Submitted:
04 April 2024
Posted:
05 April 2024
Read the latest preprint version here
Abstract
Keywords:
1. Introduction
2. History of CRISPR/Cas System
3. Nomenclature of CRISPR/Cas System
4. Cas Proteins of the CRISPR System
4.1. Cas 1 and Cas 2 Proteins
Mechanism and Application of Cas 1 and Cas 2 Proteins
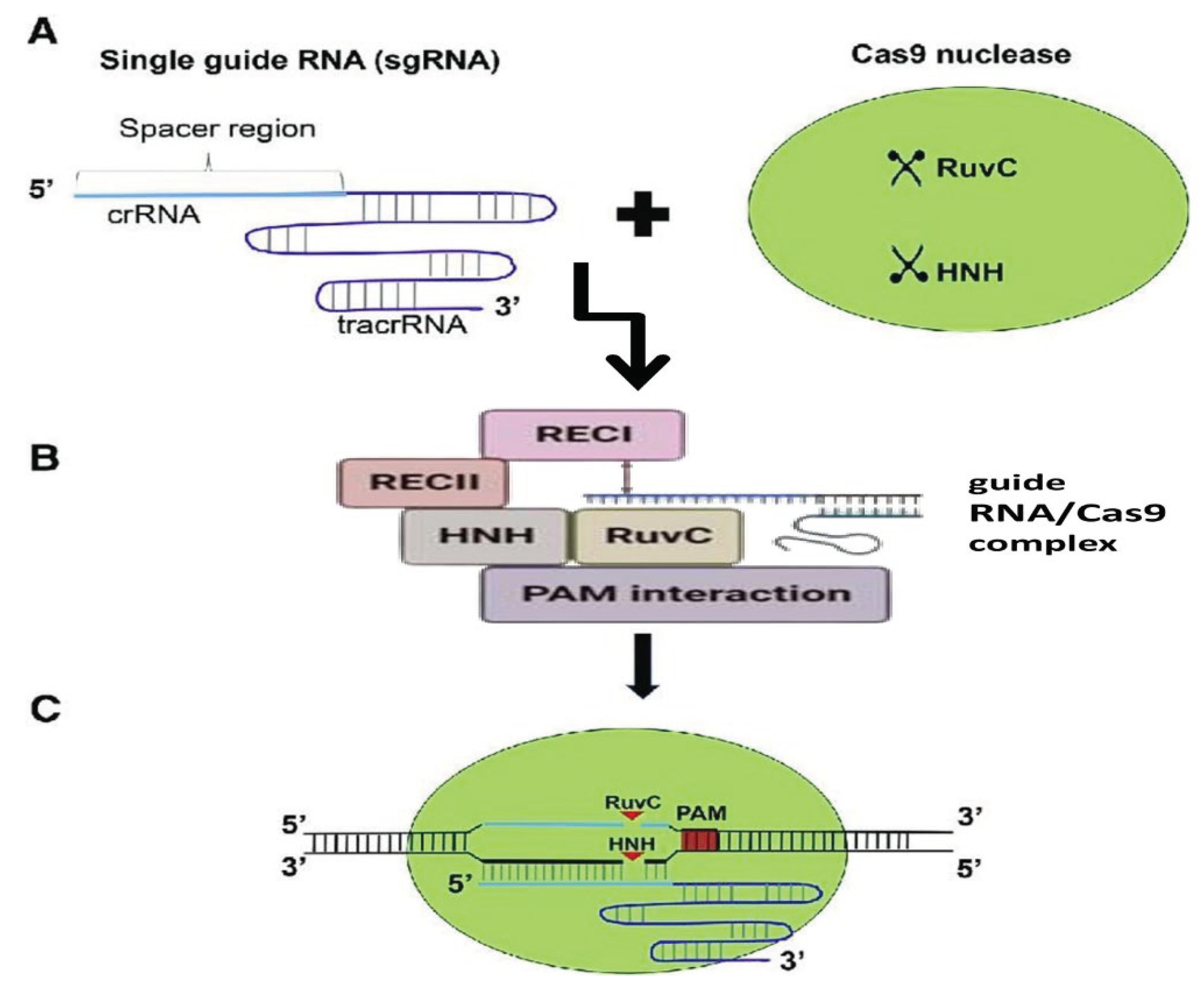
4.2. Cas9 Protein
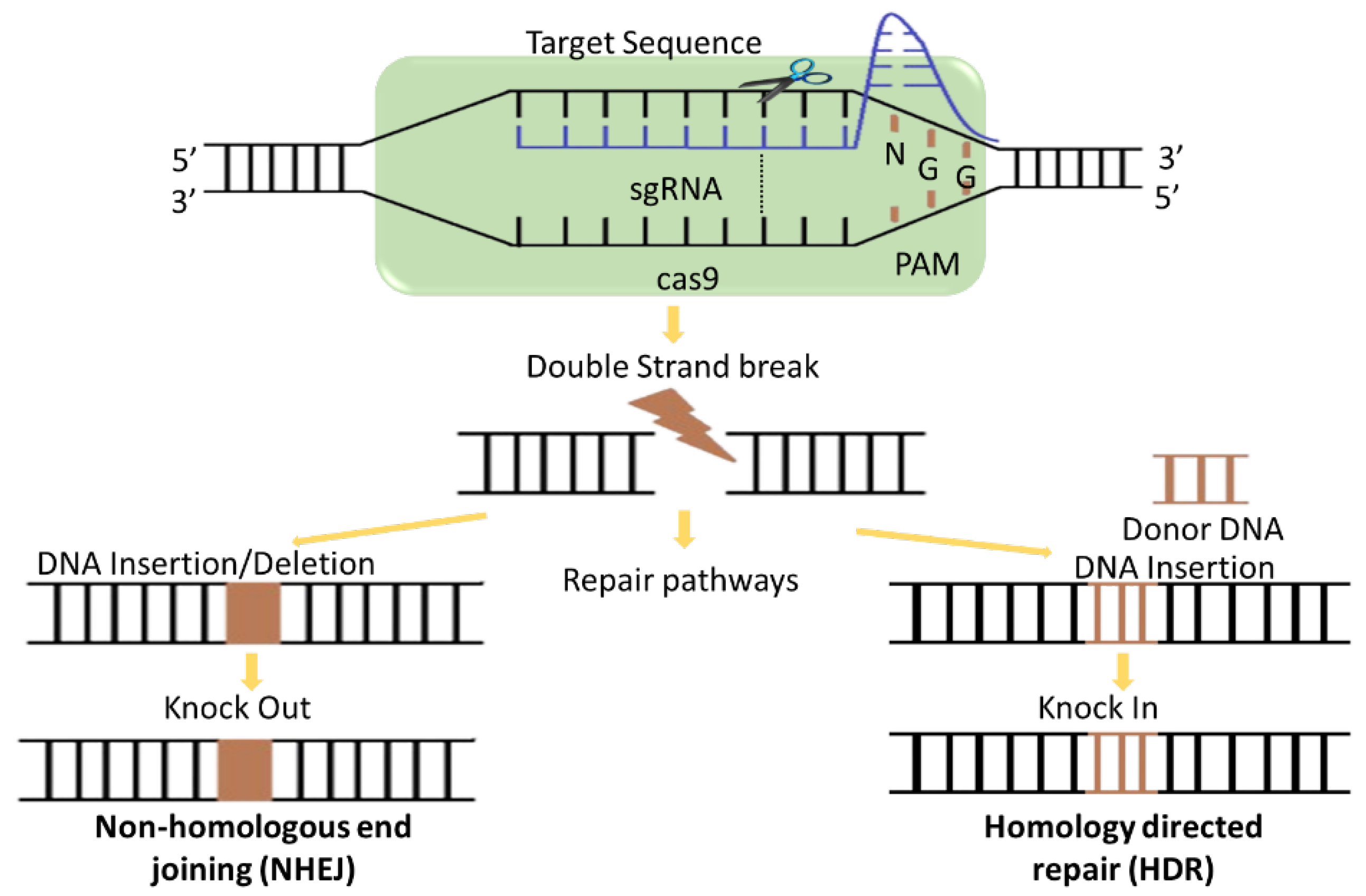
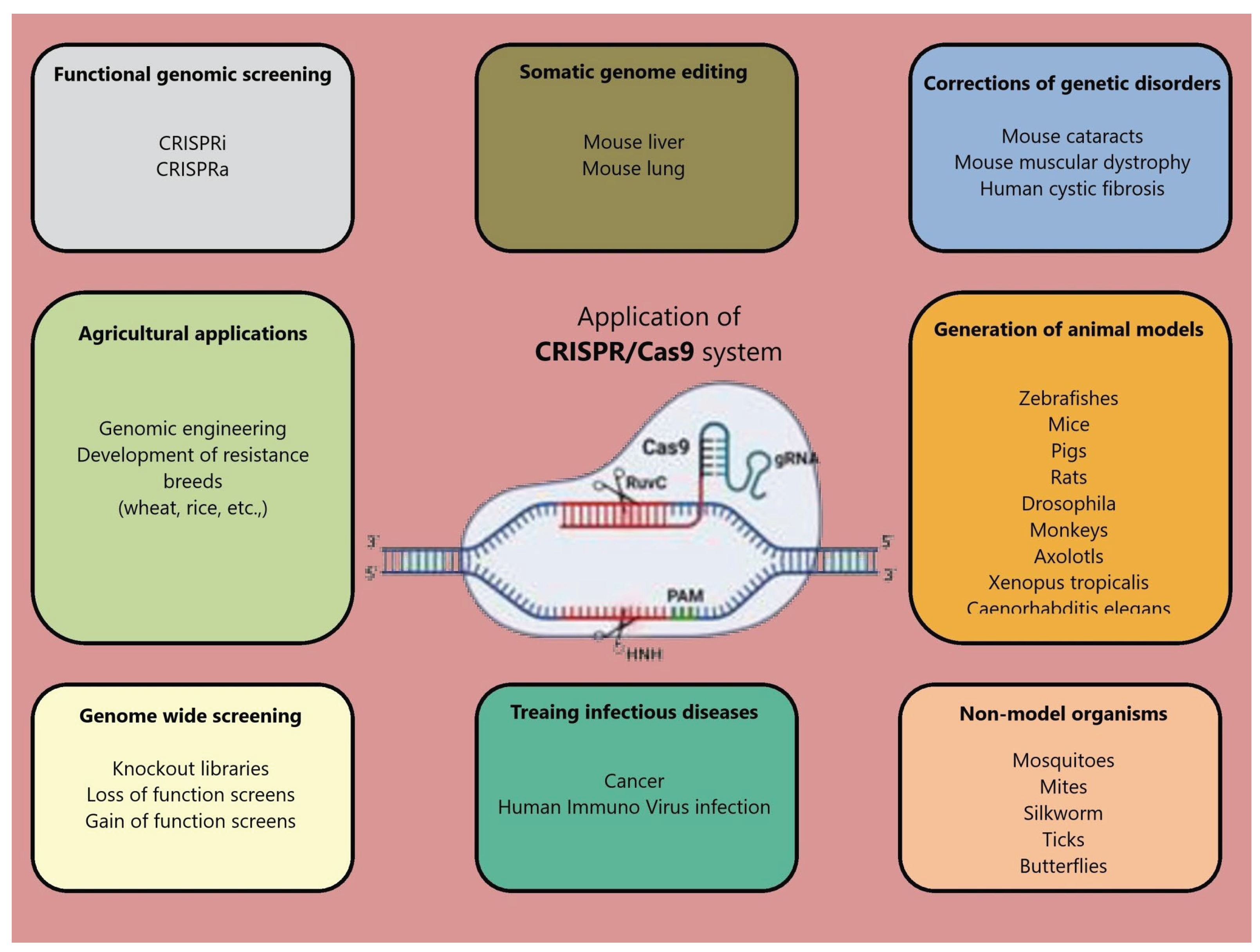
4.2.1. Mechanism of Cas9 Protein and Application
4.2.2. Pros and Cons of Cas9 Protein
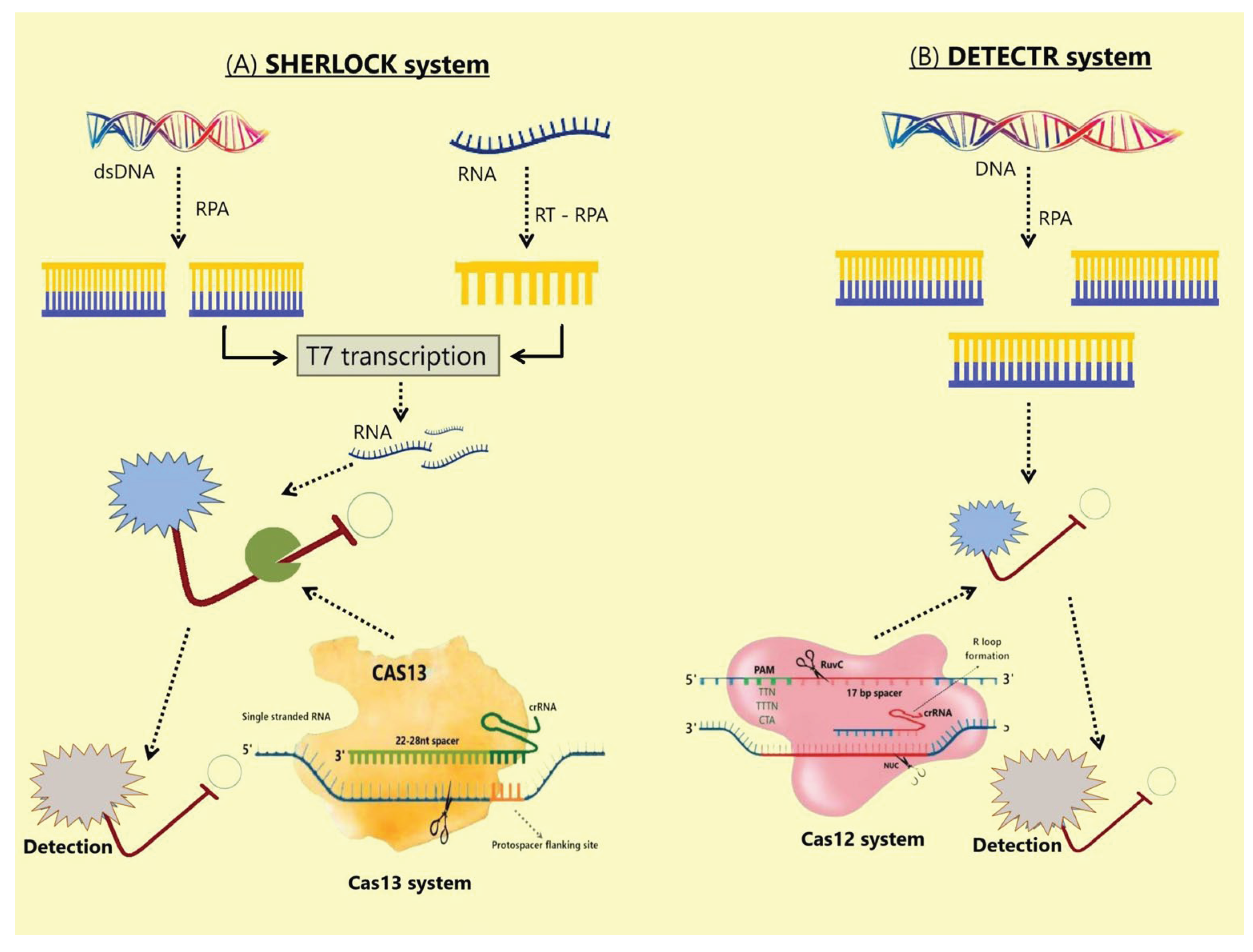
4.3. Cas12 Protein
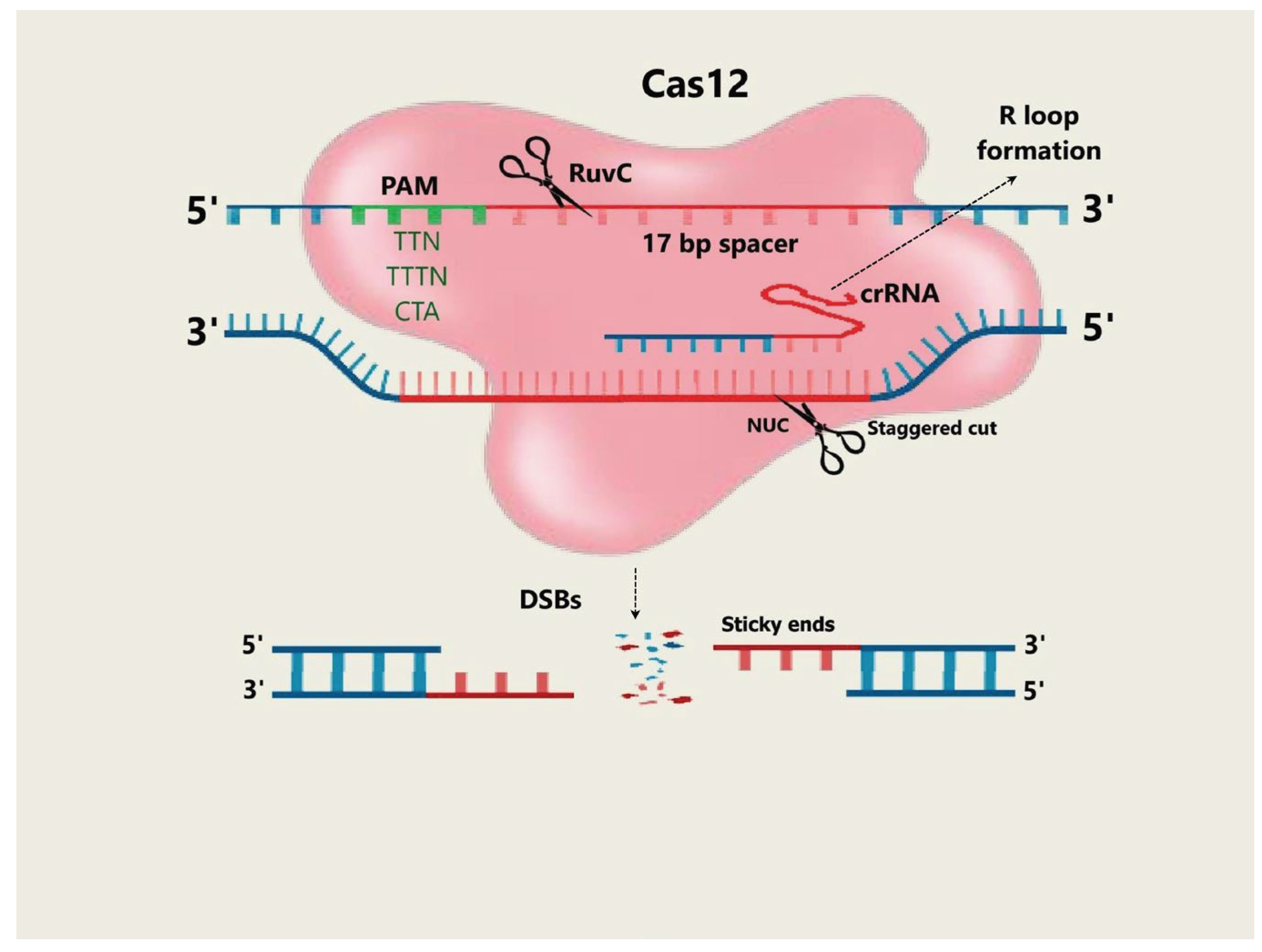
4.3.1. Mechanism and Application of Cas12 Protein
4.2.2. Pros and Cons of Cas12 Protein
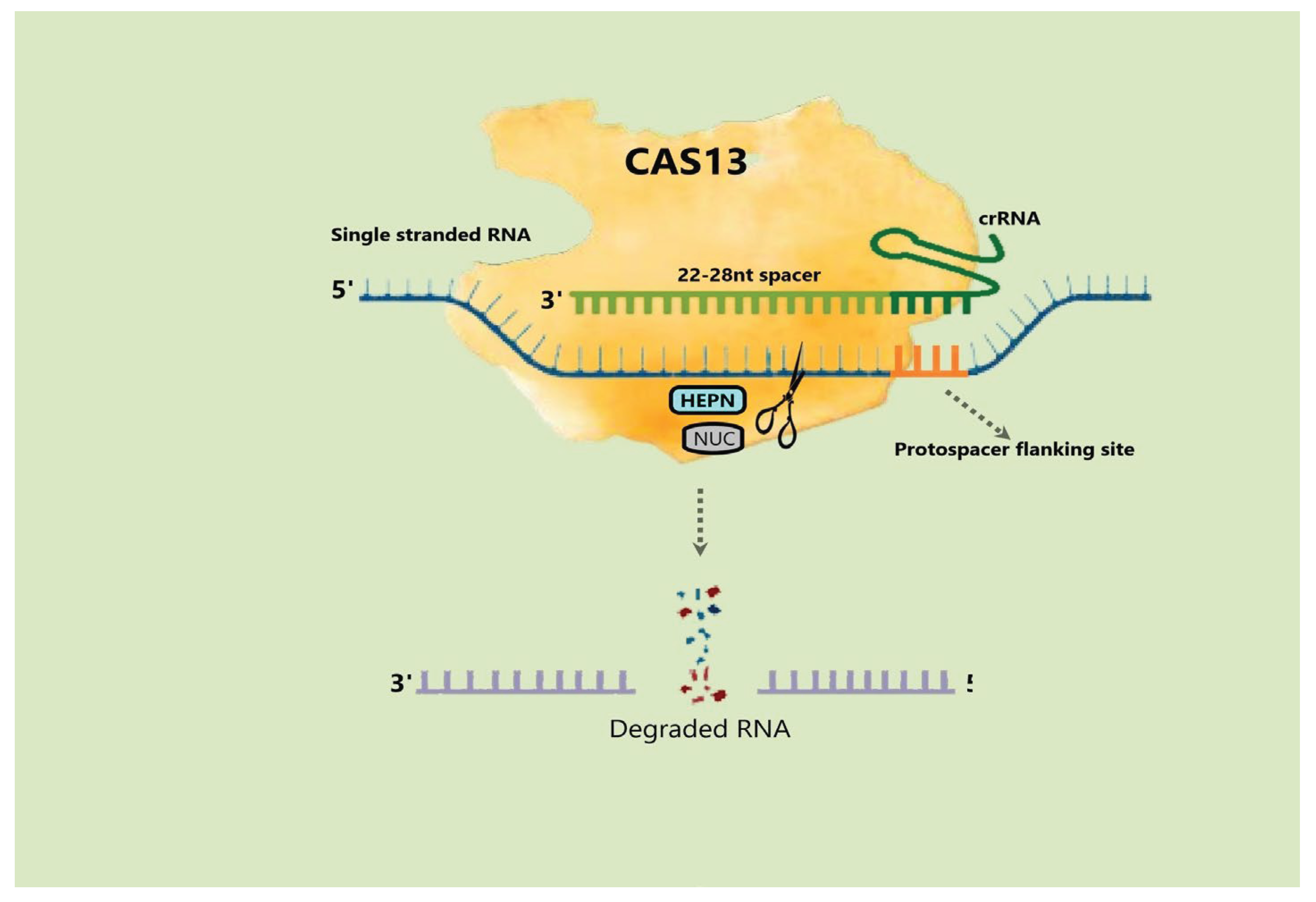
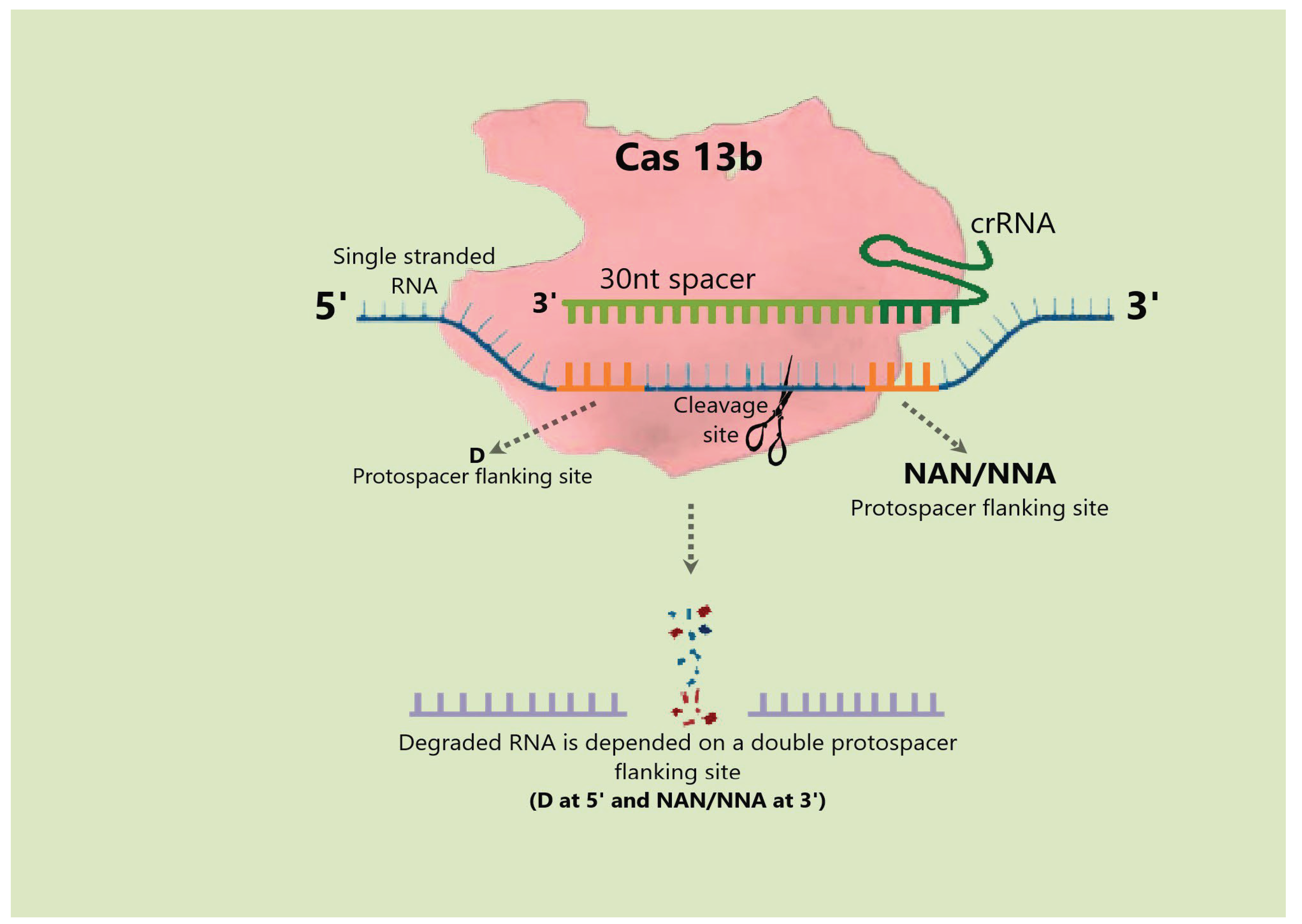
4.4. Cas13 Protein
4.4.1. Mechanism and Applications of Cas13 Protein
4.4.2. Pros and Cons of Cas13 Protein
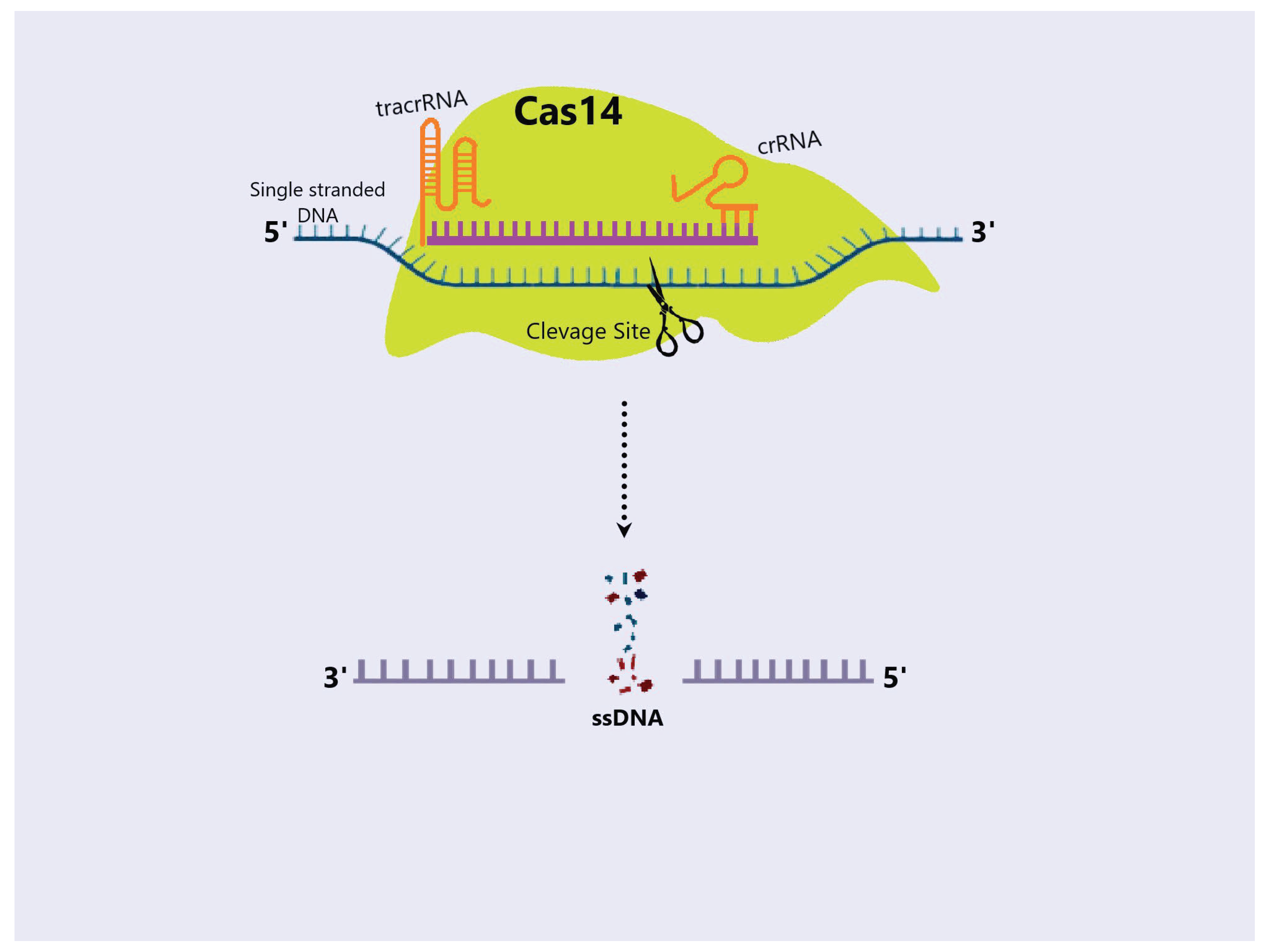
4.5. Cas14 Protein
4.5.1. Mechanism and Application of Cas14 Protein
4.5.2. Pros and Cons of Cas14 Protein
5. CRISPR/Cas Technology for Crop Improvement
6. Application against Abiotic Stress
7. Application against Biotic Stress
8. Prime Technology
Application of the Prime Technology in Plants
9. Conclusion and Future Perspectives
Author Contributions
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Clarke, J. L., & Zhang, P. (2013). Plant biotechnology for food security and bioeconomy. Plant molecular biology, 83(1-2), 1–3. [CrossRef]
- Gill, S. S., & Tuteja, N. (2010). Reactive oxygen species and antioxidant machinery in abiotic stress tolerance in crop plants. Plant physiology and biochemistry: PPB, 48(12), 909–930. [CrossRef]
- Li, R., Char, S.N., Yang, B. (2019). Creating Large Chromosomal Deletions in Rice Using CRISPR/Cas9. In: Qi, Y. (eds) Plant Genome Editing with CRISPR Systems. Methods in Molecular Biology, vol 1917. Humana, New York, NY. [CrossRef]
- Marshall, M.(2024). Scientists are attempting to rapidly domesticate wild plant species by editing specific genes, but they face major technical challenges — and concerns about exploitation of Indigenous knowledge, Nature 625, 230-232. [CrossRef]
- Akram, F., Sahreen, S., Aamir, F., Haq, I. U., Malik, K., Imtiaz, M., Naseem, W., Nasir, N., & Waheed, H. M. (2023). An Insight into Modern Targeted Genome-Editing Technologies with a Special Focus on CRISPR/Cas9 and its Applications. Molecular biotechnology, 65(2), 227–242. [CrossRef]
- Yu, D., Zhou, T., Xu, N., Sun, X., Song, S., Liu, H., Sun, Z., Lv, Q., Chen, J., Tan, Y., Sheng, X., Li, L. and Yuan, D. (2024), Novel CRISPR/Cas9 system assisted by fluorescence marker and pollen killer for high-efficiency isolation of transgene-free edited plants in rice. Plant Biotechnol. J. [CrossRef]
- Rasheed, A., Gill, R. A., Hassan, M. U., Mahmood, A., Qari, S., Zaman, Q. U., Ilyas, M., Aamer, M., Batool, M., Li, H., & Wu, Z. (2021). A Critical Review: Recent Advancements in the Use of CRISPR/Cas9 Technology to Enhance Crops and Alleviate Global Food Crises. Current issues in molecular biology, 43(3), 1950–1976. [CrossRef]
- Yang, B.(2020). Grand Challenges in Genome Editing in Plants. Frontiers in genome editing, 2, 2. [CrossRef]
- Zhou, Y., Xu, S., Jiang, N., Zhao, X., Bai, Z., Liu, J., Yao, W., Tang, Q., Xiao, G., Lv, C., Wang, K., Hu, X., Tan, J., & Yang, Y. (2022). Engineering of rice varieties with enhanced resistances to both blast and bacterial blight diseases via CRISPR/Cas9. Plant biotechnology journal, 20(5), 876–885. [CrossRef]
- Naik, B. J., Shimoga, G., Kim, S. C., Manjulatha, M., Subramanyam Reddy, C., Palem, R. R., Kumar, M., Kim, S. Y., & Lee, S. H. (2022). CRISPR/Cas9 and Nanotechnology Pertinence in Agricultural Crop Refinement. Frontiers in plant science, 13, 843575. [CrossRef]
- Rao, Y., Yang, X., Pan, C., Wang, C., & Wang, K. (2022). Advance of Clustered Regularly Interspaced Short Palindromic Repeats-Cas9 System and Its Application in Crop Improvement. Frontiers in plant science, 13, 839001. [CrossRef]
- Mao, Y., Zhang, H., Xu, N., Zhang, B., Gou, F., & Zhu, J. K. (2013). Application of the CRISPR-Cas system for efficient genome engineering in plants. Molecular plant, 6(6), 2008–2011. [CrossRef]
- Nekrasov, V., Staskawicz, B., Weigel, D., Jones, J. D., & Kamoun, S. (2013). Targeted mutagenesis in the model plant Nicotiana benthamiana using Cas9 RNA-guided endonuclease. Nature biotechnology, 31(8), 691–693. [CrossRef]
- Qi, Y., Zhang, L., Wang, L., Song, J., and Wang, J. (2020). CRISPR-Cas9 targeted editing for the fragrant gene Badh2 in rice. Sci. Agric. Sin. 53, 1501–1509. [CrossRef]
- Brooks, C., Nekrasov, V., Lippman, Z. B., & Van Eck, J. (2014). Efficient gene editing in tomato in the first generation using the clustered regularly interspaced short palindromic repeats/CRISPR-associated9 system. Plant physiology, 166(3), 1292–1297. [CrossRef]
- Feng, C., Yuan, J., Wang, R., Liu, Y., Birchler, J. A., and Han, F. (2016). Efficient targeted genome modification in maize using CRISPR-Cas9 system. J. Genet. Genomics 43, 37–43. [CrossRef]
- Wang, X., Tu, M., Wang, D., Liu, J., Li, Y., Li, Z., Wang, Y., & Wang, X. (2018). CRISPR/Cas9-mediated efficient targeted mutagenesis in grape in the first generation. Plant biotechnology journal, 16(4), 844–855. [CrossRef]
- Wang, S., Zhang, S., Wang, W., Xiong, X., Meng, F., & Cui, X. (2015). Efficient targeted mutagenesis in potato by the CRISPR/Cas9 system. Plant cell reports, 34(9), 1473–1476. [CrossRef]
- Tiwari JK, Jeevalatha A, Tuteja N, Khurana, SMP. Genome editing (CRISPR-Cas)-mediated virus resistance in potato (Solanum tuberosum L.). 1210; 49. [CrossRef]
- Braatz, J., Harloff, H. J., Mascher, M., Stein, N., Himmelbach, A., & Jung, C. (2017). CRISPR-Cas9 Targeted Mutagenesis Leads to Simultaneous Modification of Different Homoeologous Gene Copies in Polyploid Oilseed Rape (Brassica napus). Plant physiology, 174(2), 935–942. [CrossRef]
- Gostimskaya, I.(2022). CRISPR-Cas9: A History of Its Discovery and Ethical Considerations of Its Use in Genome Editing. Biochemistry. Biokhimiia, 87(8), 777–788. [CrossRef]
- Xu, Y., & Li, Z. (2020). CRISPR-Cas systems: Overview, innovations and applications in human disease research and gene therapy. Computational and structural biotechnology journal, 18, 2401–2415. [CrossRef]
- Kim, E., Koo, T., Park, S. et al. In vivo genome editing with a small Cas9 orthologue derived from Campylobacter jejuni. Nat Commun 8, 14500 (2017). [CrossRef]
- Liu, Z., Dong, H., Cui, Y., Cong, L., & Zhang, D. (2020). Application of different types of CRISPR/Cas-based systems in bacteria. Microbial cell factories, 19(1), 172. [CrossRef]
- Ceasar, S. A., Maharajan, T., Hillary, V. E., & Ajeesh Krishna, T. P. (2022). Insights to improve the plant nutrient transport by CRISPR/Cas system. Biotechnology advances, 59, 107963. [CrossRef]
- Rodríguez-Rodríguez, D. R., Ramírez-Solís, R., Garza-Elizondo, M. A., Garza-Rodríguez, M. L., & Barrera-Saldaña, H. A. (2019). Genome editing: A perspective on the application of CRISPR/Cas9 to study human diseases (Review). International journal of molecular medicine, 43(4), 1559–1574. [CrossRef]
- Zhang, B., Ye, Y., Ye, W., Perčulija, V., Jiang, H., Chen, Y., Li, Y., Chen, J., Lin, J., Wang, S., Chen, Q., Han, Y. S., & Ouyang, S. (2019). Two HEPN domains dictate CRISPR RNA maturation and target cleavage in Cas13d. Nature communications, 10(1), 2544. [CrossRef] [PubMed]
- Li, R., Char, S.N., Yang, B. (2019). Creating Large Chromosomal Deletions in Rice Using CRISPR/Cas9. In: Qi, Y. (eds) Plant Genome Editing with CRISPR Systems. Methods in Molecular Biology, vol 1917. Humana, New York, NY. [CrossRef]
- Cofsky, J. C., Karandur, D., Huang, C. J., Witte, I. P., Kuriyan, J., & Doudna, J. A. (2020). CRISPR-Cas12a exploits R-loop asymmetry t Ceasar, S. A., Maharajan, T., Hillary, V. E., & Ajeesh Krishna, T. P. (2022). Insights to improve the plant nutrient transport by CRISPR/Cas system. Biotechnology advances, 59, 107963. https://doi.org/10.1016/j.biotechadv.2022.107963o form double-strand breaks. eLife, 9, e55143. [CrossRef]
- Jiang, Y., Hu, M., Liu, A. A., Lin, Y., Liu, L., Yu, B., Zhou, X., & Pang, D. W. (2021). Detection of SARS-CoV-2 by CRISPR/Cas12a-Enhanced Colorimetry. ACS sensors, 6(3), 1086–1093. [CrossRef]
- Hillary, V. E., & Ceasar, S. A. (2022). Prime editing in plants and mammalian cells: Mechanism, achievements, limitations, and future prospects. BioEssays: news and reviews in molecular, cellular and developmental biology, 44(9), e2200032. [CrossRef]
- Hillary, V. E., Ignacimuthu, S., & Ceasar, S. A. (2021). Potential of CRISPR/Cas system in the diagnosis of COVID-19 infection. Expert review of molecular diagnostics, 21(11), 1179–1189. [CrossRef]
- Al-Shayeb, B., Sachdeva, R., Chen, L. X., Ward, F., Munk, P., Devoto, A., Castelle, C. J., Olm, M. R., Bouma-Gregson, K., Amano, Y., He, C., Méheust, R., Brooks, B., Thomas, A., Lavy, A., Matheus-Carnevali, P., Sun, C., Goltsman, D. S. A., Borton, M. A., Sharrar, A., … Banfield, J. F. (2020). Clades of huge phages from across Earth’s ecosystems. Nature, 578(7795), 425–431. [CrossRef]
- Liu, L., Li, X., Wang, J., Wang, M., Chen, P., Yin, M., Li, J., Sheng, G., & Wang, Y. (2017). Two Distant Catalytic Sites Are Responsible for C2c2 RNase Activities. Cell, 168(1-2), 121–134.e12. [CrossRef]
- Wang, B., Zhang, T., Yin, J., Yu, Y., Xu, W., Ding, J., Patel, D. J., & Yang, H. (2021). Structural basis for self-cleavage prevention by tag:anti-tag pairing complementarity in type VI Cas13 CRISPR systems. Molecular cell, 81(5), 1100–1115.e5. [CrossRef]
- Ai, Y., Liang, D., & Wilusz, J. E. (2022). CRISPR/Cas13 effectors have differing extents of off-target effects that limit their utility in eukaryotic cells. Nucleic acids research, 50(11), e65. [CrossRef]
- Lotfi, M., & Rezaei, N. (2020). CRISPR/Cas13: A potential therapeutic option of COVID-19. Biomedicine & pharmacotherapy = Biomedecine & pharmacotherapie, 131, 110738. [CrossRef]
- Myhrvold, C., Freije, C. A., Gootenberg, J. S., Abudayyeh, O. O., Metsky, H. C., Durbin, A. F., Kellner, M. J., Tan, A. L., Paul, L. M., Parham, L. A., Garcia, K. F., Barnes, K. G., Chak, B., Mondini, A., Nogueira, M. L., Isern, S., Michael, S. F., Lorenzana, I., Yozwiak, N. L., MacInnis, B. L., … Sabeti, P. C. (2018). Field-deployable viral diagnostics using CRISPR-Cas13. Science (New York, N.Y.), 360(6387), 444–448. [CrossRef]
- Alam, M. S., Kong, J., Tao, R., Ahmed, T., Alamin, M., Alotaibi, S. S., Abdelsalam, N. R., & Xu, J. H. (2022). CRISPR/Cas9 Mediated Knockout of the OsbHLH024 Transcription Factor Improves Salt Stress Resistance in Rice (Oryza sativa L.). Plants (Basel, Switzerland), 11(9), 1184. [CrossRef]
- Han, X., Chen, Z., Li, P., Xu, H.; Liu, K., Zha, W., Li, S., Chen, J., Yang, G., Huang, J., et al. Development of Novel Rice Germplasm for Salt-Tolerance at Seedling Stage Using CRISPR-Cas9. Sustainability 2022, 14, 2621. [CrossRef]
- Santosh Kumar, V. V., Verma, R. K., Yadav, S. K., Yadav, P., Watts, A., Rao, M. V., & Chinnusamy, V. (2020). CRISPR-Cas9 mediated genome editing of drought and salt tolerance (OsDST) gene in indica mega rice cultivar MTU1010. Physiology and molecular biology of plants: an international journal of functional plant biology, 26(6), 1099–1110. [CrossRef]
- Yue, E., Cao, H., & Liu, B. (2020). OsmiR535, a Potential Genetic Editing Target for Drought and Salinity Stress Tolerance in Oryza sativa. Plants (Basel, Switzerland), 9(10), 1337. [CrossRef]
- Vlčko, T., & Ohnoutková, L. (2020). Allelic Variants of CRISPR/Cas9 Induced Mutation in an Inositol Trisphosphate 5/6 Kinase Gene Manifest Different Phenotypes in Barley. Plants (Basel, Switzerland), 9(2), 195. [CrossRef]
- Ogata, T., Ishizaki, T., Fujita, M., & Fujita, Y. (2020). CRISPR/Cas9-targeted mutagenesis of OsERA1 confers enhanced responses to abscisic acid and drought stress and increased primary root growth under nonstressed conditions in rice. PloS one, 15(12), e0243376. [CrossRef]
- Usman, B., Nawaz, G., Zhao, N., Liao, S., Liu, Y., & Li, R. (2020). Precise Editing of the OsPYL9 Gene by RNA-Guided Cas9 Nuclease Confers Enhanced Drought Tolerance and Grain Yield in Rice (Oryza sativa L.) by Regulating Circadian Rhythm and Abiotic Stress Responsive Proteins. International journal of molecular sciences, 21(21), 7854. [CrossRef]
- Ding, Y., Shi, Y., & Yang, S. (2020). Molecular Regulation of Plant Responses to Environmental Temperatures. Molecular plant, 13(4), 544–564. https://doi.org/10.1016/j.molp.2020.02.004. [CrossRef]
- Zeng, Y., Wen, J., Zhao, W., Wang, Q., & Huang, W. (2020). Rational Improvement of Rice Yield and Cold Tolerance by Editing the Three Genes OsPIN5b, GS3, and OsMYB30 With the CRISPR-Cas9 System. Frontiers in plant science, 10, 1663. [CrossRef]
- Wan, D. Y., Guo, Y., Cheng, Y., Hu, Y., Xiao, S., Wang, Y., & Wen, Y. Q. (2020). CRISPR/Cas9-mediated mutagenesis of VvMLO3 results in enhanced resistance to powdery mildew in grapevine (Vitis vinifera). Horticulture research, 7, 116. [CrossRef]
- Nekrasov, V., Staskawicz, B., Weigel, D., Jones, J. D., & Kamoun, S. (2013). Targeted mutagenesis in the model plant Nicotiana benthamiana using Cas9 RNA-guided endonuclease. Nature biotechnology, 31(8), 691–693. [CrossRef]
- Santillán Martínez, M.I., Bracuto, V., Koseoglou, E. et al. CRISPR/Cas9-targeted mutagenesis of the tomato susceptibility gene PMR4 for resistance against powdery mildew. BMC Plant Biol 20, 284 (2020). [CrossRef]
- Wang, F., Wang, C., Liu, P., Lei, C., Hao, W., Gao, Y., Liu, Y. G., & Zhao, K. (2016). Enhanced Rice Blast Resistance by CRISPR/Cas9-Targeted Mutagenesis of the ERF Transcription Factor Gene OsERF922. PloS one, 11(4), e0154027. [CrossRef]
- Ma, J., Chen, J., Wang, M., Ren, Y., Wang, S., Lei, C., Cheng, Z., & Sodmergen (2018). Disruption of OsSEC3A increases the content of salicylic acid and induces plant defense responses in rice. Journal of experimental botany, 69(5), 1051–1064. [CrossRef]
- Nawaz, G., Usman, B., Peng, H., Zhao, N., Yuan, R., Liu, Y., & Li, R. (2020). Knockout of Pi21 by CRISPR/Cas9 and iTRAQ-Based Proteomic Analysis of Mutants Revealed New Insights into M. oryzae Resistance in Elite Rice Line. Genes, 11(7), 735. [CrossRef]
- Zhou, Y., Xu, S., Jiang, N., Zhao, X., Bai, Z., Liu, J., Yao, W., Tang, Q., Xiao, G., Lv, C., Wang, K., Hu, X., Tan, J., & Yang, Y. (2022). Engineering of rice varieties with enhanced resistances to both blast and bacterial blight diseases via CRISPR/Cas9. Plant biotechnology journal, 20(5), 876–885. [CrossRef]
- Thomazella, D. P. T., Seong, K., Mackelprang, R., Dahlbeck, D., Geng, Y., Gill, U. S., Qi, T., Pham, J., Giuseppe, P., Lee, C. Y., Ortega, A., Cho, M. J., Hutton, S. F., & Staskawicz, B. (2021). Loss of function of a DMR6 ortholog in tomato confers broad-spectrum disease resistance. Proceedings of the National Academy of Sciences of the United States of America, 118(27), e2026152118. [CrossRef]
- Chu, C., Huang, R., Liu, L., Tang, G., Xiao, J., Yoo, H., & Yuan, M. (2022). The rice heavy-metal transporter OsNRAMP1 regulates disease resistance by modulating ROS homeostasis. Plant, cell & environment, 45(4), 1109–1126. [CrossRef]
- Gupta, A., Liu, B., Chen, Q. J., & Yang, B. (2023). High-efficiency prime editing enables new strategies for broad-spectrum resistance to bacterial blight of rice. Plant biotechnology journal, 21(7), 1454–1464. [CrossRef]
- Anzalone, A. V., Randolph, P. B., Davis, J. R., Sousa, A. A., Koblan, L. W., Levy, J. M., Chen, P. J., Wilson, C., Newby, G. A., Raguram, A., & Liu, D. R. (2019). Search-and-replace genome editing without double-strand breaks or donor DNA. Nature, 576(7785), 149–157. [CrossRef]
- Hassan, M. M., Zhang, Y., Yuan, G., De, K., Chen, J. G., Muchero, W., Tuskan, G. A., Qi, Y., & Yang, X. (2021). Construct design for CRISPR/Cas-based genome editing in plants. Trends in plant science, 26(11), 1133–1152. [CrossRef]
- Tong, Y., Jørgensen, T. S., Whitford, C. M., Weber, T., & Lee, S. Y. (2021). A versatile genetic engineering toolkit for E. coli based on CRISPR-prime editing. Nature communications, 12(1), 5206. [CrossRef]

| Class (C)&Type (T) | Protein(P) & Target (T) |
Spacer acquisition strategy |
Name of CRISPR/Cas system |
Pre-Crispr Processing |
Self vs. Non-self Discrimination |
Effectors of CRISPR System |
Host Organism |
Reference |
|---|---|---|---|---|---|---|---|---|
| C – 1 T – III |
P – Cas3 T-ssDNAssDNA |
Cas1/Cas2 | Cas7, Cas5, Cas8, and Cas3 |
Cas6 |
- | Cas3, Cascade, and crRNA |
E. coli |
[61] |
| C – 1 T – I |
P – Cas10 T- A |
Cas4Cas1/Cas2 | Cas7, Cas5, and Cas1 |
Cas6 |
- | Cmr/Csm, crRNA, and Cas10 |
S.epidermics |
[61] |
| C – 1 T - IV |
P – Csf1 T - - |
- | Cas7, Cas5, and | - | - | - | - | [61] |
| C – 2 T – II |
P – Cas9 T – dsDNA |
Cas1/Cas2/ Cas4 |
Csf1Cas9 |
RNase III, and tracr RNA | - | Cas9, tracrRNA, and crRNA | S. thermophilus and S. pyogenes | [61] |
| C – 2 T – V |
P – Cpf1 T – ssDNA & dsDNA |
Cas1/Cas2/ Cas4 |
Cas12 |
Cpf1 |
PAM |
Cpf1, crRNA and tracrRNA |
F.novicida |
[61] |
| C – 2 T - VI |
P – C2c2 T - ssRNA |
Cas1/Cas2 |
Cas13 |
- | - | C2c1, and crRNA | - | [61] |
| ProteinName | Host organism | sgRNAsize | PAM sequence | Target | Cut site |
References |
|---|---|---|---|---|---|---|
| Cas9 | S.pyogenes | 20 | 5ʹ-NGG-3ʹ | dsDNA | 5ʹofPAM | [61] |
| Cas9 | S.pyogenes | - | 5ʹ-NAC,NTG,NTT,andNCG-3ʹ | DNA | 5ʹofPAM | [61] |
| Cas9 | F.novicida | 20 | 5ʹ-NGG-3ʹ | DNA | 5ʹofPAM | [61] |
| Cas9 | S.aureus | 21 | 5ʹ-NNGRRT-3ʹ | DNA | 5ʹofPAM | [61] |
| Cas9 | Neisseriameningitidis | 24 | 5ʹ-NNNNGATT-3ʹ | DNA | 5ʹofPAM | [61] |
| Cas9 | S. thermophilus | 20 | 5ʹ-NNAGAAW5ʹ | DNA | 5ʹofPAM | [61] |
| Cas9 | S. thermophilus | 20 | 5ʹ-NGGNG-3 | DNA | 5ʹofPAM | [61] |
| Cas9 | Campylobacterjejuna | 22 | NNNNACAC and NNNRYAC | DNA | 5ʹofPAM | [61] |
| C2c1 | Alicyclobacillusacidoterrestris | 20 | T-rich PAM | DNA | 5ʹofPAM | [61] |
| Cpf1 | PrevotellaandFrancisella | 20 | TTTV | DNA | 5ʹofPAM | [61] |
| Cpf1 | Acidaminococcussp. | 24 | 5ʹ-TTTN-3ʹ | DNA | 3ʹofPAM | [61] |
| Cas12a | Acidaminococcussp. | - | Thymine-rich PAM | DNA | 5ʹofPAM | [61] |
| Cas13 | Lb | 28 | Non-Gnucleotideatthe3ʹproto-spacerflankingsite (PFS) | ssRNA | NA | [61] |
| Cas14 | Uncultivatedarchaea | NA | NA | ssDNA | NA | [61] |
| Name | Cas | CRISPR/Cas | PAM | PAM location | Resources | Reference |
|---|---|---|---|---|---|---|
| SpRY SpG Cas9-NRNH HypaCas9 evoCas9 Sniper-Cas9 xCas9 SpCas9-NG eSpCas9 SpCas9-HF SaCas9-KKH Modified SpCas9 FnCas9variant SpCas9 SaCas9 FnCas9 NmCas9 CjCas9 St1Cas9 St1Cas9 FnCas12a LbCas12a AsCas12a LsCas13# Cas14 |
Cas9 Cas9 Cas9 Cas9 Cas9 Cas9 Cas9 Cas9 Cas9 Cas9 Cas9 Cas9 Cas9 Cas9 Cas9 Cas9 Cas9 Cas9 Cas9 Cas9 Cas12a(cpf1) Cas12a(cpf1) Cas12a(cpf1) Cas13(C2c2) Cas14 |
Type II Type II Type II Type II Type II Type II Type II Type II Type II Type II Type II Type II Type II Type II Type II Type II Type II Type II Type II Type II Type II Type II Type II TypeVI NA |
NRN or NYN NGN NRNH NGG NGG NGG NG NG NGG NGG NNNRRT NGA or NAG YG NGG NNGRRT NGG NNNNGATT NNNNRYAC NNAGAAW NGGNG TTTN or YTN TTTV TTTV NA NA |
3′ 3′ 3′ 3′ 3′ 3′ 3′ 3′ 3′ 3′ 3′ 3′ 3′ 3′ 3′ 3′ 3′ 3′ 3′ 3′ 5′ 5′ 5′ NA NA. |
Engineered SpCas9 Engineered SpCas9 Engineered SpCas9 Mutated SpCas9-HF Mutated SpCas9 Engineered SpCas9 Engineered SpCas9 Engineered SpCas9 Engineered SpCas9 Engineered SpCas9 Engineered SaCas9 Engineered SpCas9 Modified FnCas9 Streptococcus pyogenes Streptococcus aureus Francisella Novicida Neisseria meningitidis Campylobacter jejuni Streptococcusthermophilus Streptococcusthermophilus Francisella novicida Lachnospiracea bacterium Acidaminococcus sp. Leptotrichia shahii Archaea |
[31] [31] [31] [31] [31] [31] [31] [31] [31] [31] [31] [31] [31] [31] [31] [31] [31] [31] [31] [31] [31] [31] [31] [31] [31] |
| Parameter | Cas9 | Cas12 | Cas13 | Cas14 |
|---|---|---|---|---|
|
Size of Protein (Amino Acid) |
~1000-1600 | ~1300 | ~1400 | ~400-700 |
| Target | DNA | DNA | RNA | DNA |
| RNA | Two RNA molecules |
Single RNA molecules |
Two RNA molecules |
Single RNA molecules |
| Nuclease Site | 2 nuclease domains HNH and RuVc | Single nuclease RucV-Nuc |
Target RNA domain HEPN |
DNA binding domain RuVc |
| Pattern of cut | Blunt | Sticky-ended | Degraded | NA |
| Spacer Size | 16-20nt | 16-25nt | 25-35nt | NA |
| Protospacer restriction | PAM | PAM | PFS | PAM |
| Single guide molecular size (Nucleotides, nt) | 17-24nt | 42-44nt | -64nt | -140nt |
|
Non-specifically cut nucleic acids (DNA or RNA) |
DNA(SS) | DNA(SS) | RNA(SS) | DNA(SS) |
| Name of the crops | Types of CRISPR- Cas | Resistance to stress | Reference |
|---|---|---|---|
| Rice | Cas 9 | Drought tolerance, Pest resistance Salinity tolerance |
[62] |
| Cas 12 | Salinity Tolerance | [62] | |
| Cas13 | Enhancing resistance to specific stress conditions | [62] | |
| Wheat | Cas 9 | Drought tolerance, Disease resistance Salinity tolerance, Abiotic Stress |
[62] |
| Cas 12 | Salt tolerance, Drought tolerance Pest resistance |
[62] | |
| Cas 13 | It is used to target genes related to specific stress response | [62] | |
| Maize | Cas 9 | Drought tolerance, Disease resistance | [62] |
| Cas 12 | Pest resistance | [62] | |
| Cas 13 | To enhance stress resistance. | [62] | |
| Arabidopsis | Cas 9 | Drought tolerance, Salt tolerance Stress-related traits. |
[62] |
| Cas 12 | Drought tolerance, Disease resistance. | [62] | |
| Cas 13 | Potential for RNA-based genome editing and may be applied to enhance stress resistance | [62] | |
| Cotton | Cas 9 | Pest resistance | [62] |
| Cas 12 | Disease Resistance | [62] | |
| Cas 13 | RNA Virus Resistance | [62] | |
| Soybean | Cas 9 | Drought tolerance, Pest resistance Disease resistance. |
[62] |
| Cas 12 | Drought tolerance, Disease resistance. | [62] | |
| Cas 13 | RNA Virus Resistance | [62] | |
| Tomato | Cas 9 | Drought tolerance, Pest resistance Disease resistance. |
[62] |
| Cas 12 | Drought tolerance | [62] | |
| Cas 13 | RNA Virus Resistance | [62] | |
| Potato | Cas 9 | Drought tolerance,Pest resistance Disease resistance. |
[62] |
| Cas 12 | Disease resistance | [62] | |
| Cas 13 | The potential for RNA-based genome editing, and may be applied to enhance stress resistance | [62] | |
| Citrus | Cas 9 | Disease resistance | [62] |
| Cas 12 | Disease resistance, Drought tolerance. | [62] | |
| Cas 13 | It holds potential for RNA-based genome editing, which may be applied to enhance stress resistance | [62] | |
| Grape | Cas 9 | Disease resistance | [62] |
| Cas 12 | Disease resistance and improving grape quality traits. | [62] | |
| Cas 13 | Enhance stress resistance and control grapevine diseases. | [62] |
| Stress | Crop | The name of the target gene | References |
|---|---|---|---|
| Salinity | Rice (Oryza sativa) | BASIC HELIX-LOOP-HELIX 024 (OsbHLH024) | [39] |
| Rice (Oryza sativa) | RESPONSE REGULAT ORS 22 (OsRR22) | [40] | |
| Rice (Oryza sativa) | RELATED TO ABI3/VP1 2 (OsRAV2) | [62] | |
| Rice (Oryza sativa) | DROUGHT AND SALT TOLERANCE (OsDST) | [41] | |
| Rice (Oryza sativa) | NAM, ATAF and CUC 041 (OsNAC041) | [51] | |
| Rice (Oryza sativa) | OsmiR535 | [42] | |
| Barley (Hordeum vulgare) | INOSITOLTRISPHOSPHATE 5/6 KINASES 1 (HvITPK1) | [43] | |
| Tomato (Solanum lycopersicum) | HYBRID PROLINE-RICH PROTEIN 1 (SlHyPRP1) | [62] | |
| Tomato (Solanum lycopersicum) | Auxin Response Factor 4 (SlARF4) | [62] | |
| Drought | Rice (Oryza sativa) | ENHANCED RESPONSE TO ABA1 (OsERA1) | [44] |
| Rice (Oryza sativa) | OsDST | [41] | |
| Rice (Oryza sativa) | PYRABACTIN RESISTANCE-LIKE 9 (OsPYL9) | [45] | |
| Rice (Oryza sativa) | SEMI-ROLLED LEAF 1 (SRL1) and SEMI-ROLLED LEAF 2 (SRL2) | [47] | |
| Maize (Zea mays) | AUXIN-REGULATED GENE INVOLVED IN ORGAN SIZE 8 (ZmARGOS8) | [62] | |
| Wheat (Triticum aestivum) | DEHYDRATION RESPONSIVE ELEMENT BINDING PROTEIN 2 (TaDREB2) | [23] | |
| Wheat (Triticum aestivum) | ETHYLENE-RESPONSE FACTOR 3 (TaERF3) | [23] | |
| Tomato (Solanum lycopersicum) | GA-INSENSITIVE DWARF1 1 (SlGID1) | [62] | |
| Tomato (Solanum lycopersicum) | LATERAL ORGAN BOUNDARIES DOMAIN 40 (SlLBD40) | [24] | |
| Arsenic Caesium | Rice (Oryza sativa) | HIGH-AFFINITY POTASSIUM TRANSPORTER 1 (OSHAK1) | [62] |
| Rice (Oryza sativa) | ARSENITE-RESPONSIVE MYB1 (OsARM1) | [62] | |
| Low temperature | Rice (Oryza sativa) | PIN-FORMED 5b (OsPIN5b) | [62] |
| Rice (Oryza sativa) | GRAIN SIZE (GS3) | [62] | |
| Rice (Oryza sativa) | V-MYB AVIAN MYELOBLASTOSIS VIRAL ONCOGENE HOMOLOG 30(OsMYB30) | [62] | |
| High temperature | Rice (Oryza sativa) | PYRABACTIN RESISTANCE-LIKE 1/4/6 (OsPYL1/4/6) | [62] |
| Tomato (Solanum lycopersicum) | MITOGEN-ACTIVATED PROTEIN KINASES 3 (SlMAPK3) | [62] | |
| Cadmium | Rice (Oryza sativa) | NATURAL RESISTANCE-ASSOCIATED MACROPHAGE PROTEIN 5(OsNRAMP5) | [56] |
| Rice (Oryza sativa) | LOW-AFFINITY CATION TRANSPORTER 1 (OsLCT1) | [62] | |
| Rice (Oryza sativa) | NATURAL RESISTANCE-ASSOCIATED MACROPHAGE PROTEIN 1(OsNRAMP1) | [56] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




