRef] [PubMed]
- Chu, D. K. W.; Peiris, J. S. M.; Chen, H.; Guan, Y.; Poon, L. L. M. Genomic Characterizations of Bat Coronaviruses (1A, 1B and HKU8) and Evidence for Co-Infections in Miniopterus Bats. J. Gen. Virol. 2008, 89(5), 1282–1287. [Google Scholar] [CrossRef] [PubMed]
- Kohl, C.; Nitsche, A.; Kurth, A. Update on Potentially Zoonotic Viruses of European Bats. Vaccines 2021, 9(7). [Google Scholar] [CrossRef] [PubMed]
- Drexler, J. F.; Gloza-Rausch, F.; Glende, J.; Corman, V. M.; Muth, D.; Goettsche, M.; Seebens, A.; Niedrig, M.; Pfefferle, S.; Yordanov, S.; Zhelyazkov, L.; Hermanns, U.; Vallo, P.; Lukashev, A.; Müller, M. A.; Deng, H.; Herrler, G.; Drosten, C. Genomic Characterization of Severe Acute Respiratory Syndrome-Related Coronavirus in European Bats and Classification of Coronaviruses Based on Partial RNA-Dependent RNA Polymerase Gene Sequences. J. Virol. 2010, 84(21), 11336–11349. [Google Scholar] [CrossRef] [PubMed]
- Vasenkov, D.; Desmet, J. F.; Popov, I.; Sidorchuk, N. Bats Can Migrate Farther than It Was Previously Known: A New Longest Migration Record by Nathusius’ Pipistrelle Pipistrellus Nathusii (Chiroptera: Vespertilionidae). Mammalia 2022, 86(5), 524–526. [Google Scholar] [CrossRef]
- Cerri, A.; Bolatti, E. M.; Zorec, T. M.; Montani, M. E.; Rimondi, A.; Hosnjak, L.; Casal, P. E.; Domenica, V. Di; Barquez, R. M.; Poljak, M.; Giri, A. A. Coronavirus Cross-Species Transmission. 2023, 11 (5).
- Chan, J. F. W.; To, K. K. W.; Tse, H.; Jin, D. Y.; Yuen, K. Y. Interspecies Transmission and Emergence of Novel Viruses: Lessons from Bats and Birds. Trends Microbiol. 2013, 21, 544–555. [Google Scholar] [CrossRef]
- Bergner, L. M.; Mollentze, N.; Orton, R. J.; Tello, C.; Broos, A.; Biek, R.; Streicker, D. G. Characterizing and Evaluating the Zoonotic Potential of Novel Viruses Discovered in Vampire Bats. Viruses 2021, 13(2). [Google Scholar] [CrossRef] [PubMed]
- Printz, L.; Jung, K. Urban Areas in Rural Landscapes – the Importance of Green Space and Local Architecture for Bat Conservation. Front. Ecol. Evol. 2023, 11, 1–11. [Google Scholar] [CrossRef]
- Botvinkin, A. D.; Poleschuk, E. M.; Kuzmin, I. V.; Borisova, T. I.; Gazaryan, S. V.; Yager, P.; Rupprecht, C. E. Novel Lyssaviruses Isolated from Bats in Russia. Emerg. Infect. Dis. 2003, 9(12), 1623–1625. [Google Scholar] [CrossRef] [PubMed]


| Type of surveillance | Bat species | Total number per species (%) |
Bat species distribution per Region |
Virus family investigated |
||
|---|---|---|---|---|---|---|
| Latium | Tuscany | pan-CoV 21 | pan-Herpes 22,23 | |||
| Passive | Hypsugo savii | 18 (7.5) | 11 | 7 | 18 | 3+ |
| Pipistrellus kuhlii | 23 (9.6) | 9 | 14 | 23 | 3+ | |
| Pipistrellus Sp | 10 (4.2) | 1 | 9 | 10 | 0 | |
| Myotis daubentonii | 1 (0.4) | 0 | 1 | 1 | 0 | |
| Plecotus austriacus | 1 (0.4) | 0 | 1 | 1 | 0 | |
| Not identified | 4 (1.7) | 0 | 4 | 4 | 0 | |
| Tadarida teniotis | 71 (29.7) | 71 | 0 | 71 | 71+ | |
| Active | Miniopterus schreibersii | 111 (46.6) | 0 | 111* | 111 | 64 |
| Total | 239 (100) | 92 | 147 | 239 | 141 | |
| Nucleotide characteristics | Protein characteristics | ||||
|---|---|---|---|---|---|
| Accession number | identity | Querycov | Accession number | identity | Querycov |
| OP776451.1 | 78.08% | 75% | UZC49592.1 | 88.2% | 102/147 |
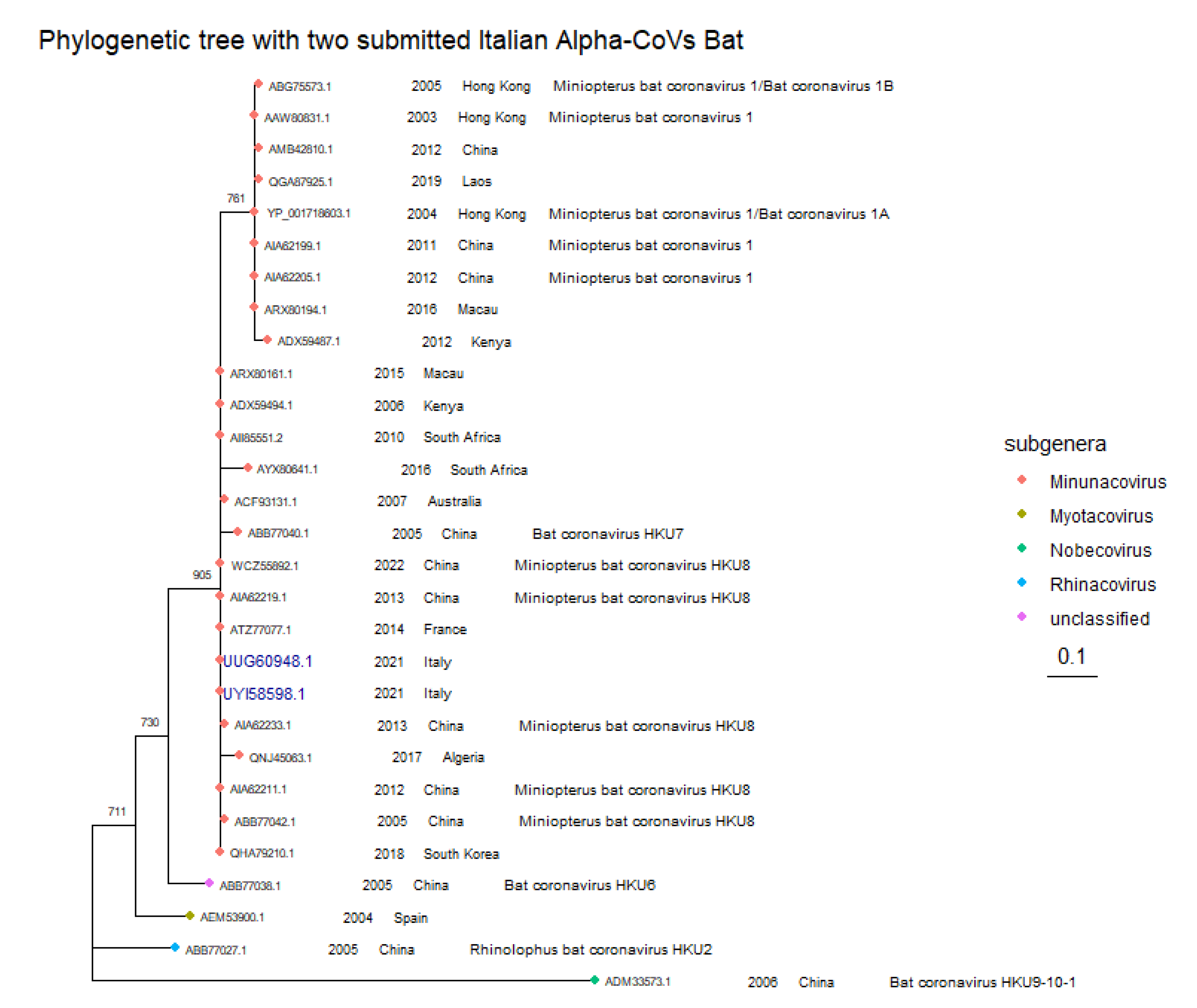
| OP627105.1 | 100% | 100% | UYI58598.1 | 100% | 147/147 |
| AN* | % identity | Similarity score | Starting position | Amino Acids |
|---|---|---|---|---|
| AMY98779.1 | 48 | 97/150 | INS 13 | VRET |
| INS 145 | VI | |||
| AMY98857.1 | 54.0 |
100/150 |
INS 13 | LRDS |
| INS 137 | VN | |||
| AMY98776.1 | 48.1 |
100/160 |
INS 13 | LREE |
| INS 22 | DD | |||
| INS 118 | HAHFVDPEFR | |||
| INS 131 | FGERELSR | |||
| AMY98771.1 | 58.0 |
111/150 |
INS 13 | VATD |
| INS 139 | ER | |||
| AMY98783.1 | 78.5 | 126/144 | / | / |
| AMY98784.1 | 100 | 144/144 | / | / |
| AMY98785.1 | 89.6 | 137/144 | / | / |
| AMY98774.1 | 47.8 | 101/159 | INS 5 | FPENVDM |
| DEL 10 | GPG | |||
| INS 22 | AD | |||
| DEL 121 | PR | |||
| DEL 126 | LS | |||
| INS 137 | VEYLPG |
| Protein AN* | Virus Name | %Identity | Similarity score | Starting position | Amino Acids | Corresponding nucleic AN* |
|---|---|---|---|---|---|---|
| CCE57225.1:640-788 | Murid betaherpesvirus 1 | 52.7 | 106/150 | INS 7 | EG | HE610455.1 |
| INS 138 | PEA | |||||
| AAK71288.1:10-157 | Baboon cytomegalovirus | 58.1 | 106/148 | INS 9 | G | AF387664.1 |
| INS 16 | QV | |||||
| INS138 | P | |||||
| AAW57296.1:328-477 | Murid betaherpesvirus 2 | 50.7 | 103/150 | DEL 7 | ND | AY728086.1 |
| INS 11 | PEVSR | |||||
| INS 128 | KVI | |||||
| YP_008492977.1:583-733 | Suid betaherpesvirus 2 | 53.0 | 100/151 | INS 10 | DVTGI | NC_022233.1 |
| INS 132 | LS | |||||
| YP_007969814.1:627-764 | Elephantid betaherpesvirus 1 | 43.2 | 88/148 | INS 13 | LREE | NC_020474.2 |
| DEL 117 | QNVLPRSDVL | |||||
| AAP57912.1:454-594 | Human betaherpesvirus 5 | 52.0 | 98/148 | INS 7 | PGGE | AY304055.1 |
| DEL 138 | DVALKVI |
| AN* | % identity | Similarity score | Starting position | Amino Acids |
|---|---|---|---|---|
| AFM85234.1 | 66.7 | 57/72 | / | / |
| AFM85236.1 | 69.4 | 63/72 | / | / |
| AMA67369.1 | 68.1 | 61/72 | / | / |
| ATA58242.1 | 59.0 | 59/78 | DEL 51 | KK |
| INS 60 | PHAPAA | |||
| ATU31554.1 | 57.5 | 52/73 | * | * |
| BBB06458.1 | 70.8 | 63/72 | / | / |
| ATU31556.1 | 64.4 | 61/73 | * | * |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




