Submitted:
26 January 2024
Posted:
26 January 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results
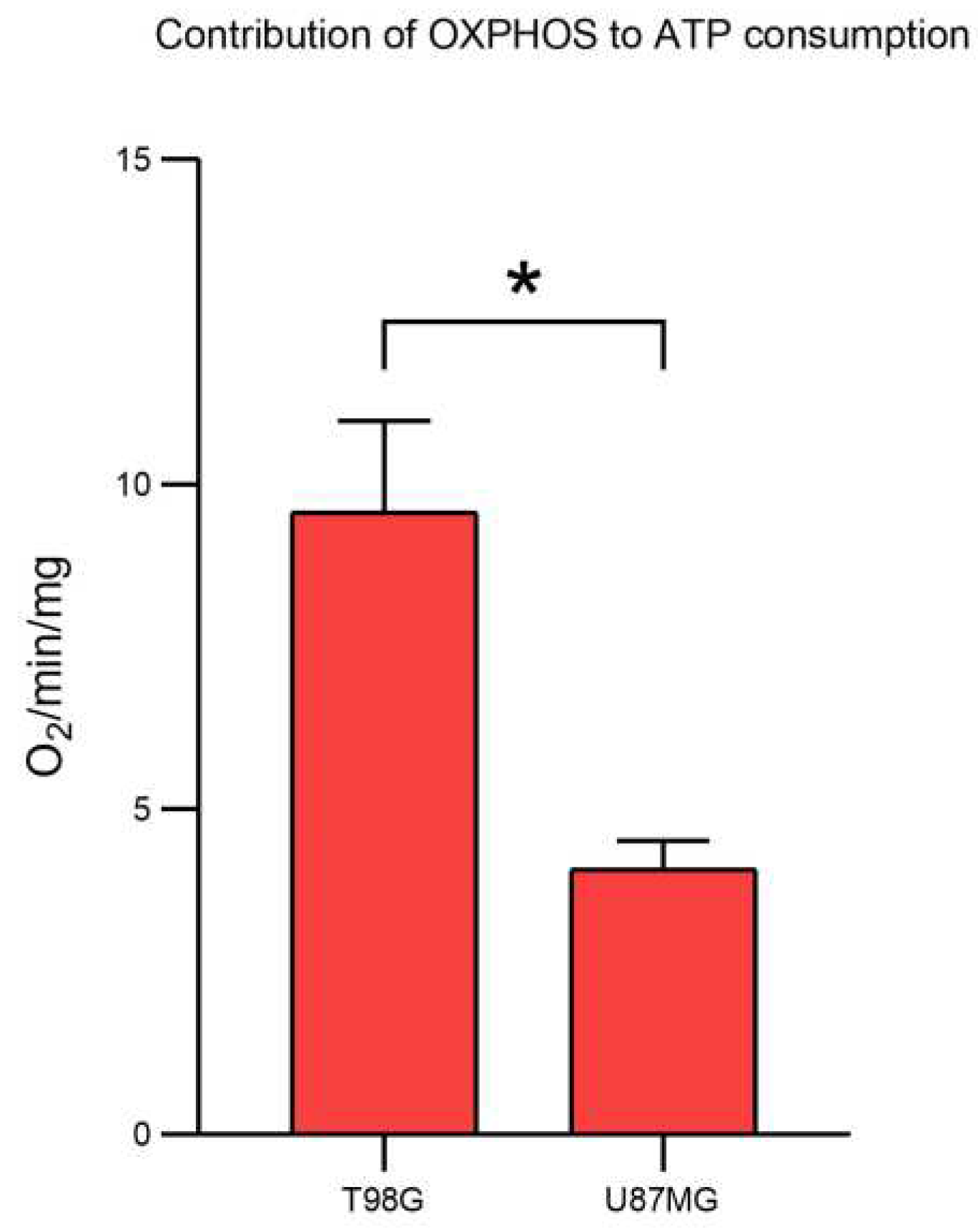
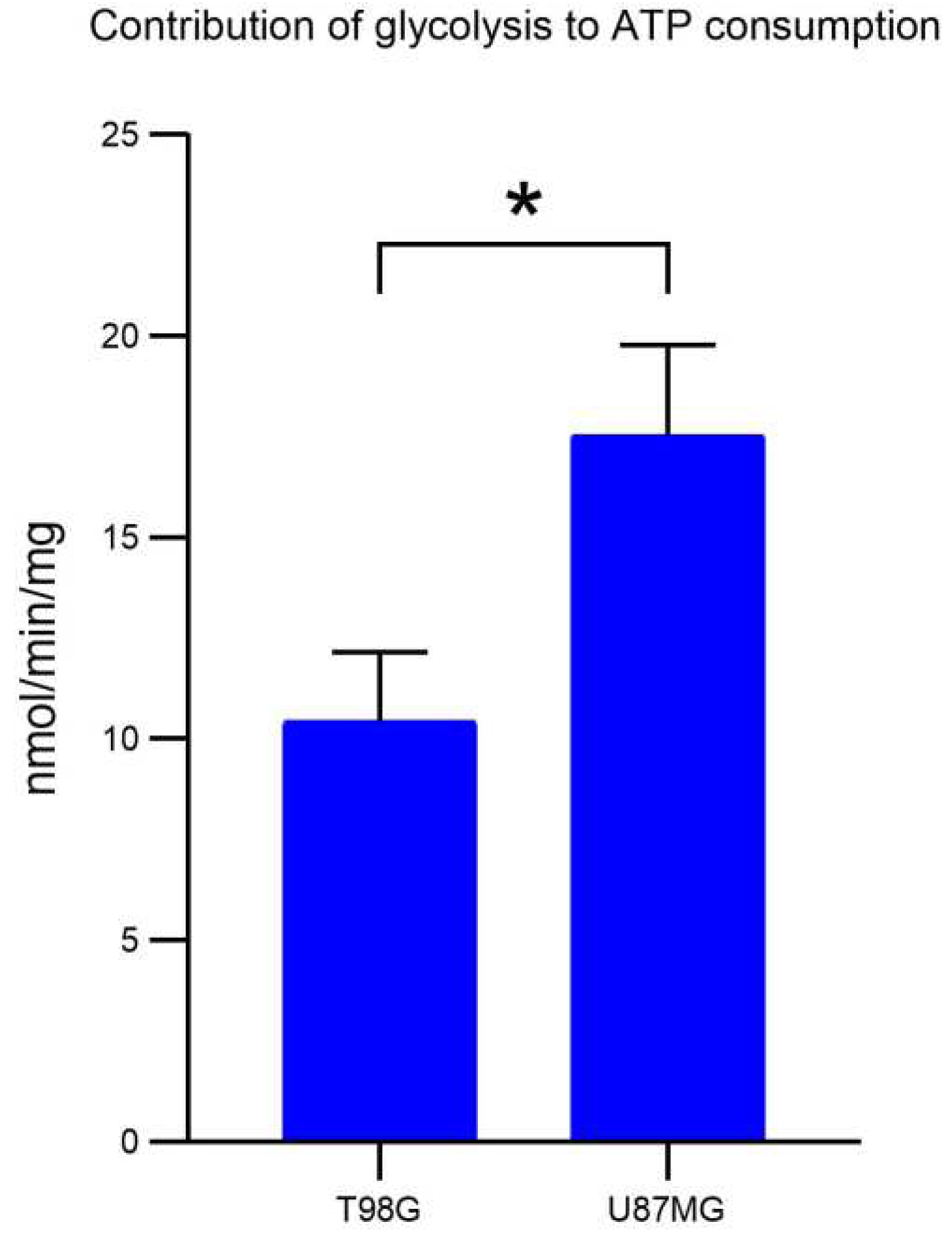
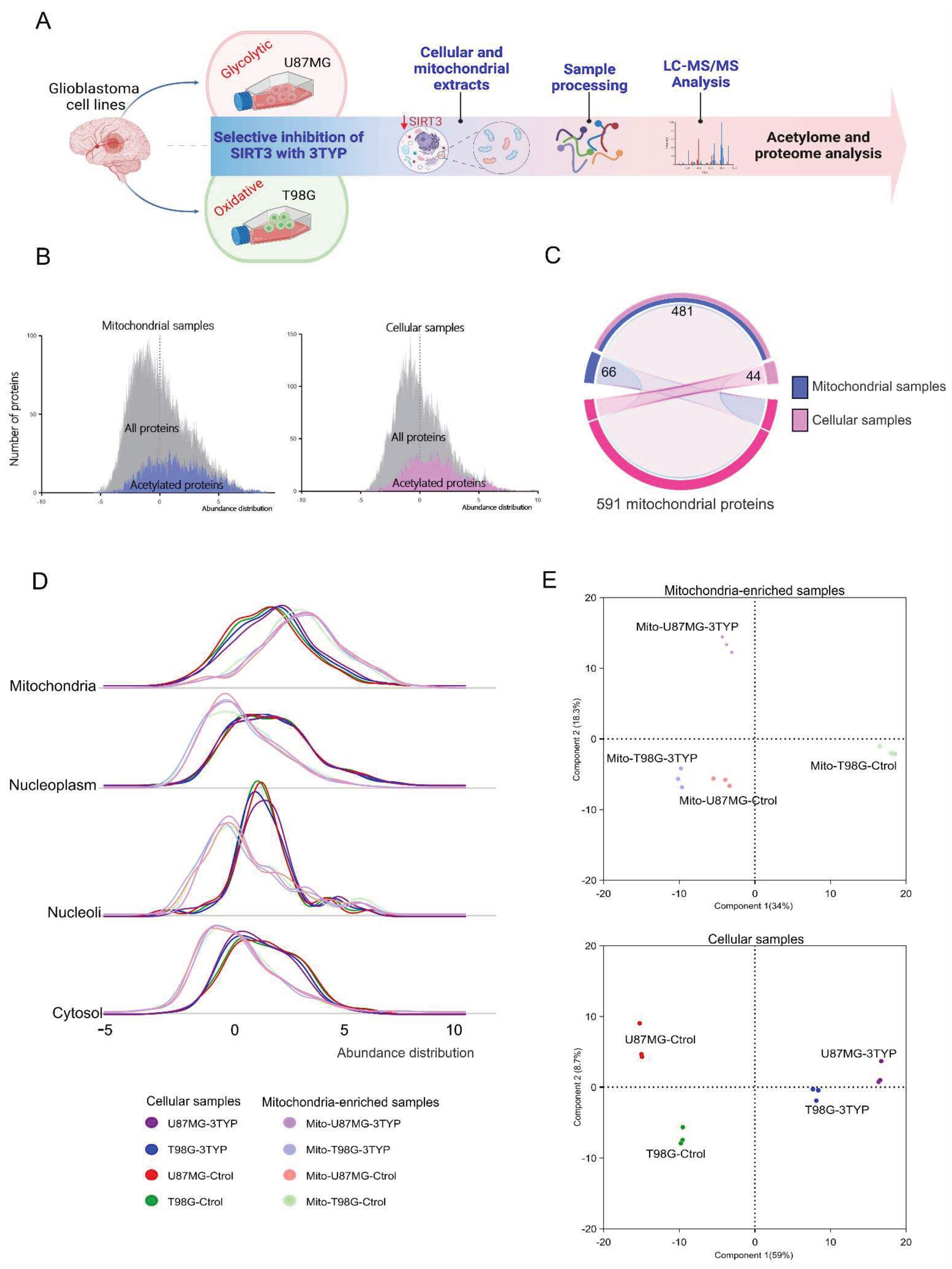
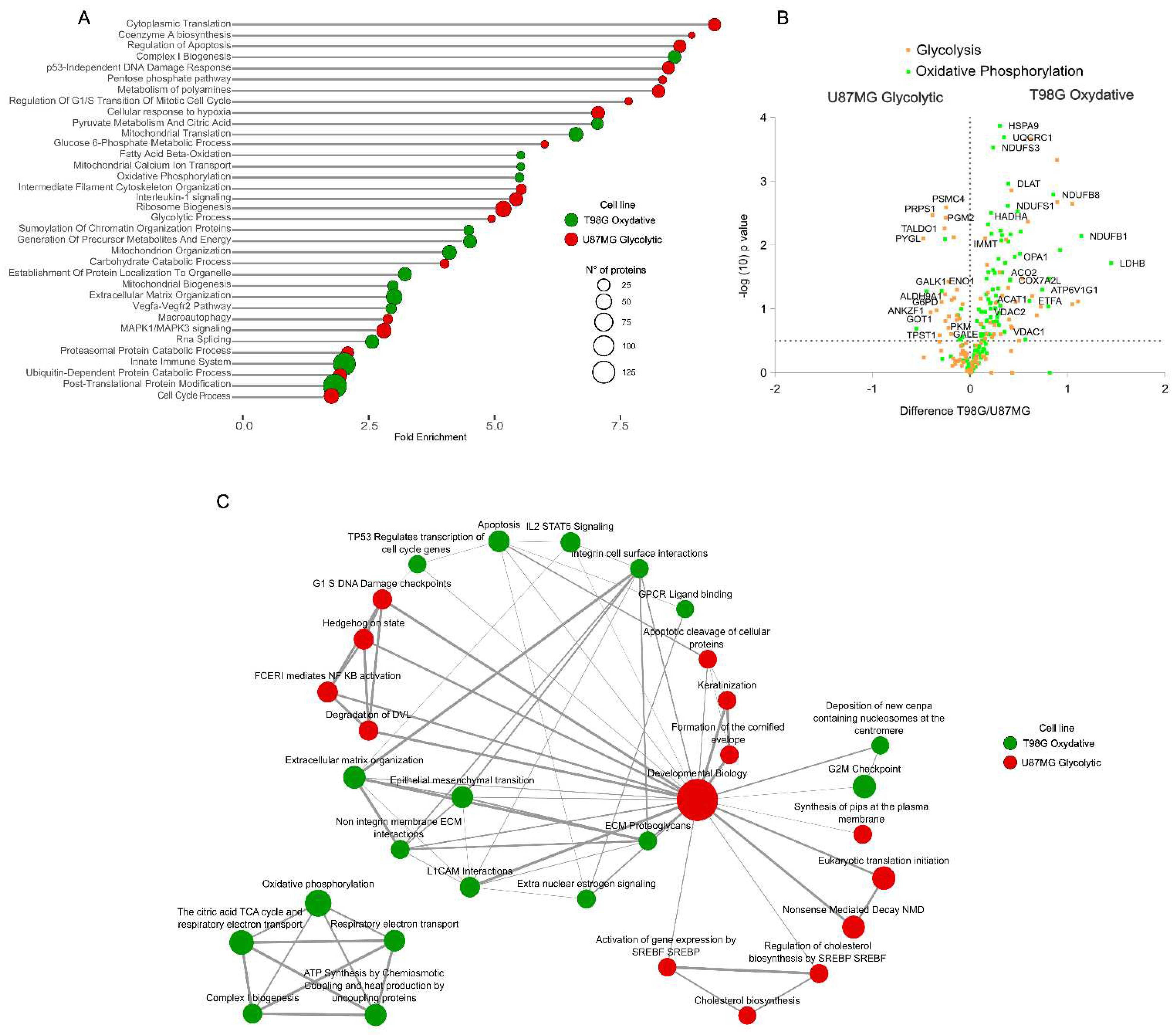
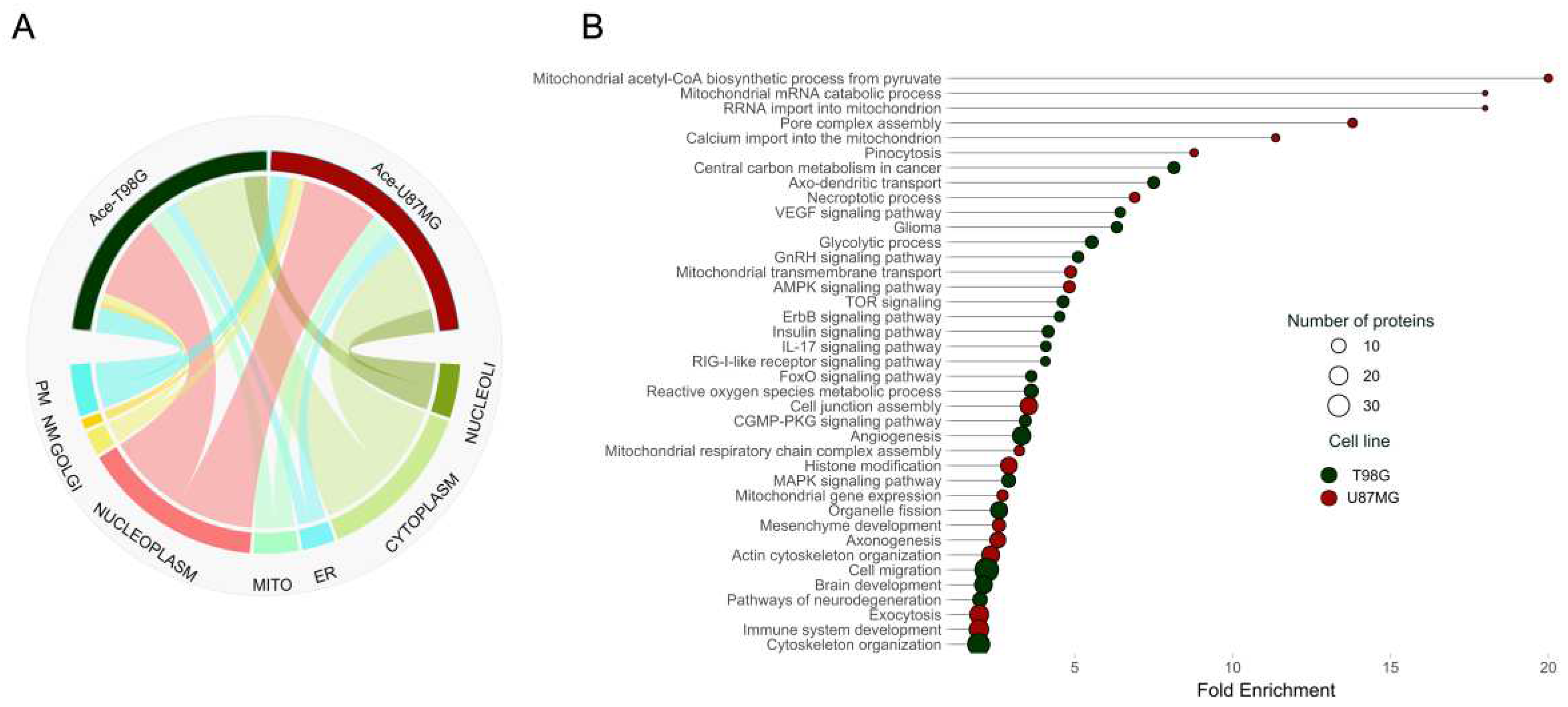
2.1. Proteome Dynamics Supporting Phenotypic Features of Glioblastoma Cells.
2.2. Acetylation Profile in Oxidative and Glycolytic Glioblastoma Cell Lines.
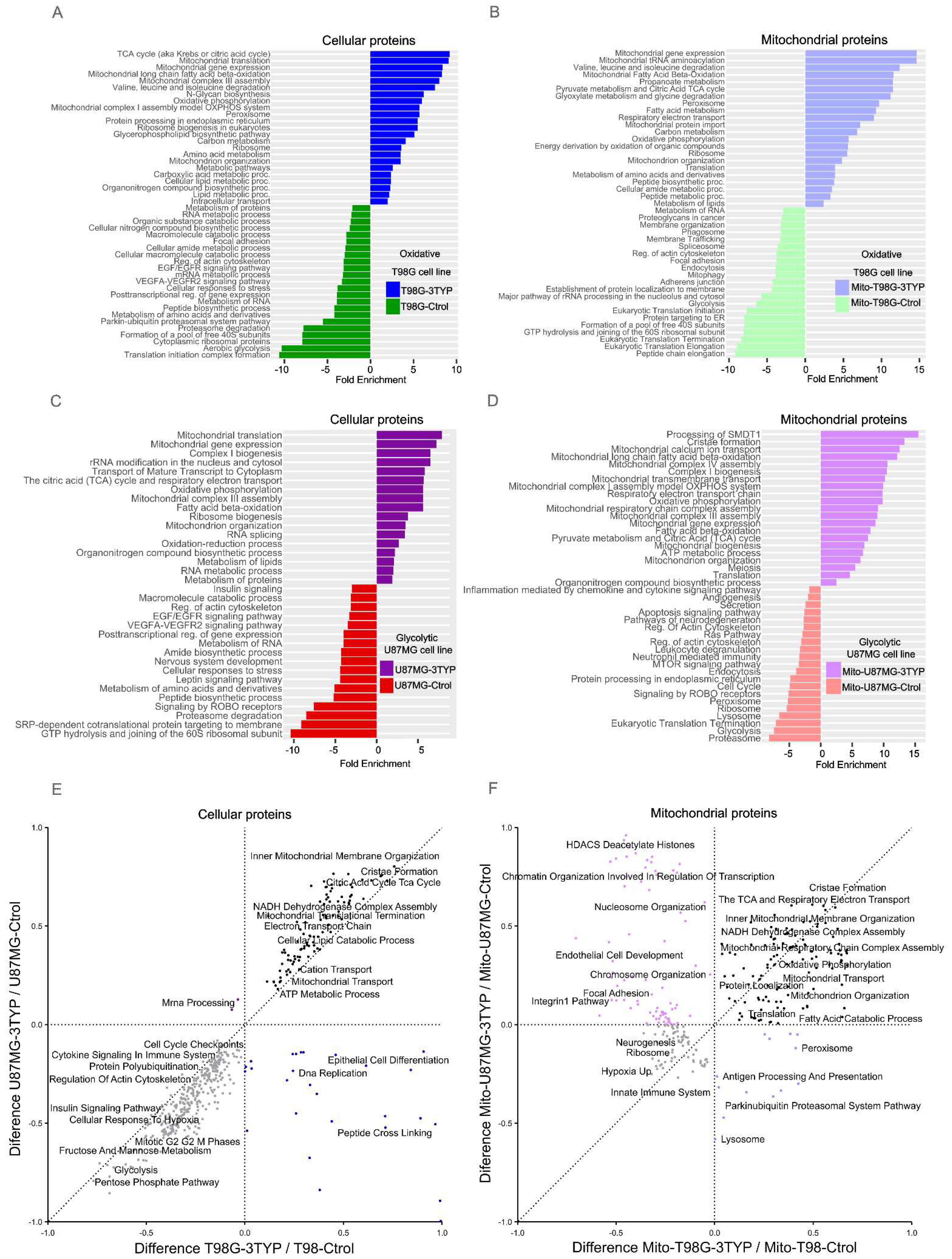
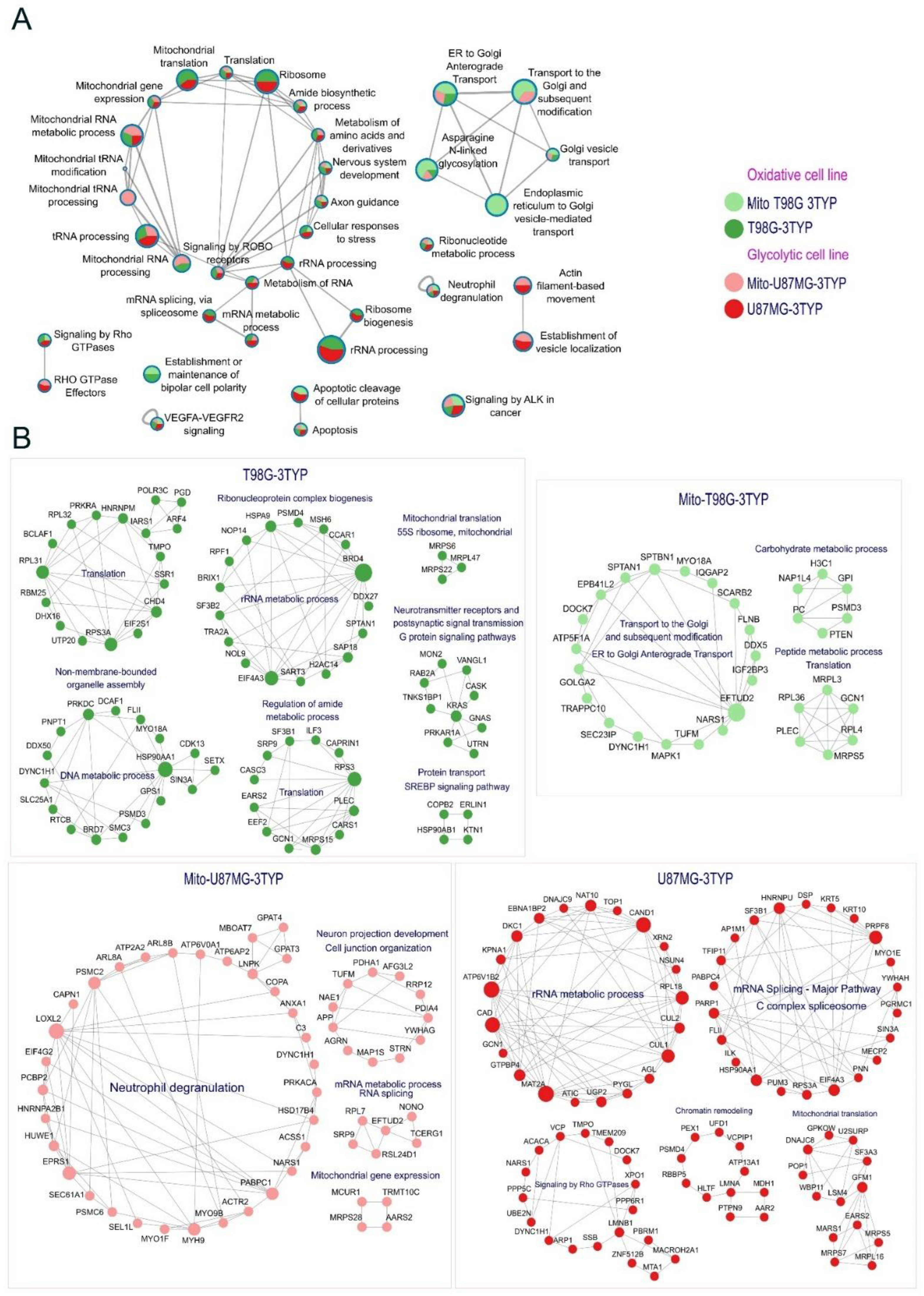
2.3. Mitochondrial Pathway Proteins are Overexpressed in Response to SIRT3 Inhibition in Glycolytic and Oxidative Glioblastoma Cell Lines.
2.4. SIRT3 Regulates the Synthesis of Proteins.
3. Materials and Methods
Cell Culture
SIRT3 Selective Inhibition Treatment with a 3-TYP Reagent
Protein Extraction
Mitochondrial Purification
Chemical Acetylation with Deuterated N-acetoxysuccinimide (NAS-d3)
LC-MS/MS and Data Analysis
4. Conclusions
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
Appendix A
References
- Francisco S (2016) Epidemiology 1. 134:. [CrossRef]
- Wilson TA, Karajannis MA, Harter DH (2014) Glioblastoma multiforme: State of the art and future therapeutics. Surg Neurol Int 5:. [CrossRef]
- Pet F Cover image: Primary diagnosis of a right frontal glioblastoma following acquisition of axial slices of T1-weighted gadolinium-enhanced MRI (left) and 18F-FDG PET (right). See page 158, chapter 9 for details.
- Duraj T, García-romero N, Carrión-navarro J, et al (2021) Beyond the warburg effect: Oxidative and glycolytic phenotypes coexist within the metabolic heterogeneity of glioblastoma. Cells 10:1–23. [CrossRef]
- Vlashi E, Lagadec C, Vergnes L, et al (2011) Metabolic state of glioma stem cells and nontumorigenic cells. Proc Natl Acad Sci U S A 108:16062–16067. [CrossRef]
- Shibao S, Minami N, Koike N, et al (2018) Metabolic heterogeneity and plasticity of glioma stem cells in a mouse glioblastoma model. Neuro Oncol 20:343–354. [CrossRef]
- Ramão A, Gimenez M, Laure HJ, et al (2012) Changes in the expression of proteins associated with aerobic glycolysis and cell migration are involved in tumorigenic ability of two glioma cell lines. Proteome Sci 10:53. [CrossRef]
- Wang Q, Zhang Y, Yang C, et al (2010) Acetylation of metabolic enzymes coordinates carbon source utilization and metabolic flux. Science (80- ) 327:1004–1007. [CrossRef]
- Arenas A, Chen J, Kuang L, et al (2020) Lysine acetylation regulates the RNA binding, subcellular localization and inclusion formation of FUS. Hum Mol Genet 29:2684–2697. [CrossRef]
- Fujita Y, Fujiwara K, Zenitani S, Yamashita T (2015) Acetylation of NDPK-D regulates its subcellular localization and cell survival. PLoS One 10:1–18. [CrossRef]
- Song EH, Oh W, Ulu A, et al (2015) Acetylation of the RhoA GEF Net1A controls its subcellular localization and activity. J Cell Sci 128:913–922. [CrossRef]
- Rardin MJ, Newman JC, Held JM, et al (2013) Label-free quantitative proteomics of the lysine acetylome in mitochondria identifies substrates of SIRT3 in metabolic pathways. Proc Natl Acad Sci U S A 110:6601–6606. [CrossRef]
- Gil J, Rezeli M, Lutz EG, et al (2021) An observational study on the molecular profiling of primary melanomas reveals a progression dependence on mitochondrial activation. Cancers (Basel) 13:. [CrossRef]
- Gil J, Betancourt LH, Pla I, et al (2019) Clinical protein science in translational medicine targeting malignant melanoma. Cell Biol Toxicol 35:293–332. [CrossRef]
- Gil J, Ramírez-Torres A, Chiappe D, et al (2017) Lysine acetylation stoichiometry and proteomics analyses reveal pathways regulated by sirtuin 1 in human cells. J Biol Chem 292:18129–18144. [CrossRef]
- Kuras M, Woldmar N, Kim Y, et al (2021) Proteomic Workflows for High-Quality Quantitative Proteome and Post-Translational Modification Analysis of Clinically Relevant Samples from Formalin-Fixed Paraffin-Embedded Archives. J Proteome Res 20:1027–1039. [CrossRef]
- Betancourt LH, Gil J, Sanchez A, et al (2021) The Human Melanoma Proteome Atlas-Complementing the melanoma transcriptome. Clin Transl Med 11:. [CrossRef]
- Elwell CE, Leung TS, Editors DKH Oxygen Transport to Tissue XXXVII.
- Uhlén M, Fagerberg L, Hallström BM, et al (2015) Tissue-based map of the human proteome. Science (80- ) 347:. [CrossRef]
- Cerchia L, Esposito CL, Jacobs AH, et al (2009) Differential SELEX in human glioma cell lines. PLoS One 4:e7971. [CrossRef]
- Choudhary C, Kumar C, Gnad F, et al (2009) Lysine acetylation targets protein complexes and co-regulated major cellular functions. Science (80- ) 325:834–840. [CrossRef]
- Allfrey VG, Faulkner R, Mirsky AE (1964) Acetylation and methylation of histones and their possible role in the regulation of RNA synthesis. Proc Natl Acad Sci U S A 51:786–794. [CrossRef]
- Access O (2021) immunosuppression in glioblastoma. 1111–1124. [CrossRef]
- Song H, Li C, Labaff AM, et al (2012) NIH Public Access. [CrossRef]
- Li P, Ge J, Li H Lysine acetyltransferases and lysine deacetylases as targets for cardiovascular disease. Nat Rev Cardiol 1:. [CrossRef]
- Manuscript A (2011) NIH Public Access. 22:263–268. [CrossRef]
- Kong F, Ma L, Wang X, et al (2022) Regulation of epithelial - mesenchymal transition by protein lysine acetylation. Cell Commun Signal 1–14. [CrossRef]
- Bradley RK, Anczuków O (2023) RNA splicing dysregulation and the hallmarks of cancer. Nat Rev Cancer 23:135–155. [CrossRef]
- Zhang J, Xiang H, Liu J, et al (2020) Mitochondrial Sirtuin 3: New emerging biological function and therapeutic target. Theranostics 10:8315–8342. [CrossRef]
- Zeng Z, Yang Y, Dai X, et al (2016) Polydatin ameliorates injury to the small intestine induced by hemorrhagic shock via SIRT3 activation-mediated mitochondrial protection. Expert Opin Ther Targets 20:645–652. [CrossRef]
- Alhazzazi TY, Kamarajan P, Verdin E, Kapila YL (2011) SIRT3 and cancer: Tumor promoter or suppressor? Biochim Biophys Acta - Rev Cancer 1816:80–88. [CrossRef]
- Wei Z, Song J, Wang G, et al Deacetylation of serine hydroxymethyl-transferase 2 by SIRT3 promotes colorectal carcinogenesis. Nat Commun. [CrossRef]
- Alhazzazi TY, Kamarajan P, Joo N, Huang J (2011) Sirtuin-3 ( SIRT3 ), a Novel Potential Therapeutic Target for Oral Cancer. 3:1670–1678. [CrossRef]
- Kim HS, Patel K, Muldoon-Jacobs K, et al (2010) SIRT3 Is a Mitochondria-Localized Tumor Suppressor Required for Maintenance of Mitochondrial Integrity and Metabolism during Stress. Cancer Cell 17:41–52. [CrossRef]
- Srivastava SP, Li J, Kitada M, et al (2018) SIRT3 de fi ciency leads to induction of abnormal glycolysis in diabetic kidney with fi brosis. [CrossRef]
- Yang Y, Cimen H, Han MJ, et al (2010) NAD+-dependent deacetylase SIRT3 regulates mitochondrial protein synthesis by deacetylation of the ribosomal protein MRPL10. J Biol Chem 285:7417–7429. [CrossRef]
- Haase M, Fitze G, Haase M, Fitze G (2015) HSP90AB1: Helping the good and the bad. Gene. [CrossRef]
- Lu Q, Guo P, Li H, et al (2022) Ecotoxicology and Environmental Safety Targeting the lncMST-EPRS / HSP90AB1 complex as novel therapeutic strategy for T-2 toxin-induced growth retardation. Ecotoxicol Environ Saf 247:114243. [CrossRef]
- Boyault C, Khochbin S (2005) Regulatory cross-talk between lysine acetylation and ubiquitination : role in the control of protein stability. 408–415. [CrossRef]
- Pan Z, Bao Y, Hu M, et al (2023) Role of NAT10-mediated ac4C-modi fi ed HSP90AA1 RNA acetylation in ER stress-mediated metastasis and lenvatinib resistance in hepatocellular carcinoma. 1–14. [CrossRef]
- Cai S, Liu X, Zhang C, et al (2016) Biochemical and Biophysical Research Communications Autoacetylation of NAT10 is critical for its function in rRNA transcription activation. Biochem Biophys Res Commun 1–6. [CrossRef]
- Li J, Wang T, Xia J, et al (2019) Enzymatic and nonenzymatic protein acetylations control glycolysis process in liver diseases. 1–15. [CrossRef]
- Tao Y, Yin H, Zhou L, et al This article has been accepted for publication and undergone full peer review but has not been through the copyediting , typesetting , pagination and proofreading process which may lead to differences between this version and the Version of Record . Pleas.
- Perez-Riverol Y, Bai J, Bandla C, et al (2022) The PRIDE database resources in 2022: A hub for mass spectrometry-based proteomics evidences. Nucleic Acids Res 50:D543–D552. [CrossRef]
- Tyanova S, Temu T, Sinitcyn P, et al (2016) The Perseus computational platform for comprehensive analysis of (prote)omics data. Nat. Methods 13:731–740. [CrossRef]
- Zhou Y, Zhou B, Pache L, et al (2019) Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat Commun 10:. [CrossRef]
- Mi H, Muruganujan A, Ebert D, et al (2019) PANTHER version 14: More genomes, a new PANTHER GO-slim and improvements in enrichment analysis tools. Nucleic Acids Res 47:D419–D426. [CrossRef]
- Raj RM, Sreeja A (2018) Analysis of computational gene prioritization approaches. Procedia Comput Sci 143:395–410. [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



