Submitted:
08 March 2024
Posted:
08 March 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. RNA Folding Engines
2.2. RNA Classes & Genuses Assayed
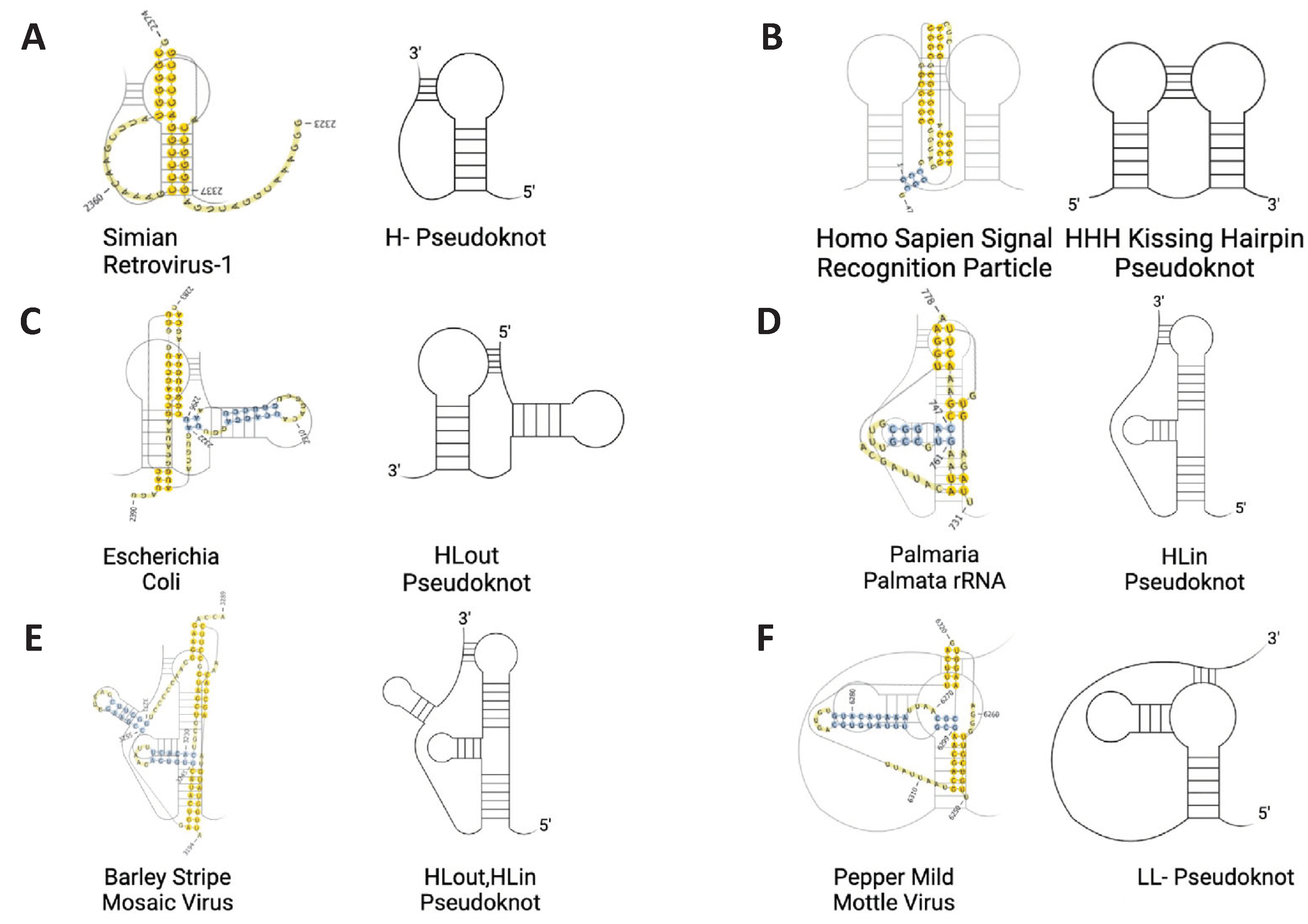
- 1)
- H-Pseudoknot
- 2)
- HHH-Pseudoknot (kissing hairpin)
- 3)
- HLout- Pseudoknot
- 4)
- HLin-Pseudoknot
- 5)
- HLout,HLin Pseudoknot
- 6)
- LL- Pseudoknot
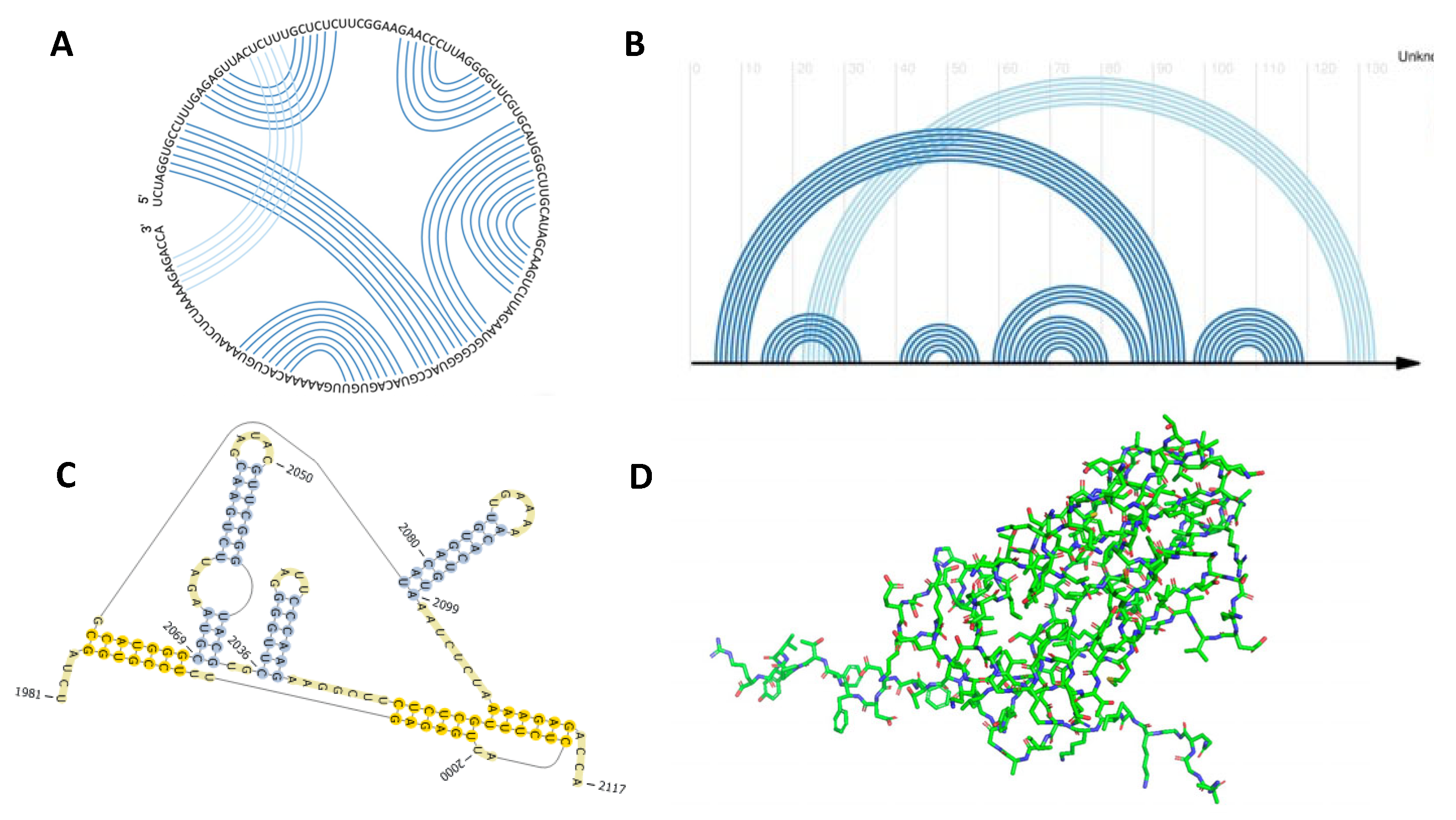
2.3. Generating Viral Pseudoknotted RNA PseudoBase++ Dataset
2.4. Performance Metrics
- A large set of well-established reference/accepted structures to compare against experimentally derived data.
- Tests for statistical significance.
- Different RNA families/ RNA types (see Supplemental Table S1) should be used than those used to train the methods being benchmarked.
3. Results
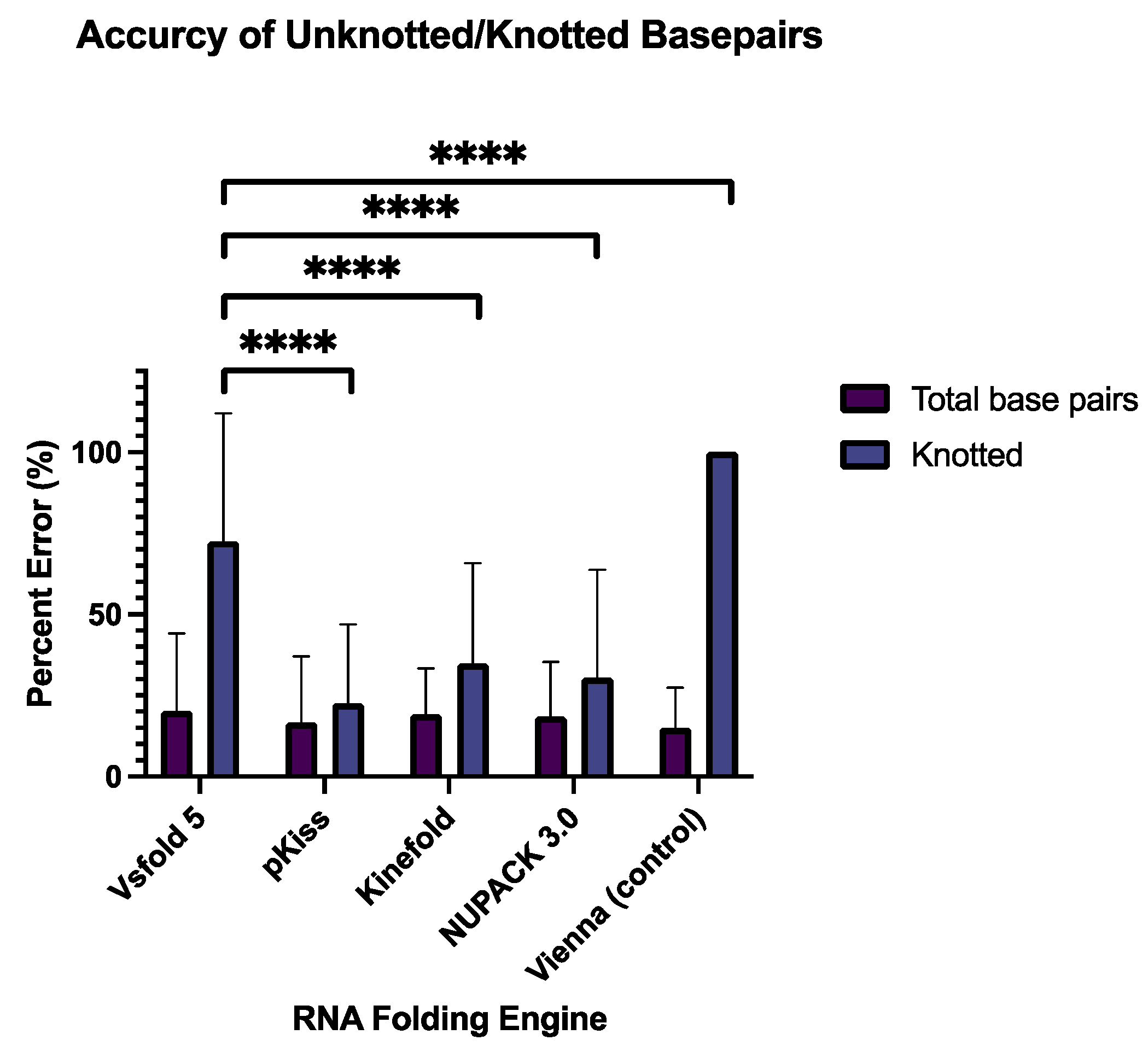
3.1. Assessment of Percent Error of in Total Base Pairs and Knotted Base Pairs
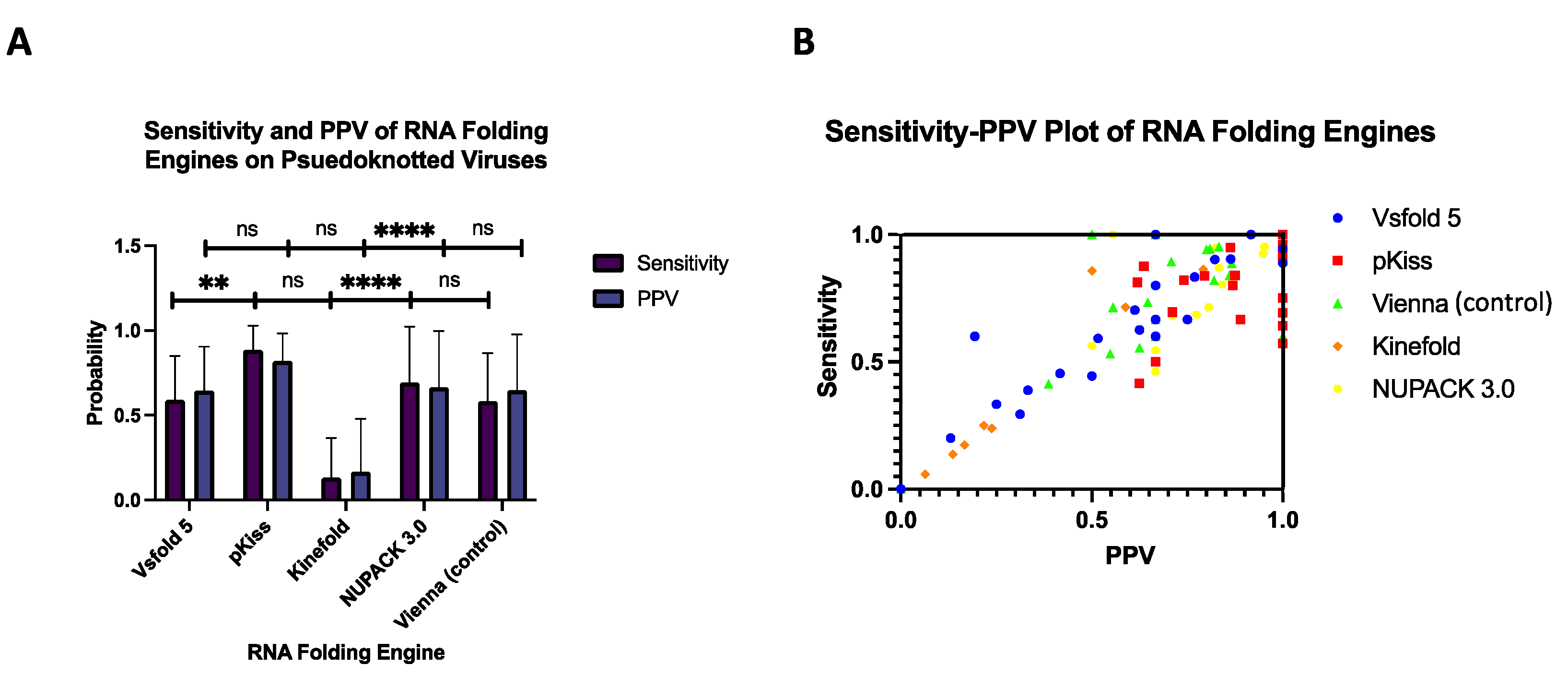
3.2. Sensitivity and Positive Predictive Value (PPV) of Folding Engines
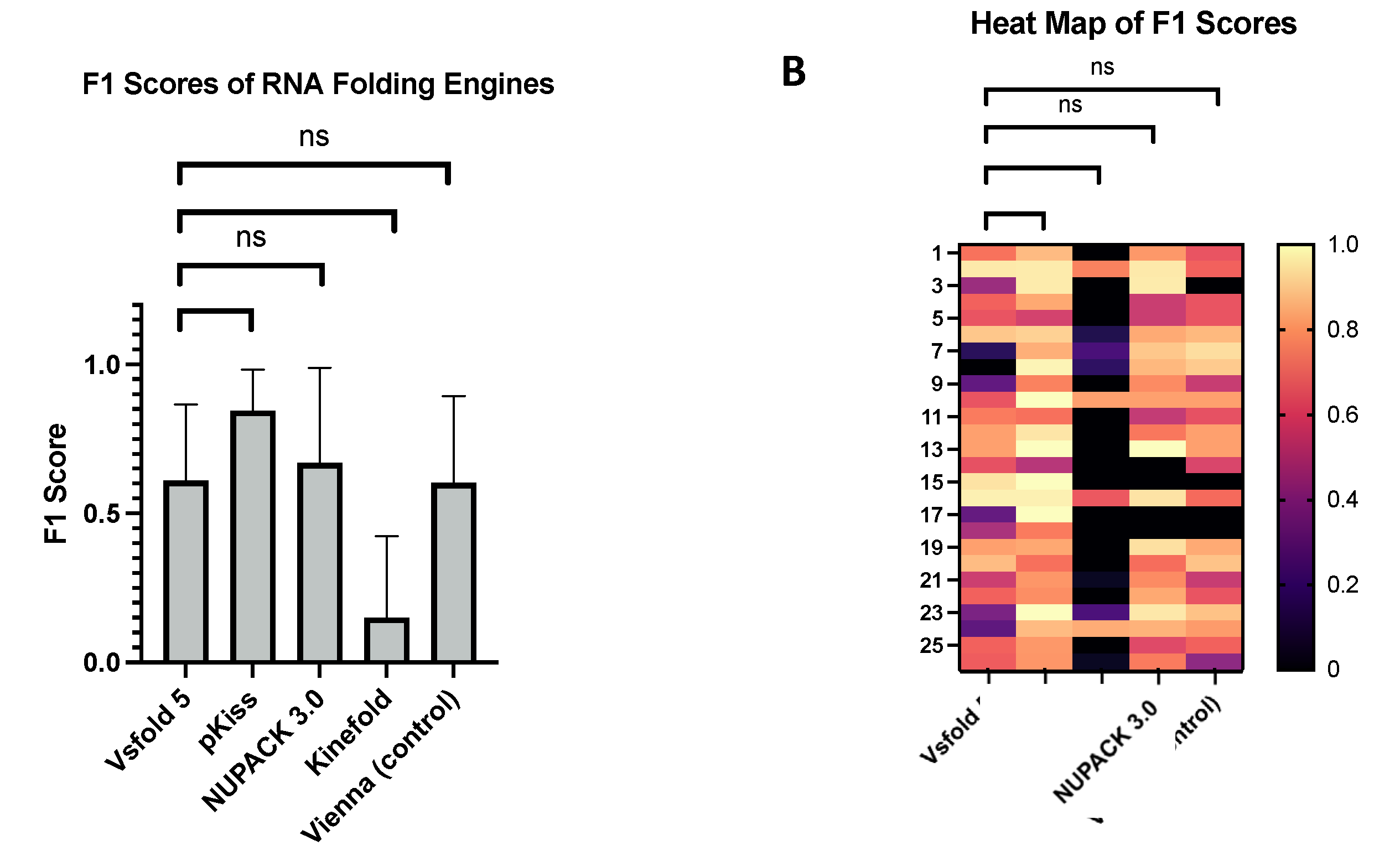
3.3. Quality of Prediction Software Assessed via F1-Scoring
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Lewin AS, Hauswirth WW. Ribozyme gene therapy: applications for molecular medicine. Trends Mol Med. 2001, 7, 221–228. [Google Scholar] [CrossRef]
- Walter NG, Engelke DR. Ribozymes: catalytic RNAs that cut things, make things, and do odd and useful jobs. Biologist (London). 2002, 49, 199–203. [Google Scholar]
- Zuber J, Schroeder SJ, Sun H, Turner DH, Mathews DH. Nearest neighbor rules for RNA helix folding thermodynamics: improved end effects. Nucleic Acids Res. 2022, 50, 5251–5262. [Google Scholar] [CrossRef]
- Wu L, Belasco JG. Let me count the ways: mechanisms of gene regulation by miRNAs and siRNAs. Mol Cell. 2008, 29, 1–7. [Google Scholar] [CrossRef]
- Leamy KA, Assmann SM, Mathews DH, Bevilacqua PC. Bridging the gap between in vitro and in vivo RNA folding. Q Rev Biophys. 2016, 49, e10. [Google Scholar]
- Brierley I, Pennell S, Gilbert RJC. Viral RNA pseudoknots: versatile motifs in gene expression and replication. Nat Rev Microbiol. 2007, 5, 598–610. [Google Scholar] [CrossRef]
- Xayaphoummine A, Bucher T, Thalmann F, Isambert H. Prediction and statistics of pseudoknots in RNA structures using exactly clustered stochastic simulations. Proceedings of the National Academy of Sciences. 2003, 100, 15310–15315. [Google Scholar] [CrossRef]
- Pu H, Li J, Li D, Han C, Yu J. Identification of an Internal RNA Element Essential for Replication and Translational Enhancement of Tobacco Necrosis Virus AC. PLoS One. 2013, 8, e57938. [Google Scholar]
- Taylor, JM. Structure and Replication of Hepatitis Delta Virus RNA. In: Hepatitis Delta Virus. Springer Berlin Heidelberg; p. 1–23.
- Tinoco I, Bustamante C. How RNA folds. J Mol Biol. 1999, 293, 271–281. [Google Scholar] [CrossRef]
- Nikolova EN, Zhou H, Gottardo FL, Alvey HS, Kimsey IJ, Al-Hashimi HM. A historical account of Hoogsteen base-pairs in duplex DNA. Biopolymers. 2013, 99, 955–968. [Google Scholar] [CrossRef] [PubMed]
- Gernot Akemann, Jinho Baik, Philippe Di Francesco, Henri Orland, Graziano Vernizzi. The Oxford Handbook of Random Matrix Theory. Oxford University Press; 2011. 872–897 p.
- Lim CS, Brown CM. Know Your Enemy: Successful Bioinformatic Approaches to Predict Functional RNA Structures in Viral RNAs. Front Microbiol. 2018, 8. [Google Scholar]
- MANS RMW, PLEIJ CWA, BOSCH L. tRNA-like structures. Eur J Biochem. 1991, 201, 303–324. [Google Scholar] [CrossRef]
- Felden B, Florentz C, Giegé R, Westhof E. Solution Structure of the 3′-End of Brome Mosaic Virus Genomic RNAs. J Mol Biol. 1994, 235, 508–531. [Google Scholar] [CrossRef]
- Lai D, Proctor JR, Zhu JYA, Meyer IM. R- chie : a web server and R package for visualizing RNA secondary structures. Nucleic Acids Res. 2012, 40, e95–e95. [Google Scholar] [CrossRef] [PubMed]
- Byun Y, Han K. PseudoViewer3: generating planar drawings of large-scale RNA structures with pseudoknots. Bioinformatics. 2009, 25, 1435–1437. [Google Scholar] [CrossRef]
- Larson SB, Lucas RW, McPherson A. Crystallographic Structure of the T=1 Particle of Brome Mosaic Virus. J Mol Biol. 2005, 346, 815–831. [Google Scholar] [CrossRef]
- Choudhary K, Deng F, Aviran S. Comparative and integrative analysis of RNA structural profiling data: current practices and emerging questions. Quantitative Biology. 2017, 5, 3–24. [Google Scholar] [CrossRef] [PubMed]
- Lavender CA, Gorelick RJ, Weeks KM. Structure-Based Alignment and Consensus Secondary Structures for Three HIV-Related RNA Genomes. PLoS Comput Biol. 2015, 11, e1004230. [Google Scholar]
- Janssen S, Giegerich R. The RNA shapes studio. Bioinformatics. 2015, 31, 423–425. [Google Scholar] [CrossRef]
- BINDEWALD E, SHAPIRO BA. RNA secondary structure prediction from sequence alignments using a network of k -nearest neighbor classifiers. RNA. 2006, 12, 342–352. [Google Scholar] [CrossRef]
- Taufer M, Licon A, Araiza R, Mireles D, van Batenburg FHD, Gultyaev AP, et al. PseudoBase++: an extension of PseudoBase for easy searching, formatting and visualization of pseudoknots. Nucleic Acids Res. 2009, 37, D127–35. [Google Scholar] [CrossRef]
- Legendre A, Angel E, Tahi F. Bi-objective integer programming for RNA secondary structure prediction with pseudoknots. BMC Bioinformatics. 2018, 19, 13. [Google Scholar]
- Sato K, Kato Y. Prediction of RNA secondary structure including pseudoknots for long sequences. Brief Bioinform. 2022, 23. [Google Scholar]
- Dawson WK, Fujiwara K, Kawai G. Prediction of RNA Pseudoknots Using Heuristic Modeling with Mapping and Sequential Folding. PLoS One. 2007, 2, e905. [Google Scholar]
- Xayaphoummine A, Bucher T, Isambert H. Kinefold web server for RNA/DNA folding path and structure prediction including pseudoknots and knots. Nucleic Acids Res. 2005, 33 (Suppl_2), W605–W610. [Google Scholar] [CrossRef] [PubMed]
- Zadeh JN, Steenberg CD, Bois JS, Wolfe BR, Pierce MB, Khan AR, et al. NUPACK: Analysis and design of nucleic acid systems. J Comput Chem. 2011, 32, 170–173. [Google Scholar] [CrossRef] [PubMed]
- Mark E. Fornace, Jining Huang, Cody T. Newman, Nicholas J. Porubsky, Marshall B. Pierce, Niles A. Pierce. NUPACK: Analysis and Design of Nucleic Acid Structures, Devices, and Systems. ChemRxiv Cambridge. 2022.
- Hofacker IL, Fontana W, Stadler PF, Bonhoeffer LS, Tacker M, Schuster P. Fast folding and comparison of RNA secondary structures. Monatshefte f�r Chemie Chemical Monthly. 1994, 125, 167–188. [Google Scholar] [CrossRef]
- Wayment-Steele HK, Kladwang W, Strom AI, Lee J, Treuille A, Becka A, et al. RNA secondary structure packages evaluated and improved by high-throughput experiments. Nat Methods. 2022, 19, 1234–1242. [Google Scholar] [CrossRef]
- Zammit A, Helwerda L, Olsthoorn RCL, Verbeek FJ, Gultyaev AP. A database of flavivirus RNA structures with a search algorithm for pseudoknots and triple base interactions. Bioinformatics. 2021, 37, 956–962. [Google Scholar] [CrossRef] [PubMed]
- Mlera L, Melik W, Bloom ME. The role of viral persistence in flavivirus biology. Pathog Dis. 2014, 71, 137–163. [Google Scholar] [CrossRef]
- Creager ANH. Tobacco Mosaic Virus and the History of Molecular Biology. Annu Rev Virol. 2022, 9, 39–55. [Google Scholar] [CrossRef] [PubMed]
- Peselis A, Serganov A. Structure and function of pseudoknots involved in gene expression control. WIREs RNA. 2014, 5, 803–822. [Google Scholar] [CrossRef]
- Lucas A, Dill KA. Statistical mechanics of pseudoknot polymers. J Chem Phys. 2003, 119, 2414–2421. [Google Scholar] [CrossRef]
- Du Z, Holland JA, Hansen MR, Giedroc DP, Hoffman DW. Base-pairings within the RNA pseudoknot associated with the simian retrovirus-1 gag-pro frameshift site. J Mol Biol. 1997, 270, 464–470. [Google Scholar] [CrossRef]
- Zwieb, C. SRPDB (Signal Recognition Particle Database). Nucleic Acids Res. 2000, 28, 171–172. [Google Scholar] [CrossRef]
- De Rijk P, Robbrecht E, de Hoog S, Caers A, Van de Peer Y, De Wachter R. Database on the structure of large subunit ribosomal RNA. Nucleic Acids Res. 1999, 27, 174–178. [Google Scholar] [CrossRef]
- Van de Peer, Y. The European Small Subunit Ribosomal RNA database. Nucleic Acids Res. 2000, 28, 175–176. [Google Scholar] [CrossRef]
- Chiu JKH, Chen YPP. Conformational Features of Topologically Classified RNA Secondary Structures. PLoS One. 2012, 7, e39907. [Google Scholar]
- Ferré-D’Amaré AR, Zhou K, Doudna JA. Crystal structure of a hepatitis delta virus ribozyme. Nature. 1998, 395, 567–574. [Google Scholar] [CrossRef]
- Tomita K, Ishitani R, Fukai S, Nureki O. Complete crystallographic analysis of the dynamics of CCA sequence addition. Nature. 2006, 443, 956–960. [Google Scholar] [CrossRef] [PubMed]
- Liberati A, Altman DG, Tetzlaff J, Mulrow C, Gøtzsche PC, Ioannidis JPA, et al. The PRISMA statement for reporting systematic reviews and meta-analyses of studies that evaluate health care interventions: explanation and elaboration. J Clin Epidemiol. 2009, 62, e1–34. [Google Scholar] [CrossRef]
- Mathews DH. How to benchmark RNA secondary structure prediction accuracy. Methods. 2019, 162–16, 60–67.
- Wang Y, Liu Y, Wang S, Liu Z, Gao Y, Zhang H, et al. ATTfold: RNA Secondary Structure Prediction With Pseudoknots Based on Attention Mechanism. Front Genet. 2020 Dec 15;11.
- An JY, Meng FR, Yan ZJ. An efficient computational method for predicting drug-target interactions using weighted extreme learning machine and speed up robot features. BioData Min. 2021, 14, 3. [Google Scholar]
- Sun Y, Liu F, Fan C, Wang Y, Song L, Fang Z, et al. Characterizing sensitivity and coverage of clinical WGS as a diagnostic test for genetic disorders. BMC Med Genomics. 2021, 14, 102. [Google Scholar]
- Fu L, Cao Y, Wu J, Peng Q, Nie Q, Xie X. UFold: fast and accurate RNA secondary structure prediction with deep learning. Nucleic Acids Res. 2022, 50, e14–e14. [Google Scholar] [CrossRef]
- Sikka J, Satya K, Kumar Y, Uppal S, Shah RR, Zimmermann R. Learning Based Methods for Code Runtime Complexity Prediction. In 2020. p. 313–25.
- Bellaousov S, Mathews DH. ProbKnot: Fast prediction of RNA secondary structure including pseudoknots. RNA. 2010, 16, 1870–1880. [Google Scholar] [CrossRef]
- Li H, Zhu D, Zhang C, Han H, Crandall KA. Characteristics and Prediction of RNA Structure. Biomed Res Int. 2014, 2014, 1–10. [Google Scholar]
- MATHEWS, DH. Using an RNA secondary structure partition function to determine confidence in base pairs predicted by free energy minimization. RNA. 2004, 10, 1178–1190. [Google Scholar] [CrossRef] [PubMed]
- Do CB, Woods DA, Batzoglou S. CONTRAfold: RNA secondary structure prediction without physics-based models. Bioinformatics. 2006, 22, e90–8. [Google Scholar] [CrossRef] [PubMed]
- Xia T, SantaLucia J, Burkard ME, Kierzek R, Schroeder SJ, Jiao X, et al. Thermodynamic Parameters for an Expanded Nearest-Neighbor Model for Formation of RNA Duplexes with Watson−Crick Base Pairs. Biochemistry. 1998, 37, 14719–14735. [Google Scholar] [CrossRef] [PubMed]
- Jabbari H, Wark I, Montemagno C. RNA secondary structure prediction with pseudoknots: Contribution of algorithm versus energy model. PLoS One. 2018, 13, e0194583. [Google Scholar]
- Nussinov R, Pieczenik G, Griggs JR, Kleitman DJ. Algorithms for Loop Matchings. SIAM J Appl Math. 1978, 35, 68–82. [Google Scholar] [CrossRef]
- Michael Zuker, David Sankoff. RNA SECONDARY STRUCTURES AND THEIR PREDICTION. Bull Math Biol. 1984, 46, 591–621. [Google Scholar] [CrossRef]
- Andronescu M, Bereg V, Hoos HH, Condon A. RNA STRAND: The RNA Secondary Structure and Statistical Analysis Database. BMC Bioinformatics. 2008, 9, 340. [Google Scholar]
| Name of the folding engine | Method of Prediction | Thermodynamic model parameters | Pseudoknots enforced | Auxiliary parameters enforced | References |
|---|---|---|---|---|---|
| Chiba Institute of Technology’s Vsfold5: RNA Secondary Structure Prediction Server | MEA | Jacobson-Stockmayer (standard parameters) | Yes | Kuhn length, Mg2+ binding, contiguous stems, minimum stem length, length of leading stem | [26] |
| Kuhn length, Mg2+ binding, contiguous stems, minimum stem length, length of leading stem | |||||
| Universitat Bielefeld’s BiBiServ’s (pKiss) | MFE | Turner model | Yes | H-type penalty, K-type penalty, maximal pseudoknot size, minimal hairpin length, lonely base pairs | [21] |
| InstitutCurie’s Kinefold | MFE | Turner model | No | Co-transcriptional Fold, Simulated molecular time, Tracing and Forcing Helices | [27] |
| NUPACK 3.0 | MFE | Turner model | No | ||
| Mg2+ binding, dangling end options, input of multiple interacting | [28,29] | ||||
| Vienna RNAfold | MFE | Turner model | No | Avoiding isolated base pairs, incorporation of G–Quadruplex formation into the structure prediction algorithm, dangling end options, addition of modified base pairs | [30] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




