Submitted:
01 April 2024
Posted:
02 April 2024
You are already at the latest version
Abstract
Keywords:
Introduction
Protein Language Models: Harnessing Evolutionary Information for Protein Design
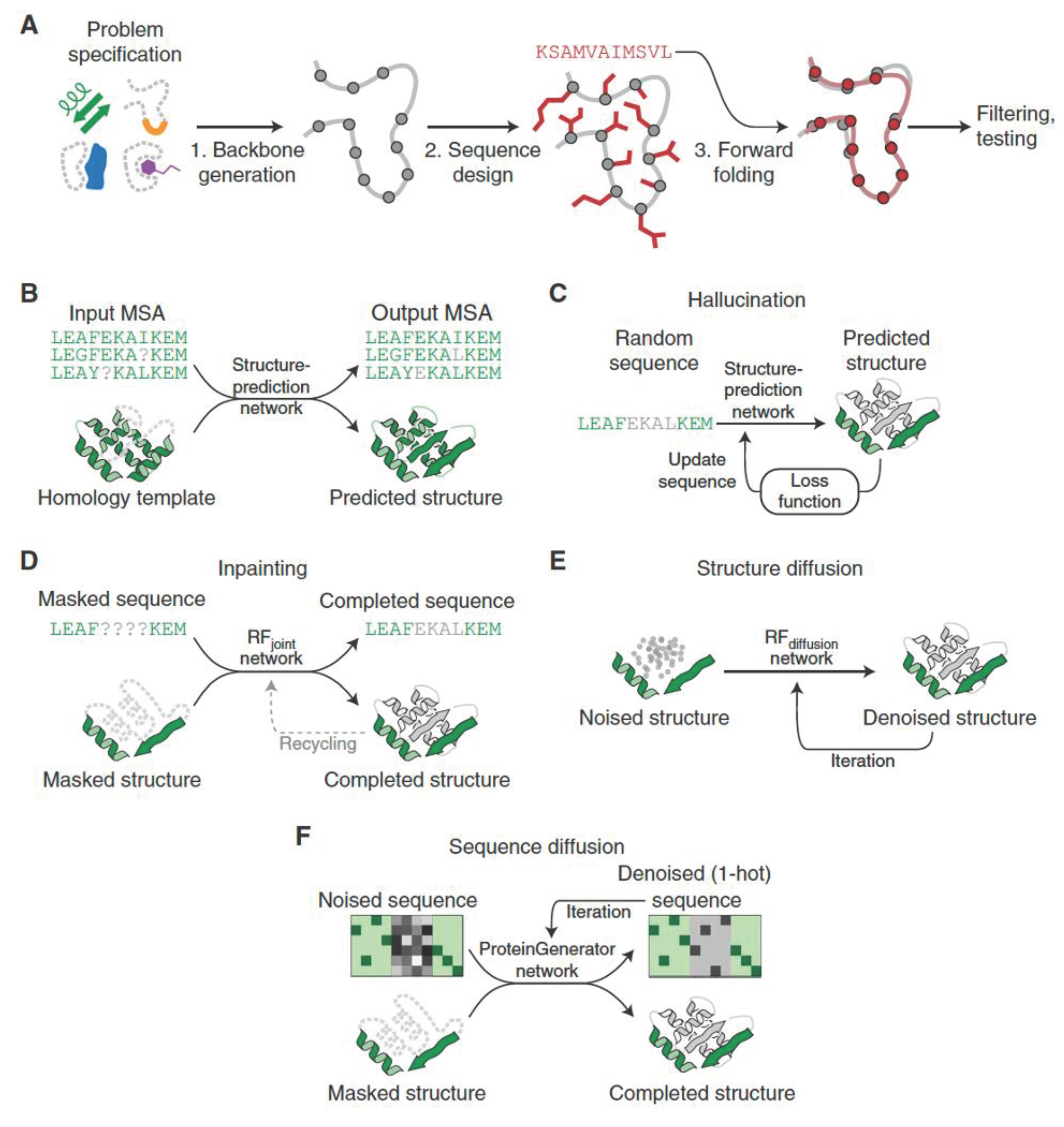
De Novo Protein Design Workflow Using Deep Learning
Structure-Prediction Neural Networks for Protein Design
The RFdiffusion Approach
Generalized Biomolecular Modeling and Design with Rosettafold All-Atom
Applications and Successes of Ai-Driven Protein Design
Challenges and Future Directions
References
- Anishchenko, I. et al. (2021) ‘De novo protein design by deep network hallucination’, Nature, 600(7889), pp. 547–552. [CrossRef]
- Baek, M. et al. (2021) ‘Accurate prediction of protein structures and interactions using a three-track neural network’, Science, 373(6557), pp. 871–876. [CrossRef]
- Cao, L. et al. (2022) ‘Design of protein-binding proteins from the target structure alone’, Nature, 605(7910), pp. 551–560. [CrossRef]
- Chothia, C. (1984) ‘Principles that determine the structure of proteins’, Annual review of biochemistry, 53, pp. 537–572. [CrossRef]
- Chu, A.E., Lu, T. and Huang, P.-S. (2024) ‘Sparks of function by de novo protein design’, Nature biotechnology, 42(2), pp. 203–215. [CrossRef]
- Dou, J. et al. (2018) ‘De novo design of a fluorescence-activating β-barrel’, Nature, 561(7724), pp. 485–491. [CrossRef]
- Feig, M. and Sugita, Y. (2019) ‘Whole-Cell Models and Simulations in Molecular Detail’, Annual review of cell and developmental biology, 35, pp. 191–211. [CrossRef]
- Ferruz, N. et al. (2023) ‘From sequence to function through structure: Deep learning for protein design’, Computational and structural biotechnology journal, 21, pp. 238–250. [CrossRef]
- Ferruz, N., Schmidt, S. and Höcker, B. (2022) ‘ProtGPT2 is a deep unsupervised language model for protein design’, Nature communications, 13(1), p. 4348. [CrossRef]
- Gainza, P. et al. (2020) ‘Deciphering interaction fingerprints from protein molecular surfaces using geometric deep learning’, Nature methods, 17(2), pp. 184–192. [CrossRef]
- Gainza, P. et al. (2023) ‘De novo design of protein interactions with learned surface fingerprints’, Nature, 617(7959), pp. 176–184. [CrossRef]
- Glasscock, C.J. et al. (2023) ‘Computational design of sequence-specific DNA-binding proteins’, bioRxiv : the preprint server for biology [Preprint]. [CrossRef]
- Hie, B.L. et al. (2024) ‘Efficient evolution of human antibodies from general protein language models’, Nature biotechnology, 42(2), pp. 275–283. [CrossRef]
- Hsu, C., Fannjiang, C. and Listgarten, J. (2024) ‘Generative models for protein structures and sequences’, Nature biotechnology, 42(2), pp. 196–199. [CrossRef]
- Huang, P.-S., Boyken, S.E. and Baker, D. (2016) ‘The coming of age of de novo protein design’, Nature, 537(7620), pp. 320–327. [CrossRef]
- Jiang, L. et al. (2008) ‘De novo computational design of retro-aldol enzymes’, Science, 319(5868), pp. 1387–1391. [CrossRef]
- Jumper, J. et al. (2021) ‘Highly accurate protein structure prediction with AlphaFold’, Nature, 596(7873), pp. 583–589. [CrossRef]
- Korendovych, I.V. and DeGrado, W.F. (2020) ‘protein design, a retrospective’, Quarterly reviews of biophysics, 53, p. e3. [CrossRef]
- Krishna, R. et al. (2024) ‘Generalized biomolecular modeling and design with RoseTTAFold All-Atom’, Science, p. eadl2528. [CrossRef]
- Li, Y. et al. (2023) ‘Denoising Diffusion Probabilistic Models and Transfer Learning for citrus disease diagnosis’, Frontiers in plant science, 14, p. 1267810. [CrossRef]
- Lu, H. et al. (2022) ‘Machine learning-aided engineering of hydrolases for PET depolymerization’, Nature, 604(7907), pp. 662–667. [CrossRef]
- Madani, A. et al. (2023) ‘Large language models generate functional protein sequences across diverse families’, Nature biotechnology, 41(8), pp. 1099–1106. [CrossRef]
- Martínez-Mauricio, K.L., García-Jacas, C.R. and Cordoves-Delgado, G. (2024) ‘Examining evolutionary scale modeling-derived different-dimensional embeddings in the antimicrobial peptide classification through a KNIME workflow’, Protein science: a publication of the Protein Society, 33(4), p. e4928. [CrossRef]
- Notin, P. et al. (2024) ‘Machine learning for functional protein design’, Nature biotechnology, 42(2), pp. 216–228. [CrossRef]
- Papaleo, E. et al. (2016) ‘The Role of Protein Loops and Linkers in Conformational Dynamics and Allostery’, Chemical reviews, 116(11), pp. 6391–6423. [CrossRef]
- Polizzi, N.F. and DeGrado, W.F. (2020) ‘A defined structural unit enables de novo design of small-molecule-binding proteins’, Science, 369(6508), pp. 1227–1233. [CrossRef]
- Rives, A. et al. (2021) ‘Biological structure and function emerge from scaling unsupervised learning to 250 million protein sequences’, Proceedings of the National Academy of Sciences of the United States of America, 118(15). [CrossRef]
- Rocklin, G.J. et al. (2017) ‘Global analysis of protein folding using massively parallel design, synthesis, and testing’, Science, 357(6347), pp. 168–175. [CrossRef]
- Schubach, M. et al. (2024) ‘CADD v1.7: using protein language models, regulatory CNNs and other nucleotide-level scores to improve genome-wide variant predictions’, Nucleic acids research, 52(D1), pp. D1143–D1154. [CrossRef]
- Scott, A.J. et al. (2021) ‘Constructing ion channels from water-soluble α-helical barrels’, Nature chemistry, 13(7), pp. 643–650. [CrossRef]
- Sesterhenn, F. et al. (2020) ‘De novo protein design enables the precise induction of RSV-neutralizing antibodies’, Science, 368(6492). [CrossRef]
- Singer, J.M. et al. (2022) ‘Large-scale design and refinement of stable proteins using sequence-only models’, PloS one, 17(3), p. e0265020. [CrossRef]
- Trippe, B.L. et al (2022) ‘Diffusion probabilistic modeling of protein backbones in 3D for the motif-scaffolding problem’, arXiv, 2206.04119. [CrossRef]
- Walls, A.C. et al. (2020) ‘Elicitation of Potent Neutralizing Antibody Responses by Designed Protein Nanoparticle Vaccines for SARS-CoV-2’, Cell, 183(5), pp. 1367–1382.e17. [CrossRef]
- Wang, J. et al. (2022) ‘Scaffolding protein functional sites using deep learning’, Science, 377(6604), pp. 387–394. [CrossRef]
- Watson, J.L. et al. (2023) ‘De novo design of protein structure and function with RFdiffusion’, Nature, 620(7976), pp. 1089–1100. [CrossRef]
- Woolfson, D.N. (2021) ‘A Brief History of De Novo Protein Design: Minimal, Rational, and Computational’, Journal of molecular biology, 433(20), p. 167160. [CrossRef]
- Wu, K.E. et al. (2024) ‘Protein structure generation via folding diffusion’, Nature communications, 15(1), p. 1059. [CrossRef]
- Xie, K. et al. (2022) ‘Inpainting the metal artifact region in MRI images by using generative adversarial networks with gated convolution’, Medical physics, 49(10), pp. 6424–6438. [CrossRef]
- Yang, K.K., Wu, Z. and Arnold, F.H. (2019) ‘Machine-learning-guided directed evolution for protein engineering’, Nature methods, 16(8), pp. 687–694. [CrossRef]
- Zhang, Y. et al. (2023) ‘Attention is all you need: utilizing attention in AI-enabled drug discovery’, Briefings in bioinformatics, 25(1). [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



