Submitted:
19 April 2024
Posted:
22 April 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results
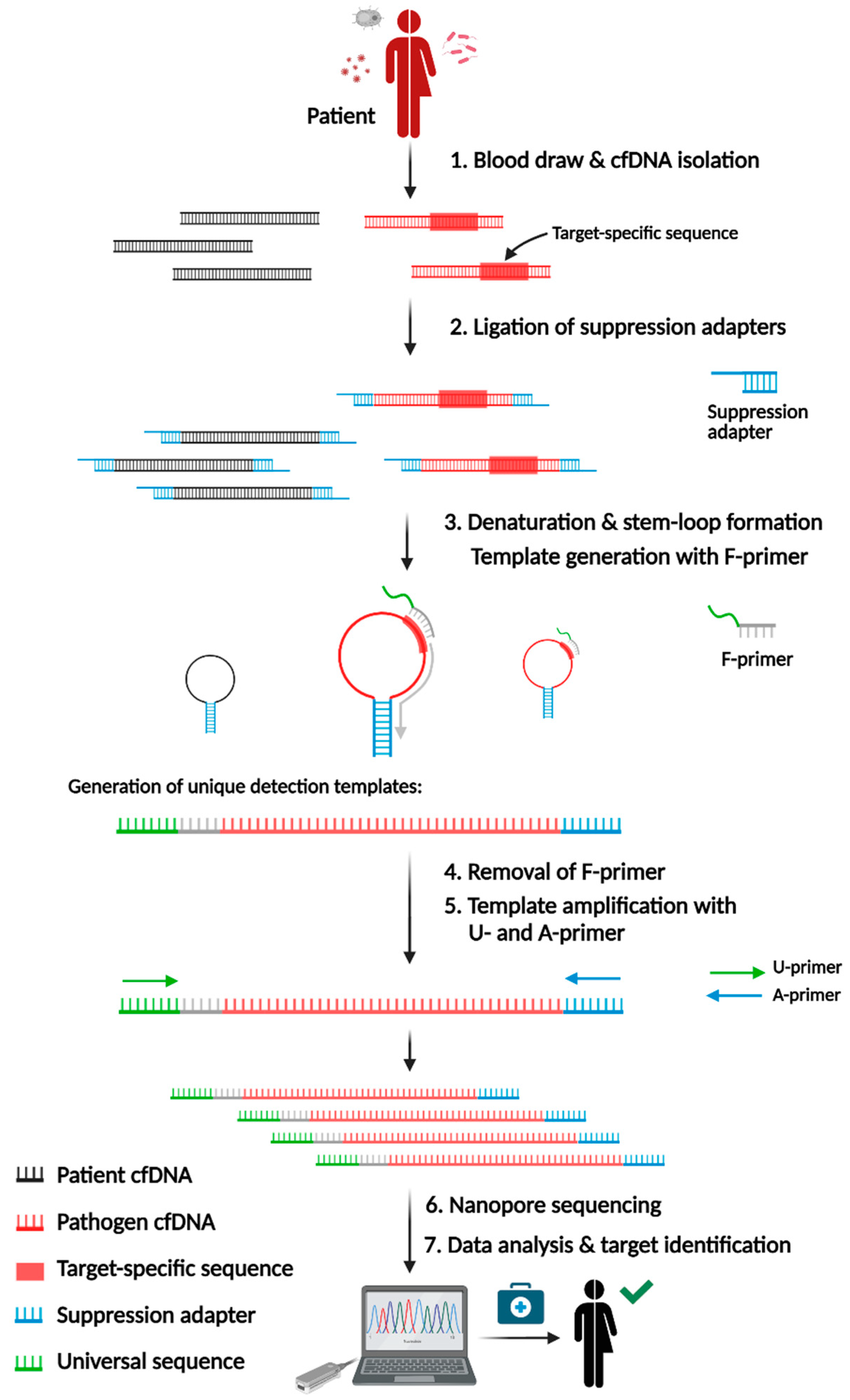
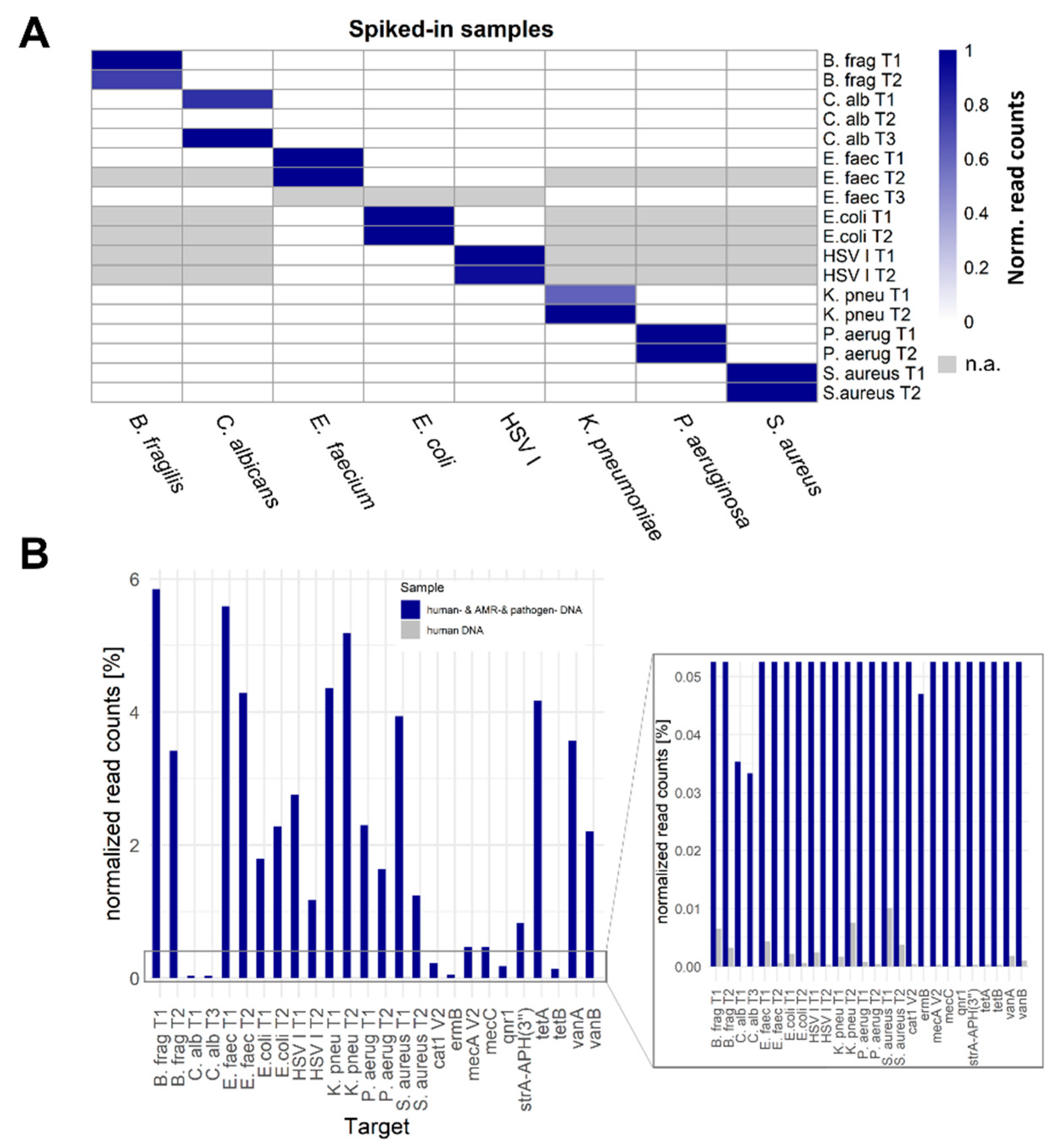
2.1. SUPSETS Procedure
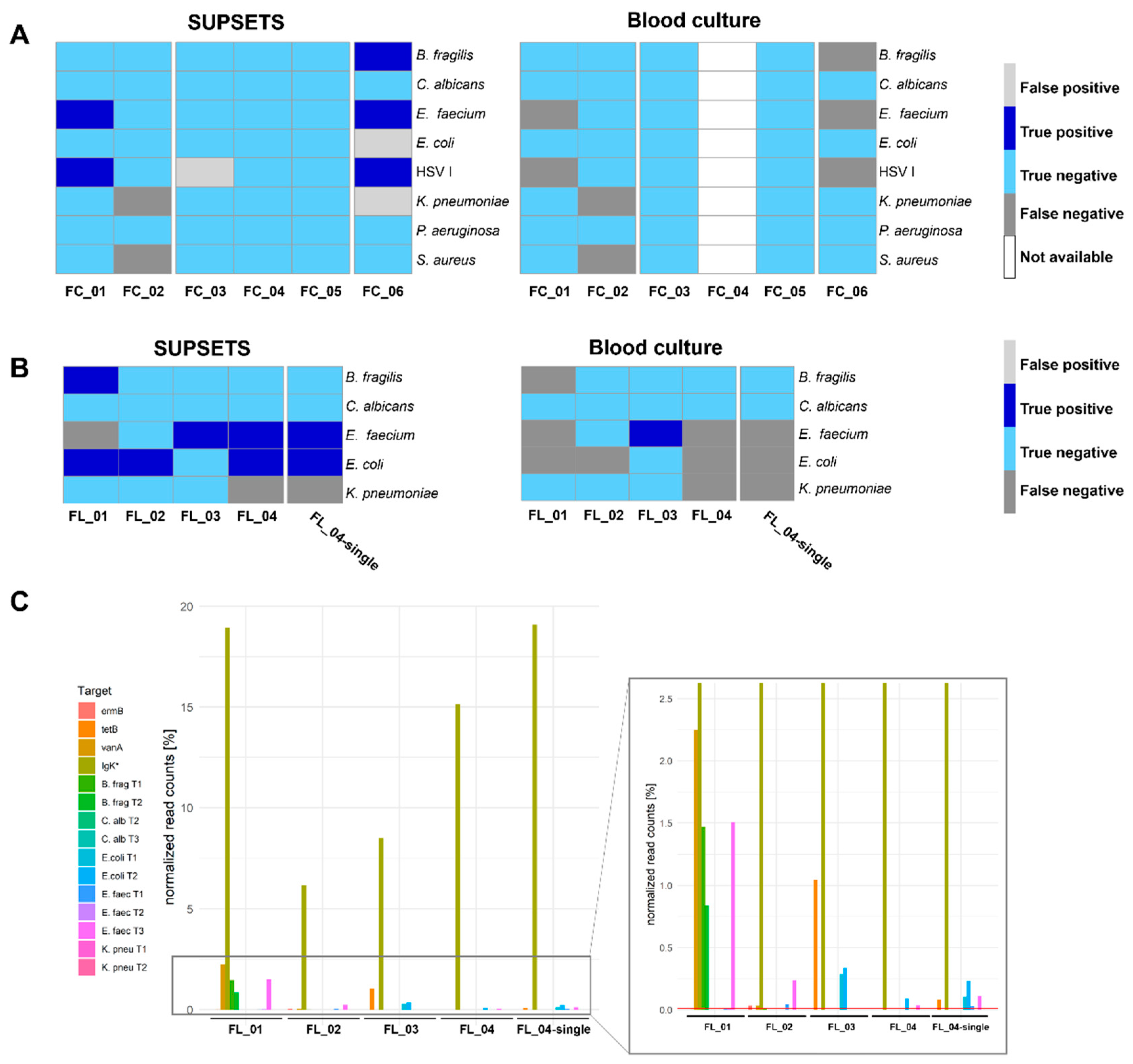
2.2. SUPSETS Validation on Selected Clinical Sepsis Samples
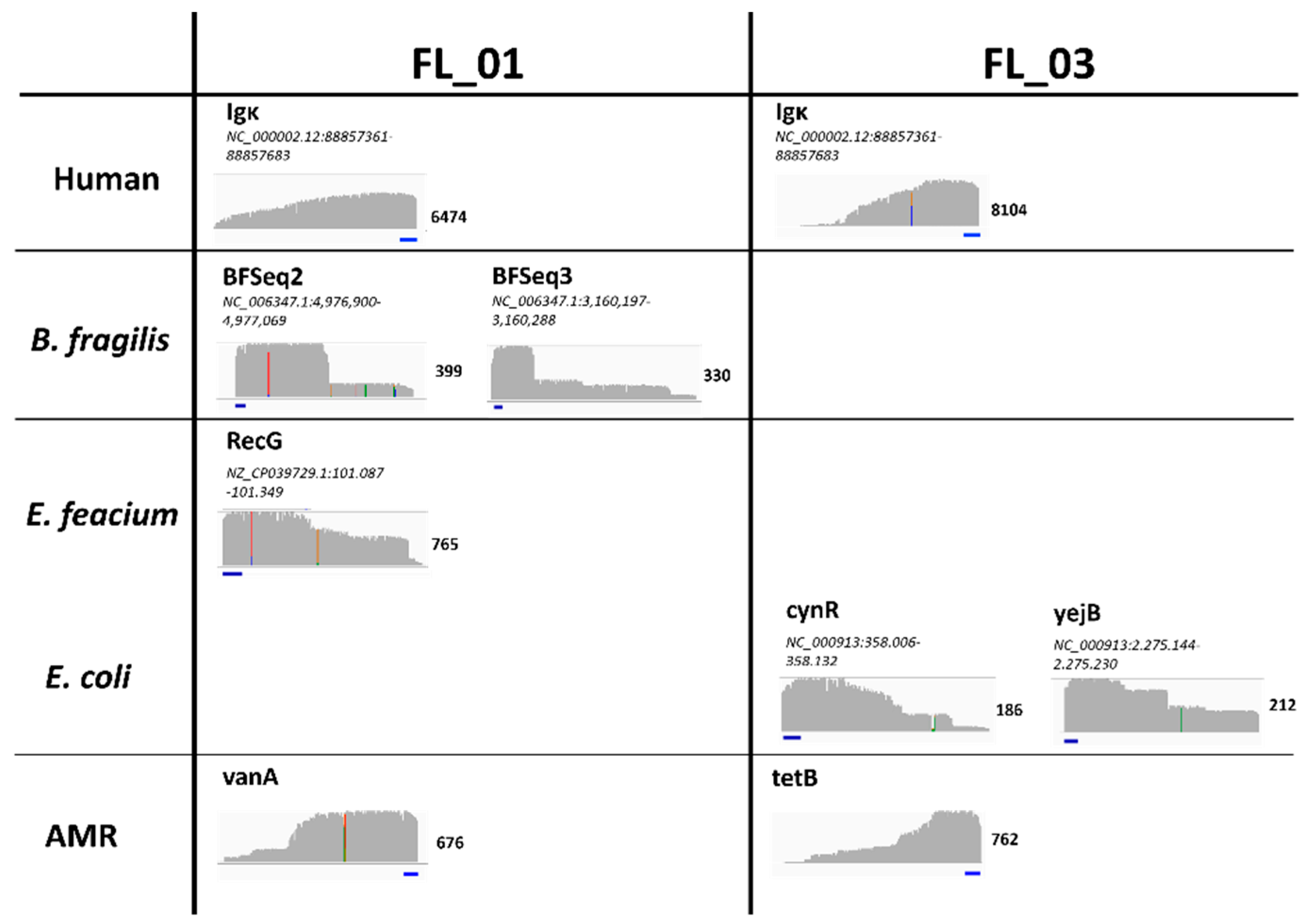
2.3. First Read Detection of SUPSETS within Selected Sepsis Samples
3. Discussion
4. Materials and Methods
4.1. Ethics Approval and Consent to Participate for Clinical Samples
4.2. Microbiology and Preparation of Microbial DNA
4.3. Identification of Species- or Genus-Specific Nucleotide Sequences and Primer Design
4.4. One Step PCR for Primer Testing
4.5. SUPSETS Workflow
4.6. Analysis of Sequencing Results
4.7. Clinical Sample Preparation
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Singer, M.; Deutschman, C.S.; Seymour, C.; Shankar-Hari, M.; Annane, D.; Bauer, M.; Bellomo, R.; Bernard, G.R.; Chiche, J.D.; Coopersmith, C.M.; et al. The Third International Consensus Definitions for Sepsis and Septic Shock (Sepsis-3). JAMA 2016, 315, 801. [Google Scholar] [CrossRef]
- Rudd, K.E.; Johnson, S.C.; Agesa, K.M.; Shackelford, K.A.; Tsoi, D.; Kievlan, D.R.; Colombara, D. V.; Ikuta, K.S.; Kissoon, N.; Finfer, S.; et al. Global, Regional, and National Sepsis Incidence and Mortality, 1990–2017: Analysis for the Global Burden of Disease Study. The Lancet 2020, 395, 200–211. [Google Scholar] [CrossRef] [PubMed]
- Kotfis, K.; Wittebole, X.; Jaschinski, U.; Solé-Violán, J.; Kashyap, R.; Leone, M.; Nanchal, R.; Fontes, L.E.; Sakr, Y.; Vincent, J.L. A Worldwide Perspective of Sepsis Epidemiology and Survival According to Age: Observational Data from the ICON Audit. J Crit Care 2019, 51, 122–132. [Google Scholar] [CrossRef] [PubMed]
- Camacho-Gonzalez, A.; Spearman, P.W.; Stoll, B.J. Neonatal Infectious Diseases. Evaluation of Neonatal Sepsis. Pediatr Clin North Am 2013, 60, 367–389. [Google Scholar] [CrossRef]
- Mayr, F.B.; Yende, S.; Angus, D.C. Epidemiology of Severe Sepsis. Virulence 2014, 5, 4–11. [Google Scholar] [CrossRef] [PubMed]
- Schmidt, K.F.R.; Huelle, K.; Reinhold, T.; Prescott, H.C.; Gehringer, R.; Hartmann, M.; Lehmann, T.; Mueller, F.; Reinhart, K.; Schneider, N.; et al. Healthcare Utilization and Costs in Sepsis Survivors in Germany–Secondary Analysis of a Prospective Cohort Study. J Clin Med 2022, 11, 1142. [Google Scholar] [CrossRef]
- van den Berg, M.; van Beuningen, F.E.; ter Maaten, J.C.; Bouma, H.R. Hospital-Related Costs of Sepsis around the World: A Systematic Review Exploring the Economic Burden of Sepsis. J Crit Care 2022, 71, 154096. [Google Scholar] [CrossRef]
- Rhee, C.; Klompas, M. Sepsis Trends: Increasing Incidence and Decreasing Mortality, or Changing Denominator? J Thorac Dis 2020, 12, S89. [Google Scholar] [CrossRef]
- Murray, C.J.; Ikuta, K.S.; Sharara, F.; Swetschinski, L.; Robles Aguilar, G.; Gray, A.; Han, C.; Bisignano, C.; Rao, P.; Wool, E.; et al. Global Burden of Bacterial Antimicrobial Resistance in 2019: A Systematic Analysis. Lancet 2022, 399, 629. [Google Scholar] [CrossRef]
- O’Neill, J. Tackling Drug-Resistant Infections Globally: Final Report And Recommendations. The Review On Antimicrobial Resistance 2016.
- Bloos, F.; Rüddel, H.; Thomas-Rüddel, D.; Schwarzkopf, D.; Pausch, C.; Harbarth, S.; Schreiber, T.; Gründling, M.; Marshall, J.; Simon, P.; et al. Effect of a Multifaceted Educational Intervention for Anti-Infectious Measures on Sepsis Mortality: A Cluster Randomized Trial. Intensive Care Med 2017, 43, 1602–1612. [Google Scholar] [CrossRef] [PubMed]
- Patel, M. Utility of Blood Culture in Sepsis Diagnostics. Journal of The Academy of Clinical Microbiologists 2016, 18, 74. [Google Scholar] [CrossRef]
- Rhodes, A.; Evans, L.E.; Alhazzani, W.; Levy, M.M.; Antonelli, M.; Ferrer, R.; Kumar, A.; Sevransky, J.E.; Sprung, C.L.; Nunnally, M.E.; et al. Surviving Sepsis Campaign: International Guidelines for Management of Sepsis and Septic Shock: 2016. Intensive Care Med 2017, 43, 304–377. [Google Scholar] [CrossRef] [PubMed]
- Scheer, C.S.; Fuchs, C.; Gründling, M.; Vollmer, M.; Bast, J.; Bohnert, J.A.; Zimmermann, K.; Hahnenkamp, K.; Rehberg, S.; Kuhn, S.O. Impact of Antibiotic Administration on Blood Culture Positivity at the Beginning of Sepsis: A Prospective Clinical Cohort Study. Clinical Microbiology and Infection 2019, 25, 326–331. [Google Scholar] [CrossRef] [PubMed]
- Vincent, J.L.; Brealey, D.; Libert, N.; Abidi, N.E.; O’Dwyer, M.; Zacharowski, K.; Mikaszewska-Sokolewicz, M.; Schrenzel, J.; Simon, F.; Wilks, M.; et al. Rapid Diagnosis of Infection in the Critically Ill, a Multicenter Study of Molecular Detection in Bloodstream Infections, Pneumonia, and Sterile Site Infections. Crit Care Med 2015, 43, 2283–2291. [Google Scholar] [CrossRef] [PubMed]
- Sinha, M.; Jupe, J.; Mack, H.; Coleman, T.P.; Lawrence, S.M.; Fraley, S.I. Emerging Technologies for Molecular Diagnosis of Sepsis. Clin Microbiol Rev 2018, 31. [Google Scholar] [CrossRef] [PubMed]
- Kaleta, E.J.; Clark, A.E.; Johnson, D.R.; Gamage, D.C.; Wysocki, V.H.; Cherkaoui, A.; Schrenzel, J.; Wolk, D.M. Use of PCR Coupled with Electrospray Ionization Mass Spectrometry for Rapid Identification of Bacterial and Yeast Bloodstream Pathogens from Blood Culture Bottles. J Clin Microbiol 2011, 49, 345–353. [Google Scholar] [CrossRef] [PubMed]
- Mandel, P. and M.P. Les Acides Nucléiques Du Plasma Sanguin Chez l’Homme. Available online: https://pubmed.ncbi.nlm.nih.gov/18875018/ (accessed on 26 June 2020).
- Stawski, R.; Stec-Martyna, E.; Chmielecki, A.; Nowak, D.; Perdas, E. Current Trends in Cell-Free DNA Applications. Scoping Review of Clinical Trials. Biology (Basel) 2021, 10, 906. [Google Scholar] [CrossRef]
- Hartwig, C.; Drechsler, S.; Vainshtein, Y.; Maneth, M.; Schmitt, T.; Ehling-Schulz, M.; Osuchowski, M.; Sohn, K. From Gut to Blood: Spatial and Temporal Pathobiome Dynamics during Acute Abdominal Murine Sepsis. Microorganisms 2023, 11, 627. [Google Scholar] [CrossRef]
- Grumaz, S.; Grumaz, C.; Vainshtein, Y.; Stevens, P.; Glanz, K.; Decker, S.O.; Hofer, S.; Weigand, M.A.; Brenner, T.; Sohn, K. Enhanced Performance of Next-Generation Sequencing Diagnostics Compared with Standard of Care Microbiological Diagnostics in Patients Suffering from Septic Shock. Crit Care Med 2019, 47, e394–e402. [Google Scholar] [CrossRef]
- Mikheyev, A.S.; Tin, M.M.Y. A First Look at the Oxford Nanopore MinION Sequencer. Mol Ecol Resour 2014, 14, 1097–1102. [Google Scholar] [CrossRef] [PubMed]
- Jain, M.; Olsen, H.E.; Paten, B.; Akeson, M. The Oxford Nanopore MinION: Delivery of Nanopore Sequencing to the Genomics Community. Genome Biol 2016, 17, 239. [Google Scholar] [CrossRef] [PubMed]
- Quick, J.; Ashton, P.; Calus, S.; Chatt, C.; Gossain, S.; Hawker, J.; Nair, S.; Neal, K.; Nye, K.; Peters, T.; et al. Rapid Draft Sequencing and Real-Time Nanopore Sequencing in a Hospital Outbreak of Salmonella. Genome Biol 2015, 16, 114. [Google Scholar] [CrossRef] [PubMed]
- Schmidt, K.; Mwaigwisya, S.; Crossman, L.C.; Doumith, M.; Munroe, D.; Pires, C.; M. Khan, A.; Woodford, N.; Saunders, N.J.; Wain, J.; et al. Identification of Bacterial Pathogens and Antimicrobial Resistance Directly from Clinical Urines by Nanopore-Based Metagenomic Sequencing. Journal of Antimicrobial Chemotherapy 2017, 72, 104–114. [Google Scholar] [CrossRef] [PubMed]
- Charalampous, T.; Kay, G.L.; Richardson, H.; Aydin, A.; Baldan, R.; Jeanes, C.; Rae, D.; Grundy, S.; Turner, D.J.; Wain, J.; et al. Nanopore Metagenomics Enables Rapid Clinical Diagnosis of Bacterial Lower Respiratory Infection. Nature Biotechnology 2019 37:7 2019, 37, 783–792. [Google Scholar] [CrossRef] [PubMed]
- Grumaz, C.; Hoffmann, A.; Vainshtein, Y.; Kopp, M.; Grumaz, S.; Stevens, P.; Decker, S.O.; Weigand, M.A.; Hofer, S.; Brenner, T.; et al. Rapid Next-Generation Sequencing–Based Diagnostics of Bacteremia in Septic Patients. Journal of Molecular Diagnostics 2020, 22, 405–418. [Google Scholar] [CrossRef] [PubMed]
- Broude, N.E.; Storm, N.; Malpel, S.; Graber, J.H.; Lukyanov, S.; Sverdlov, E.; Smith, C.L. PCR Based Targeted Genomic and CDNA Differential Display. Genet Anal 1999, 15, 51–63. [Google Scholar] [CrossRef] [PubMed]
- Broude, N.E.; Zhang, L.; Woodward, K.; Englert, D.; Cantor, C.R. Multiplex Allele-Specific Target Amplification Based on PCR Suppression. Proc Natl Acad Sci U S A 2001, 98, 206–211. [Google Scholar] [CrossRef] [PubMed]
- Heath, K.E.; Day, I.N.M.; Humphries, S.E. Universal Primer Quantitative Fluorescent Multiplex (UPQFM) PCR: A Method to Detect Major and Minor Rearrangements of the Low Density Lipoprotein Receptor Gene. J Med Genet 2000, 37, 272–280. [Google Scholar] [CrossRef]
- Grumaz, S.; Stevens, P.; Grumaz, C.; Decker, S.O.; Weigand, M.A.; Hofer, S.; Brenner, T.; von Haeseler, A.; Sohn, K. Next-Generation Sequencing Diagnostics of Bacteremia in Septic Patients. Genome Med 2016, 8, 73. [Google Scholar] [CrossRef]
- Huang, Q.; Fu, A.; Wang, Y.; Zhang, J.; Zhao, W.; Cheng, Y. Microbiological Diagnosis of Endophthalmitis Using Nanopore Targeted Sequencing. Clin Exp Ophthalmol 2021, 49, 1060–1068. [Google Scholar] [CrossRef] [PubMed]
- Brenner, T.; Decker, S.O.; Grumaz, S.; Stevens, P.; Bruckner, T.; Schmoch, T.; Pletz, M.W.; Bracht, H.; Hofer, S.; Marx, G.; et al. Next-Generation Sequencing Diagnostics of Bacteremia in Sepsis (Next GeneSiS-Trial): Study Protocol of a Prospective, Observational, Noninterventional, Multicenter, Clinical Trial. Medicine 2018, 97. [Google Scholar] [CrossRef] [PubMed]
- Nieman, A.E.; Savelkoul, P.H.M.; Beishuizen, A.; Henrich, B.; Lamik, B.; MacKenzie, C.R.; Kindgen-Milles, D.; Helmers, A.; Diaz, C.; Sakka, S.G.; et al. A Prospective Multicenter Evaluation of Direct Molecular Detection of Blood Stream Infection from a Clinical Perspective. BMC Infect Dis 2016, 16, 1–9. [Google Scholar] [CrossRef]
- Marimón, J.M.; Valiente, A.; Ercibengoa, M.; García-Arenzana, J.M.; Pérez-Trallero, E. Erythromycin Resistance and Genetic Elements Carrying Macrolide Efflux Genes in Streptococcus Agalactiae. Antimicrob Agents Chemother 2005, 49, 5069–5074. [Google Scholar] [CrossRef] [PubMed]
- Seymour, C.W.; Gesten, F.; Prescott, H.C.; Friedrich, M.E.; Iwashyna, T.J.; Phillips, G.S.; Lemeshow, S.; Osborn, T.; Terry, K.M.; Levy, M.M. Time to Treatment and Mortality during Mandated Emergency Care for Sepsis. New England Journal of Medicine 2017, 376, 2235–2244. [Google Scholar] [CrossRef] [PubMed]
- Loose, M.; Malla, S.; Stout, M. Real-Time Selective Sequencing Using Nanopore Technology. Nat Methods 2016, 13, 751–754. [Google Scholar] [CrossRef] [PubMed]
- Clarke, J.; Wu, H.C.; Jayasinghe, L.; Patel, A.; Reid, S.; Bayley, H. Continuous Base Identification for Single-Molecule Nanopore DNA Sequencing. Nature Nanotechnology 2009, 4, 265–270. [Google Scholar] [CrossRef] [PubMed]
- Al-Wadees, A.A.N.; Al-Khayyat, A.N.; Yaqoob, Q.A. The Outcome of Sepsis Patients Admitted to the Intensive Care Unit: Experience of 100 Cases. Medical Archives 2021, 75, 35. [Google Scholar] [CrossRef] [PubMed]
- Why Do I Need to Put a Light Shield on My Flow Cell? | Oxford Nanopore Technologies Help Center. Available online: https://help.nanoporetech.com/en/articles/8304478-why-do-i-need-to-put-a-light-shield-on-my-flow-cell (accessed on 21 December 2023).
- Alcock, B.P.; Huynh, W.; Chalil, R.; Smith, K.W.; Raphenya, A.R.; Wlodarski, M.A.; Edalatmand, A.; Petkau, A.; Syed, S.A.; Tsang, K.K.; et al. CARD 2023: Expanded Curation, Support for Machine Learning, and Resistome Prediction at the Comprehensive Antibiotic Resistance Database. Nucleic Acids Res 2023, 51, D690–D699. [Google Scholar] [CrossRef]
- Gourlé, H.; Karlsson-Lindsjö, O.; Hayer, J.; Bongcam-Rudloff, E. Simulating Illumina Metagenomic Data with InSilicoSeq. Bioinformatics 2019, 35, 521–522. [Google Scholar] [CrossRef]
- Wood, D.E.; Lu, J.; Langmead, B. Improved Metagenomic Analysis with Kraken 2. Genome Biol 2019, 20. [Google Scholar] [CrossRef]
- Camacho, C.; Coulouris, G.; Avagyan, V.; Ma, N.; Papadopoulos, J.; Bealer, K.; Madden, T.L. BLAST+: Architecture and Applications. BMC Bioinformatics 2009, 10, 421. [Google Scholar] [CrossRef]
- GitHub - Nanoporetech/Qcat: Qcat Is a Python Command-Line Tool for Demultiplexing Oxford Nanopore Reads from FASTQ Files. Available online: https://github.com/nanoporetech/qcat (accessed on 22 February 2021).
- Untergasser, A.; Cutcutache, I.; Koressaar, T.; Ye, J.; Faircloth, B.C.; Remm, M.; Rozen, S.G. Primer3--New Capabilities and Interfaces. Nucleic Acids Res 2012, 40. [Google Scholar] [CrossRef]
- Matz, M.; Usman, N.; Shagin, D.; Bogdanova, E.; Lukyanov, S. Ordered Differential Display: A Simple Method for Systematic Comparison of Gene Expression Profiles. Nucleic Acids Res 1997, 25, 2541–2542. [Google Scholar] [CrossRef]
- De Coster, W.; D’Hert, S.; Schultz, D.T.; Cruts, M.; Van Broeckhoven, C. NanoPack: Visualizing and Processing Long-Read Sequencing Data. Bioinformatics 2018, 34, 2666–2669. [Google Scholar] [CrossRef]
- Li, H. Minimap2: Pairwise Alignment for Nucleotide Sequences. Bioinformatics 2018, 34, 3094–3100. [Google Scholar] [CrossRef]
- Li, H.; Handsaker, B.; Wysoker, A.; Fennell, T.; Ruan, J.; Homer, N.; Marth, G.; Abecasis, G.; Durbin, R. The Sequence Alignment/Map Format and SAMtools. Bioinformatics 2009, 25, 2078–2079. [Google Scholar] [CrossRef] [PubMed]
- Thorvaldsdóttir, H.; Robinson, J.T.; Mesirov, J.P. Integrative Genomics Viewer (IGV): High-Performance Genomics Data Visualization and Exploration. Brief Bioinform 2013, 14, 178–192. [Google Scholar] [CrossRef] [PubMed]
| Sample | SUPSETS | Blood culture | Clinical specimen |
|---|---|---|---|
| FC_06 | vanA | - | Vancomycine |
| FL_01 | vanA | - | Vancomycine |
| FL_02 | vanA, ermB | - | Vancomycine |
| FL_03 | tetB | - | - |
| FL_04-single | tetB | - | - |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




