Submitted:
30 January 2024
Posted:
31 January 2024
You are already at the latest version
Abstract
Keywords:
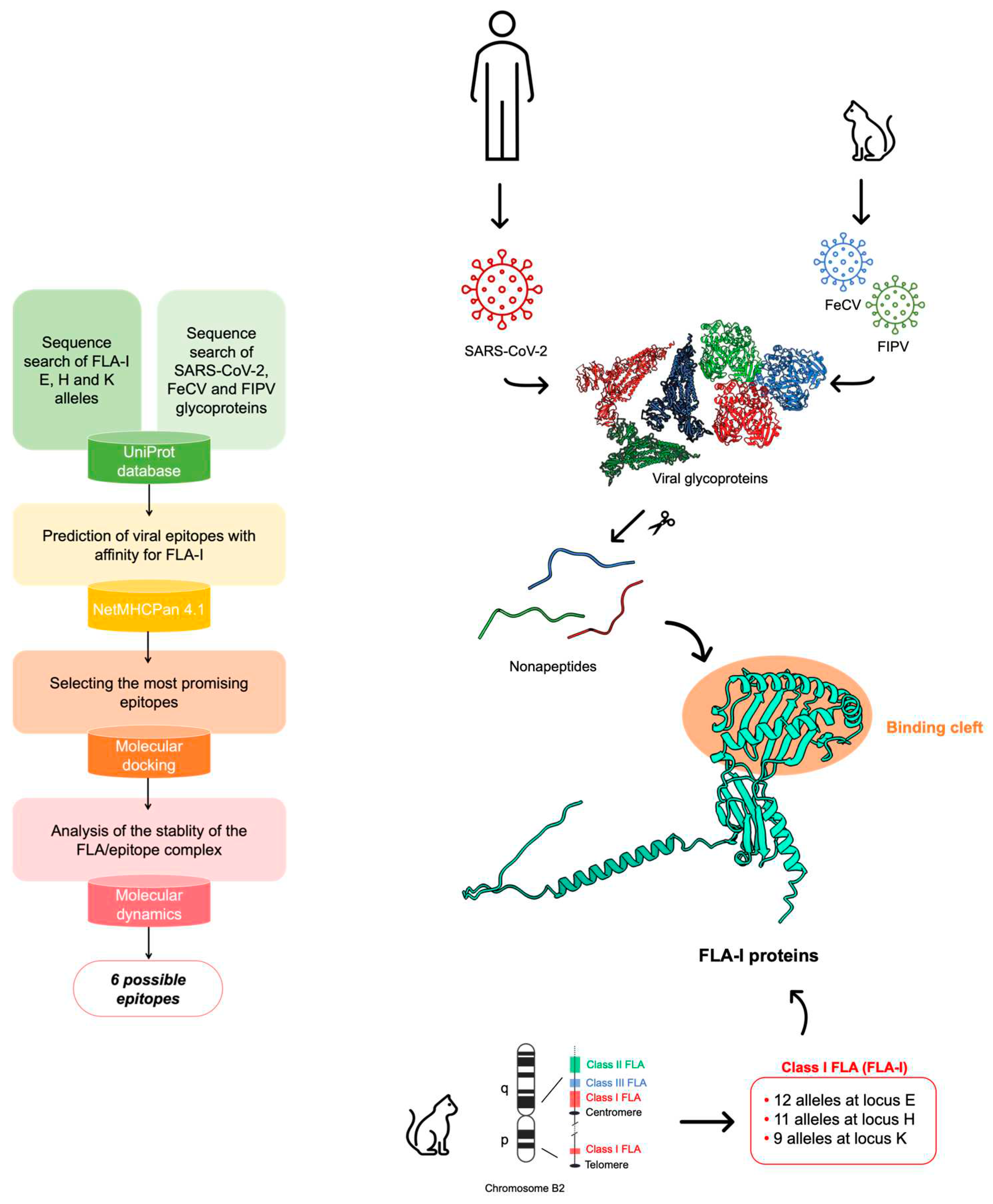
1. Introduction
2. Materials and Methods
Epitope mapping, sequence alignment and peptide search
Molecular docking
Protein structure prediction with AlphaFold2
Molecular dynamics
3. Results
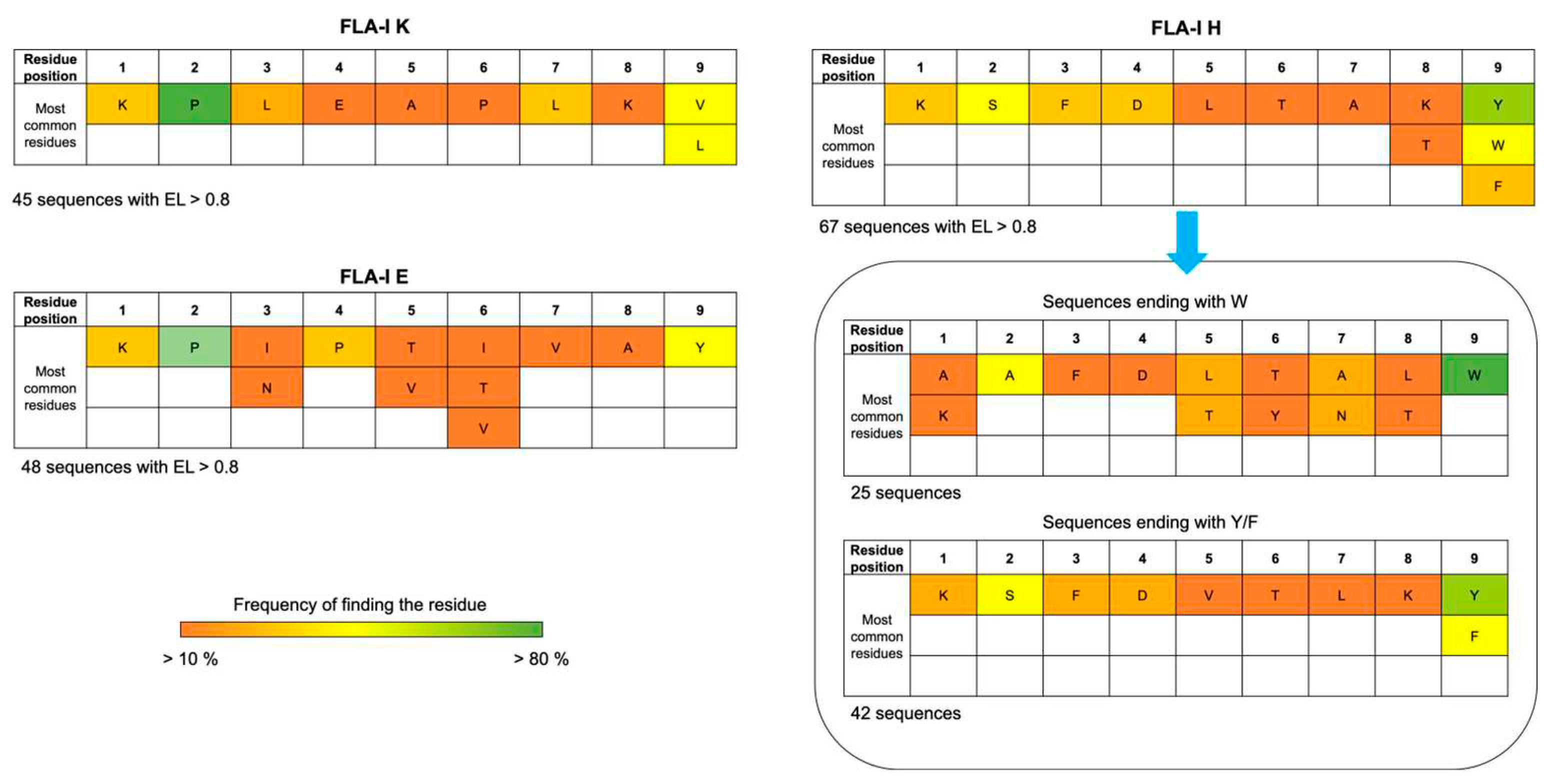
Epitope mapping
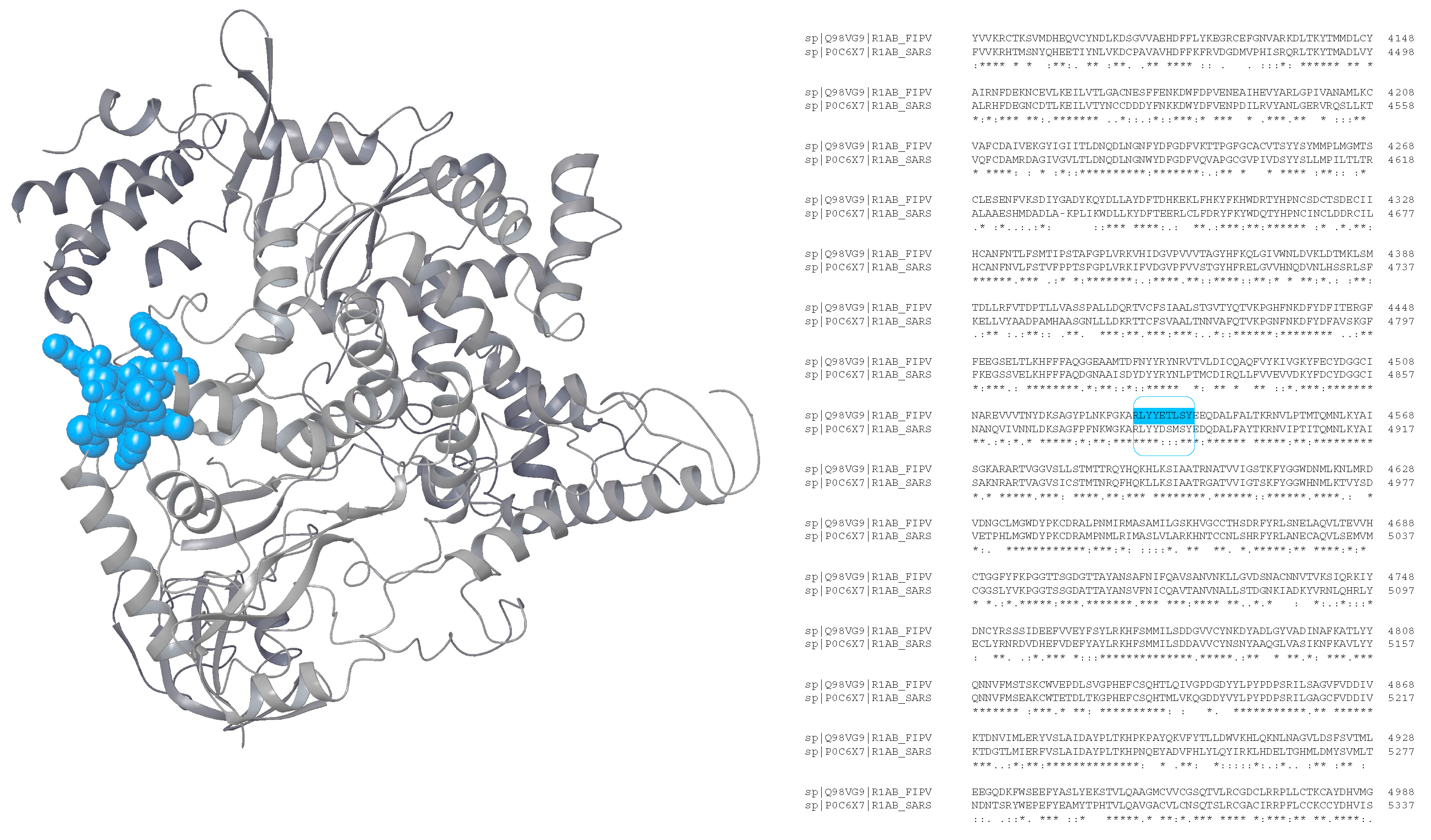
Sequence alignment
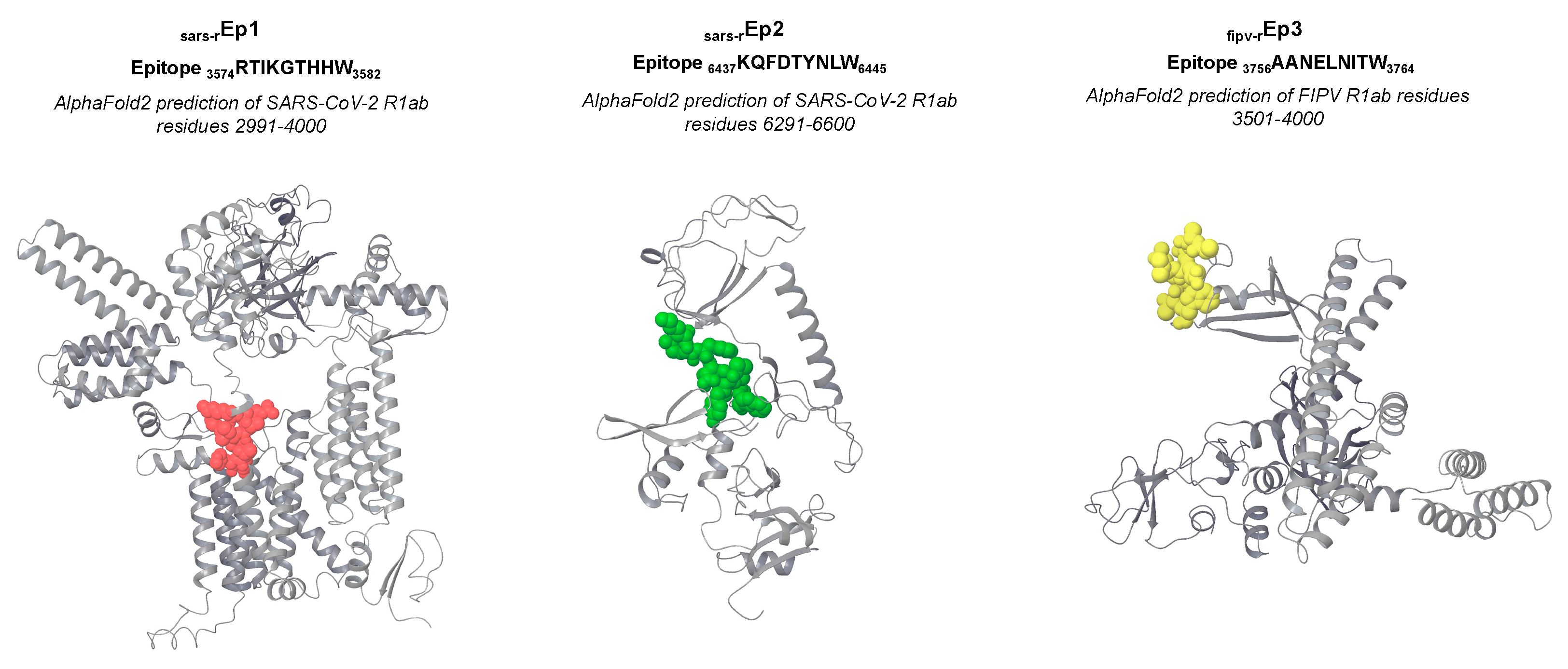
Molecular docking
| Epitope | Alias |
| SARS-CoV-2 R1a 3574RTIKGTHHW3582 | sars-rEp1 |
| SARS-CoV-2 R1ab 6437KQFDTYNLW6445 | sars-rEp2 |
| FIPV R1ab 3756AANELNITW3764 | fipv-rEp3 |
| FIPV R1ab 4533RLYYETLSY4541 | fipv-rEp4 |
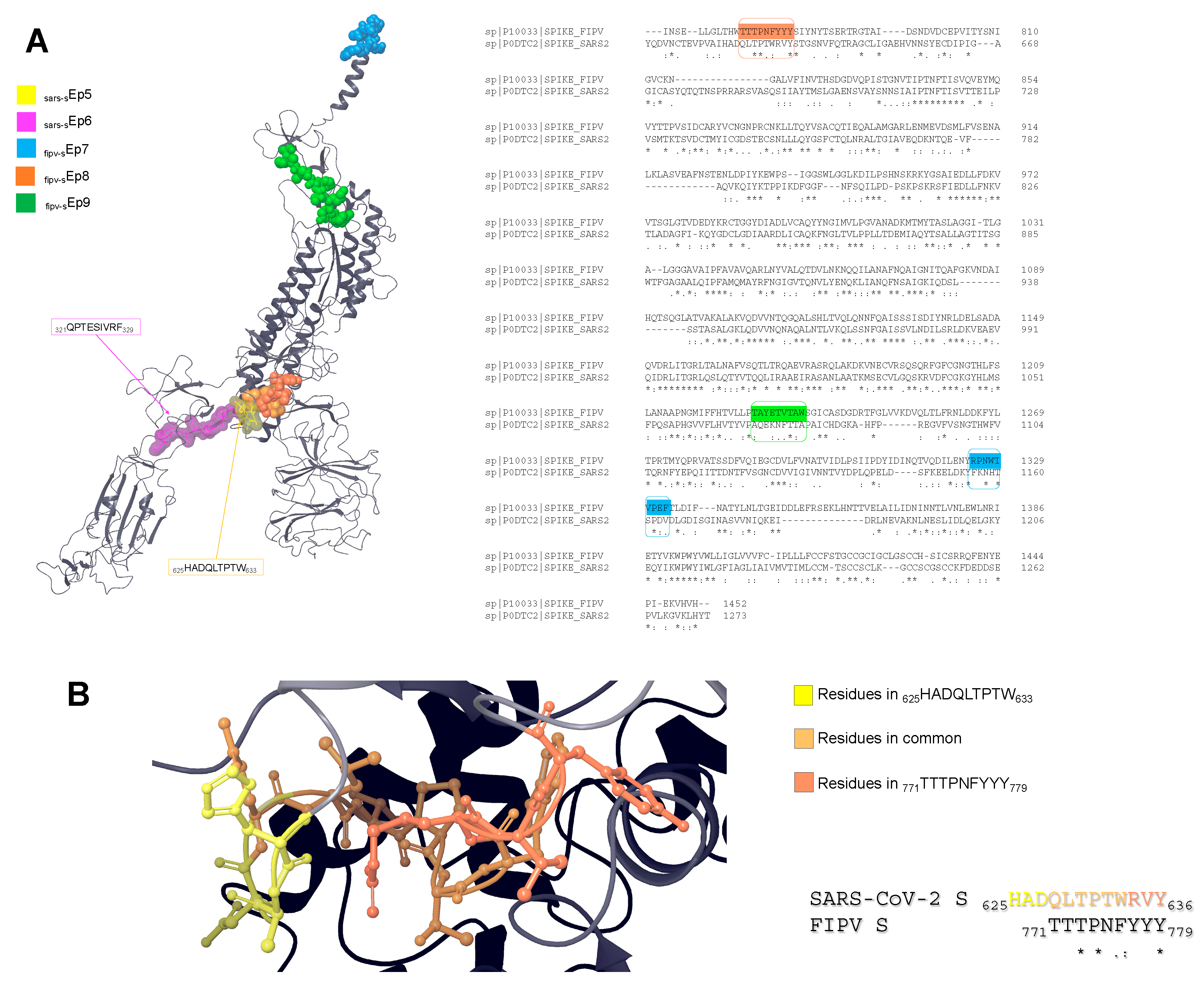
| SARS-CoV-2 S 625HADQLTPTW633 | sars-sEp5 |
| SARS-CoV-2 S 321QPTESIVRF329 | sars-sEp6 |
| FIPV S 1325RPNWTVPEF1333 | fipv-sEp7 |
| FIPV S 771TTTPNFYYY779 | fipv-sEp8 |
| FIPV S 1228TAYETVTAW1236 | fipv-sEp9 |
Analysis of possible surface exposition
Molecular dynamics
- sars-rEp1 with FLA-I H*00501 for the outstanding docking score;
- fipv-rEp4 with FLA-I H*00501 for the docking score and the possible localization on the viral glycoprotein surface;
- sars-sEp5 with FLA-I E*01001 and sars-sEp6 with FLA-I E*00701 for the docking scores and the position on the viral glycoprotein surface;
- fipv-sEp8 with FLA-I H*00501 and fipv-sEp9 with H*00401 for the docking scores and the possible localization on the viral glycoprotein surface.
Peptide search
4. Discussion and conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Conflicts of Interest
References
- A. Kipar, M.L. Meli, Feline infectious peritonitis: still an enigma?, Vet Pathol 51(2) (2014) 505-26. [CrossRef]
- S.J. Coggins, J.M. Norris, R. Malik, M. Govendir, E.J. Hall, B. Kimble, M.F. Thompson, Outcomes of treatment of cats with feline infectious peritonitis using parenterally administered remdesivir, with or without transition to orally administered GS-441524, Journal of Veterinary Internal Medicine (2023). [CrossRef]
- A. Sweet, N. Andre, G. Whittaker, RNA in-situ hybridization for pathology-based diagnosis of feline infectious peritonitis (FIP): current diagnostics for FIP and comparison to the current gold standard, Qeios (2022). [CrossRef]
- C.W. Olsen, W.V. Corapi, C.K. Ngichabe, J.D. Baines, F.W. Scott, Monoclonal antibodies to the spike protein of feline infectious peritonitis virus mediate antibody-dependent enhancement of infection of feline macrophages, Journal of virology 66(2) (1992) 956-965. [CrossRef]
- N.C. Pedersen, An overview of feline enteric coronavirus and infectious peritonitis virus infections, Feline Practice 23 (1995) 7-20.
- T. Hayashi, K. Doi, K. Fujiwara, Role of circulating antibodies and thymus-dependent lymphocytes in production of effusive type feline infectious peritonitis after oral infection, Molecular biology and pathogenesis of coronaviruses (1984) 383-384. [CrossRef]
- J. Shi, Z. Wen, G. Zhong, H. Yang, C. Wang, B. Huang, R. Liu, X. He, L. Shuai, Z. Sun, Y. Zhao, P. Liu, L. Liang, P. Cui, J. Wang, X. Zhang, Y. Guan, W. Tan, G. Wu, H. Chen, Z. Bu, Susceptibility of ferrets, cats, dogs, and other domesticated animals to SARS-coronavirus 2, Science 368(6494) (2020) 1016-1020. [CrossRef]
- T.H.C. Sit, C.J. Brackman, S.M. Ip, K.W.S. Tam, P.Y.T. Law, E.M.W. To, V.Y.T. Yu, L.D. Sims, D.N.C. Tsang, D.K.W. Chu, R. Perera, L.L.M. Poon, M. Peiris, Infection of dogs with SARS-CoV-2, Nature 586(7831) (2020) 776-778. [CrossRef]
- M. Buonocore, C. Marino, M. Grimaldi, A. Santoro, M. Firoznezhad, O. Paciello, F. Prisco, A.M. D'Ursi, New putative animal reservoirs of SARS-CoV-2 in Italian fauna: A bioinformatic approach, Heliyon 6(11) (2020) e05430. [CrossRef]
- S.J. O'Brien, N. Yuhki, Comparative genome organization of the major histocompatibility complex: lessons from the Felidae, Immunological Reviews 167(1) (1999) 133-144. [CrossRef]
- R.P. Sutmuller, R. Offringa, C.J. Melief, Revival of the regulatory T cell: new targets for drug development, Drug Discov Today 9(7) (2004) 310-6. [CrossRef]
- B.M. Gonzales-Portillo, J.Y. Lee, A.A. Vandenbark, H. Offner, C.V. Borlongan, Major histocompatibility complex Class II-based therapy for stroke, Brain Circ 7(1) (2021) 37-40. [CrossRef]
- J.Y. Lee, V. Castelli, B. Bonsack, A.B. Coats, L. Navarro-Torres, J. Garcia-Sanchez, C. Kingsbury, H. Nguyen, A.A. Vandenbark, R. Meza-Romero, H. Offner, C.V. Borlongan, A Novel Partial MHC Class II Construct, DRmQ, Inhibits Central and Peripheral Inflammatory Responses to Promote Neuroprotection in Experimental Stroke, Transl Stroke Res 11(4) (2020) 831-836. [CrossRef]
- J. Brown, C. Kingsbury, J.Y. Lee, A.A. Vandenbark, R. Meza-Romero, H. Offner, C.V. Borlongan, Spleen participation in partial MHC class II construct neuroprotection in stroke, CNS Neurosci Ther 26(7) (2020) 663-669. [CrossRef]
- G. Benedek, A.A. Vandenbark, N.J. Alkayed, H. Offner, Partial MHC class II constructs as novel immunomodulatory therapy for stroke, Neurochemistry international 107 (2017) 138-147. [CrossRef]
- G. Benedek, W. Zhu, N. Libal, A. Casper, X. Yu, R. Meza-Romero, A.A. Vandenbark, N.J. Alkayed, H. Offner, A novel HLA-DRα1-MOG-35-55 construct treats experimental stroke, Metabolic brain disease 29 (2014) 37-45. [CrossRef]
- P. Correale, L. Mutti, F. Pentimalli, G. Baglio, R.E. Saladino, P. Sileri, A. Giordano, HLA-B*44 and C*01 Prevalence Correlates with Covid19 Spreading across Italy, Int J Mol Sci 21(15) (2020). [CrossRef]
- R. Liang, Y. Sun, Y. Liu, J. Wang, Y. Wu, Z. Li, L. Ma, N. Zhang, L. Zhang, X. Wei, Z. Qu, N. Zhang, C. Xia, Major Histocompatibility Complex Class I (FLA-E*01801) Molecular Structure in Domestic Cats Demonstrates Species-Specific Characteristics in Presenting Viral Antigen Peptides, J Virol 92(6) (2018). [CrossRef]
- T.W. Beck, J. Menninger, W.J. Murphy, W.G. Nash, S.J. O'Brien, N. Yuhki, The feline major histocompatibility complex is rearranged by an inversion with a breakpoint in the distal class I region, Immunogenetics 56(10) (2005) 702-709. [CrossRef]
- J.C. Holmes, S.G. Holmer, P. Ross, A.S. Buntzman, J.A. Frelinger, P.R. Hess, Polymorphisms and tissue expression of the feline leukocyte antigen class I loci FLAI-E, FLAI-H, and FLAI-K, Immunogenetics 65(9) (2013) 675-89. [CrossRef]
- A.M. Aranyos, S.R. Roff, R. Pu, J.L. Owen, J.K. Coleman, J.K. Yamamoto, An initial examination of the potential role of T-cell immunity in protection against feline immunodeficiency virus (FIV) infection, Vaccine 34(12) (2016) 1480-8. [CrossRef]
- M. Omori, R. Pu, T. Tanabe, W. Hou, J. Coleman, M. Arai, J. Yamamoto, Cellular immune responses to feline immunodeficiency virus (FIV) induced by dual-subtype FIV vaccine, Vaccine 23(3) (2004) 386-398. [CrossRef]
- R. Pu, M. Omori, S. Okada, S.L. Rine, B.A. Lewis, E. Lipton, J.K. Yamamoto, MHC-restricted protection of cats against FIV infection by adoptive transfer of immune cells from FIV-vaccinated donors, Cell Immunol 198(1) (1999) 30-43. [CrossRef]
- B. Reynisson, B. Alvarez, S. Paul, B. Peters, M. Nielsen, NetMHCpan-4.1 and NetMHCIIpan-4.0: improved predictions of MHC antigen presentation by concurrent motif deconvolution and integration of MS MHC eluted ligand data, Nucleic Acids Res 48(W1) (2020) W449-W454. [CrossRef]
- C. UniProt, UniProt: the Universal Protein Knowledgebase in 2023, Nucleic Acids Res 51(D1) (2023) D523-D531. [CrossRef]
- H.G. Rammensee, Chemistry of peptides associated with MHC class I and class II molecules, Curr Opin Immunol 7(1) (1995) 85-96. [CrossRef]
- P. Zhou, B. Jin, H. Li, S.Y. Huang, HPEPDOCK: a web server for blind peptide-protein docking based on a hierarchical algorithm, Nucleic Acids Res 46(W1) (2018) W443-W450. [CrossRef]
- Y. Zhang, M.F. Sanner, AutoDock CrankPep: combining folding and docking to predict protein-peptide complexes, Bioinformatics 35(24) (2019) 5121-5127. [CrossRef]
- R.N. Kirchdoerfer, A.B. Ward, Structure of the SARS-CoV nsp12 polymerase bound to nsp7 and nsp8 co-factors, Nat Commun 10(1) (2019) 2342. [CrossRef]
- J. Jumper, R. Evans, A. Pritzel, T. Green, M. Figurnov, O. Ronneberger, K. Tunyasuvunakool, R. Bates, A. Zidek, A. Potapenko, A. Bridgland, C. Meyer, S.A.A. Kohl, A.J. Ballard, A. Cowie, B. Romera-Paredes, S. Nikolov, R. Jain, J. Adler, T. Back, S. Petersen, D. Reiman, E. Clancy, M. Zielinski, M. Steinegger, M. Pacholska, T. Berghammer, S. Bodenstein, D. Silver, O. Vinyals, A.W. Senior, K. Kavukcuoglu, P. Kohli, D. Hassabis, Highly accurate protein structure prediction with AlphaFold, Nature 596(7873) (2021) 583-589. [CrossRef]
- M. Mirdita, K. Schütze, Y. Moriwaki, L. Heo, S. Ovchinnikov, M. Steinegger, ColabFold: making protein folding accessible to all, Nature methods 19(6) (2022) 679-682. [CrossRef]
- K. Wang, Z. Jia, L. Bao, L. Wang, L. Cao, H. Chi, Y. Hu, Q. Li, Y. Zhou, Y. Jiang, Q. Zhu, Y. Deng, P. Liu, N. Wang, L. Wang, M. Liu, Y. Li, B. Zhu, K. Fan, W. Fu, P. Yang, X. Pei, Z. Cui, L. Qin, P. Ge, J. Wu, S. Liu, Y. Chen, W. Huang, Q. Wang, C.F. Qin, Y. Wang, C. Qin, X. Wang, Memory B cell repertoire from triple vaccinees against diverse SARS-CoV-2 variants, Nature 603(7903) (2022) 919-925. [CrossRef]
- S. Di Micco, R. Rahimova, M. Sala, M.C. Scala, G. Vivenzio, S. Musella, G. Andrei, K. Remans, L. Mammri, R. Snoeck, G. Bifulco, F. Di Matteo, V. Vestuto, P. Campiglia, J.A. Marquez, A. Fasano, Rational design of the zonulin inhibitor AT1001 derivatives as potential anti SARS-CoV-2, Eur J Med Chem 244 (2022) 114857. [CrossRef]
- S. Weingarten-Gabbay, S. Klaeger, S. Sarkizova, L.R. Pearlman, D.Y. Chen, K.M.E. Gallagher, M.R. Bauer, H.B. Taylor, W.A. Dunn, C. Tarr, J. Sidney, S. Rachimi, H.L. Conway, K. Katsis, Y. Wang, D. Leistritz-Edwards, M.R. Durkin, C.H. Tomkins-Tinch, Y. Finkel, A. Nachshon, M. Gentili, K.D. Rivera, I.P. Carulli, V.A. Chea, A. Chandrashekar, C.C. Bozkus, M. Carrington, M.C.-. Collection, T. Processing, N. Bhardwaj, D.H. Barouch, A. Sette, M.V. Maus, C.M. Rice, K.R. Clauser, D.B. Keskin, D.C. Pregibon, N. Hacohen, S.A. Carr, J.G. Abelin, M. Saeed, P.C. Sabeti, Profiling SARS-CoV-2 HLA-I peptidome reveals T cell epitopes from out-of-frame ORFs, Cell 184(15) (2021) 3962-3980 e17. [CrossRef]
- A. Tarke, C.H. Coelho, Z. Zhang, J.M. Dan, E.D. Yu, N. Methot, N.I. Bloom, B. Goodwin, E. Phillips, S. Mallal, J. Sidney, G. Filaci, D. Weiskopf, R. da Silva Antunes, S. Crotty, A. Grifoni, A. Sette, SARS-CoV-2 vaccination induces immunological T cell memory able to cross-recognize variants from Alpha to Omicron, Cell 185(5) (2022) 847-859 e11. [CrossRef]
- P. Gattinger, K. Niespodziana, K. Stiasny, S. Sahanic, I. Tulaeva, K. Borochova, Y. Dorofeeva, T. Schlederer, T. Sonnweber, G. Hofer, R. Kiss, B. Kratzer, D. Trapin, P.A. Tauber, A. Rottal, U. Kormoczi, M. Feichter, M. Weber, M. Focke-Tejkl, J. Loffler-Ragg, B. Muhl, A. Kropfmuller, W. Keller, F. Stolz, R. Henning, I. Tancevski, E. Puchhammer-Stockl, W.F. Pickl, R. Valenta, Neutralization of SARS-CoV-2 requires antibodies against conformational receptor-binding domain epitopes, Allergy 77(1) (2022) 230-242. [CrossRef]
- S. Zhang, S. Qiao, J. Yu, J. Zeng, S. Shan, L. Tian, J. Lan, L. Zhang, X. Wang, Bat and pangolin coronavirus spike glycoprotein structures provide insights into SARS-CoV-2 evolution, Nat Commun 12(1) (2021) 1607. [CrossRef]
- M.C. Horzinek, H. Lutz, N. Pedersen, Antigenic relationships among homologous structural polypeptides of porcine, feline, and canine coronaviruses, Infection and Immunity 37(3) (1982) 1148-1155. [CrossRef]
- L. Schrödinger, Schrödinger Release 2023-2: Maestro, New York, NY, 2023.
- S.F. Altschul, W. Gish, W. Miller, E.W. Myers, D.J. Lipman, Basic local alignment search tool, J Mol Biol 215(3) (1990) 403-10. [CrossRef]
- M.J. Abraham, T. Murtola, R. Schulz, S. Páll, J.C. Smith, B. Hess, E. Lindahl, GROMACS: High performance molecular simulations through multi-level parallelism from laptops to supercomputers, SoftwareX 1 (2015) 19-25. [CrossRef]
- J. Huang, A.D. MacKerell Jr, CHARMM36 all-atom additive protein force field: Validation based on comparison to NMR data, Journal of computational chemistry 34(25) (2013) 2135-2145. [CrossRef]
| Extended conformation | Helix conformation | ||||||
|---|---|---|---|---|---|---|---|
| Organism | Viral glycoprotein | Epitope | FLA-I Receptor | Docking score (kcal/mol) | Number of poses with RMSD < 6 Å | Docking score (kcal/mol) | Number of poses with RMSD < 6 Å |
| SARS-CoV-2 | R1a | RTIKGTHHW | FLA-I H*00501 | -22.6 | 41 | -22.5 | 40 |
| SARS-CoV-2 | R1ab | KQFDTYNLW | FLA-I K*00701 | -20.8 | 27 | -19.1 | 51 |
| FIPV | R1ab | AANELNITW | FLA-I H*00501 | -20.5 | 18 | -19 | 31 |
| FIPV | R1ab | RLYYETLSY | FLA-I H*00501 | -19.8 | 22 | -20.2 | 43 |
| SARS-CoV-2 | R1ab | KQFDTYNLW | FLA-I H*00501 | -19.3 | 22 | -17 | 32 |
| SARS-CoV-2 | R1a | RTIKGTHHW | FLA-I K*00701 | -19.2 | 38 | -18.7 | 35 |
| SARS-CoV-2 | R1a | RTIKGTHHW | FLA-I H*00401 | -18.7 | 15 | -19.6 | 7 |
| FIPV | R1ab | YNLDIPHKL | FLA-I K*00701 | -18.7 | 19 | -18.3 | 26 |
| SARS-CoV-2 | R1a | RTIKGTHHW | FLA-I H*008012 | -18.4 | 7 | -18.1 | 8 |
| SARS-CoV-2 | R1a | VPFWITIAY | FLA-I E*00701 | -17.1 | 6 | -18.1 | 16 |
| FIPV | R1ab | AANELNITW | FLA-I H*00401 | -17 | 19 | -18.1 | 25 |
| SARS-CoV-2 | R1a | VPFWITIAY | FLA-I K*00801 | -16.8 | 20 | -15.2 | 40 |
| FIPV | R1ab | RLYYETLSY | FLA-I H*00401 | -16.7 | 22 | -18 | 23 |
| FIPV | S | RPNWTVPEF | FLA-I E*00701 | -16.6 | 21 | -15.4 | 27 |
| FIPV | S | TTTPNFYYY | FLA-I H*00501 | -16.5 | 6 | -16.3 | 44 |
| SARS-CoV-2 | R1a | VPMEKLKTL | FLA-I E*00701 | -16.4 | 29 | -15.8 | 22 |
| SARS-CoV-2 | R1a | RTIKVFTTV | FLA-I E*01101 | -16.4 | 2 | -14.7 | 21 |
| SARS-CoV-2 | S | HADQLTPTW | FLA-I E*01001 | -16.3 | 55 | -14.5 | 19 |
| FIPV | S | RPNWTVPEF | FLA-I E*00101 | -16.2 | 29 | -13.2 | 21 |
| SARS-CoV-2 | R1a | VPFWITIAY | FLA-I E*00101 | -16.1 | 30 | -17.9 | 39 |
| FIPV | R1ab | QNFDTYMLW | FLA-I H*00501 | -16 | 12 | -19.2 | 33 |
| SARS-CoV-2 | S | HADQLTPTW | FLA-I H*008012 | -16 | 9 | -14.5 | 3 |
| SARS-CoV-2 | S | HADQLTPTW | FLA-I E*00101 | -15.9 | 17 | -14.5 | 19 |
| SARS-CoV-2 | S | HADQLTPTW | FLA-I H*00401 | -15.9 | 11 | -16 | 23 |
| SARS-CoV-2 | S | QPTESIVRF | FLA-I E*00701 | -15.6 | 14 | -16.3 | 17 |
| FIPV | S | TAYETVTAW | FLA-I H*00401 | -15.5 | 7 | -15.5 | 10 |
| SARS-CoV-2 | R1a | VPFWITIAY | FLA-I E*00501 | -15.2 | 3 | -15.2 | 7 |
| SARS-CoV-2 | R1a | VPMEKLKTL | FLA-I E*00101 | -15.1 | 16 | -14.5 | 20 |
| SARS-CoV-2 | R1a | VPFWITIAY | FLA-I H*008012 | -14.8 | 16 | -14.6 | 27 |
| SARS-CoV-2 | R1a | VPMEKLKTL | FLA-I K*00801 | -14.8 | 31 | -16.2 | 15 |
| SARS-CoV-2 | S | QPTESIVRF | FLA-I E*00501 | -14.5 | 16 | -14.5 | 2 |
| SARS-CoV-2 | R1a | VPMEKLKTL | FLA-I E*00501 | -14.4 | 9 | -14.2 | 23 |
| SARS-CoV-2 | S | QPTESIVRF | FLA-I E*00101 | -14.4 | 32 | -15.9 | 17 |
| SARS-CoV-2 | R1a | VPFWITIAY | FLA-I E*00501 | -14.3 | 7 | -14.2 | 24 |
| SARS-CoV-2 | R1a | LPSLATVAY | FLA-I E*00101 | -14.1 | 20 | -14.4 | 36 |
| FIPV | R1ab | YPYGSGMVV | FLA-I K*00801 | -13.8 | 35 | -14.8 | 39 |
| FIPV | S | RPNWTVPEF | FLA-I E*00501 | -13.1 | 1 | -11.7 | 5 |
| FIPV | S | TAYETVTAW | FLA-I H*008012 | -13.1 | 11 | -13.6 | 38 |
| FIPV | R1ab | RPIPDVPAY | FLA-I E*00501 | -13 | 16 | -11.3 | 26 |
| SARS-CoV-2 | S | QPTESIVRF | FLA-I E*00501 | -12.7 | 1 | -13.4 | 6 |
| FIPV | R1ab | RPIPDVPAY | FLA-I E*00701 | -12.5 | 11 | -13.7 | 34 |
| SARS-CoV-2 | R1a | LPSLATVAY | FLA-I E*00501 | -12.3 | 40 | -13.2 | 7 |
| FIPV | R1ab | RPIPDVPAY | FLA-I E*00101 | -12.2 | 23 | -10.5 | 20 |
| FIPV | S | TAYETVTAW | FLA-I H*00601 | -11.7 | 18 | -13.7 | 20 |
| SARS-CoV-2 | R1a | VPMEKLKTL | FLA-I E*00501 | -11.4 | 22 | -13.2 | 12 |
| FIPV | R1ab | RPIPDVPAY | FLA-I E*00501 | -11.1 | 5 | -10 | 15 |
| SARS-CoV-2 | R1a | LPSLATVAY | FLA-I E*00501 | -10.1 | 11 | -12.5 | 21 |
| fipv-rEp4 (FIPV R1ab 4533RLYYETLSY4541 – UniProt ID: Q98VG9) | ||
|---|---|---|
| Retrieved in: | ||
| Organism | Protein | UniProt ID |
| Porcine transmissible gastroenteritis coronavirus (strain Purdue) (TGEV) | R1ab | P0C6Y5 |
| Canine coronavirus | R1ab | A0A0D5ZXX1 |
| Mink coronavirus strain WD1133 | R1ab | D9J202 |
| Swine enteric coronavirus | R1ab | A0A0U2LWJ9 |
| Transmissible gastroenteritis virus | R1ab | C8YR34 |
| fipv-sEp8 (FIPV S 771TTTPNFYYY779 – UniProt ID: P10033) | ||
| Retrieved in: | ||
| Organism | Protein | UniProt ID |
| Canine coronavirus (strain BGF10) | S | Q7T6T3 |
| Canine coronavirus K378 | S | Q65984 |
| Canine coronavirus strain Insavc-1 | S | P36300 |
| Porcine transmissible gastroenteritis coronavirus (strain Miller) (TGEV) | S | P33470 |
| Porcine transmissible gastroenteritis coronavirus (strain FS772/70) (TGEV) | S | P18450 |
| Porcine transmissible gastroenteritis coronavirus (strain NEB72-rt) (TGEV) | S | Q01977 |
| Porcine transmissible gastroenteritis coronavirus (strain Purdue) (TGEV) | S | P07946 |
| fipv-sEp9 (FIPV S 771TAYETVTAW779 – UniProt ID: P10033) | ||
| Retrieved in: | ||
| Organism | Protein | UniProt ID |
| Canine coronavirus (strain BGF10) | S | Q7T6T3 |
| Canine coronavirus K378 | S | Q65984 |
| Canine coronavirus strain Insavc-1 | S | P36300 |
| Porcine transmissible gastroenteritis coronavirus (strain Miller) (TGEV) | S | P33470 |
| Porcine transmissible gastroenteritis coronavirus (strain FS772/70) (TGEV) | S | P18450 |
| Porcine transmissible gastroenteritis coronavirus (strain NEB72-rt) (TGEV) | S | Q01977 |
| Porcine transmissible gastroenteritis coronavirus (strain Purdue) (TGEV) | S | P07946 |
| Mink coronavirus strain WD1133 | S | D9J204 |
| Porcine respiratory coronavirus (strain RM4) | S | P24413 |
| Porcine respiratory coronavirus (86/137004/isolate British) | S | P27655 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




