Submitted:
28 March 2024
Posted:
29 March 2024
You are already at the latest version
Abstract
Keywords:
Highlights
- Widely-held views that ageing is caused by pathogenic gene function has limited our scope-of-view that an over-activation of wild-type gene function may explain proximate causes of ageing
- Genetic alleles that drive reproductive fitness are preferentially selected for, despite their (hyperfunctional) pleiotropic effects that contribute to ageing pathology later in life
- C. elegans represent a candidate model to explore therapeutic mechanisms to counter pleiotropic expression changes in ageing, but current experimental approaches are limited in their design and should be re-addressed
Introduction
Ageing-induced genetic pleiotropy – an overlooked aspect of healthspan research?
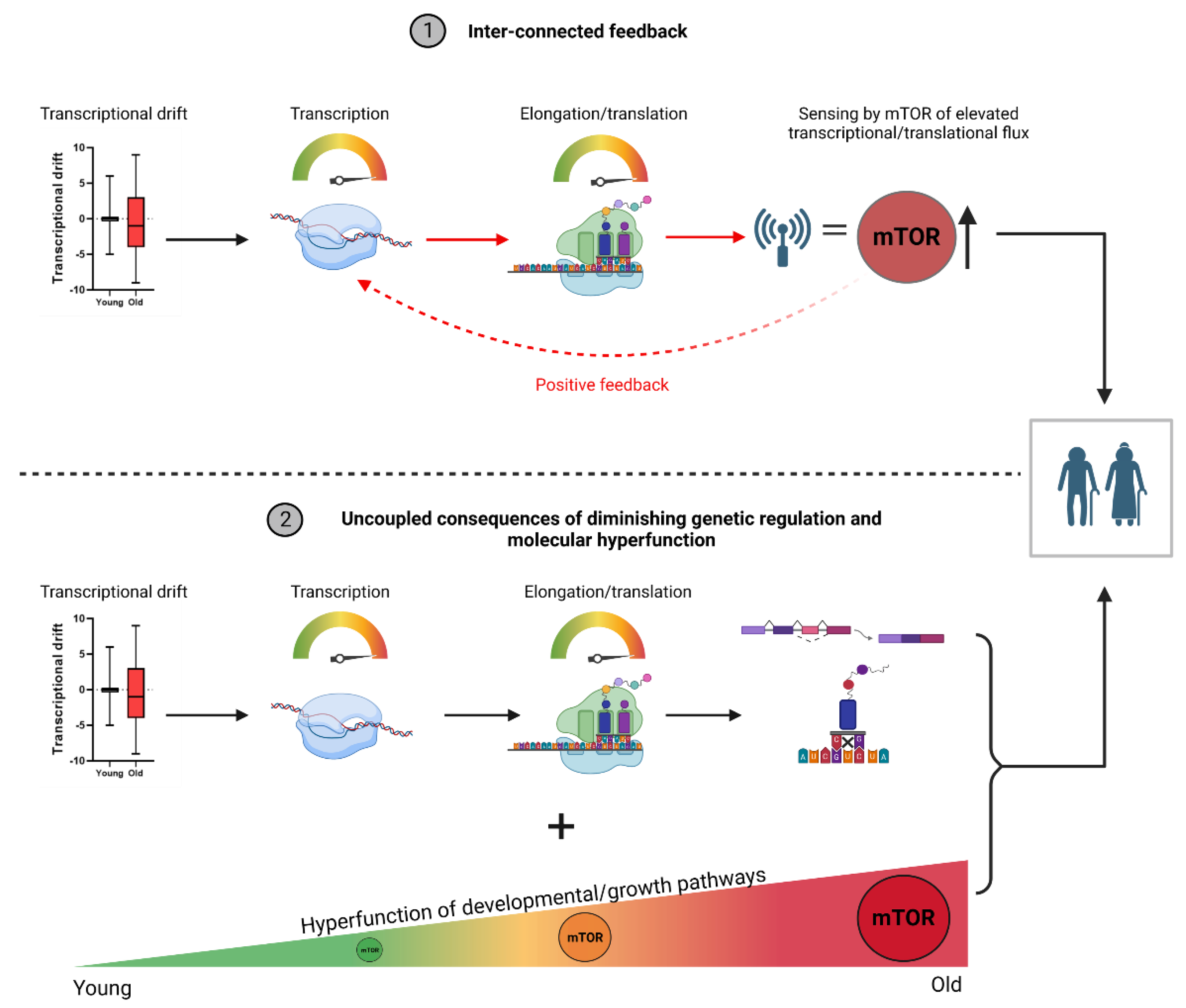
Molecular hyperfunction as a candidate theory of ageing
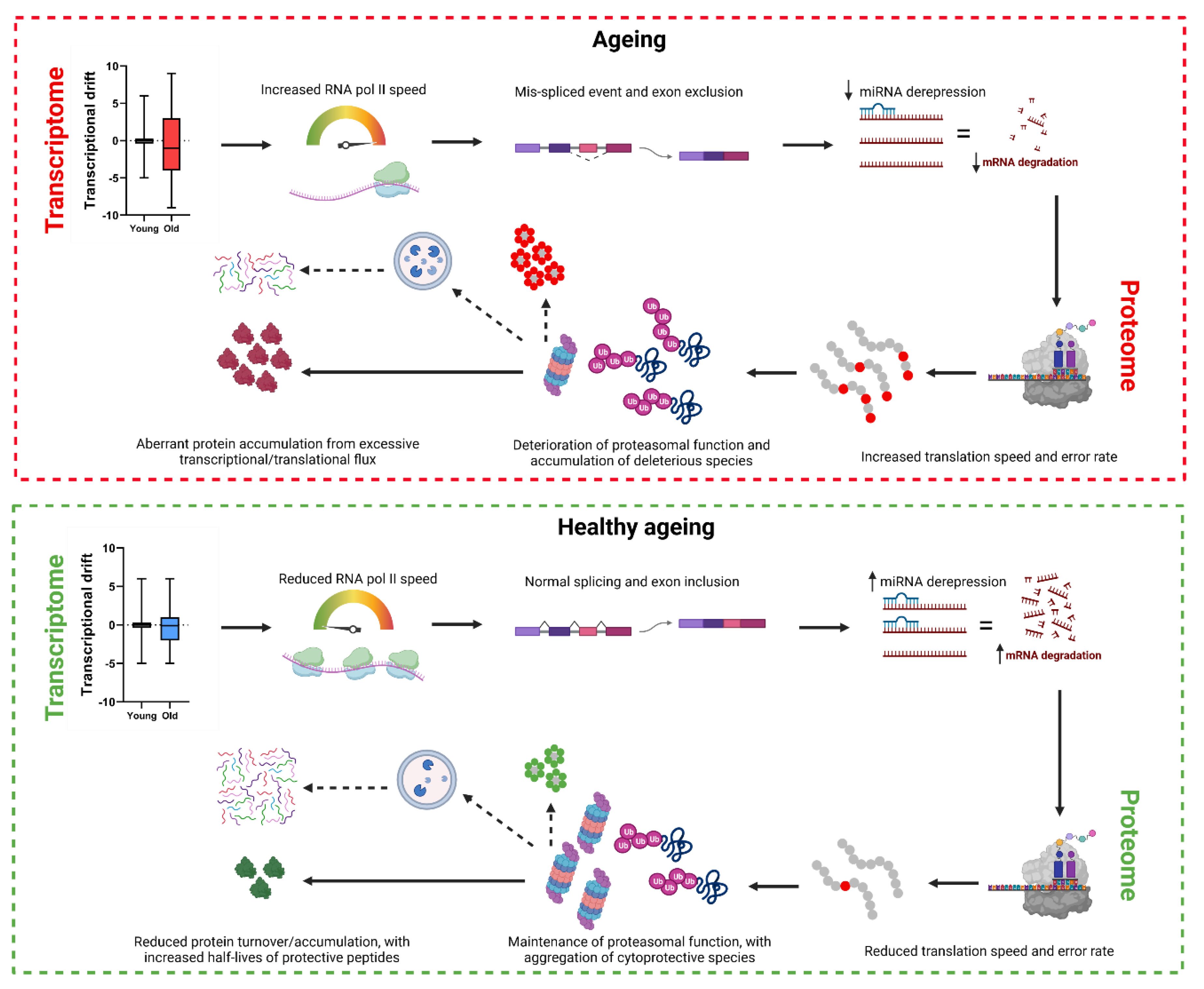
Transcriptomic remodeling towards a hyperfunctional state in ageing
Progressive protein biosynthesis is a feature of the C. elegans proteome
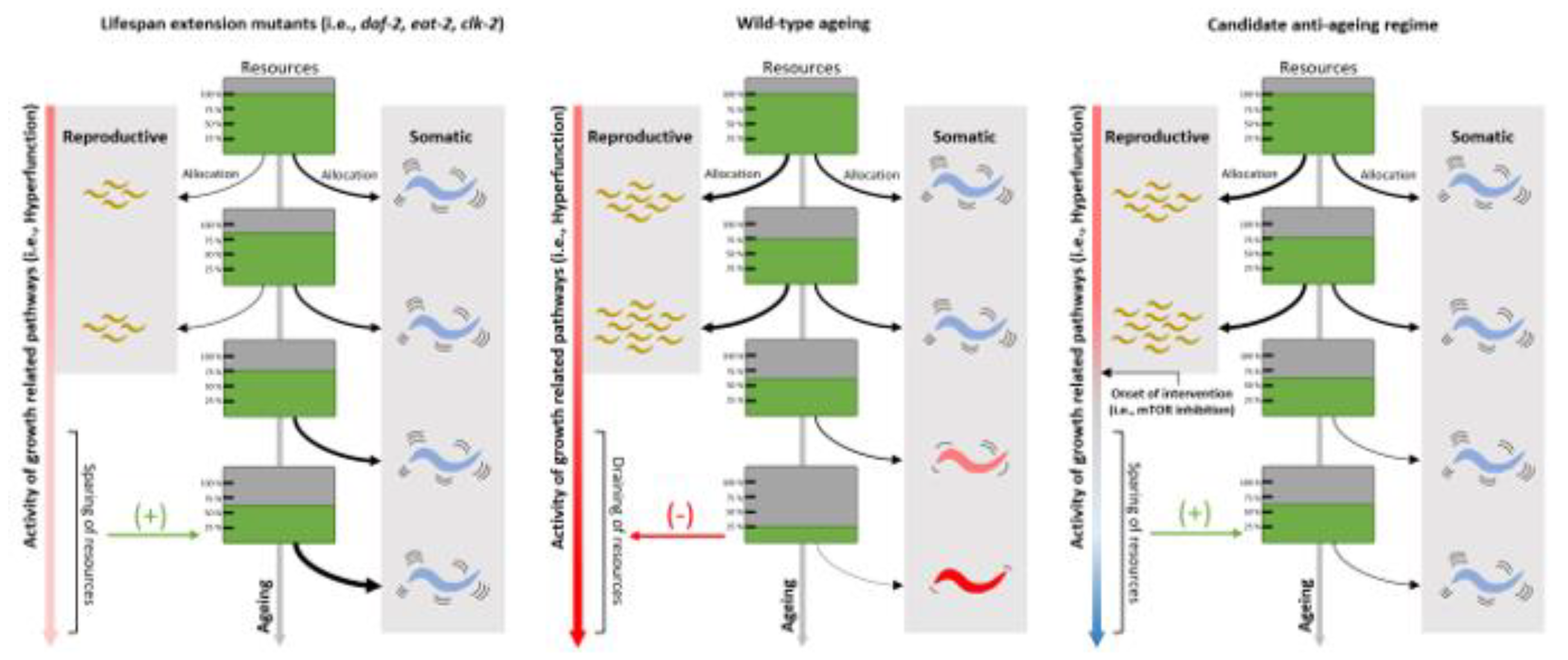
Attenuation of protein accumulation, transcriptional drift and elongation speeds from life-long mutations in the Insulin signalling pathway
Lifespan-extending mutants for healthspan research – a complex story
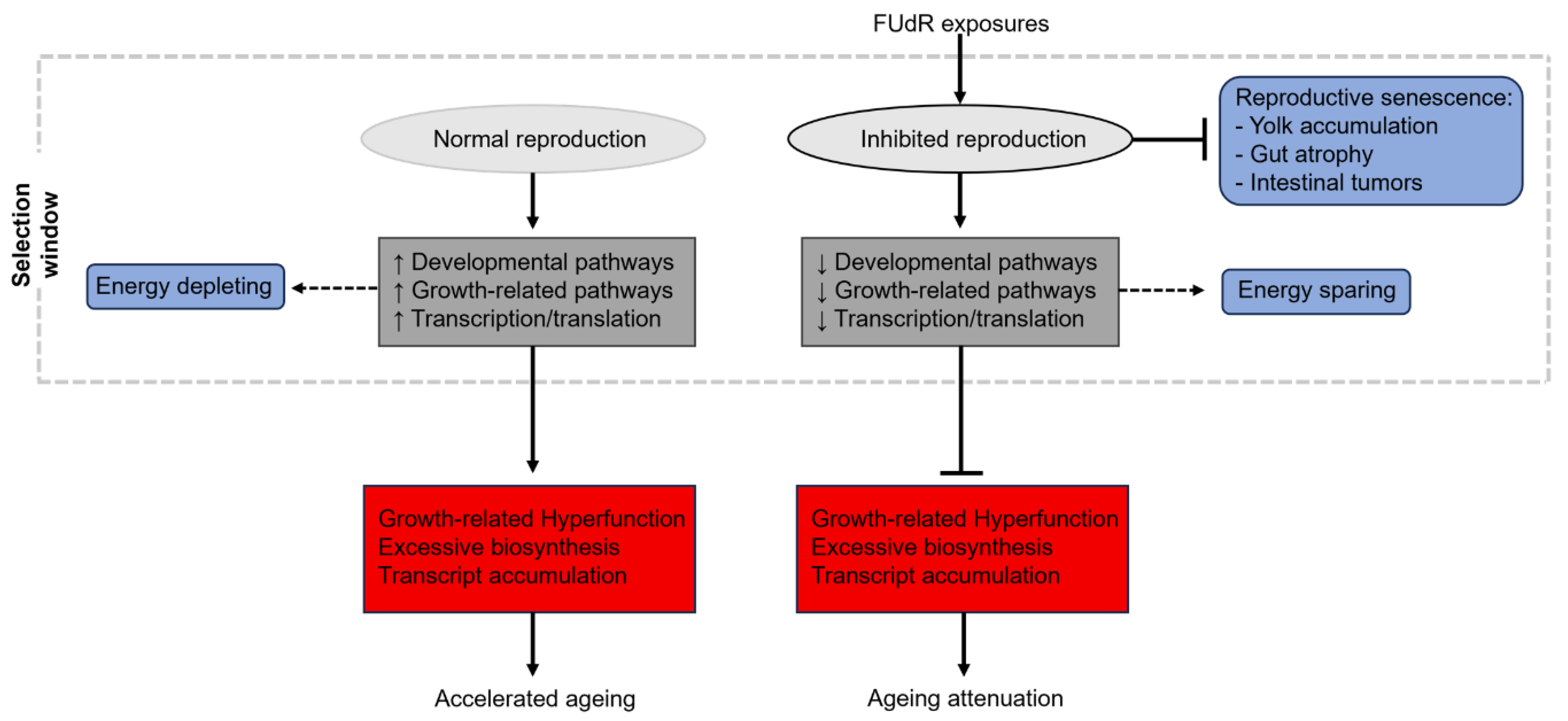
Re-thinking genetic and therapeutic screening in C. elegans for ageing research
Lifespan extension or gero-compression?
Protein-degron systems represent a new candidate approach for antagonistic pleiotropy and hyperfunction research
Next steps: using C. elegans to map the ageing pleiotropome on a systems and functional level
Conclusions and future directions
Acknowledgements
Contributions
Competing interests
References
- Garmany, A., S. Yamada, and A. Terzic, Longevity leap: mind the healthspan gap. NPJ Regen Med 2021, 6, 57. [Google Scholar] [CrossRef] [PubMed]
- Kirkwood, T.B. , Evolution of ageing. Nature 1977, 270, 301–4. [Google Scholar] [CrossRef] [PubMed]
- Harman, D. , Aging: a theory based on free radical and radiation chemistry. J Gerontol 1956, 11, 298–300. [Google Scholar] [CrossRef] [PubMed]
- Williams, G.C. , Pleiotropy, Natural-Selection, and the Evolution of Senescence. Evolution 1957, 11, 398–411. [Google Scholar] [CrossRef]
- de Magalhaes, J.P. and G.M. Church, Genomes optimize reproduction: aging as a consequence of the developmental program. Physiology (Bethesda) 2005, 20, 252–9. [Google Scholar] [PubMed]
- Gems, D. , The hyperfunction theory: An emerging paradigm for the biology of aging. Ageing Res Rev 2022, 74, 101557. [Google Scholar] [CrossRef] [PubMed]
- Blagosklonny, M.V. , Why the disposable soma theory cannot explain why women live longer and why we age. Aging-Us 2010, 2, 884–887. [Google Scholar] [CrossRef] [PubMed]
- Gladyshev, V.N. , The Free Radical Theory of Aging Is Dead. Long Live the Damage Theory! Antioxidants & Redox Signaling 2014, 20, 727–731. [Google Scholar]
- Lemaitre, J.F.; et al. A unified framework for evolutionary genetic and physiological theories of aging. PLoS Biol 2024, 22, e3002513. [Google Scholar] [CrossRef] [PubMed]
- Min, B.; et al. Age-associated bimodal transcriptional drift reduces intergenic disparities in transcription. Aging (Albany NY) 2018, 10, 789–807. [Google Scholar] [CrossRef] [PubMed]
- Rangaraju, S.; et al. Suppression of transcriptional drift extends C. elegans lifespan by postponing the onset of mortality. Elife 2015, 4, e08833. [Google Scholar] [CrossRef] [PubMed]
- Perez-Gomez, A., J. N. Buxbaum, and M. Petrascheck, The aging transcriptome: read between the lines. Current Opinion in Neurobiology 2020, 63, 170–175. [Google Scholar] [CrossRef] [PubMed]
- Blagosklonny, M.V. , The hyperfunction theory of aging: three common misconceptions. Oncoscience 2021, 8, 103–107. [Google Scholar] [CrossRef] [PubMed]
- Blagosklonny, M.V. , Aging and immortality: quasi-programmed senescence and its pharmacologic inhibition. Cell Cycle 2006, 5, 2087–102. [Google Scholar] [CrossRef] [PubMed]
- Blagosklonny, M.V. , Cell senescence, rapamycin and hyperfunction theory of aging. Cell Cycle 2022, 21, 1456–1467. [Google Scholar] [CrossRef] [PubMed]
- Herndon, L.A.; et al. Stochastic and genetic factors influence tissue-specific decline in ageing. Nature 2002, 419, 808–814. [Google Scholar] [CrossRef] [PubMed]
- Narayan, V.; et al. Deep Proteome Analysis Identifies Age-Related Processes in C. elegans. Cell Syst 2016, 3, 144–159. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; et al. Ageing induces tissue-specific transcriptomic changes in Caenorhabditis elegans. EMBO J 2022, 41, e109633. [Google Scholar] [CrossRef] [PubMed]
- Budovskaya, Y.V.; et al. An elt-3/elt-5/elt-6 GATA transcription circuit guides aging in C. elegans. Cell 2008, 134, 291–303. [Google Scholar] [CrossRef] [PubMed]
- de la Guardia, Y.; et al. Run-on of germline apoptosis promotes gonad senescence in C. elegans. Oncotarget 2016, 7, 39082–39096. [Google Scholar] [CrossRef] [PubMed]
- Takemon, Y.; et al. Proteomic and transcriptomic profiling reveal different aspects of aging in the kidney. Elife 2021, 10. [Google Scholar] [CrossRef] [PubMed]
- Kenyon, C.; et al. A C-Elegans Mutant That Lives Twice as Long as Wild-Type. Nature 1993, 366, 461–464. [Google Scholar] [CrossRef] [PubMed]
- Bansal, A.; et al. Uncoupling lifespan and healthspan in Caenorhabditis elegans longevity mutants. Proc Natl Acad Sci U S A 2015, 112, E277–86. [Google Scholar] [CrossRef] [PubMed]
- Boeck, M.E.; et al. The time-resolved transcriptome of C. elegans. Genome Res 2016, 26, 1441–1450. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; et al. A parthenogenetic quasi-program causes teratoma-like tumors during aging in wild-type C. elegans. NPJ Aging Mech Dis 2018, 4, 6. [Google Scholar] [CrossRef] [PubMed]
- Ezcurra, M.; et al. C. elegans Eats Its Own Intestine to Make Yolk Leading to Multiple Senescent Pathologies. Curr Biol 2018, 28, 3352. [Google Scholar] [CrossRef] [PubMed]
- Gangloff, Y.G.; et al. Disruption of the mouse mTOR gene leads to early postimplantation lethality and prohibits embryonic stem cell development. Molecular and Cellular Biology 2004, 24, 9508–9516. [Google Scholar] [CrossRef] [PubMed]
- Vellai, T.; et al. Genetics: influence of TOR kinase on lifespan in C. elegans. Nature 2003, 426, 620. [Google Scholar] [CrossRef] [PubMed]
- Peng, Y.; et al. PI3K/Akt/mTOR Pathway and Its Role in Cancer Therapeutics: Are We Making Headway? Frontiers in Oncology 2022, 12. [Google Scholar] [CrossRef] [PubMed]
- Long, X.; et al. TOR deficiency in C. elegans causes developmental arrest and intestinal atrophy by inhibition of mRNA translation. Curr Biol 2002, 12, 1448–61. [Google Scholar] [CrossRef]
- Abdellatif, M.; et al. Fine-Tuning Cardiac Insulin-Like Growth Factor 1 Receptor Signaling to Promote Health and Longevity. Circulation 2022, 145, 1853–1866. [Google Scholar] [CrossRef] [PubMed]
- Wu, D.; et al. An antagonistic pleiotropic gene regulates the reproduction and longevity tradeoff. Proc Natl Acad Sci U S A 2022, 119, e2120311119. [Google Scholar] [CrossRef] [PubMed]
- Salminen, A. , Aryl hydrocarbon receptor (AhR) reveals evidence of antagonistic pleiotropy in the regulation of the aging process. Cellular and Molecular Life Sciences 2022, 79. [Google Scholar] [CrossRef] [PubMed]
- Lopez-Otin, C.; et al. Hallmarks of aging: An expanding universe. Cell 2023, 186, 243–278. [Google Scholar] [CrossRef]
- Blagosklonny, M.V. , TOR-driven aging Speeding car without brakes. Cell Cycle 2009, 8, 4055–4059. [Google Scholar] [CrossRef] [PubMed]
- de Magalhaes, J.P. , Ageing as a software design flaw. Genome Biology 2023, 24. [Google Scholar] [CrossRef] [PubMed]
- Gems, D. and J.P. de Magalha, The hoverfly and the wasp: A critique of the hallmarks of aging as a paradigm. Ageing Research Reviews 2021, 70. [Google Scholar]
- Chen, D.; et al. Longevity determined by developmental arrest genes in Caenorhabditis elegans. Aging Cell 2007, 6, 525–33. [Google Scholar] [CrossRef] [PubMed]
- Liu, W.Q.; et al. Large-scale across species transcriptomic analysis identifies genetic selection signatures associated with longevity in mammals. Embo Journal 2023, 42. [Google Scholar] [CrossRef] [PubMed]
- Vintila, A.R.; et al. Mitochondrial sulfide promotes life span and health span through distinct mechanisms in developing versus adult treated Caenorhabditis elegans. Proc Natl Acad Sci U S A 2023, 120, e2216141120. [Google Scholar] [CrossRef] [PubMed]
- Debes, C.; et al. Ageing-associated changes in transcriptional elongation influence longevity. Nature 2023, 616, 814–821. [Google Scholar] [CrossRef] [PubMed]
- Yang, M.M.; et al. The Translational Regulation in mTOR Pathway. Biomolecules 2022, 12. [Google Scholar] [CrossRef] [PubMed]
- Ham, S.; et al. Systematic transcriptome analysis associated with physiological and chronological aging in Caenorhabditis elegans. Genome Res 2022, 32, 2003–2014. [Google Scholar] [CrossRef] [PubMed]
- O'Brien, J.; et al. Overview of MicroRNA Biogenesis, Mechanisms of Actions, and Circulation. Front Endocrinol (Lausanne) 2018, 9, 402. [Google Scholar] [CrossRef] [PubMed]
- Kato, M.; et al. Age-associated changes in expression of small, noncoding RNAs, including microRNAs, in C. elegans. RNA 2011, 17, 1804–20. [Google Scholar] [CrossRef] [PubMed]
- Ye, P.Y.; et al. An mTORC1-Mdm2-Drosha axis for miRNA biogenesis in response to glucose- and amino acid-deprivation. Molecular Cancer Research 2016, 14. [Google Scholar] [CrossRef]
- Johnson, S.C.; et al. mTOR inhibition alleviates mitochondrial disease in a mouse model of Leigh syndrome. Science 2013, 342, 1524–8. [Google Scholar] [CrossRef] [PubMed]
- Ramanathan, A. and S.L. Schreiber, Direct control of mitochondrial function by mTOR. Proceedings of the National Academy of Sciences of the United States of America 2009, 106, 22229–22232. [Google Scholar] [CrossRef] [PubMed]
- Migliavacca, E.; et al. Mitochondrial oxidative capacity and NAD(+) biosynthesis are reduced in human sarcopenia across ethnicities. Nat Commun 2019, 10, 5808. [Google Scholar] [CrossRef] [PubMed]
- Sharifi, S.; et al. Reducing the metabolic burden of rRNA synthesis promotes healthy longevity in Caenorhabditis elegans. Nat Commun 2024, 15, 1702. [Google Scholar] [CrossRef] [PubMed]
- Walther, D.M.; et al. Widespread Proteome Remodeling and Aggregation in Aging. Cell 2015, 161, 919–932. [Google Scholar] [CrossRef] [PubMed]
- Welker, N.C., J. W. Habig, and B.L. Bass, Genes misregulated in C. elegans deficient in Dicer, RDE-4, or RDE-1 are enriched for innate immunity genes. RNA 2007, 13, 1090–102. [Google Scholar] [CrossRef] [PubMed]
- Gygi, S.P.; et al. Correlation between protein and mRNA abundance in yeast. Molecular and Cellular Biology 1999, 19, 1720–1730. [Google Scholar] [CrossRef] [PubMed]
- Washburn, M.P.; et al. Protein pathway and complex clustering of correlated mRNA and protein expression analyses in. Proceedings of the National Academy of Sciences of the United States of America 2003, 100, 3107–3112. [Google Scholar] [CrossRef] [PubMed]
- Ideker, T.; et al. Integrated genomic and proteomic analyses of a systematically perturbed metabolic network. Science 2001, 292, 929–934. [Google Scholar] [CrossRef] [PubMed]
- Pan, K.Z.; et al. Inhibition of mRNA translation extends lifespan in Caenorhabditis elegans. Aging Cell 2007, 6, 111–9. [Google Scholar] [CrossRef] [PubMed]
- Takauji, Y.; et al. Restriction of protein synthesis abolishes senescence features at cellular and organismal levels. Sci Rep 2016, 6, 18722. [Google Scholar] [CrossRef] [PubMed]
- Depuydt, G.; et al. Increased Protein Stability and Decreased Protein Turnover in the Caenorhabditis elegans Ins/IGF-1 daf-2 Mutant. J Gerontol A Biol Sci Med Sci 2016, 71, 1553–1559. [Google Scholar] [CrossRef] [PubMed]
- Ellis, R.J. and A.P. Minton, Protein aggregation in crowded environments. Biological Chemistry 2006, 387, 485–497. [Google Scholar] [CrossRef] [PubMed]
- Dhondt, I.; et al. FOXO/DAF-16 Activation Slows Down Turnover of the Majority of Proteins in C. elegans. Cell Rep 2016, 16, 3028–3040. [Google Scholar] [CrossRef] [PubMed]
- Buttgereit, F. and M.D. Brand, A Hierarchy of Atp-Consuming Processes in Mammalian-Cells. Biochemical Journal 1995, 312, 163–167. [Google Scholar] [CrossRef] [PubMed]
- Li, S.T.; et al. DAF-16 stabilizes the aging transcriptome and is activated in mid-aged Caenorhabditis elegans to cope with internal stress. Aging Cell 2019, 18, e12896. [Google Scholar] [CrossRef] [PubMed]
- Stout, G.J.; et al. Insulin/IGF-1-mediated longevity is marked by reduced protein metabolism. Mol Syst Biol 2013, 9, 679. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.J.; et al. A demographic analysis of the fitness cost of extended longevity in. Journals of Gerontology Series a-Biological Sciences and Medical Sciences 2007, 62, 126–135. [Google Scholar] [CrossRef] [PubMed]
- Austad, S.N. and J.M. Hoffman, Is antagonistic pleiotropy ubiquitous in aging biology? Evolution Medicine and Public Health 2018, 287–294. [Google Scholar] [CrossRef] [PubMed]
- Hahm, J.H.; et al. C. elegans maximum velocity correlates with healthspan and is maintained in worms with an insulin receptor mutation. Nat Commun 2015, 6, 8919. [Google Scholar] [CrossRef] [PubMed]
- Venz, R.; et al. End-of-life targeted degradation of DAF-2 insulin/IGF-1 receptor promotes longevity free from growth-related pathologies. Elife 2021, 10. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; et al. A demographic analysis of the fitness cost of extended longevity in Caenorhabditis elegans. J Gerontol A Biol Sci Med Sci 2007, 62, 126–35. [Google Scholar] [CrossRef] [PubMed]
- Mitchell, D.H.; et al. Synchronous growth and aging of Caenorhabditis elegans in the presence of fluorodeoxyuridine. J Gerontol 1979, 34, 28–36. [Google Scholar] [CrossRef] [PubMed]
- Lexell, J., C. C. Taylor, and M. Sjostrom, What is the cause of the ageing atrophy? Total number, size and proportion of different fiber types studied in whole vastus lateralis muscle from 15- to 83-year-old men. J Neurol Sci 1988, 84, 275–94. [Google Scholar] [CrossRef] [PubMed]
- Hosono, R.; et al. Age-Dependent Changes in Mobility and Separation of the Nematode Caenorhabditis-Elegans. Experimental Gerontology 1980, 15, 285–289. [Google Scholar] [CrossRef] [PubMed]
- Mitchell, W.K.; et al. Sarcopenia, dynapenia, and the impact of advancing age on human skeletal muscle size and strength; a quantitative review. Front Physiol 2012, 3, 260. [Google Scholar] [CrossRef] [PubMed]
- Fleg, J.L.; et al. Accelerated longitudinal decline of aerobic capacity in healthy older adults. Circulation 2005, 112, 674–82. [Google Scholar] [CrossRef] [PubMed]
- Shur, N.F.; et al. Age-related changes in muscle architecture and metabolism in humans: The likely contribution of physical inactivity to age-related functional decline. Ageing Res Rev 2021, 68, 101344. [Google Scholar] [CrossRef] [PubMed]
- Gallagher, D.; et al. Weight stability masks sarcopenia in elderly men and women. Am J Physiol Endocrinol Metab 2000, 279, E366–75. [Google Scholar] [CrossRef] [PubMed]
- Pradhan, S.; et al. Environmental Programming of Adult Foraging Behavior in C. elegans. Curr Biol 2019, 29, 2867–2879. [Google Scholar] [CrossRef] [PubMed]
- Dillin, A.; et al. Rates of behavior and aging specified by mitochondrial function during development. Science 2002, 298, 2398–401. [Google Scholar] [CrossRef]
- Bazopoulou, D.; et al. Developmental ROS individualizes organismal stress resistance and lifespan. Nature 2019, 576, 301–305. [Google Scholar] [CrossRef] [PubMed]
- Dillin, A., D. K. Crawford, and C. Kenyon, Timing requirements for insulin/IGF-1 signaling in. Science 2002, 298, 830–834. [Google Scholar] [CrossRef] [PubMed]
- Lind, M.I.; et al. Cost-free lifespan extension via optimization of gene expression in adulthood aligns with the developmental theory of ageing. Proc Biol Sci 2021, 288, 20201728. [Google Scholar] [CrossRef] [PubMed]
- Harrison, D.E.; et al. Rapamycin fed late in life extends lifespan in genetically heterogeneous mice. Nature 2009, 460, 392–U108. [Google Scholar] [CrossRef] [PubMed]
- Mao, K.; et al. Late-life targeting of the IGF-1 receptor improves healthspan and lifespan in female mice. Nat Commun 2018, 9, 2394. [Google Scholar] [CrossRef] [PubMed]
- Bitto, A.; et al. Transient rapamycin treatment can increase lifespan and healthspan in middle-aged mice. Elife 2016, 5. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.R., S. C. Ding, and J. Wang, Digital health for aging populations. Nature Medicine 2023, 29, 1623–1630. [Google Scholar] [CrossRef] [PubMed]
- Scott, A.J., M. Ellison, and D.A. Sinclair, The economic value of targeting aging. Nat Aging 2021, 1, 616–623. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.B.; et al. Extended Twilight among Isogenic C. elegans Causes a Disproportionate Scaling between Lifespan and Health. Cell Syst 2016, 3, 333–345. [Google Scholar] [CrossRef] [PubMed]
- Zhu, T.Y.; et al. Single-worm quantitative proteomics reveals aging heterogeneity in isogenic Caenorhabditis elegans. Aging Cell 2023, e14055. [Google Scholar] [CrossRef] [PubMed]
- Rando, T.A. and T. Wyss-Coray, Asynchronous, contagious and digital aging. Nat Aging 2021, 1, 29–35. [Google Scholar] [CrossRef] [PubMed]
- Dong, X., B. Milholland, and J. Vijg, Evidence for a limit to human lifespan. Nature 2016, 538, 257–259. [Google Scholar] [CrossRef] [PubMed]
- Nishimura, K.; et al. An auxin-based degron system for the rapid depletion of proteins in nonplant cells. Nat Methods 2009, 6, 917–22. [Google Scholar] [CrossRef] [PubMed]
- Phanindhar, K. and R.K. Mishra, Auxin-inducible degron system: an efficient protein degradation tool to study protein function. Biotechniques 2023, 74, 186–198. [Google Scholar] [CrossRef] [PubMed]
- Roy, C.; et al. DAF-2/insulin IGF-1 receptor regulates motility during aging by integrating opposite signaling from muscle and neuronal tissues. Aging Cell 2022, 21, e13660. [Google Scholar] [CrossRef] [PubMed]
- Smith, H.J.; et al. Neuronal mTORC1 inhibition promotes longevity without suppressing anabolic growth and reproduction in C. elegans. PLoS Genet 2023, 19, e1010938. [Google Scholar] [CrossRef] [PubMed]
- Yee, Z.; et al. Inhibition of mTOR decreases insoluble proteins burden by reducing translation in C. elegans. Biogerontology 2021, 22, 101–118. [Google Scholar] [CrossRef] [PubMed]
- Roux, A.E.; et al. Individual cell types in C. elegans age differently and activate distinct cell-protective responses. Cell Rep 2023, 42, 112902. [Google Scholar] [CrossRef] [PubMed]
- Ruiz, J.R.; et al. Association between muscular strength and mortality in men: prospective cohort study. BMJ 2008, 337, a439. [Google Scholar] [CrossRef] [PubMed]
- Dickinson, J.M.; et al. Mammalian target of rapamycin complex 1 activation is required for the stimulation of human skeletal muscle protein synthesis by essential amino acids. J Nutr 2011, 141, 856–62. [Google Scholar] [CrossRef] [PubMed]
- Terzis, G.; et al. Resistance exercise-induced increase in muscle mass correlates with p70S6 kinase phosphorylation in human subjects. Eur J Appl Physiol 2008, 102, 145–52. [Google Scholar] [CrossRef] [PubMed]
- Phillips, B.E.; et al. Molecular networks of human muscle adaptation to exercise and age. PLoS Genet 2013, 9, e1003389. [Google Scholar] [CrossRef]
- Plaza-Diaz, J.; et al. Impact of Physical Activity and Exercise on the Epigenome in Skeletal Muscle and Effects on Systemic Metabolism. Biomedicines 2022, 10. [Google Scholar] [CrossRef]
- AgeUK. Later life in the United Kingdom. 2019; Available from: https://www.ageuk.org.uk/globalassets/age-uk/documents/reports-and-publications/later_life_uk_factsheet.pdf.
- Kern, D.G.a.C. Biological Constraint as a Cause of Aging. PrePrints 2022. [Google Scholar]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



