Submitted:
12 April 2024
Posted:
16 April 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Reagents and Antibodies
2.2. Animals
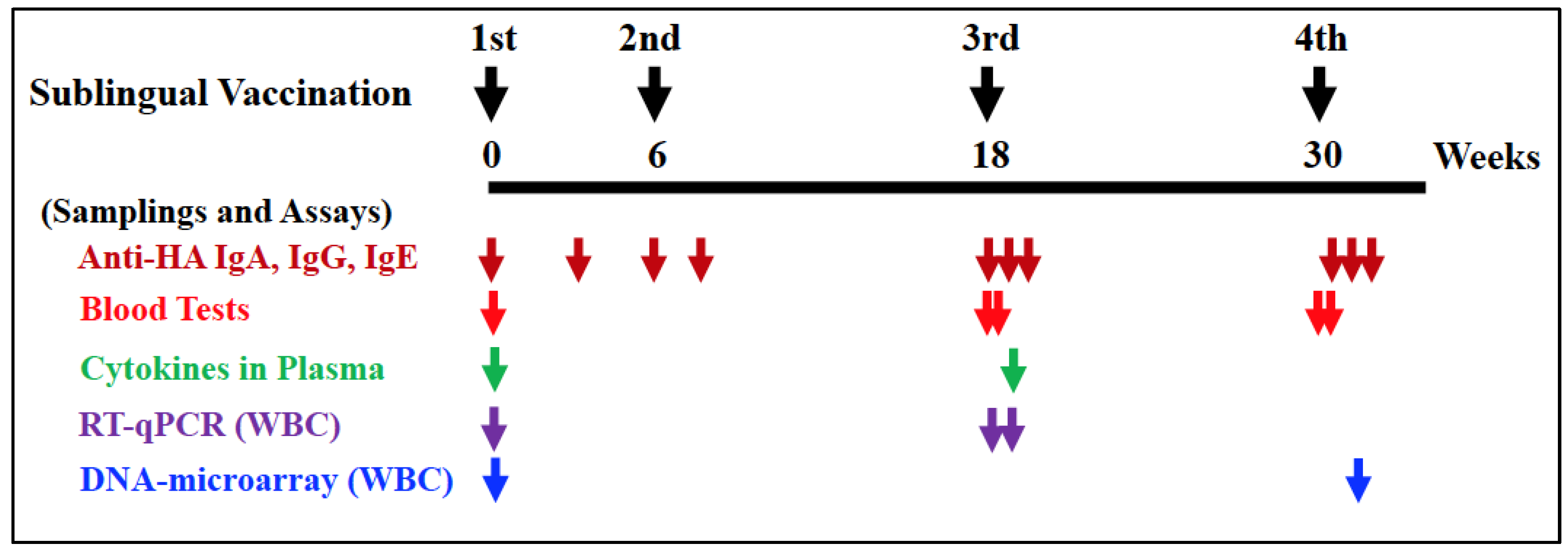
2.3. Vaccination and Sampling
2.4. ELISA for HA-Specific Antibodies
2.5. Blood Tests
2.6. Cytokine Assays
2.7. WBC Isolation
2.8. RNA Isolation
2.9. Gene Expression Analyses Using Quantitative Reverse Transcription PCR
2.10. DNA Microarray Analyses
2.11. Bioinformatic Analysis of Microarray Data
3. Results
3.1. The Complete Blood Count, the Biochemical Blood Test, and CRP
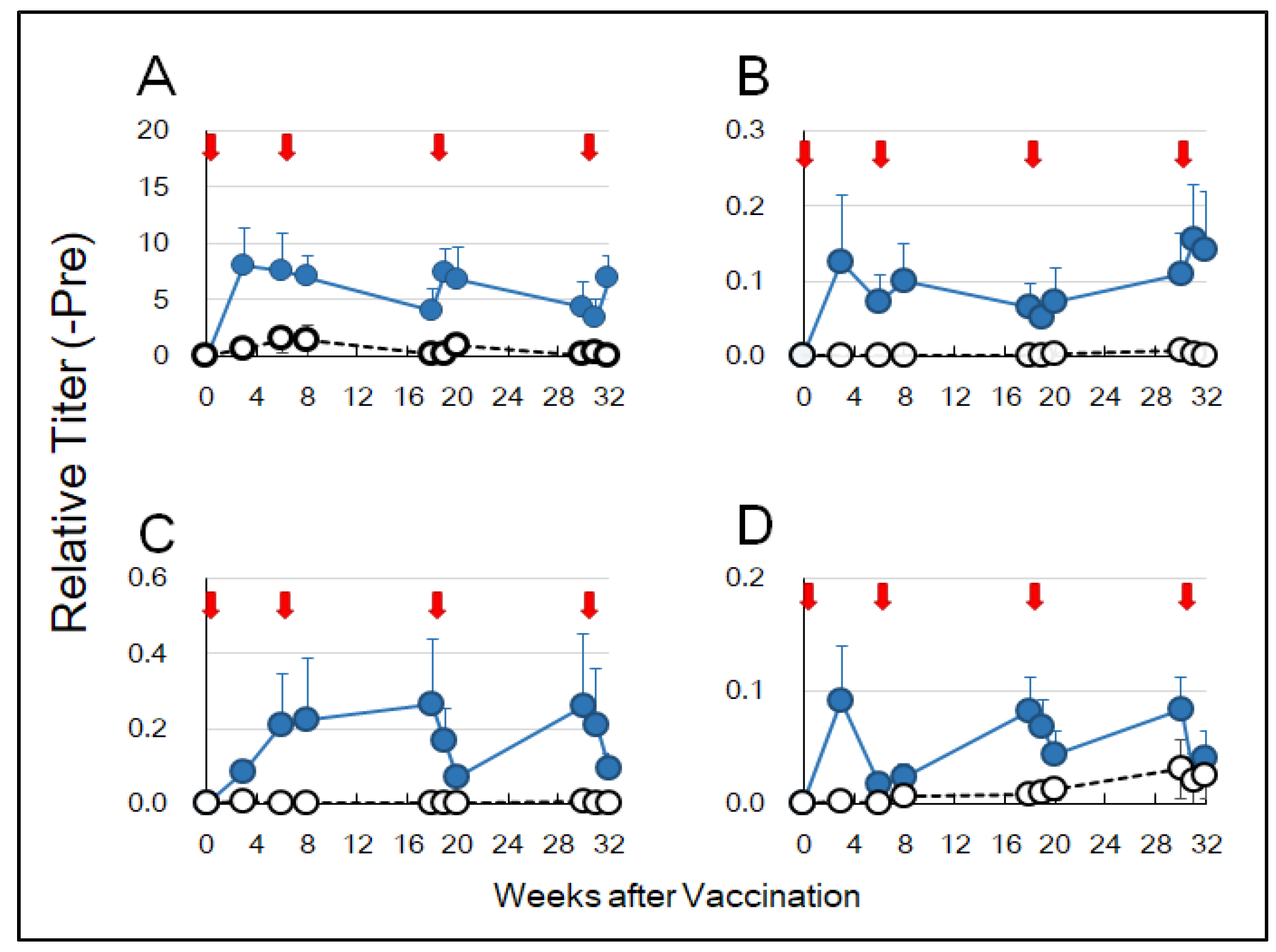
3.2. HA-Specific Antibodies
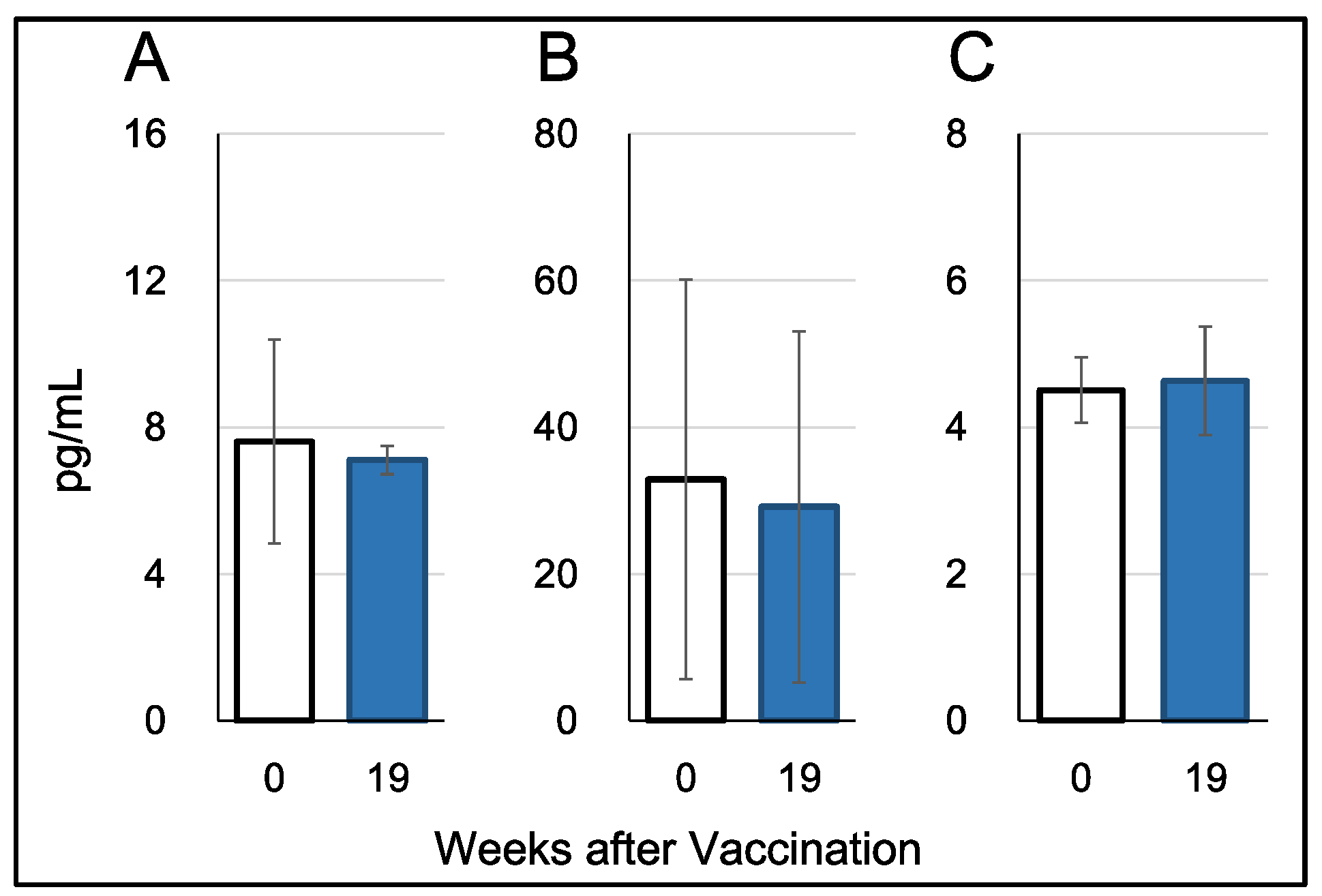
3.3. Production of Inflammation-Associating Cytokines
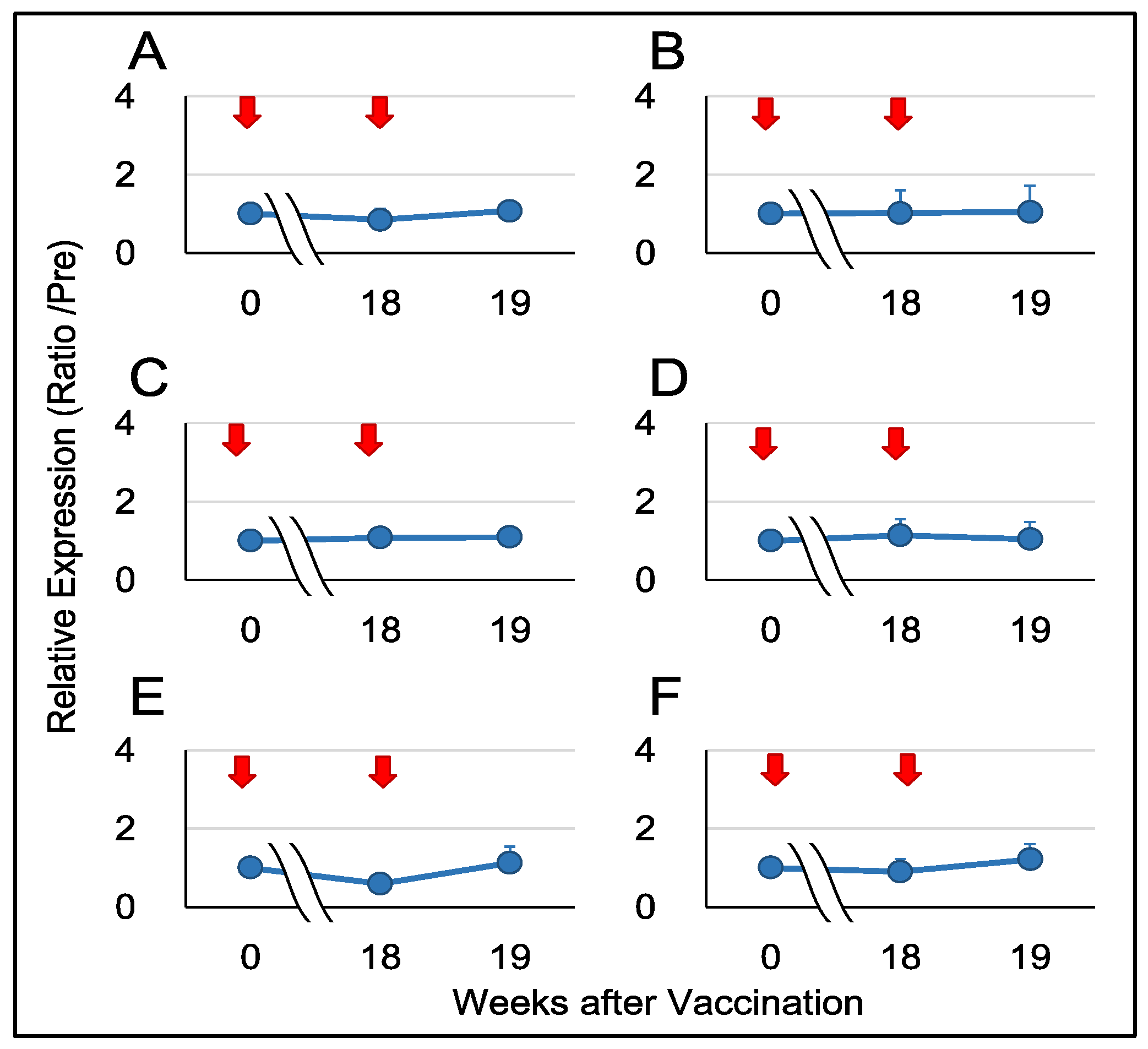
3.4. Gene Expression of Immuno-Proinflammatory Factors
3.5. Microarray Analyses and Bioinformatics
3.5.1. Upregulated Genes
| Gene symbol | Fold change | Product; Description; Function [Reference] | Expected effect* |
|---|---|---|---|
| CLEC4G | 2.2 | C-type lectin; Receptor of PAMP [16]; Activation of inflammatory reactions [17]; Transcript variants [18] | ↑ |
| PF4V1 | 2.2 | Also known as CXCL4L1; Chemoattractant of T and NK cells [19] | ↑ |
| KLF1 | 2.1 | Zn-finger transcription factor; Transcription of β-globin gene; Upregulation of CD274 in Tregs [20] | ↓ |
| OLFM1 | 2.1 | Nervous system; Inhibition of non-canonical NF-κB pathway in CRC [21] | ↓ |
| GNG11 | 2.0 | G protein gamma family; Downregulated in splenic marginal zone lymphomas [22] | ↓ |
3.5.2. Downregulated Genes
| Gene symbol | Fold change | Product; Description; Function [Reference] | Expected effect* |
|---|---|---|---|
| NEURL1B | 0.45 | Ubiquitin protein ligase; Development of the nervous system [23]; Downregulated in CRC tissues [24]; Downregulated by Poly(I:C) adjuvant [8] | ↑ |
| CHST15 | 0.45 | Sulfotransferase; Expressed in B-cell lineage; Upregulation of RAG1 [25] | ↓ |
| MOB3A | 0.48 | Protein kinase activator; Inhibition of GC through Hippo pathway [26] | ↑ |
| ANXA6 | 0.48 | Expressed in T cells; Component of T-cell plasma membrane; Stimulation of helper T cells [27] | ↓ |
| DNAJA3 | 0.48 | Mitochondrial HSP; Stimulation of ATPase activity of Hsp70; Activation of NF-κB [28] | ↓ |
| HSPH1 | 0.49 | HSP; Replacement of Hsp70-bound ADP with ATP; Holdase and disaggregase activity [29]; Stimulation of NF-κB signaling in activated B-cell diffuse large B-cell lymphoma [30] | ↓ |
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- World Health Organization. (2023) Influenza (Seasonal) 2023. Available from: https://www.who.int/news-room/fact-sheets/detail/influenza-(seasonal).
- World Health Organization. (2022) Vaccines against influenza: WHO position paper – May 2022. Available from: https://www.who.int/publications/i/item/who-wer9719.
- L. Colombo, S. Hadigal, J. Nauta, et al. (2024) Influvac Tetra: clinical experience on safety, efficacy, and immunogenicity. Expert Rev Vaccines. 23:88-101. [CrossRef]
- R. Mudgal, S. Nehul, S. Tomar. (2020) Prospects for mucosal vaccine: shutting the door on SARS-CoV-2. Human Vaccine Immnunother. 16:2921–2931. [CrossRef]
- F. Lemiale, W.P. Kong, L.M. Akyürek, et al. (2003) Enhanced mucosal immunoglobulin A response of intranasal adenoviral vector human immunodeficiency virus vaccine and localization in the central nervous system. J Virol. 77:10078-10087. [CrossRef]
- E. Sasaki, H. Momose, Y. Hiradate, et al. (2018) Establishment of a novel safety assessment method for vaccine adjuvant development. Vaccine, 36:7112–7118. [CrossRef]
- T. Yamamoto, M. Tanji, F. Mitsunaga, et al. (2023) SARS-CoV-2 sublingual vaccine with RBD antigen and poly(I:C) adjuvant: Preclinical study in cynomolgus macaques. Biol Methods Protoc. bpad 017. [CrossRef]
- T. Yamamoto, F. Mitsunaga, K. Wasaki, et al. (2023) Mechanism Underlying the Immune Responses of a Sublingual Vaccine for SARS-CoV-2 with RBD Antigen and Adjuvant, Poly(I:C) or AddaS03, in Non-human Primates. Arch. Microbiol. Immunol. 7:150-164. [CrossRef]
- Iwasaki, S.B. Omer. (2020) Why and How Vaccines Work. Cell. 15: 290-295. [CrossRef]
- T. Zhao, Y. Cai, Y. Jiang, et al. (2023) Vaccine adjuvants: mechanisms and platforms. Signal Transduct Target Ther. 8:283. [CrossRef]
- Ainai, E. van Riet, R. Ito, et al. (2020) Human immune responses elicited by an intranasal inactivated H5 influenza vaccine. Microbiol Immunol. 64:313-325. [CrossRef]
- R.A. Hoffman, P.C. Kung, W.P. Hansen, et al. (1980) Simple and rapid measurement of human T lymphocytes and their subclasses in peripheral blood. Proc Natl Acad Sci U S A. 77:4914-4917. [CrossRef]
- A.R. Jeong, S. Nakamura, F. Mitsunaga. (2008) Gene expression profile of Th1 and Th2 cytokines and their receptors in human and nonhuman primates. J Med Primatol. 37:290-296. [CrossRef]
- J. Ye, G. Coulouris, I. Zaretskaya, et al. (2012) Primer-BLAST: A Tool to Design Target-Specific Primers for Polymerase Chain Reaction. BMC Bioinformatics. 13:134. [CrossRef]
- B.G. Gabrielsson, L.E. Olofsson, A. Sjögren, et al. (2005) Evaluation of Reference Genes for Studies of Gene Expression in Human Adipose Tissue. Obesity Research. 13:649-652. [CrossRef]
- D. Hoffmann, S. Mereiter, Y.J. Oh, et al. (2021) Identification of lectin receptors for conserved SARS-CoV-2 glycosylation sites. EMBO J. 40:e108375. [CrossRef]
- Q. Lu, J. Liu, S. Zhao, et al. (2021) SARS-CoV-2 exacerbates proinflammatory responses in myeloid cells through C-type lectin receptors and Tweety family member 2. Immunity. 54:1304-1319. [CrossRef]
- Dominguez-Soto, L. Aragoneses-Fenoll, E. Martin-Gayo, et al. (2007) The DC-SIGN-related lectin LSECtin mediates antigen capture and pathogen binding by human myeloid cells. Blood. 109:5337-45. [CrossRef]
- S. Struyf, L. Salogni, M.D. Burdick, et al. (2011) Angiostatic and chemotactic activities of the CXC chemokine CXCL4L1 (platelet factor-4 variant) are mediated by CXCR3. Blood. 117:480-8. [CrossRef]
- S. Teruya, T. Okamura, T. Komai, et al. (2018) Egr2-independent, Klf1-mediated induction of PD-L1 in CD4+ T cells. Sci Rep. 8:7021. [CrossRef]
- W. Shi, Z. Ye, L. Zhuang, et al. (2016) Olfactomedin 1 negatively regulates NF-κB signalling and suppresses the growth and metastasis of colorectal cancer cells. J Pathol. 240:352-365. [CrossRef]
- E. Ruiz-Ballesteros, M. Mollejo, A. Rodriguez, et al. (2005) Splenic marginal zone lymphoma: proposal of new diagnostic and prognostic markers identified after tissue and cDNA microarray analysis. Blood. 106:1831-8. [CrossRef]
- G. Rullinkov, R. Tamme, A. Sarapuu, et al. (2009) Neuralized-2: expression in human and rodents and interaction with Delta-like ligands. Biochem Biophys Res Commun. 389:420-5. [CrossRef]
- J. Liu, Z. Liu, X. Zhang, et al. (2020) Aberrant methylation and microRNA-target regulation are associated with downregulated NEURL1B: a diagnostic and prognostic target in colon cancer. Cancer Cell Int. 20:342. [CrossRef]
- L.K. Verkoczy, P.A. Marsden, N.L. Berinstein (1998) hBRAG, a novel B cell lineage cDNA encoding a type II transmembrane glycoprotein potentially involved in the regulation of recombination activating gene 1 (RAG1). Eur J Immunol. 28:2839-53. [CrossRef]
- L. Dong, Y. Cao, H. Yang, et al. (2023) The hippo kinase MST1 negatively regulates the differentiation of follicular helper T cells. Immunology. 168:511-525. [CrossRef]
- R. Cornely, A.H. Pollock, C. Rentero, et al. (2016) Annexin A6 regulates interleukin-2-mediated T-cell proliferation. Immunol Cell Biol. 94:543-53. [CrossRef]
- Y. Momiuchi, K. Kumada, T. Kuraishi, et al. (2015) The Role of the Phylogenetically Conserved Cochaperone Protein Droj2/DNAJA3 in NF-κB Signaling. J Biol Chem. 290:23816-25. [CrossRef]
- Bracher, J. Verghese (2015) The nucleotide exchange factors of Hsp70 molecular chaperones. Front Mol Biosci. 2:10. [CrossRef]
- Boudesco, E. Verhoeyen, L. Martin, et al. (2018) HSP110 sustains chronic NF-κB signaling in activated B-cell diffuse large B-cell lymphoma through MyD88 stabilization. Blood. 132:510-520. [CrossRef]
- E. Sasaki, H. Momose, Y. Hiradate, et al. (2018) Modeling for influenza vaccines and adjuvants profile for safety prediction system using gene expression profiling and statistical tools. PLoS One. 13: e0191896. [CrossRef]
- E. Sasaki, H. Asanuma, H. Momose, et al. (2020) Immunogenicity and Toxicity of Different Adjuvants Can Be Characterized by Profiling Lung Biomarker Genes After Nasal Immunization. Front Immunol. 11:2171. [CrossRef]
- F. Dotiwala, A.K. Upadhyay (2023) Next Generation Mucosal Vaccine Strategy for Respiratory Pathogens. Vaccines. 11:1585. [CrossRef]
- J. Mestas, C.C. Hughes. (2004) Of mice and not men: differences between mouse and human immunology. J Immunol. 172:2731-2738. [CrossRef]
- J.L. Naour, L. Galluzzi, L. Zitvogel, et al. (2020) Trial watch: TLR3 agonists in cancer therapy. Oncoimmunology. 9:1771143. [CrossRef]
- J. De Waele, T. Verhezen, S. van der Heijden, et al. (2021) A systematic review on poly(I:C) and poly-ICLC in glioblastoma: adjuvants coordinating the unlocking of immunotherapy. J Exp Clin Cancer Res. 40:213. [CrossRef]
- K. Tewari, B.J. Flynn, S.B. Boscardin, et al. (2010) Poly(I:C) is an effective adjuvant for antibody and multi-functional CD4+ T cell responses to Plasmodium falciparum circumsporozoite protein (CSP) and αDEC-CSP in non human primates. Vaccine. 28:7256-7266. [CrossRef]
- J. Fučíková, D. Rožková, H. Ulčová, et al. (2011) Poly I: C-activated dendritic cells that were generated in CellGro for use in cancer immunotherapy trials. J Transl Med. 9:223. [CrossRef]
- H. Kato, O. Takeuchi, S. Sato S, et al. (2006) Differential roles of MDA5 and RIG-I helicases in the recognition of RNA viruses. Nature. 441:101-105. [CrossRef]
- H. Sultan, J. Wu, T. Kumai, et al. (2018) Role of MDA5 and interferon-I in dendritic cells for T cell expansion by anti-tumor peptide vaccines in mice. Cancer Immunol Immunother. 67: 1091-1103. [CrossRef]
- H. Kraan, H. Vrieling, C. Czerkinsky, et al. (2014) Buccal and sublingual vaccine delivery. J Control Release. 190:580-92. [CrossRef]
- L. Mascarell, V. Lombardi, A. Louise, et al. (2008) Oral dendritic cells mediate antigen-specific tolerance by stimulating TH1 and regulatory CD4+ T cells. J Allergy Clin Immunol. 122:603-9.e5. [CrossRef]
- Hervouet, C. Luci, S. Bekri, et al. (2014) Antigen-bearing dendritic cells from the sublingual mucosa recirculate to distant systemic lymphoid organs to prime mucosal CD8 T cells. Mucosal Immunol. 7:280-91. [CrossRef]
- A.L. Paris, E. Colomb, B. Verrier, et al. (2021) Sublingual vaccination and delivery systems. J Control Release. 332:553-562. [CrossRef]
- M.L.B. Hillaire, N.J. Nieuwkoop, A.C.M. Boon, et al. (2013) Binding of DC-SIGN to the hemagglutinin of influenza A viruses supports virus replication in DC-SIGN expressing cells. PLoS One. 8:e56164. [CrossRef]
- L. Tang, J. Yang, W. Liu, et al. (2009) Liver sinusoidal endothelial cell lectin, LSECtin, negatively regulates hepatic T-cell immune response. Gastroenterology. 137:1498-508.e1-5. [CrossRef]
- P. Moingeon, L. Mascarell (2012) Induction of tolerance via the sublingual route: mechanisms and applications. Clin Dev Immunol. 2012:623474. [CrossRef]
- R.G. Hibbert, P. Teriete, G.J. Grundy, et al. (2005) The structure of human CD23 and its interactions with IgE and CD21. J Exp Med. 202:751-60. [CrossRef]
- M. Brandhofer, A. Hoffmann, X. Blanchet, et al. (2022) Heterocomplexes between the atypical chemokine MIF and the CXC-motif chemokine CXCL4L1 regulate inflammation and thrombus formation. Cell Mol Life Sci. 79:512. [CrossRef]
- K. Dutchak, S. Garnett, M. Nicoll, et al. (2022) MOB3A Bypasses BRAF and RAS Oncogene-Induced Senescence by Engaging the Hippo Pathway. Mol Cancer Res. 20:770-781. [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




