Submitted:
08 March 2024
Posted:
11 March 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Material and Methods
2.1. Sample Collection
2.2. Chemicals and Apparatus
2.3. Centrifugation-Assisted Ultrafiltration (CUF) of Milk Samples
2.4. Two-Dimensional Chromatographic Separation (2D-LC) and Ultraviolet Detection (DAD)
3. Results and Discussion
3.1. 2D-LC-DAD Method Development
3.2. Sample Treatment
3.3. 2D-LC-DAD Analytical Performance
3.3.1. Linearity and Sensitivity
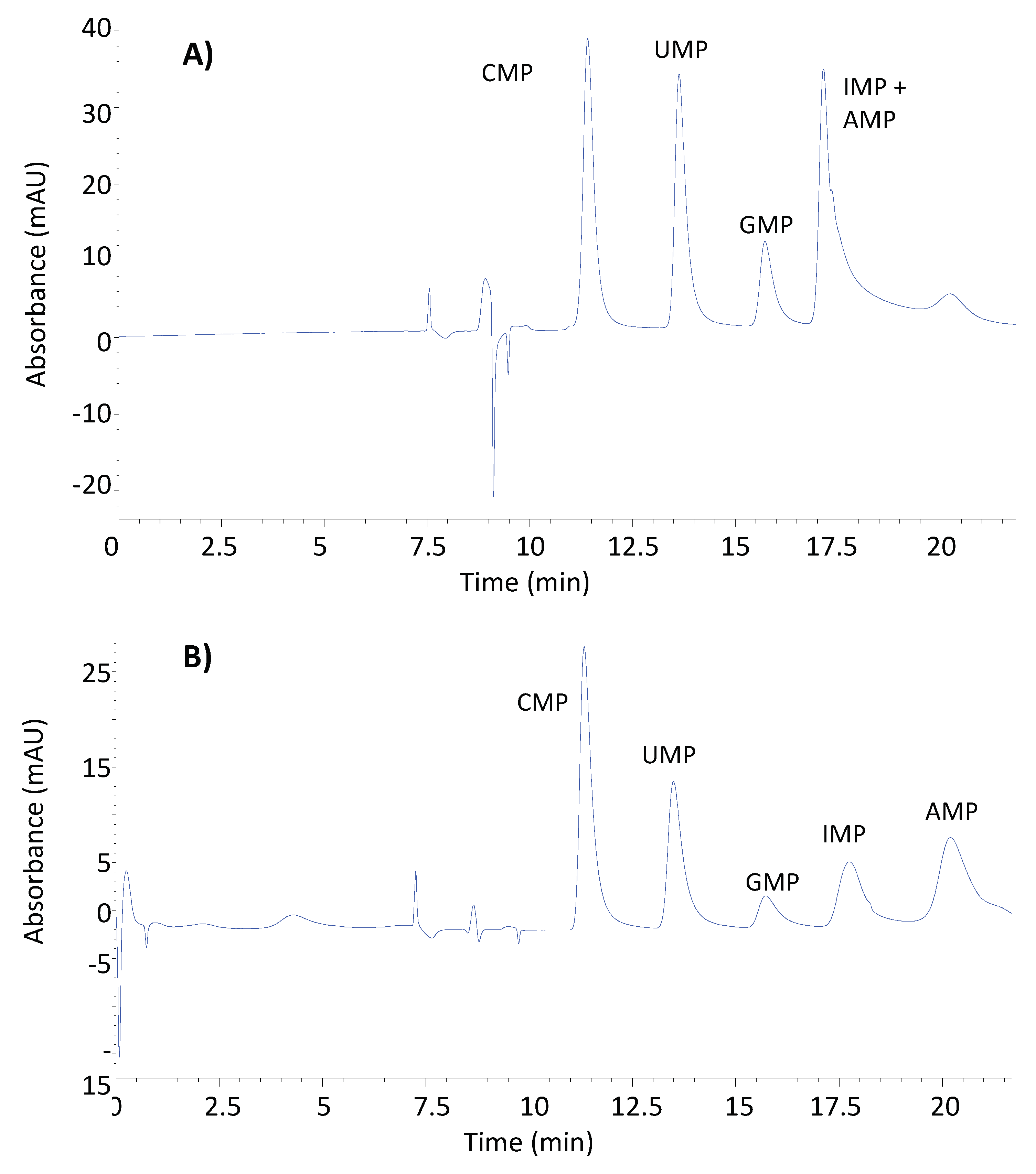
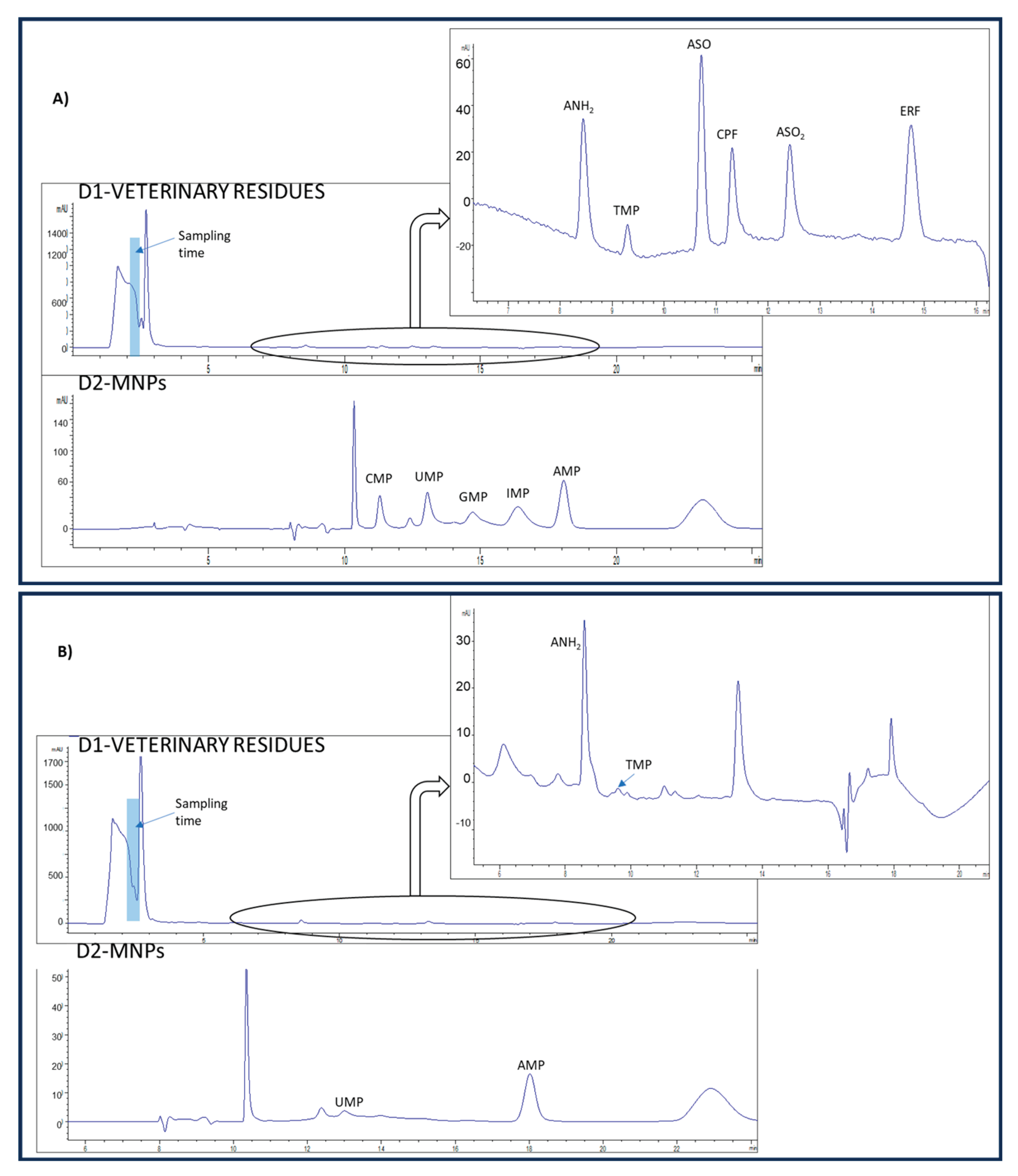
3.3.2. Selectivity
3.3.3. Recovery
3.3.4. Precision
3.4. Analysis of Sheep’s Milk Samples
4. Conclusions
Supplementary Materials
Acknowledgments
References
- L. G. Davis and M. D. Sintes-Zydowicz; Nucleotides: nutrition and health effects. Comprehensive Reviews in Food Science and Food Safety 2009, 8, 144–156. [Google Scholar]
- (M. C. Martínez-Costa et al.; Nucleotide content in breast milk and formulas: effect of processing and potential role in infancy. Journal of Pediatric Gastroenterology and Nutrition 2009, 48, S61–S69. [Google Scholar]
- Fernández-Cruz, A.R.; et al. ; Veterinary drugs residues in animal-derived foods: a review of the last decade. Journal of Food Science and Technology 2020, 57, 1647–1660. [Google Scholar]
- European Commission. Commission Regulation (EU) No 37/2010 of 22 December 2009 on pharmacologically active substances and their classification regarding maximum residue limits in foodstuffs of animal origin. Official Journal of the European Union, vol. L15/1-L15/72, 2010. 22 December.
- J. Fink-Gremmels and E. J. Wassenberg-Brandt van Edelveen; Antibiotics in animal feed. Antimicrobial Resistance in Bacteria of Animal Origin, F. M. Aarestrup ed., Washington DC: ASM Press, 2006; pp. 73–108. [Google Scholar]
- J. Echevarria et al., Anthelmintic drugs for sheep farms: residues in milk and cheese. Food Additives & Contaminants: Part A 2011, 28, 1108–1114. [Google Scholar]
- M. A. Bakkali et al.; Determination of albendazole and its main metabolites in ovine milk by liquid chromatography-tandem mass spectrometry. Journal of Chromatography B: Analytical Technologies in the Biomedical and Life Sciences 2009, 878, 3053–3058. [Google Scholar]
- M. -L Rauhala and T.A Pakkanen; Determination of nucleotides and nucleosides in milks by a novel UPLC method. Food Chemistry 2009, 115, 560–567. [Google Scholar]
- D. R Stoll; Two-dimensional liquid chromatography: a state of the art review. Analytical Chemistry 2020, 92, 280–301. [Google Scholar]
- P. J Schoenmakers et al.; Comprehensive two-dimensional liquid chromatography: basic possibilities and practical applications. Journal of Chromatography A 2019, 1589, 2–13. [Google Scholar]
- S. Fekete et al.; Recent developments and future trends in ultra-high performance liquid chromatography; Journal of Pharmaceutical and Biomedical Analysis 2015, 113, 82–92.
- Commission Decision 2002/657/EC of 12 August 2002 implementing CouncilDirective 96/23/EC concerning the performance of analytical methods and the interpretation of results. Off. J. Eur. Comm. L. 2002, 8.
- 13. COMMISSION REGULATION (EU) No 37/2010 of 22 December 2009 on pharmacologically active substances and their classification regarding maximum residue limits in foodstuffs of animal origin, Off. J. Eur. Comm. L. (2009) 15. Commission Implementing Regulation (EU) 2023/981 of 17 May 2023. Off. J. Eur. Comm. L. 2023, 134.
| Chromatographic separation |
Analyte |
atr (min) |
bk | cα |
|---|---|---|---|---|
| Gradient elution method 1 | ANH2 | 3.7 | 2.4 | - |
| TMP | 5.3 | 3.8 | 1.62 | |
| ASO | 9.2 | 7.4 | 1.93 | |
| CPF | 14.7 | 12.4 | 1.68 | |
| ASO2 | 16.0 | 13.5 | 1.10 | |
| ERF | 16.9 | 14.4 | 1.06 | |
| Gradient elution method 2 | ANH2 | 6.1 | 4.5 | - |
| TMP | 7.0 | 5.4 | 1.18 | |
| ASO | 11.4 | 9.4 | 1.63 | |
| CPF | 12.4 | 10.3 | 1.10 | |
| ASO2 | 13.5 | 11.3 | 1.10 | |
| ERF | 14.0 | 11.7 | 1.04 | |
| Gradient elution method 3 | ANH2 | 8.4 | 6.6 | - |
| TMP | 9.3 | 7.5 | 1.15 | |
| ASO | 10.5 | 8.9 | 1.20 | |
| CPF | 11.4 | 9.4 | 1.10 | |
| ASO2 | 12.5 | 10.4 | 1.11 | |
| ERF | 15.2 | 12.8 | 1.24 |
| Chromatographic. separation |
Analyte |
atr (min) |
bk | cα |
|---|---|---|---|---|
| Isocratic elution method | CMP | 11.2 | 9.2 | |
| UMP | 13.0 | 10.8 | 1.5 | |
| GMP | 14.8 | 12.5 | 1.4 | |
| IMP | 16.5 | 14.0 | 1.2 | |
| AMP | 17.7 | 15.1 | 1.2 |
| Chromatographic separation |
Analyte |
aLinear range (mg L-1) |
Slope (L mg1) | br |
cMQL (mg L-1) |
dMRL (mg L-1) |
Recovery (%) ± eSD | Repeatability (fCV %) |
Reproducibility (fCV %) |
||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| gL1 | hL2 | iL3 | gL1 | hL2 | iL3 | gL1 | hL2 | iL3 | |||||||
| 1D | ANH2 | 0.01-0.5 | 0.2759 | 0.9982 | 0.01 | e0.1 | 70 ± 4 | 78 ± 4 | 80 ± 4 | 6 | 5 | 5 | 13 | 8 | 7 |
| TMP | 0.025-0.5 | 0.1460 | 0.9957 | 0.025 | 0.05 | 84 ± 3 | 90 ± 4 | 95 ± 1 | 4 | 4 | 1 | 4 | 5 | 2 | |
| ASO | 0.01-0.5 | 0.3513 | 0.9995 | 0.01 | e0.1 | 95 ± 5 | 119 ± 2 | 108 ± 2 | 5 | 2 | 2 | 6 | 4 | 3 | |
| CPF | 0.025-0.5 | 0.6130 | 0.9879 | 0.025 | f0.1 | 106 ± 10 | 120 ± 5 | 114 ± 7 | 9 | 4 | 6 | 11 | 7 | 6 | |
| ASO2 | 0.025-0.5 | 0.5744 | 0.9789 | 0.025 | e0.1 | 108 ± 4 | 118 ± 3 | 109 ± 3 | 3 | 3 | 3 | 4 | 4 | 3 | |
| ERF | 0.025-0.5 | 0.3569 | 0.9997 | 0.025 | f0.1 | 84 ± 4 | 86 ± 3 | 82 ± 2 | 5 | 4 | 3 | 7 | 5 | 4 | |
| 2D | CMP | 0.25-100 | 13.893 | 0.9984 | 0.25 | - | 82 ± 10 | 83 ± 4 | 93 ± 6 | 12 | 5 | 6 | 14 | 8 | 8 |
| UMP | 0.5-100 | 11.617 | 0.9838 | 0.5 | - | 80 ± 11 | 91 ± 6 | 117 ± 10 | 14 | 7 | 8 | 15 | 10 | 9 | |
| GMP | 0.75-100 | 6.542 | 0.9967 | 0.75 | - | 81 ± 8 | 89 ± 3 | 90 ± 4 | 10 | 4 | 4 | 12 | 7 | 6 | |
| IMP | 0.5-100 | 13.964 | 0.9983 | 0.5 | - | 85 ± 2 | 83 ± 3 | 85 ± 2 | 3 | 4 | 3 | 4 | 5 | 3 | |
| AMP | 0.25-100 | 19.880 | 0.9997 | 0.25 | - | 80 ± 5 | 119 ± 12 | 118 ± 4 | 7 | 10 | 4 | 10 | 10 | 7 | |
| Chromatographic separation |
Analyte | Sample 1 | Sample 2 | Sample 3 | |||
|---|---|---|---|---|---|---|---|
|
aR (%) ± bSD |
cConc. (mg L-1) ± bSD |
aR (%) ± bSD |
cConc. (mg L-1) ± bSD |
aR (%) ± bSD |
cConc. (mg L-1) ± bSD |
||
| 1D | ANH2 | 68 ± 10 | 0.675 ± 0.015 | 78 ± 12 | 0.578 ± 0.022 | 68 ± 3 | nd |
| TMP | 85 ± 13 | 0.085 ± 0.005 | 82 ± 5 | 0.015 ± 0.001 | 87 ± 12 | nd | |
| ASO | 122 ± 2 | nd | 123 ± 3 | nd | 122 ± 4 | nd | |
| CPF | 123 ± 3 | nd | 116 ± 4 | nd | 120 ± 7 | nd | |
| ASO2 | 120 ± 6 | nd | 113 ± 2 | nd | 103 ± 4 | nd | |
| ERF | 79 ±2 | nd | 84 ± 9 | nd | 80 ± 9 | nd | |
| 2D | CMP | 88 ± 3 | nd | 93 ± 11 | nd | 96 ± 2 | nd |
| UMP | 122 ± 6 | 0.7 ± 0.1 | 119 ± 7 | <MQL | 115 ± 2 | 1.6 ± 0.2 | |
| GMP | 80 ± 2 | nd | 88 ± 8 | nd | 91 ± 1 | nd | |
| IMP | 81 ± 3 | nd | 83 ± 7 | nd | 84 ± 1 | nd | |
| AMP | 84 ± 2 | 16 ± 1 | 84 ± 1 | 15.2 ± 0.3 | 86 ± 2 | 15.4 ± 0.8 | |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




